Abstract
Purpose
The human epidermal growth factor receptor 2 (HER2) status in breast cancer is important for prognostic prediction and the determination of optimal treatment. Current methods rely on protein expression, as determined by immunohistochemistry (IHC), as well as gene amplification as determined by in situ hybridisation (ISH). We explored whether quantitative droplet digital PCR (ddPCR) can be used for the detection and absolute quantitation of HER2 mRNA.
Methods
Digital droplet PCR (ddPCR) was performed for HER2 mRNA on 178 formalin-fixed paraffin-embedded (FFPE) breast cancer specimens. HER2 positive, equivocal and negative cases as defined by standard criteria were included and both core biopsies and tissue sections were assessed.
Results
HER2 positive cases contained significantly higher levels of HER2 mRNA (169–1,000,000 copies/µl) by ddPCR compared with equivocal (112–139 copies/µl, p = 0.025) and negative cases (6.2–644 copies/µl. p < 0.001). A continuum of transcript quantity was observed but a cutoff of 490 copies/µl distinguished between HER2 positive and negative cases. Results were consistent between core biopsy and tissue sections.
Conclusions
ddPCR can be used to quantify HER2 mRNA transcripts in FFPE breast cancer specimens. Our results highlight the potential of ddPCR on FFPE tissue to be used to accurately quantify HER2 transcripts. Validation in large cohorts will be required to determine a clinically applicable cutoff.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The human epidermal growth factor receptor 2 gene (HER2) is responsible for the formation of the HER2 receptor on the surface of epithelial cells. HER2 is overexpressed in 15–20% cases of breast cancer and is important in diagnosis. It is a major negative prognostic factor associated with a shorter survival time, a higher percentage of cancer recurrences and a lower response to chemotherapy and hormone therapy [1]. Determination of HER2 status is also important in the treatment of breast cancer as some patients with HER2-positive disease respond well to the anti-HER2 humanised monoclonal antibody, trastuzumab [2]. HER2 status is generally assessed by protein expression by immunohistochemistry (IHC) and/or gene amplification by in situ hybridisation (ISH). In general there is good but variable concordance between IHC and gene amplification for HER2 status [3–6].
In addition to HER2 protein and gene amplification, a number of techniques have been applied to assess HER2 mRNA. These include quantitative (Q) or reverse transcription (RT) polymerase chain reaction (PCR) and RNA in situ hybridisation (RNA-ISH) [7–9]. Most of these RNA-based studies have reported high concordance with both IHC and gene amplification, ranging from 86.0 to 99.4% [7–10]. However, this is not universal as a recent large multi-centre RT-PCR study has shown. This study reported “positive agreement of less than 50%” with IHC and ISH and a high false-negative rate for HER2 positive cases [11]. Further, all these previously reported PCR-based studies have provided only relative, semi-quantitative assessments of HER2 transcripts compared to a universal reference gene. Therefore, these mRNA methods have not gained widespread acceptance for clinical use.
Droplet digital PCR (ddPCR) is another RNA-based method, which in contrast to the other methods, can provide absolute quantification of the number of mRNA transcripts [12]. This is based on partitioning the PCR reaction mixture into thousands of droplets so that each droplet contains either 1 or 0 molecules of target RNA. The droplets then undergo thermal cycling to generate an amplified product. A positive fluorescence signal is present only in the droplets that contain a target molecule. It is an objective method that is capable of quantifying mRNA expression with high precision [13, 14]. ddPCR can also be used to quantify DNA [12, 15–17]. In studies comparing HER2 DNA in breast and gastric cancers by ddPCR and conventional methods (i.e., IHC and ISH) there was variable concordance (90.6–100% and 67.6–96% respectively). A more recent study comparing HER2 DNA in breast cancer cell lines by ddPCR, QPCR and next-generation sequencing (NGS) reported that each method demonstrated slightly different HER2 copy numbers that were likely associated with platform-specific biases [18]. To date, no equivalent study has been undertaken for mRNA quantitation on breast cancer. In this study, we have therefore assessed ddPCR for its ability to quantitate HER2 gene expression (mRNA transcript) in formalin fixed paraffin embedded (FFPE) tissue of breast cancer.
Materials and methods
Samples
A total of 178 FFPE breast cancer specimens from patients with breast cancer diagnosed between 2011 and 2013 were selected to provide sufficient representation of all categories from the PathWest Laboratory Medicine database at Queen Elizabeth II Medical Centre, Western Australia. Cases (median age = 62, range 27–95) with a confirmed diagnosis of carcinoma of the breast based on histopathology of resected material were included. All cases were required to have sufficient FFPE tumour tissue available for study. Exclusions were history of a prior breast cancer or any other clinically active malignancy. Healthy controls (median age = 38, range 26–48) were included if the breast biopsy or specimen was shown to have normal breast histology with no evidence of malignancy on histopathological review. The healthy control cases were patients who had undergone cosmetic breast reduction surgery and no other clinically relevant history and no history of malignancy. The clinico-pathological details of the cases are listed in Table 1. Six ‘normal’ breast specimens collected from women undergoing breast reduction surgery and six blood specimens (EDTA anticoagulated tubes) from healthy volunteers were tested as biological negative controls. Ethical approval for this study was obtained from the Sir Charles Gardiner Hospital Human Research Ethics Committee (HREC Number 2013-051, July 2013–16). For this type of study, formal consent was not required and a waiver of consent was obtained for all participants included in the study.
Pathology assessment
All FFPE samples were assessed by standard haematoxylin and eosin (H&E) staining, IHC and silver-enhanced in situ hybridisation (SISH). IHC and SISH were performed on a BenchMark XT-automated IHC/ISH slide stainer (Ventana, USA) using the Clone c-erbB-2 antibody (Dako, Denmark) and the INFORM HER2 Dual ISH DNA Probe Cocktail Assay (Ventana, USA) respectively. Both were visualised by light microscopy and reviewed by expert breast pathologists. HER2 status was categorised, in accordance with 2013 ASCO/CAP guidelines, as (Table 1):
-
(i)
positive (IHC 3+ or SISH showing ≥6 signals/cell)
-
(ii)
equivocal (IHC 2+ and/or SISH showing ≥4 and <6 signals/cell); or
-
(iii)
negative (IHC 0/1+ or SISH showing <4 signals/cell) [19].
Cell culture
To determine whether ddPCR could be performed reproducibly in FFPE, we compared the results for two cell lines that were fresh and also made into a cell pellet (formalin fixed and embedded in paraffin). We tested two breast cancer cell lines (MCF7 and MDA-MB-231), both fresh and FFPE. MCF7 cells are known to express HER2 and MDA-MB-231 cells are considered negative for HER2.The MCF-7 (HER2 positive) and MDA-MB-231 (HER2 negative) breast cancer cell lines were obtained from the American Type Culture Collection (ATCC, USA) and grown in RPMI-1640 (ThermoScientific, USA) with 10% Foetal Bovine Serum and 1% Penicillin and Streptomycin (ThermoScientific, USA). Fresh cells were harvested by trypsinisation (ThermoScientific, USA), washed in phosphate buffered saline, counted using a hematocytometer and processed as per the human breast biopsies. The cell lines were also processed into a cell block, fixed in formalin and embedded in paraffin using standard tissue processing methods. For this, cells were trypsinised, washed and pelleted via centrifugation at 800 g for 10 min. Expired human plasma (Transfusion Medicine Laboratory, PathWest, WA) was added to the cell pellet in a 1:1 ratio and mixed with a glass pipette. Following this, three drops of Liquid Fib (Diagnostica Stago S.A.S., France) were added and the suspension incubated at room temperature for 2 min. Pelleted cells were dislodged, fixed in 10% formalin (Amber Scientific, Australia) for a minimum of 2 h and processed into paraffin using a Leica Peloris II Tissue processor (Leica Biosystems, USA) according to standard histological practice. Sections of the embedded cells were cut at 4 and 10 μm and processed as per the human breast biopsies.
RNA isolation
For each case, a single, 2 mm core biopsy approximately 5 mm deep was taken from a region of the tissue with >70% tumour cells using a sterile 2-mm biopsy punch (Kai Industries, Japan). Five HER2-negative cases were randomly selected as biological replicates; for these 3 × 2 mm cores were taken from different areas of the tumour each with >70% tumour cells. Four and 10 µm sections were cut from six cases to test the limit of detection. Tissue sections or core biopsies were deparaffinised with a series of xylene and ethanol washes. Total RNA was then extracted using the MasterPure Complete DNA and RNA Purification Kit (Epicentre Biotechnologies, USA) according to the manufacturer’s instructions. In brief, each sample was incubated at 65 °C overnight with proteinase K. Following digestion, a series of protein precipitation and isopropanol washes were carried out to remove proteins and precipitate the nucleic acids. A DNase I step was included to ensure the removal of all genomic and contaminating DNA from the total nucleic acid preparations. RNA was quantitatively and qualitatively assessed using the Qubit Fluorometer (Life Technologies, USA) and NanoDrop Spectrophotometer (ThermoScientific, USA) respectively.
Complementary DNA preparation
Total RNA was reverse transcribed into complementary DNA (cDNA) using iScript Advanced cDNA Synthesis Kit (BioRad, USA) according to the manufacturer’s instructions. In brief, each 20µL reaction mix contained 1000 ng of RNA, 5 × advanced reaction mix and advanced reverse transcriptase. The complete reaction mix was amplified using the Veriti 96-Well Thermal Cycler (Life Technologies, USA) as per the manufacture’s cycling conditions. The samples were reverse transcribed for 30 min at 42 °C followed by enzyme inactivation for 5 min at 85 °C.
Droplet digital PCR
The ddPCR reaction was carried out as previously described [12, 13], with one exception. Given that FFPE RNA and DNA can be highly degraded, we omitted the restriction enzyme treatment step. Three cases were randomly selected as technical replicates and cDNA from the same core was assayed by ddPCR in triplicate. Briefly, each 20 µL reaction mixture contained 100 ng of cDNA, 2× ddPCR probe mastermix (Bio-Rad, USA) and ERBB2/6-carboxyfluorescein-minor groove binder (FAM-MGB) primer/probe set (HER2: Hs01001580_m1: ThermoFisher Scientific, USA). No template and RNA only negative controls were included in each assay. Droplet formation was carried out using a QX100 droplet generator (BioRad, USA). Each sample was partitioned into an emulsion of approximately 20,000 uniformly sized nanoliter droplets. On average, each droplet contained one template and sufficient mastermix reagent and primer/probe to carry out a PCR reaction. The droplet emulsion was transferred to a 96-well plate (Eppendorf, Germany), heat-sealed at 180 °C for 5 s with foil. Thermal cycling was performed using the C1000 Touch Thermal Cycler—BioRad, USA) as per the manufacturer’s cycling conditions. Each plate was incubated at 95 °C for 10 min, followed by 40 cycles of 94 °C for 30 s and 60 °C for 1 min, with a last 10 min incubation at 98 °C. After the PCR, the RNA content of the droplets was quantified using the QX200 Droplet Reader (BioRad, USA). ddPCR data were analysed using QuantaSoft software (version 1.6.6; Bio-Rad). The fraction of positive and negative droplets was determined and data were fitted to a Poisson distribution to determine the absolute concentration (copies/µL) of the target sequence.
Statistical analyses
A Kruskal–Wallis test was performed to assess whether there were statistically significant differences between HER2 mRNA levels and the IHC result, and HER2 mRNA levels between 4 and 10 µm sections and the core biopsies. A two-tailed Pearson correlation was used to measure the relationship between ddPCR and SISH. Principal Components Analysis (PCA) was performed to provide a meaningful visual representation of the structure of cases and variables. ClustVis using singular value decomposition (SVD) with imputation was used to calculate principal components [20]. Receiver operating characteristic (ROC) analysis was performed to evaluate the discriminatory ability of HER2 mRNA copy number, to establish a ‘cutoff’ value for ddPCR and calculate the positive predictive value (PPV), negative predictive value (NPV), sensitivity, specificity and accuracy. Intra-class correlation coefficients (ICCs) and Pearson correlations were calculated to provide a statistical means of testing the similarity between 4, 10 µm and core biopsies. P values presented are two-sided and were considered significant at an alpha level of 0.05. Statistical analyses were carried out using SPSS version 22.0.
Results
To determine whether ddPCR and the criteria of HER2 copy number can be used on fresh as well as FFPE specimens, we tested two breast cancer cell lines (MCF7 and MDA-MB-231) in each condition. Comparison of the fresh and FFPE specimens by ddPCR showed no significant difference in HER2 copy number (p > 0.01; Table 2).
The mean patient age was 62 years (range, 27–95 years) and the majority of cases were invasive ductal carcinomas (91%) with all three Nottingham Grades represented (Table 1). Of the 178 cases, 55 (31%) were HER2 positive, 117 (66%) HER2 negative and 6 (3%) equivocal based on the ASCO/CAP Guideline Recommendations [1]. There was a 96% concordance between the HER2 results for IHC and gene amplification. The six discordant cases were weakly positive (1+) by IHC but showed gene amplification (≥6 signals/cell) by SISH and were classified as HER2 positive.
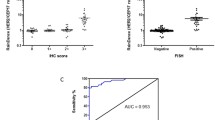
RNA quality was assessed using Bioanalyzer and Nanodrop analyses. The RNA was of sufficient quality in all the cases whether from 4 to 10 µm thickness sections or the 2 mm core biopsies to be reverse transcribed and amplified using industry validated primers specific for a conserved region in the HER2 gene. In the 2 mm cores, the HER2 mRNA levels ranged from 6.2 to 1,000,000 copies/µl. The HER2 positive (n = 55) cases had statistically significant higher levels (32–1,000,000 copies/µl, mean = 22,392, median = 1098) than HER2 equivocal (n = 6) (54–1048 copies/µl, mean = 125, median = 125, p = 0.025) and negative (n = 117) cases (6–644 copies/µl, mean = 138, median = 107, p < 0.001) (Fig. 1). In addition, a strong, positive correlation was observed between HER2 mRNA levels and gene amplification (R = 0.723, p < 0.001, Fig. 2). The ddPCR approach showed that discordant cases had a wide range of mRNA levels (169–859 copies/µl, Fig. 3). However when visualised using a waterfall plot, the data appeared as a continuum with no clear distinction between HER2 positive and HER2 negative cases (Fig. 3).
HER2 mRNA concentration by ddPCR categorised by HER2 status. The HER2 mRNA concentration is plotted in descending order (natural logarithmic scale). HER2 positive cases (based on standard criteria) are displayed as light grey bars and HER2 negative cases are displayed as black bars. Equivocal cases are marked pink with a ‘x’. Discordant cases (by IHC and SISH) are marked with ‘o’. The dashed line indicates 490 copies/µL HER2 mRNA concentration threshold. Cases to the left of the dashed line had a result greater than 490 copies/µL HER2 mRNA
Technical replicates, performed to assess assay robustness and reproducibility, showed a standard deviation range from 1 to 165 across samples with low (15 copies/µl) to high (5033 copies/µl) HER2 mRNA expression (Supplementary Data). Biological replicates from HER2 negative cases (ranging from 0 to 2+ by IHC and 1.7–2.8 signals per cell by ISH) were analysed. Cores taken from different areas of each tumour varied with standard deviations ranging from 29 to 316 copies/µl (Fig. 4). The ‘no template’ and ‘RNA only’ technical, negative controls were consistently negative as expected. The HER2 mRNA expression ranged from 1 to 32 copies/µl (mean 13, standard deviation 14) in normal breast tissue and from 0 to 6 copies/µl in normal blood (mean 2, standard deviation 3).
Principal component analysis (PCA), used to visualise patterns within the dataset (Fig. 5), showed that 77.8 and 16.4% of the total variance was due to ddPCR (PCA1) and SISH (PCA2), respectively. PCA revealed that HER2 positive, equivocal and negative cases group closely together and were discrete from each other. This suggests that cases that were negative by ddPCR were also negative by SISH and so on for positive and equivocal cases. HER2 mRNA levels were significantly higher in IHC 3+ stained cases compared with equivocal (2 +), weak (1 +) and negative (0) cases (p < 0.001) (Fig. 6). In addition, mRNA concentrations were significantly different between equivocal and weak (p = 0.015) or negative cases (p < 0.001).
Stratification of ddPCR HER2 mRNA values by IHC scores. The concentration of HER2 mRNA was significantly higher in IHC 3+ cases compared with equivocal (2+), weak (1+) and negative cases (***p < 0.001). In addition, HER2 mRNA was significantly higher in equivocal cases compared with weak (**p = 0.015) and negative cases (***p < 0.001). Dashed line indicates 490 copies/µL HER2 mRNA concentration threshold
ROC curve analysis was performed to estimate HER2 gene transcript level that could discriminate HER2 positive from HER2 negative cases within this cohort. At a level of 490 copies/µl (dashed line, Fig. 3), ddPCR reached 85% sensitivity, 98% specificity, 85% PPV, 98% NPV and was 94% accurate (Area Under the Curve (AUC), 0.973; 95% CI, 0.951–0.995; p < 0.001). Comparison of AUCs for ddPCR (AUC, 0.973; 95% CI, 0.951–0.995; p < 0.001) and IHC (AUC, 0.935; 95% CI, 0.898–0.972; p < 0.001) showed ddPCR to be the superior, as the AUC was greater and the confidence interval was narrower. When this 490 copies/µl cutoff was applied, 8 HER2 positive cases had HER2 mRNA expression levels below this cutoff (range 112–425 copies/µl) and 1 HER2 negative case had 644 copies/µl and was above the cutoff. Both equivocal cases were classified as negative based on ddPCR results. A similar trend was reported regardless of whether cores or sections were used demonstrating the potential of this approach for routine diagnostic use.
Having determined that 2 mm core biopsies could discriminate HER2 positive from HER2 negative cases, we then applied this approach to the tissue sections to determine the sensitivity and diagnostic potential of this method. No significant differences were seen in HER2 transcript levels between the core biopsies and either the 4 or 10 µm sections (Fig. 7). We observed strong, statistically significant correlations between 4 and 10 µm sections (R = 0.995, p < 0.001), 4 µm sections and cores and (R = 0.981, p = 0.003) 10 µm sections and cores (R = 0.968, p = 0.007). The ICC (0.971, p < 0.001 with 95% confidence 0.852 to 0.997) supported this and demonstrates no significant difference based on the amount of tissue assessed. We then applied the 490 copies/µl cut off and found that it was 100% successful in discriminating HER2 positive from HER2 negative cases. The HER2 equivocal case analysed in the three sample types was consistently negative by ddPCR (Fig. 7).
Discussion
HER2 status is important in determining the prognosis and for the management of breast cancer. This is the first comprehensive report demonstrating that ddPCR can provide an absolute quantitation of HER2 gene expression (mRNA) in FFPE breast cancer specimens. Overall, the results were accurate and technically reproducible and this was independent of the amount of tissue analysed. This is in agreement with other ddPCR studies (both DNA and RNA based) of breast and gastric cancers and therefore shows the robustness of this approach in routine specimens [12, 15–18].
RNA in FFPE specimens has varying levels of degradation due to its inherent instability as a single-stranded molecule. As a result, RNA can be difficult to analyse using traditional qPCR approaches. Further, qPCR is a semi-quantitative method and cannot provide an absolute quantification of RNA transcripts. We opted to use the more accurate method of ddPCR in this study for the reason that is quantitative, reproducible and generates a precise number of transcripts without the need for external calibrators. In addition, ddPCR has been optimised for low amounts of target RNA and is ideal for analysis of commonly available FFPE specimens. We show that 6 to 1000,000 copies/µl of HER2 transcript could be detected in breast cancers with clinically defined HER2 status ranging from negative to positive. Only one other study has used ddPCR to study HER2 transcript expression breast cancer [12]. This study, unlike ours, used a semi-quantitative approach and HER2 mRNA expression was normalised relative to standard reference genes. This is a limitation as reference genes are known to fluctuate significantly under pathological conditions [21]. By optimising each step of our ddPCR assay, we have harnessed the absolute quantitative power and technical reproducibility of this approach to accurately enumerate the number of HER2 mRNA transcripts without the need for normalisation against reference gene.
Biological replicates showed that there is intra-tumoural heterogeneity in HER2 transcript levels, consistent with other reports [22, 23]. Our results show that biological heterogeneity only had an impact on the overall HER2 result for one case. This case was categorised as HER2 negative by IHC and SISH but the ddPCR results favoured a positive result (mean = 540, standard deviation = 146). Overall, this finding is similar to another report and suggests that a single representative core or section of a tumour mass is sufficient for an accurate HER2 determination [24].
To assess the potential of ddPCR for application in routine diagnostics, we tested this approach using 4 and 10 µm sections which we estimate to have had 103 fewer cells than the core biopsies [25, 26]. We confirmed that despite this difference in number of assayed cells similar HER2 transcript levels were measured. This result was highly reproducible in technical replicates for both sections and cores, in keeping with the findings of another ddPCR study [27]. This further supports ddPCR as a robust assay that upon repeat testing generates the same (or very similar) result. In addition, we showed that, using two breast cancer cell lines, there was no reduction in the number of copies of HER2 following formalin fixation and embedding into paraffin. This confirmed that FFPE material was suitable for quantifying HER2 copies and could be applied to the routine processed clinical material.
HER2 status is determined by analysis of small amounts of tissue: IHC bases the result on as little as 30% of the tumour and SISH by counting signals in 20–60 cells [1]. The approach that we have used has the capacity to analyse ~20-times more cells in the same amount of material as conventional testing. This will increase sensitivity in accuracy determining HER2 status using a semi-automated, quantitative method. Despite a continuum of HER2 mRNA expression in the breast cancer cases, ddPCR results strongly correlated with gene amplification. When the 490 copies/µl cutoff (determined by ROC analyses) was applied, 8 HER2 positive cases had HER2 mRNA expression levels below this cutoff (range 112–425 copies/µl) and 1 HER2 negative case had 644 copies/µl and was above the cutoff. Both equivocal cases were classified as negative based on ddPCR results. A similar trend was reported regardless of whether cores or sections were used demonstrating the potential of this approach for routine diagnostic use. Quality control measures suggest that the false negative results in HER2-positive cases were not due to RNA quantity or quality. However, we cannot rule out sampling error and tumour heterogeneity. We anticipate that testing this approach in a larger cohort will refine the cut-off level to eliminate false negative results. Another explanation could be that the HER2 status was in error. We therefore reviewed the IHC and SISH data of each of the HER2-positive cases with <490 copies. All of these 8 “false-negative” cases were confirmed HER2 positive based on gene amplification by SISH (>6 copies). Of these, two had an IHC score of 3+ with 209 and 425 copies/µl of HER2 transcript. The other 6 ddPCR “false negative” cases had equivocal (2+) or weak (1+) IHC staining results and contained lower ranges of HER2 transcripts (186–325 and 169–396 copies/µl respectively). A possible biological explanation for these results is that increased HER2 gene copy number (via gene amplification or polysomy 17) does not universally lead to increased gene expression (transcripts) and translation to HER2 protein (IHC). Also of interest were the six cases with discrepant IHC and SISH results. ddPCR supported the positive result obtained by SISH for two of these cases suggesting that they may best be classified as HER2 positive. In contrast, HER2 mRNA levels were below the 490 copies/µl cutoff for the remaining four cases suggesting that these cases may be best classified as negative. A possible explanation for this could be that although gene amplification occurred (as indicated by the high SISH values) subsequent gene transcription and translation did not (as per the low ddPCR and IHC result).
In summary, this study demonstrates the potential utility of ddPCR as an accurate objective and practical method for assessing HER2 status in FFPE. We have shown that ddPCR can be applied to standard sections and core biopsies and gives a precise numerical measure without requiring a reference gene and normalisation. The result generated is more representative of the tumour since the number of cells analysed is more than 20-times greater than ISH. Validation in large, independent cohorts will be required to determine whether an absolute number of HER2 transcripts can discriminate between HER2-positive and negative breast cancer. An increased sample number will refine the cutoff, maximise both the positive and negative predictive values and subsequently minimise both the false positive and false negative rates (4.5 and 0.5% in the current study). This is essential before ddPCR for HER2 could be considered as a robust alterative to the current gold standard methods (IHC and ISH).
References
Wolff AC, Hammond MEH, Hicks DG, Dowsett M, McShane LM, Allison KH, et al. Recommendations for human epidermal growth factor receptor 2 testing in breast cancer: American Society of Clinical Oncology/College of American Pathologists clinical practice guideline update. J Clin Oncol. 2013;31(31):3997–4013. doi:10.1200/Jco.2013.50.9984.
Puglisi F, Fontanella C, Amoroso V, Bianchi GV, Bisagni G, Falci C, et al. Current challenges in HER2-positive breast cancer. Crit Rev Onc/Hem. 2016;98:211–21. doi:10.1016/j.critrevonc.2015.10.016.
Press MF, Sauter G, Bernstein L, Villalobos IE, Mirlacher M, Zhou JY, et al. Diagnostic evaluation of HER-2 as a molecular target: an assessment of accuracy and reproducibility of laboratory testing in large, prospective, randomized clinical trials. Clin Can Res. 2005;11(18):6598–607. doi:10.1158/1078-0432.CCR-05-0636.
Perez EA, Suman VJ, Davidson NE, Martino S, Kaufman PA, Lingle WL, et al. HER2 testing by local, central, and reference laboratories in specimens from the North Central Cancer Treatment Group N9831 intergroup adjuvant trial. J Clin Oncol. 2006;24(19):3032–8. doi:10.1200/JCO.2005.03.4744.
Roche PC, Suman VJ, Jenkins RB, Davidson NE, Martino S, Kaufman PA, et al. Concordance between local and central laboratory HER2 testing in the breast intergroup trial N9831. JNCI. 2002;94(11):855–7.
Varga Z, Tubbs RR, Moch H. Concomitant detection of HER2 protein and gene alterations by immunohistochemistry (IHC) and silver enhanced in situ hybridization (SISH) identifies HER2 positive breast cancer with and without gene amplification. PLoS One. 2014;9(8):e105961. doi:10.1371/journal.pone.0105961.
Wang Z, Portier BP, Gruver AM, Bui S, Wang H, Su N, et al. Automated quantitative RNA in situ hybridization for resolution of equivocal and heterogeneous ERBB2 (HER2) status in invasive breast carcinoma. JMD. 2013;15(2):210–9. doi:10.1016/j.jmoldx.2012.10.003.
Denkert C, Loibl S, Kronenwett R, Budczies J, von Torne C, Nekljudova V, et al. RNA-based determination of ESR1 and HER2 expression and response to neoadjuvant chemotherapy. Ann Oncol. 2013;24(3):632–9. doi:10.1093/annonc/mds339.
Alba J, Gutierrez J, Coupe VM, Fernandez B, Vazquez-Boquete A, Alba J, et al. HER2 status determination using RNA-ISH–a rapid and simple technique showing high correlation with FISH and IHC in 141 cases of breast cancer. Histol Histopathol. 2012;27(8):1021–7.
Olsson H, Jansson A, Holmlund B, Gunnarsson C. Methods for evaluating HER2 status in breast cancer: comparison of IHC, FISH, and real-time PCR analysis of formalin-fixed paraffin-embedded tissue. Pathol Lab Med Intl. 2013;5:31–7. doi:10.2147/PLMI.S44976.
Dabbs DJ, Klein ME, Mohsin SK, Tubbs RR, Shuai Y, Bhargava R. High false-negative rate of HER2 quantitative reverse transcription polymerase chain reaction of the Oncotype DX test: an independent quality assurance study. J Clin Oncol. 2011;29(32):4279–85. doi:10.1200/JCO.2011.34.7963.
Heredia NJ, Belgrader P, Wang S, Koehler R, Regan J, Cosman AM, et al. Droplet digital PCR quantitation of HER2 expression in FFPE breast cancer samples. Methods. 2013;59(1):S20–3. doi:10.1016/j.ymeth.2012.09.012.
Hindson BJ, Ness KD, Masquelier DA, Belgrader P, Heredia NJ, Makarewicz AJ, et al. High-throughput droplet digital PCR system for absolute quantitation of DNA copy number. Anal Chem. 2011;83(22):8604–10. doi:10.1021/ac202028g.
Dodd DW, Gagnon KT, Corey DR. Digital quantitation of potential therapeutic target RNAs. Nucl Acid Therap. 2013;23(3):188–94. doi:10.1089/nat.2013.0427.
Zhu Y, Lu D, Lira ME, Xu Q, Du Y, Xiong J, et al. Droplet digital polymerase chain reaction detection of HER2 amplification in formalin fixed paraffin embedded breast and gastric carcinoma samples. Exp Mol Pathol. 2015;. doi:10.1016/j.yexmp.2015.11.027.
Kinugasa H, Nouso K, Tanaka T, Miyahara K, Morimoto Y, Dohi C, et al. Droplet digital PCR measurement of HER2 in patients with gastric cancer. Br J Cancer. 2015;112(10):1652–5. doi:10.1038/bjc.2015.129.
Belgrader P, Tanner SC, Regan JF, Koehler R, Hindson BJ, Brown AS. Droplet digital PCR measurement of HER2 copy number alteration in formalin-fixed paraffin-embedded breast carcinoma tissue. Clin Chem. 2013;59(6):991–4. doi:10.1373/clinchem.2012.197855.
Lih CJ, Si H, Das B, Harrington RD, Harper KN, Sims DJ, et al. Certified DNA reference materials to compare HER2 gene amplification measurements using next-generation sequencing methods. JMD. 2016;18(5):753–61. doi:10.1016/j.jmoldx.2016.05.008.
Wolff AC, Hammond ME, Hicks DG, Dowsett M, McShane LM, Allison KH, et al. Recommendations for human epidermal growth factor receptor 2 testing in breast cancer: American Society of Clinical Oncology/College of American Pathologists clinical practice guideline update. Arch Pathol Lab Med. 2014;138(2):241–56. doi:10.5858/arpa.2013-0953-SA.
Metsalu T, Vilo J. ClustVis: a web tool for visualizing clustering of multivariate data using Principal Component Analysis and heatmap. Nucl Acids Res. 2015;43(W1):W566–70. doi:10.1093/nar/gkv468.
Ferguson BS, Nam H, Hopkins RG, Morrison RF. Impact of reference gene selection for target gene normalization on experimental outcome using real-time qRT-PCR in adipocytes. PLoS One. 2010;5(12):e15208. doi:10.1371/journal.pone.0015208.
Aurilio G, Disalvatore D, Pruneri G, Bagnardi V, Viale G, Curigliano G, et al. A meta-analysis of oestrogen receptor, progesterone receptor and human epidermal growth factor receptor 2 discordance between primary breast cancer and metastases. Eur J Cancer. 2014;50(2):277–89. doi:10.1016/j.ejca.2013.10.004.
Koren S, Bentires-Alj M. Breast tumor heterogeneity: source of fitness hurdle for therapy. Mol cell. 2015;60(4):537–46. doi:10.1016/j.molcel.2015.10.031.
Bethune GC, Mullen JB, Chang MC. Detecting intratumoral heterogeneity in routine breast-HER2 testing: low yield of testing multiple blocks. Ann Diagn Pathol. 2015;19(6):385–90. doi:10.1016/j.anndiagpath.2015.08.001.
Morrogh M, Olvera N, Bogomolniy F, Borgen PI, King TA. Tissue preparation for laser capture microdissection and RNA extraction from fresh frozen breast tissue. Biotechniques. 2007;43(1):41–2.
Spinney L. Cancer: caught in time. Nature. 2006;442(7104):736–8. doi:10.1038/442736a.
Garcia-Murillas I, Lambros M, Turner NC. Determination of HER2 amplification status on tumour DNA by digital PCR. PLoS One. 2013;8(12):e83409. doi:10.1371/journal.pone.0083409.
Acknowledgements
We would like to thank the Cancer Council of Western Australia, the Ada Bartholomew Trust and Zonta, Club of Peel, Western Australia for their generous funding support. For this type of study formal consent was not required. A waiver of consent was obtained for all participants included in the study.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there are no potential conflicts of interest.
Rights and permissions
About this article
Cite this article
Meehan, K., Clynick, B., Mirzai, B. et al. HER2 mRNA transcript quantitation in breast cancer. Clin Transl Oncol 19, 606–615 (2017). https://doi.org/10.1007/s12094-016-1573-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12094-016-1573-2