Abstract
Purpose
EBER-1 (a non-coding RNA transcribed by EBV) expression was detected in most of Epstein–Barr virus (EBV)-positive nasopharyngeal carcinoma (NPC) patients. However, the relevance between EBER-1 expression and NPC clinical outcome has not been reported. This study aims to assess the possible correlations of EBER-1 expression and clinical parameters and its potential prognostic predictive ability in NPC patient’s outcomes.
Methods
We examined EBER-1 mRNA expression in 301 NPC and 130 non-NPC tissues using in situ hybridization and did statistics.
Results
EBER-1 expression was up-regulated in NPC tissues when compared to non-NPC tissues. A receiver operating characteristic analysis revealed that EBER-1 expression could distinguish non-cancerous patients from NPC patients (p < 0.001, sensitivity: 72.5 %, specificity: 83.5 %, AUC = 0.815). A survival analysis revealed that patients with high levels of EBER-1 expression had a significantly good prognosis (Disease-free survival: p = 0.019, overall survival: p = 0.006).
Conclusion
These results indicated that EBER-1 expression is a potential prognosis factor of NPC and highly negative correlated with the progress of NPC.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In contrast to other head and neck cancer and epithelial malignancy in general, nasopharyngeal carcinoma (NPC) has a remarkably distinctive ethnic and geographic distribution, and this distribution of NPC indicates that the development of this cancer may be related to genetic and environmental factors [1–4].
A unique feature of NPC is its strong association with Epstein–Barr virus (EBV) [4–8]. EBV is a large dsDNA lymphotropic herpesvirus which was first isolated five decades ago and is widespread in human populations of all ages [9, 10], causes infectious mononucleosis and has a strong association with various malignancies such as Burkitt’s lymphoma, Hodgkin’s disease and NPC [11, 12]. A close relationship between NPC and EBV has been widely accepted since Henle et al. reported higher serum IgA titer in NPC patients [13].
EBV genomes are detected in most cases of NPC, regardless of whether they are from an endemic or non-endemic area. Moreover, EBV genomes detected in NPC tumors are monoclonal. Thus, it is generally accepted that EBV is the causative agent of NPC [6]. EBV-encoded RNAs (EBER-1 and EBER-2) are two small non-coding, non-polyadenylated RNAs that are abundantly expressed in latently EBV-infected cells. Some reports suggest that EBER-1 is transcribed by RNA polymerase II and III, whereas EBER-2 is transcribed by polymerase III only [14]. EBER-1 has been reported to be present at ten fold higher levels compared with EBER-2, presumably because of its longer half-life [1]. EBER-1 does not code any protein and its function is still not clear, but it is known to be abundantly produced in infected cells, making itself an appropriate biomarker for detection of EBV. Detection of EBER by in situ hybridization (ISH) has been established as the most sensitive and practical method for detecting EBV infection. Thus, in this work, ISH technology was used to elucidate the potential roles of EBER-1 in NPC. We used the tissue microarray including 301 NPC and 130 non-NPC patients and examined the clinical features and outcomes of NPC, and performed an analysis of the prognostic factors of NPC in terms of clinical issues.
Methods
Patients and tissue specimens
Before study initiation, ethical approval was obtained from Xiangya Hospital in Changsha, and the Central South University Ethics Review committees/Institutional Review Boards. Informed consent was obtained from all patients, 301 cases NPC and 130 non-NPC tissues which were collected between January, 2002 and October, 2004 at Xiangya Hospital. Patients’ clinical information, such as age, sex, tumor size, lymphatic invasion and TNM stage was collected and stored in a database. The survey for NPC patients’ follow-up information was on August 9, 2010. 77 NPC patients had valid follow-up data and the longest survival time was 96 months. The NPC tissue microarray was constructed as described in our previous studies [15, 16].
In situ hybridization (ISH)
The EBER-1 probe was 5′-agacaccgtcctcaccacccgggacttgta-3′. The probes were labeled with DIG-dUTP (Roche) at the 3′ end according to manufacturer protocols. A poly-d(T) probe was used as a control for total RNA preservation. The DIG oligonucleotide 3′-tailing labeling kit was purchased from Roche, as previously described [15]. A semi-quantitative scoring criterion for ISH was used in which both the staining intensity and number of positive areas were recorded. 0 was named as negative, + to ++ was named as low expression and +++ was named as high expression. The scores corresponding to the overall distribution of EBER-1 were averaged across the different tumor plugs in each case. All sections were scored independently by two pathologists who were blinded to the clinicopathological features and the clinical course.
Statistical analysis
Data were analyzed using the Chi-squared test or the Fisher exact test to assess associations between EBER-1 expression levels and the clinicopathologic parameters. The sensitivity and specificity of a test based on a scoring system for a range of cutoff points can be shown by an ROC curve. A measure of performance that combines sensitivity and specificity is the area under the ROC curve (AUC). The overall survival (OS) was defined as the time elapsed between the diagnosis and the date of death. Disease-free survival (DFS) was defined as the time elapsed between the diagnosis and the date of first treatment failure. The OS and DFS estimates over time were calculated using the Kaplan–Meier method, and the differences were compared using the log-rank test [17]. The results of the analysis were considered significant in a log-rank test if p < 0.05. Calculations were performed using the SPSS 13.0 statistical software for Windows (SPSS Inc, Chicago). A p value less than 0.05 was considered statistically significant, and all statistical tests were two sided.
Results
Expression of EBER-1 in NPC and non-cancerous NPE
Representative images of EBER-1 ISH signals are shown in Fig. 1. EBER-1 was expressed in 71.1 % (214 of 301) of NPC samples and in 11.5 % (15 of 130) of non-cancerous nasopharyngeal epithelium (NPE). The difference was significant between NPC and non-cancerous nasopharyngeal epitheliums (p < 0.001, Fig. 2).
Representative images of EBER-1 as detected by ISH. Brown denotes a positive signal. Negative expression of EBER-1 was in nasopharyngeal column epithelial cells (a) and in differentiated non-keratinizing NPC (b), weak positive in differentiated non-keratinizing NPC (c) and strong positive in undifferentiated NPC (d)
EBER-1 could distinguish non-cancerous and NPC patients
Next, we evaluated the usefulness of EBER-1 ISH signal for distinguishing between non-cancerous and NPC patients. We selected some significant data points to develop a test by setting an optimal cutoff value and using it for the calculation of the sensitivity, specificity and area under the curve (AUC). Figure 3 shows ROC curve for the scoring systems for all non-cancerous and NPC patients. The AUC of EBER-1 for predicting NPC patients was 0.815 (95 % confidence interval = 0.776–0.854, p < 0.001), with a sensitivity and specificity of 72.5 and 83.5 %, respectively.
Correlations between expression of EBER-1 and NPC clinical pathological parameters
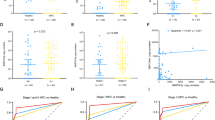
Expression of EBER-1 in NPC was not associated with some parameters such as gender and age, but higher EBER-1 expression was associated with earlier clinical stage (Fig. 4a, p < 0.0001), T stage (Fig. 4b, p = 0.007) and lower lymph node metastasis potential (Fig. 4c, p = 0.04).
EBER-1 expression proportion in different clinical stages, T, and N stages. a Higher EBER-1 expression was associated with earlier clinical stage (I, n = 19; II, n = 99; III, n = 113; IV, n = 70; p < 0.001); b Higher EBER-1 expression was associated with earlier T stage (T1, n = 40; T2, n = 139; T3, n = 63; T4, n = 59; p = 0.007); c Higher EBER-1 expression was associated with lower lymph node metastasis potential (N0, n = 108; N1, n = 102; N2, n = 80; N3, n = 11; p = 0.044)
EBER-1 could predict the clinical outcome of NPC
There were 77 NPC patients who had valid follow-up data. We then examined the correlation between EBER-1 expression and relapse and cancer-related deaths using a Kaplan–Meier survival analysis. The median disease-free survival (DFS) time of NPC patients with negative, low and high expression of EBER-1 was 35, 48 and 71 months, respectively (Fig. 5a, p = 0.019). While the median overall survival (OS) time was 35, 56 and 75 months in NPC patients with negative, low and high expression of EBER-1, respectively (Fig. 5b, p = 0.006). Both DFS and OS were significantly longer in patients with high EBER-1 expression compared to those with low or negative expression. These results strongly suggested that EBER-1 expression may involve in the development of NPC.
Discussion
EBV establishes a latent infection in B cells and epithelial cells in over 90 % of the human adult population as a lifelong and largely asymptomatic infection. But EBV infection has been linked to several types of tumors such as Hodgkin’s lymphoma, Burkitt’s lymphoma, melanoma, and gastric adenocarcinoma, as well as and NPC, which have long been known to be EBV-associated neoplasm [11]. Our previous studies also had reported that NPC is closely associated with EBV infection [18–20].
Reports have shown that EBV produces several viral oncoproteins, including six EBV-encoded nuclear antigen proteins (EBNA-1, EBNA-2, EBNA-3A, 3B, 3C and EBNA-LP) and three latent membrane proteins (LMP1, LMP2A and LMP2B), which were expressed in different latency programs [21]. Interestingly, two types of non-protein-coding RNA, EBV-encoded RNAs (EBER-1 and EBER-2) and long alternative splicing non-coding RNA at BamH1 A rightward transcripts (BARTs) [22] are expressed in all forms of latency programs [23]. Among these transcripts, EBNA1, EBNA2, EBNA3A, EBNA3C, EBNA-LP, and LMP1 are essential for growth transformation, whereas EBNA3B, LMP2A, LMP2B, BamHI A rightward transcripts, and EBERs are not essential [21]. Both EBER-1 and EBER-2 are expressed at high levels in latently infected cells and structurally highly conserved with each other; however, they have distinct biological functions in EBV-infected cells [24]. Our results suggested that EBER-1, especially the high expression of EBER-1, had a strong impact on the good prognosis of NPC. EBER-2 alone may modulate the worst prognosis changes of NPC since it is structurally strikingly similar to EBER-1. It was reported that EBER-2 alone could modulate the malignancy phenotype in lymphocyte B cells [24]. So EBER-2 alone may increase the development of NPC malignancy even if the NPC cells lose the expression of EBER-1.
By now, radiotherapy is still the most important treatment for NPC. EBV-related NPC may have the characteristic of high sensitivity to radiotherapy, then favorable prognosis [25]. In fact, other than encoding oncogene and promoting nasopharyngeal epithelium malignant transformation, some genes encoded by EBV could inhibit the phenotype of NPC cells such as Rta and Zta, two transcription factors expressed by EBV during the immediate-early stage of the lytic cycle, could inhibit cell proliferation and result in cell cycle arrest through induction of cyclin-dependent kinase inhibitors [26–29]. Multiple lines of evidence have suggested that miR-BARTs target cellular genes mainly for preventing apoptosis and escaping the host immune system. Surprisingly, not all miR-BARTs promote cell growth or inhibit apoptosis. Recently, some studies reported that miR-BARTs exert their suppressive function and “anti-cancer” activities [30]. In this sense, not only all the genes encoded by EBV have oncogenes’ function but also some have tumor suppressor genes’ function.
EBER-1 is expressed abundantly in all forms of cells latently infected with EBV (107 copies per cell), making itself an appropriate target for detection of EBV [1, 31]. We used ISH to detect EBER-1 expression and found that most of NPC patients expressed EBER-1. The difference was very significant between NPC and non-NPC patients. From the ROC curve, we could discriminate NPC and non-NPC patients through analyzing the expression of EBER-1. Simultaneously, we found that in different clinical stages, T stages and N stages, the expression of EBER-1 was the highest in clinical stage II, T2 stage and N1 stage. However, the expression of EBER-1 was the lowest in clinical stage IV, T4 stage and N3 stage. These findings maybe suggested the “hit-and-run” role for EBV in NPC and in some rare subclone of proliferating cells that has spontaneously lost EBV at some point after tumor initiation but is now capable of sustaining virus-independent growth and may have a survival advantage through avoidance of immune surveillance [32, 33]. All these results suggested that EBER-1 plays an important role in NPC carcinogenesis. However, the details of the mechanisms that EBER-1 involved in NPC development and progression are poorly understood. A growing body of evidence suggested that this small, highly expressed, non-coding RNA plays significant roles in EBV-mediated oncogenesis [31, 34]. Several reports indicated that EBERs can confer an apoptotic-resistant phenotype in immortalized epithelial cells and can support cell growth by stimulating insulin-like growth factor (IGF-1) secretion in EBV-positive gastric carcinoma and NPC cell lines [31]. Long non-coding RNAs (lncRNAs) are molecules, often longer than 200nt in length, with limited protein-coding capacity, a coding potential often less than 100 amino acids [35–37]. Accumulating evidence indicates that lncRNAs participate in many important physiological processes by modulating gene expression at the epigenetic, transcriptional, post-transcriptional levels and some participate in carcinogenesis and cancer progression [37]. In fact, EBER-1 has indeed some lncRNA characteristics. It has no exon and ORF, 905 bp in length and does not encode any amino acids. We previously reported that EBER-1 was strongly negatively correlated with another human lncRNA, LINC00312 [38], in NPC, which suggests that EBER-1 could possibly regulate the expression of LINC00312 through some genes and signaling pathways, but the exact mechanism by which this occurs requires further study. So in our future studies, we will focus on the mechanism of EBER-1 in NPC carcinogenesis.
In summary, our data reveal that EBER-1 maybe involved in the progression of human nasopharyngeal epithelial carcinogenesis and could discriminate NPC and non-NPC patients. Moreover, positive expression especially high expression of EBER-1 in NPC is closely correlated with good prognosis of NPC patients.
References
Ahmed W, Khan G. The labyrinth of interactions of Epstein–Barr virus-encoded small RNAs. Rev Med Virol. 2013;24(1):3–14. doi:10.1002/rmv.1763.
Zeng Z, Zhou Y, Zhang W, Li X, Xiong W, Liu H, et al. Family-based association analysis validates chromosome 3p21 as a putative nasopharyngeal carcinoma susceptibility locus. Genet Med. 2006;8(3):156–60. doi:10.1097/01.gim.0000196821.87655.d0.
Xiong W, Zeng ZY, Xia JH, Xia K, Shen SR, Li XL, et al. A susceptibility locus at chromosome 3p21 linked to familial nasopharyngeal carcinoma. Cancer Res. 2004;64(6):1972–4.
Zeng Z, Huang H, Zhang W, Xiang B, Zhou M, Zhou Y, et al. Nasopharyngeal carcinoma: advances in genomics and molecular genetics. Sci China Life Sci. 2011;54(10):966–75. doi:10.1007/s11427-011-4223-5.
Takada K. Role of EBER and BARF1 in nasopharyngeal carcinoma (NPC) tumorigenesis. Semin Cancer Biol. 2012;22(2):162–5. doi:10.1016/j.semcancer.2011.12.007.
Raab-Traub N. Epstein–Barr virus in the pathogenesis of NPC. Semin Cancer Biol. 2002;12(6):431–41.
Liao Q, Zeng Z, Guo X, Li X, Wei F, Zhang W, et al. LPLUNC1 suppresses IL-6-induced nasopharyngeal carcinoma cell proliferation via inhibiting the Stat3 activation. Oncogene. 2014;33(16):2098–109. doi:10.1038/onc.2013.161.
Yang Y, Liao Q, Wei F, Li X, Zhang W, Fan S, et al. LPLUNC1 inhibits nasopharyngeal carcinoma cell growth via down-regulation of the MAP kinase and cyclin D1/E2F pathways. PLoS One. 2013;8(5):e62869. doi: 10.1371/journal.pone.0062869
Epstein MA, Achong BG, Barr YM. Virus particles in cultured lymphoblasts from Burkitt’s Lymphoma. Lancet. 1964;1(7335):702–3.
Yu Z, Song Y, Gong Z, Zeng Z, Lu J, Li X, et al. The mechanism and tumorigenesis of oncogenic DNA virus integration. Prog Biochem Biophys. 2014;41(4):324–31. doi:10.3724/SP.J.1206.2012.00554.
Young LS, Rickinson AB. Epstein–Barr virus: 40 years on. Nat Rev Cancer. 2004;4(10):757–68. doi:10.1038/nrc1452.
Zhou Y, Zeng Z, Zhang W, Xiong W, Wu M, Tan Y, et al. Lactotransferrin: a candidate tumor suppressor-deficient expression in human nasopharyngeal carcinoma and inhibition of NPC cell proliferation by modulating the mitogen-activated protein kinase pathway. Int J cancer. 2008;123(9):2065–72. doi:10.1002/ijc.23727.
Henle G, Henle W. Epstein–Barr virus-specific IgA serum antibodies as an outstanding feature of nasopharyngeal carcinoma. Int J cancer. 1976;17(1):1–7.
Wang Y, Zhang X, Chao Y, Jia Y, Xing X, Luo B. New variations of Epstein–Barr virus-encoded small RNA genes in nasopharyngeal carcinomas, gastric carcinomas, and healthy donors in northern China. J Med Virol. 2010;82(5):829–36. doi:10.1002/jmv.21714.
Fan SQ, Ma J, Zhou J, Xiong W, Xiao BY, Zhang WL, et al. Differential expression of Epstein–Barr virus-encoded RNA and several tumor-related genes in various types of nasopharyngeal epithelial lesions and nasopharyngeal carcinoma using tissue microarray analysis. Hum Pathol. 2006;37(5):593–605. doi:10.1016/j.humpath.2006.01.010
Zhang W, Zeng Z, Zhou Y, Xiong W, Fan S, Xiao L, et al. Identification of aberrant cell cycle regulation in Epstein–Barr virus-associated nasopharyngeal carcinoma by cDNA microarray and gene set enrichment analysis. Acta Biochim et Biophys Sin. 2009;41(5):414–28.
Xiong W, Wu X, Starnes S, Johnson SK, Haessler J, Wang S, et al. An analysis of the clinical and biologic significance of TP53 loss and the identification of potential novel transcriptional targets of TP53 in multiple myeloma. Blood. 2008;112(10):4235–46. doi:10.1182/blood-2007-10-119123.
Zeng ZY, Zhou YH, Zhang WL, Xiong W, Fan SQ, Li XL, et al. Gene expression profiling of nasopharyngeal carcinoma reveals the abnormally regulated Wnt signaling pathway. Hum Pathol. 2007;38(1):120–33. doi:10.1016/j.humpath.2006.06.023.
Zeng Z, Zhou Y, Xiong W, Luo X, Zhang W, Li X, et al. Analysis of gene expression identifies candidate molecular markers in nasopharyngeal carcinoma using microdissection and cDNA microarray. J Cancer Res Clin Oncol. 2007;133(2):71–81. doi:10.1007/s00432-006-0136-2.
Zhou Y, Zeng Z, Zhang W, Xiong W, Li X, Zhang B, et al. Identification of candidate molecular markers of nasopharyngeal carcinoma by microarray analysis of subtracted cDNA libraries constructed by suppression subtractive hybridization. Eur J Cancer Prev. 2008;17(6):561–71. doi:10.1097/CEJ.0b013e328305a0e8.
Brooks L, Yao QY, Rickinson AB, Young LS. Epstein–Barr virus latent gene transcription in nasopharyngeal carcinoma cells: coexpression of EBNA1, LMP1, and LMP2 transcripts. J Virol. 1992;66(5):2689–97.
Zeng Z, Huang H, Huang L, Sun M, Yan Q, Song Y, et al. Regulation network and expression profiles of Epstein–Barr virus-encoded microRNAs and their potential target host genes in nasopharyngeal carcinomas. Sci China Life Sci. 2014;57(3):315–26. doi:10.1007/s11427-013-4577-y.
Cai X, Schafer A, Lu S, Bilello JP, Desrosiers RC, Edwards R, et al. Epstein–Barr virus microRNAs are evolutionarily conserved and differentially expressed. PLoS Pathog. 2006;2(3):e23. doi:10.1371/journal.ppat.0020023.
Wu Y, Maruo S, Yajima M, Kanda T, Takada K. Epstein–Barr virus (EBV)-encoded RNA 2 (EBER2) but not EBER1 plays a critical role in EBV-induced B-cell growth transformation. J Virol. 2007;81(20):11236–45. doi:10.1128/JVI.00579-07.
Shi W, Pataki I, MacMillan C, Pintilie M, Payne D, O’Sullivan B, et al. Molecular pathology parameters in human nasopharyngeal carcinoma. Cancer. 2002;94(7):1997–2006.
Cayrol C, Flemington E. G0/G1 growth arrest mediated by a region encompassing the basic leucine zipper (bZIP) domain of the Epstein–Barr virus transactivator Zta. J Biol Chem. 1996;271(50):31799–802.
Cayrol C, Flemington EK. The Epstein–Barr virus bZIP transcription factor Zta causes G0/G1 cell cycle arrest through induction of cyclin-dependent kinase inhibitors. EMBO J. 1996;15(11):2748–59.
Guo Q, Sun X, Yuan C, Zhou H, Li Y, Jie G, et al. Effect of Rta protein of Epstein–Barr virus on the cell cycle in HeLa cells. Acta Virol. 2011;55(4):311–6.
Huang SY, Hsieh MJ, Chen CY, Chen YJ, Chen JY, Chen MR, et al. Epstein–Barr virus Rta-mediated transactivation of p21 and 14-3-3sigma arrests cells at the G1/S transition by reducing cyclin E/CDK2 activity. J gen Virol. 2012;93(Pt 1):139–49. doi:10.1099/vir.0.034405-0.
Choi H, Lee H, Kim SR, Gho YS, Lee SK. Epstein–Barr virus-encoded microRNA BART15-3p promotes cell apoptosis partially by targeting BRUCE. J Virol. 2013;87(14):8135–44. doi:10.1128/JVI.03159-12.
Samanta M, Iwakiri D, Kanda T, Imaizumi T, Takada K. EB virus-encoded RNAs are recognized by RIG-I and activate signaling to induce type I IFN. EMBO J. 2006;25(18):4207–14. doi:10.1038/sj.emboj.7601314.
Ambinder RF. Gammaherpesviruses and “Hit-and-Run” oncogenesis. Am J Pathol. 2000;156(1):1–3. doi:10.1016/S0002-9440(10)64697-4.
Niller HH, Wolf H, Minarovits J. Viral hit and run-oncogenesis: genetic and epigenetic scenarios. Cancer Lett. 2011;305(2):200–17. doi:10.1016/j.canlet.2010.08.007.
Samanta M, Takada K. Modulation of innate immunity system by Epstein–Barr virus-encoded non-coding RNA and oncogenesis. Cancer Sci. 2010;101(1):29–35. doi:10.1111/j.1349-7006.2009.01377.x.
Gong Z, Zhang S, Zhang W, Huang H, Li Q, Deng H, et al. Long non-coding RNAs in cancer. Sci China Life sci. 2012;55(12):1120–4. doi:10.1007/s11427-012-4413-9.
Tang K, Wei F, Bo H, Huang HB, Zhang WL, Gong ZJ, et al. Cloning and functional characterization of a novel long non-coding RNA gene associated with hepatocellular carcinoma. Prog Biochem Biophys. 2014;41(2):153–62. doi:10.3724/Sp.J.1206.2012.00613.
Gong Z, Zhang S, Zeng Z, Wu H, Yang Q, Xiong F, et al. LOC401317, a p53-regulated long non-coding RNA, inhibits cell proliferation and induces apoptosis in the nasopharyngeal carcinoma cell line HNE2. PLoS One. 2014;9(11):e110674. doi:10.1371/journal.pone.0110674.
Zhang W, Huang C, Gong Z, Zhao Y, Tang K, Li X, et al. Expression of LINC00312, a long intergenic non-coding RNA, is negatively correlated with tumor size but positively correlated with lymph node metastasis in nasopharyngeal carcinoma. J Mol Histol. 2013;44(5):545–54. doi:10.1007/s10735-013-9503-x.
Acknowledgments
This work was supported by the National Natural Science Foundation of China (Grant Nos. 81172189, 81272298, 81372907, 81472531 and 91229122), the Hunan Province Natural Science Foundation of China (Grant Nos. 14JJ1010, 12JJ2044), the Hunan Province Science and Technology Foundation of China (Grant No. 2012FJ6073).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interests.
Informed consent
Before study initiation, ethical approval was obtained from Xiangya Hospital in Changsha, and the Central South University Ethics Review Committees/Institutional Review Boards. NPC samples, non-tumor nasopharyngeal epithelial tissues were collected at Xiangya Hospital. All of the individuals participating in this project gave informed consent, and the study was approved by the Xiangya Medical Ethics Board.
Rights and permissions
About this article
Cite this article
Zeng, Z., Fan, S., Zhang, X. et al. Epstein–Barr virus-encoded small RNA 1 (EBER-1) could predict good prognosis in nasopharyngeal carcinoma. Clin Transl Oncol 18, 206–211 (2016). https://doi.org/10.1007/s12094-015-1354-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12094-015-1354-3