Abstract
Information capacity of nucleotide sequences measures the unexpectedness of a continuation of a given string of nucleotides, thus having a sound relation to a variety of biological issues. A continuation is defined in a way maximizing the entropy of the ensemble of such continuations. The capacity is defined as a mutual entropy of real frequency dictionary of a sequence with respect to the one bearing the most expected continuations; it does not depend on the length of strings contained in a dictionary. Various genomes exhibit a multi-minima pattern of the dependence of information capacity on the string length, thus reflecting an order within a sequence. The strings with significant deviation of an expected frequency from the real one are the words of increased information value. Such words exhibit a non-random distribution alongside a sequence, thus making it possible to retrieve the correlation between a structure, and a function encoded within a sequence.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
A study of statistical properties of nucleotide sequences challenges researchers. Careful investigation of the nucleotide sequence may still bring new facts, new knowledge, and inspire a researcher with new methodology. In general, there are two approaches in such studies: the former implies the most sophisticated and advanced knowledge derived from biological issues of a sequence under consideration, the latter is mathematically oriented and aims to retrieve as much knowledge, as possible, from a sequence itself, avoiding an implementation of any additional, external issues. Here, we present some new results obtained within this latter paradigm.
Information capacity of a symbol sequence (a nucleotide sequence, in particular) is the key issue of the paper. The former, in turn, is based on the idea of a “reconstructed” frequency dictionary; such a dictionary is an ensemble of all the continuations of shorter strings meeting the linear constraints, and extreme principle (see “Reconstructed dictionary: maximum entropy”). Information capacity is deeply related to a complexity of sequence. Both are determined through the calculation of mutual entropy of a real frequency against the “reconstructed” one (see “Information capacity of a dictionary”). The methodology developed here also yields some intriguing results concerning the redundancy of genetic entities (see “Redundancy of genes is affected by splicing”), symmetry in frequency dictionaries pattern (see “Informational symmetry in genomes”) and other biologically inspired issues. The methodology provides a new approach towards the codon usage bias problem (see “Codon usage bias”).
Frequency dictionary
Consider a nucleotide sequence, of the length N. Furthermore, we shall suppose that no other symbol, but A, C, G or T is found within a sequence.Footnote 1 Any string \({\nu_1\nu_2\ldots}\nu_q\) of q nucleotides occurred within a sequence makes a word ω. All the words (of the length q) found within the sequence make its q-support. Providing each word from the support with the number n ω of its copies, one gets the finite dictionary W(q) of the sequence. Some biological issues concerning the finite dictionaries can be seen in Redundancy of genes is affected by splicing. Changing the number n ω for frequency
where N is the length of sequence, one gets the frequency dictionary W(q) of thickness q. This definition is valid, when sequence is connected into a ring; for the motivation standing behind such a connection, see “Reconstructed dictionary: maximum entropy”.
A frequency dictionary W(q) is the key entity for further studies of nucleotide sequences. Obviously, such a dictionary provides a student with the detailed knowledge of the frequencies of shorter words (i.e., thinner frequency dictionaries W(l), 1 ≤ l < q). Indeed, to get a thinner dictionary W(l), one must sum up the frequency of words differing in starting q − l, or ending q − l symbols.Footnote 2 Thus, a downward transformation of dictionary W(q) into the dictionary W(l) with 1 ≤ l < q is simple and unambiguous. The situation is getting worse, for upward transformations.
Reconstructed dictionary: maximum entropy
An upward transformation \(W(q) \mapsto W(q^{\prime})\) of a frequency dictionary W(q) into a thicker one W(q′) (as q′ > q) is ambiguous, in general; the case of unambiguity of an upward transformation is studied in section “Redundancy of genes is affected by splicing”. To begin with, let us consider an upward transformation of a given frequency dictionary W(q) into one symbol thicker entity W(q + 1). Since a word ω may have several (four, to be exact, for genetic entities) continuations into the words
then one gets a family of frequency dictionaries {W(q + 1)}, instead of the single one.
An abundance of the family {W(q + 1)} is constrained with the linear constraints
that make each specific (thicker) dictionary W′(q + 1) ∈ {W(q + 1)} generate the given frequency dictionary W(q). Meanwhile, the linear constraints (Eq. 1) fail to figure out the unique frequency dictionary W(q + 1) of the thickness q + 1. Yet, an ambiguity of a thicker dictionary exists. It should be stressed that the ensemble {W(q + 1)} must incorporate the real frequency dictionary W(q + 1) of thickness q + 1.
The key idea of the choice of a specific frequency dictionary \(\widetilde{W}(q+1)\) is to figure out the dictionary bearing the most probable continuation. Such a dictionary must meet the extreme principle:
with obvious linear constraints (Eq. 1). Here \(\tilde{f}_{\nu_1\nu_2\ldots \nu_q\nu_{q+1}} \in \widetilde{W}(q+1),\) and \(f_{\nu_{1}\nu_{2}\ldots\nu_{q}}\) belong to the real frequency dictionary W(q). A small remark should be made here. We develop a frequency dictionary W(q) from a real sequence; thus, one may consider the dictionary as a mapping of a sequence into the dictionary. On the other hand, a frequency dictionary is a set of pairs \(\left(\omega, {\frac{n_{\omega}} {N}}\right).\) Hence, any set of the couples of the shown type meeting the normalization rule
might be considered as a frequency dictionary. Meanwhile, such a dictionary might have no any real sequence behind it. The reconstructed frequency dictionary (Eq. 2) may not correspond to a real sequence.
LaGrange multiplier method (Bugaenko et al. 1996, 1998; Rui and Bin 2001; Sadovsky 2005) yields quite simple and clear solution of the problem (Eqs. 1, 2):
Equation 3 looks like a transitional probability of Markovian process; yet, this formula describes the frequency of the most expected continuation of two words (combining the given one). It has nothing to do with the properties of an original sequence. The coincidence is not eventual, but the formula is derived with neither hypothesis towards the structure of an original sequence. This coincidence just reflects the fact that the relevant Markov chain (that could be formally generated from any frequency dictionary) implements the hypothesis of the most expected continuation of a word. Of course, the order r of such a process is r = q − 1, where q is the frequency thickness.
Equation 3 could easily be extended for the case of the reconstruction of \(\widetilde{W}(q+s)\) from W(q) with s > 1 (see Bugaenko et al. 1996, 1998; Rui and Bin 2001 for details); it should be said that the extension then differs from Markov chain relations.
Information capacity of a dictionary
For a nucleotide sequence, one can always generate a series of real frequency dictionaries of increasing thickness q:
For each dictionary W(j), 1 < j ≤ q, one can develop a series of similar dictionaries
derived due to Eqs. 2, 3 from the dictionary that is one symbol thinner:
Thus, one gets a series of reconstructed dictionaries of the same thickness j, 2 ≤ j ≤ q. Comparison of real frequency dictionaries versus reconstructed ones yields the information capacity of a sequence.Footnote 3
Obviously, the issue of information capacity significantly depends on the method of comparison. An approach based on so-called “reconstruction quality” is presented in Bugaenko et al. (1996, 1998). Here, we pursue another approach based on the calculation of mutual (or relative) entropy (Gorban et al. 2005). This entity is also known as Kullback, or Kullback–Leibler divergence; it should be stressed that this entity has been implemented by the outstanding American physicist, J. W. Gibbs.
Mutual entropy\(\overline{S}\) of two frequency dictionaries W(1) and W(2) (of the same thickness q) is defined as follows:
Here ω enlists the words in two dictionaries. This definition holds true, if \(f^{(2)}_{\omega} > 0\) for any ω ∈ W(1). There might be various ways to define W(2) in Eq. 4; we shall define information capacity as the divergence between real frequency dictionary W q , and the reconstructed one \(\widetilde{W}_q\) derived from the thinner one (see Reconstructed dictionary: maximum entropy). Combining Eqs. 3 and 4, one gets
Four terms in the fraction in Eq. 5 yield four terms in the sum expansion for \(\overline{S}_q.\) Finally, the summation over “extra” indices gives
Formally speaking, a derivation of \(\widetilde{W}_q\) could be done starting from any frequency dictionary W l , 2 ≤ l < q. Further, we shall keep on the case of the derivation of \(\widetilde{W}_q\) from Wq−1. This choice is obvious: still one retrieves the complete biological knowledge from the comparison of real and derived frequency dictionaries. A study of the retrieval of the biological issues from thinner dictionaries would bring nothing, but the extra noise conspiring the valuable information.
Nonetheless, Eqs. (4–6) could easily be extended for the case of derivation of a dictionary \(\widetilde{W}(q)\) from W(l), where 2 ≤ l < q − 1. Equation 6 looks like
for this case; see details in Sadovsky (2002a, b, 2003, 2006). The Eqs. (2–6) are applicable to a symbol sequence of an arbitrary nature, if it is developed from a finite alphabet ℵ. Also, it should be stressed that, maximal possible value of \(\overline{S}_q\) defined according to Eq. 5 does not depend on q, and is equal to 2 × ln2, for a sequence from a four-letter alphabet.
Information capacity of genomes
Genomes are the symbol sequences from the four-letter alphabet \(\aleph =\{\texttt{ A, C, G, T} \}.\) The methodology based on the study of a distribution of rather short strings at the ensemble of the latter brings numerous biologically valuable data. Here we present two of them: a fractal pattern of genomes, and a pattern of distribution of information valuable words alongside a genome. We calculated the information capacity (Eq. 6) for genomes of various organisms.
The dependence of \(\overline{S}_q\) on the dictionary thickness q is always bell-shaped. It obviously results from a finiteness of a genome. Surely, the position of the maximum of Eq. 6 depends both on a structure of a sequence, and its length N. Meanwhile, the impact of the length is very strong, and there is no sense to study specifically the position and value of the maximum of Eq. 6. The information capacity defined above is scale-free; it means that the maximal possible value of \(\overline{S}_q\) does not depend on q (Sadovsky 2006). More interesting is the behaviour of \(\overline{S}_q\) for considerably small q, say, 2 ≤ q ≤ 8: it is a non-monotonic one.
Figure 1 shows a typical pattern of information capacity [that is the mutual entropy (Eq. 6)] calculated for four chromosomes of Caenorhabditis elegans. The sequences are deposited at EMBL-bank, and are labelled by their bank identifier. The curve of Eq. 6 is bell-shaped; this shape results from a finite sampling effect of the sequences. As a sequence grows up infinitely, the position of maximum shifts (infinitely) upright, and the value of the latter goes down. For C. elegans genome, the pattern of the information capacity \(\overline{S}(q)\) (see Eq. 6) for 2 ≤ q ≤ 9 is quite smooth, and exhibits a single minimum (at q = 5). Figure 2 shows the detailed patterns of the dependence of \(\overline{S}\) on q (2 ≤ q ≤ 8) for the genome of Eremothecium gossypii. Here, the non-monotonous pattern of the dependence of \(\overline{S}\) on q is evident. Probably, it results from an extended occurrence of non-coding areas observed within such genomes. As a rule, prokaryotic genomes exhibit a significant variety of the patterns, in comparison to eukaryotic ones.
The patterns of \(\overline{S}_q\) observed for various genomes are very diverse. A multi-minima pattern could be interpreted as a fractal structure observed within a genome; besides, some genomes exhibit an inverse behaviour of \(\overline{S}_q,\) for 2 ≤ q ≤ 8. The most common pattern is that the first minimum of Eq. 6 manifests at q = 3. Meanwhile, some genomes exhibit maximum at q = 3. No clear relation between such inversion in \(\overline{S}_q\) pattern, and taxonomy of a genome bearer was found (see Sadovsky 2005 for details).
Statistical semantics of genomes
Consider the definition of information capacity (Eq. 6) in more detail. Obviously, the terms with f ω = 0 and \(f_{\omega} = \tilde{f}_{\omega}\) do not contribute to the sum. The greatest contribution to Eq. 6 is provided by the terms with the most significant deviation of f ω from \(\tilde{f}_{\omega};\) such words are the information valuable words (IVW).
More exactly, let αl, αl < 1 (αr, respectively; αr > 1) be the left information value threshold (the right one, respectively). The words, that fail to match the double inequality
make the ensemble of IVWs. Sometimes, α l might be equal to α −1 r . Whether a word is of information value, or not, heavily depends on the levels of αl and αr. Obviously, αl and αr both depend on q, in general. It was found that, various genomes exhibit quite diverse patterns of the distribution of the words of their information value \(p_{\omega} = f_{\omega}/\tilde{f}_{\omega},\) and the pattern is different for different q. Further, we shall distinguish the words with p > 1 and p < 1.
The words of excessive information value are the points of increased unpredictability, within a genome. Obviously, it is scale-related depending on q, in general. Suppose, the lists of IVW of increasing length q (say, 3 ≤ q ≤ 8) are determined, for the same genome. Consider, then, a chain of embedments of shorter IVW into longer ones:
where each word ω j in Eq. 8 is of information value. Here any word of the length q − 1 is a subset of a longer word, since any shorter word is a subword of a longer one.
If all the words in Eq. 8 exhibit p > 1, then the chain is an upstream one; the chain is downstream, if p < 1, for all words ω j in it. A union of all the chains (Eq. 8) rooted from the same shortest word makes a pyramid. The pyramid is an ascending one, if all the chains in it are also ascending, and vice versa. One can develop a pyramid, that is neither ascending, nor descending; we shall not consider such entities in this paper.
Figure 3 shows the downstream pyramid developed for the Bacillus subtilis genome. Definitely, the pyramid is not unique; all the words in the latter exhibit an excess of a real frequency over the expected one; the information value thresholds were as follows: αl = αl = 1.2, for all q, 3 ≤ q ≤ 8. The words marked with asterisk in the figure are the truncations resulting from an absence of an embedment into a longer information valuable word. The pyramid development was truncated at q = 8.
Statistical semantics, then, is a relation between the distribution of IVW (defined earlier), and functional properties of a sequence under consideration. One may expect two options here: the first is a matching of such a distribution to some known structure, the second is the identification of some basically new pattern in sequences. Space limitations of the paper do not allow to study the relation in detail; here we just outline a general methodology to reveal such a relation.
To begin with, there is no simple relation between shorter IVWs and longer ones: given that IVW may be embedded into a longer IVW, or it may be not. Moreover, if an embedment takes place, then there is no guarantee, that the shorter IVW is embedded into a longer one exhibiting the same sign of p. Consider, then, the union of all the chains (Eq. 8) starting from the same ω3; the union composes a graph with cycles (of length 2). The graph is upward (downward, respectively), if all chains (Eq. 8) in it are upstream ones (downstream, respectively). The shortest word is root, and the longest one is apex. Evidently, there exist pyramids that are neither upward, nor downward, that is, those with alteration of \(p_{\omega_j}\) value, as j grows up: 3 ≤ j ≤ q *.
Consider a set Ω * q of IVW (of length q) identified in some way. Consider, then, the points of the inclusion of all the words ω ∈ Ω * q within genome, with respect to their copy abundance; let K(ω*) be the set of nucleotide numbers where the copies of the word ω* occur within a sequence. Then, \(\overline{K}_q\)
is the union of all the nucleotide numbers where the copies of IVWs take place, at sequence. So, statistical semantics of genome is the hypothesis towards the pattern of \(\overline{K}_q\) distribution.
Figure 4 shows the distribution of the inclusions of information valuable words that are apices of a family of pyramids developed for Drosophila melanogaster chromosome 2R, both for downstream and upstream pyramids. Obviously, the set Ω * q could be identified in various ways. Both the cardinality of Ω * q and abundance of the copies of the words from it, may be very high. One can pursue several approaches to specify Ω * q ; in particular, to consider the apices of a family of pyramids, only. Preliminary observations show that \(\overline{K}_q\) distribution exhibits the pattern heavily differing from a homogeneous one, or from a random distribution (say, Poisson distribution).
Another problem here is a kind of interference. Figure 4 shows the distribution of the distances between two successive apices, with no respect to the origin of the latter. Thus, an expected patterns of the targeted occurrence of the apices might be conspired with the superposition of a number of such patterns differing in various parameters, etc. Here one faces a kind of the law of large numbers: a combination of various patterns differing in particular parameter values, etc., may look like a random (or quasi-random) distribution.
Some more applications
Information capacity measured through the calculation of mutual entropy reveals numerous biologically valuable effects in nucleotide sequences (see Sadovsky 2002a, b, 2003, 2006). Meanwhile, a study of the structure of frequency dictionary of a genetic entity reveals more peculiarities and biological issues standing behind; here, we investigate a problem of codon usage bias determination (see Codon usage bias), that is a classical problem of molecular biology and genetics. Furthermore, a study of a finite dictionary brings simple, but quite clear and distinct results in definition (and in turn measurement) of a redundancy of a symbol sequence (see Redundancy of genes is affected by splicing). Doubtlessly, there is a lot more intriguing details concerning the properties of symbol sequences standing behind the dictionaries.
Redundancy of genes is affected by splicing
Consider a nucleotide sequence of the length N; let W(q) be the finite dictionary of it (see Frequency dictionary for definition). Obviously, rather short words in the sequence are quite abundant, while it goes down, as q grows up. Simultaneously, a diversity in number of copies of specific words goes down. Evidently, the word of the length q = N is unique, within the sequence. On the contrary, words are rather numerous, in copies, when q is considerably small.
There exists the specific thickness d * provided that all the words of this length are single, for any specific sequence and N. Obviously, d * is one symbol longer, that the longest repeat observed in a nucleotide sequence. Consider a finite dictionary W(d * + 1); two arbitrary words ω1 and \(\omega_2, {\omega_1}{\in}{\bf W}(d^{\ast}), {\omega_2}\in {\bf W}(d^{\ast}+1)\) either intersect over a word \(\omega^{\prime} (\omega^{\prime} {\in}{\bf W}(d^{\ast}+1))\) of the length d *, or they do not. Further, any intersecting couple is unique (see Sadovsky 2002c, 2003, 2006; Gorban et al. 1994; Popova and Sadovsky 1995 for details). Thus, W(l), l > d * could be unambiguously derived from the (finite) dictionary W(d * + 1). So, nothing is lost, when a finite dictionary is truncated at the thickness d * + 1.
The unambiguity of an extension of finite dictionary W(d * + 1) into W(l), (l > d *) provides a researcher with the redundancy measure. The truncation thickness d * is that latter. Sure, d * depends both on N, and structure of the sequence. To avoid the dependence on N, one must normalize d *.
Random non-correlated sequence (from the same alphabet ℵ) of the same length N is a perfect reference sample to normalize a sequence under consideration. This idea brings a normalization procedure for genetic entities. To compare two (or more) different genetic entities, one should calculate the ratio
for them. The point is that d * value observed for random non-correlated sequences from four-letter alphabet is tightly proportional to lnN. Actually, d * value is proportional to lnN for a sequence from any (finite) alphabet, while the proportionality factor λ (r ∼λln N) depends on the alphabet cardinality. λ = 1 for four-letter alphabet with equal frequency of each letter occurrence (Zubkov and Mikhailov 1974). Table 1 shows the pattern of r observed for some human genes, before and after the intron excision. This pattern (decrease of r due to splicing) is absolutely common for eukaryotic genes; the pattern is less evident for viral genes (with neither respect to the host genome of a virus), or for prokaryotic genes. Ratio (10) is the redundancy measure, indeed. It generalizes the common idea of a redundancy based on a two-particle entropy calculation (Shannon and Weaver 1949; Durand et al. 2004; Zvonkin and Levin 1970).
Codon usage bias
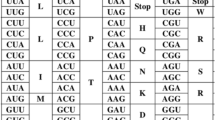
It is a common fact, that genetic code is degenerated. All amino acids (except two ones) are encoded with two or more codons called synonymous. They usually differ in the third nucleotide position. The synonymous codons manifest with different frequency, and the difference is observed both between various genomes and genes of the same genome (Nakamura 2000; Sharp et al. 1993). Codon usage bias is studied quite intensively (Carbone et al. 2003). Basically, the measure of codon usage bias should be as much independent of particular biological issues, as possible. Any biological assumptions (such as mutational bias or translational selection) should be avoided; the definition is likely to be as purely mathematical as possible. The idea of entropy suits here best of all.
Here we propose three new indices of codon usage bias. All the indices are based on mutual entropy \(\overline{S}\) calculation. They differ in the codon frequency distribution supposed to be “quasi-equilibrium” one. An index of codon usage bias is defined in a simple and concise way:
Here \(p_{\nu_1\nu_2\nu_3}\) is the frequency of a codon \({\nu_1\nu_2\nu_3},\) and \(\widetilde{p}_{\nu_1\nu_2\nu_3}\) is the frequency of reference codon distribution. This latter would be defined in three different ways, so that three independent (while related) indices (11) would be implemented, to measure the codon usage bias.
There are three clear and reasonable ways to define \(\widetilde{p}_{\nu_1\nu_2\nu_3}:\)
-
to equalize \(\widetilde{p}_{\nu_1\nu_2\nu_3}\) to the frequency \(f_{\nu_{1}\nu_{2}\nu_{3}}\) of the relevant triplet;
-
to put the frequencies of synonymous codons equal, keeping the frequency of the relevant amino acid the same
$$ \widetilde{p}_{\nu_1\nu_2\nu_3} = {\frac{1}{M}} \sum\limits_{\omega^{\prime}} f_{\omega^{\prime}}, $$where ω′ enlists M synonymous codons for the given amino acid, and
-
to put \(\widetilde{p}_{\nu_1\nu_2\nu_3}\) equal to the reconstructed frequency of the codon
$$ \tilde{p}_{\nu_1\nu_2\nu_3} = {\frac{f_{\nu_1\nu_2} \times f_{\nu_2\nu_3}}{f_{\nu_2}}} , $$(12)defined according to (3). Here \(f_{\nu_{i}\nu_{i+1}}\) is the dinucleotide frequency determined due to a downward summation of the codon frequency, and \(f_{\nu_{j}}\) is the frequency of the central nucleotide determined with respect to the codon frequency.
We have determined the indices (11) for 104 bacterial genomes, for all three patterns of the quasi-equilibrium frequency definition. Then, the data have been classified with unsupervised classification technique. Figure 5 shows the pattern of the bacterial genomes distribution at the space determined by the indices (11). Actually, the points representing individual genomes fall very close to a plane; the pattern of the distribution shown in the figure looks like a swallow-tail. Straight lines A and B are the kernels of the relevant classes implemented due to an unsupervised automated classification (see Sadovsky et al. 2007 for more details).
The classes observed due to statistical classification (as shown in Fig. 5) reveal a simple and clear biological issue: the genomes in the classes differ strongly from the point of view of C + G content. We calculated the total concentration of the nucleotides C and G over the entire genome, for everyone on them. Usually, C + G content is determined for considerably short fragments of a genome; we calculated the ratio
for entire genome. The classes identified through the development of unsupervised classification exhibit the following values of (13): average value is \(\langle\chi_{{\texttt{A}}}\rangle = 0.59075\) and \(\langle\chi_{{\texttt{B}}}\rangle = 1.30348,\) respectively, and the variances are \(\sigma_{\texttt{A}}=0.17826\) and \(\sigma_{\texttt{B}} =0.48974,\) respectively.
The correlation between unsupervised classification, and C + G content is not evident. The classes shown in Fig. 5 occupy a plane (differing from simple octant one); occupy here means that the genomes are located quite close to the plane, and the location at the indices (11–12) space might be parameterized by a single parameter χ. Meanwhile, this fact was not evident; moreover, the detailed study of the distribution shown in Fig. 5 may bring some more specific knowledge towards the fine structure of the classes observed through the implementation of unsupervised classification.
Informational symmetry in genomes
Formally speaking, each word ω, ω∈W(q) exhibits specific information value \(p_{\omega} = f_{\omega}/\tilde{f}_{\omega}.\) It is a common fact, that DNA molecule exists as a double helix, where nucleotides meet the complementary rule: A always opposes T , and C always opposes G , in reciprocal strands of the molecule. Thus, for each word ω, one always can develop so-called complementary palindrome \(\overline{\omega},\) that is read equally, in the opposite direction, with respect to the complementary rule. The couple
makes such complementary palindrome, of the length q = 6. Some complementary palindromes are perfect: same word makes both parts of that latter (e.g., ATATAT ). Obviously, a perfect complementary palindrome may have even symbols, only. The number of perfect palindromes (of the length q) is
for genetic sequences.
Surprisingly, the words coupling into a complementary palindrome exhibit similar (or close) information value. The correspondence, of course is not absolute: a variation between the values of p ω and \(p_{\overline{\omega}}\) may be rather significant. Moreover, the concordance in information values of the words within a couple of a complementary palindrome is deteriorated, as q grows up. Table 2 shows the symmetry of the triplets versus complementary palindromic triplets for 3R chromosome of D. melanjgaster. Here ω3 is a triplet, \(\overline{\omega}_3\) is the relevant complementary palindromic triplet; \(p_{\omega_3}\) and \(p_{\overline{\omega}_3}\) are their frequency, respectively.
We calculated p ω for each word ranging from q = 3 to q = 8; then, the absolute value of the difference \(\mu_{\omega} = p_{\omega} - p_{\overline{\omega}}\) was determined. Here \(\omega \div \overline{\omega}\) is a complementary palindrome. Then, \(\langle \mu \rangle,\) σμ, and maximal and minimal values of μ have been found. Table 3 shows these data observed for D. melanogaster chromosome 3R. The Table shows quite reasonable correspondence between the words within a complementary palindrome, with respect to their information value. Evidently, the relation would decay, as q grows up. One sees, that the standard deviation of the absolute values of a difference between the relevant information values observed aver a complementary palindrome increases, for thicker dictionaries. The correspondence observed within a complementary palindrome is perfect, for 3 ≤ q ≤ 6. Some discrepancies take place for longer words. Here are the palindromes manifesting the greatest deviation in the information value between the words: \(\texttt{ACCGCTA} \div \texttt{TAGCGGT} , \texttt{ TAGCGGA} \div \texttt{ TCCGCTA} , \texttt{ ACCCGTA} \div \texttt{ TACGGGT} , \hbox{and} \texttt{ CCTAGTC} \div \texttt{ GACTAGG},\) for q = 7; similar couples for q = 8 are the following: \(\texttt{ CGGGTACG} \div \texttt{ CGTACCCG} , \texttt{ AGTACCGG} \div \texttt{ CCGGTACT} , \texttt{ CGGTAGAC} \div \texttt{GTCTACCG} , \hbox{and}\, \texttt{GCGCGTAC} \div \texttt{GTACGCGC}.\)
The symmetry described earlier manifests in various patterns, for different genomes. Eukaryotic genomes seem to be rather homogeneous, from the point of view of the relevance of information value of the strings composing complementary palindromes. Here are two key factors affecting the pattern: the former is the length of a sequence, and the latter is its origin. Thus, the situation is getting worse, as one takes into consideration bacterial genomes. They are shorter, and they differ in the structure. Hence, it yields a significant decay of the symmetry observed for W(6) and thicker. Moreover, bacterial genomes manifest an inversion on the information value magnitude of the words making a complementary palindrome: quite often, the words of the length q = 7 and q = 8 making a complementary palindrome manifest the inverse values of p ω and \(p_{\bar{\omega}}.\)
The most intriguing thing in the symmetry described earlier is that it is found for a single DNA strand; formally, such strand “knows nothing” about the opposite one. The symmetry makes a constraint on the alphabet cardinality: it must me even. Yet, there is no clear idea what issue stands in origin of the symmetry: an evenness of genetic alphabet, or the symmetry. Meanwhile, they put each other together.
Discussion and conclusion
A study of statistical properties nucleotide sequences has a long-standing history. A lot has been found in biological issues encoded at the symbol sequences. Building a bridge between structure of biological macromoleculae and the function is the key problem of up-to-date biology, including bioinformatics, molecular genetics and relevant fields of mathematics and/or physics. A variety of methods, techniques and approaches implemented in this field assures a student in a visible progress, while the key problem still challenges the researchers. Evidently, the key problem may be pursued with two different approaches: the former is micro-scale, and the latter is meso- and macro-scale. A study of codon usage bias (see Carbone et al. 2003; Nakamura 2000; Sharp et al. 1993 and Codon usage bias) is a typical technique implemented within a micro-scale methodology. The works done in this area are numerous and fruitful.
The situation is worse, for the studies done within the meso- and macro-scale methodology. Here, we introduced an idea of statistical semantics of genomes, that is expected to bring something new. Statistical semantics itself is a hypothesis on the correlation between the pattern of the distribution alongside a genome of specially marked strings, and the functionally charged sites in it. Doubtlessly, a lot depends on the specific form of the hypothesis, e.g., on the method of the identification of those marked strings. Basic idea of our approach is to use as much “biologically independent” features to identify the strings, as possible. Surely, this approach does not eliminate other ones, which explore the biological features. Meanwhile, our approach provides a student with essential, fundamental constraints towards the possibility to reveal a (biologically valuable) knowledge from the study of statistics, primarily, of short strings observed within a genome.
We used the information value (that is the ratio of real frequency of a string, and its expected frequency; see (2–3) for details) to identify the specially marked strings observed over a genome. A possible way to retrieve the biologically valuable knowledge from the statistics is as following. Suppose, the set \(\overline{K}_q\) (see Eq. 9) of specially marked strings is identified. Suppose, then, that the set of biologically (or, wider, semantically) meaningful sites Σ is also identified. Each site σ j , σ j ∈ Σ could be identified with the number \(m_{\sigma_j}\) of the starting nucleotide. Thus, statistical semantics means an implementation of the distribution function revealing the relation between the sets \(\overline{K}_q\) and Σ.
Consider a set of apices identified in some way; one may expect that the distribution of these words is either random (say, following Poisson distribution), or non-random, manifesting some order. To reveal a pattern, we studied the distribution of the distances (measured as a number of nucleotides) between two next apices observed within a genome. Figure 4 shows such distribution developed for chromosome 3R of D. melanogaster; pyramids are rooted at q = 3, and truncated at q = 8. The histogram starts from q = 9; the point is that the number of couples located in a symbol next to each other is equal to 708, and the number of couples gaped with two symbols is equal to 158. It means, that such couples intersect over a string of the length q = 7 and q = 6, respectively. This increase in frequency in such tight couples demonstrates that there are numerous information valuable words of q > 9.
Evidently, a study of an individual relation through such distribution function does not make so much sense. There is no way to retrieve knowledge towards the biological issues standing behind the nucleotide sequences immediately. To do that, a researcher should go into comparative investigation of a family of genomes exhibiting various level of (biologically determined) relationship. Suppose, one has a family of quite related genomes (say, the entities of the same bacterial species, but of different strains), and all the entities are completely deciphered (i.e., any site within a genome is clearly attributed with the function). Suppose, then, that properly tuned sets of apices are determined, for these genomes. If a new genome is added to the family of entities, then one can expect that the apices of pyramids of this new entity determined in the manner similar to those from a family, would match the sites with similar functionality, with high probability.
Notes
The theory and methodology described below is applicable to a sequence from an arbitrary (finite) alphabet ℵ, say, for amino acid sequences.
An equality of these two sums stands behind the connection of a sequence into a ring.
Strictly speaking, information capacity is defined for a frequency dictionary, not for a sequence; we shall not make the difference between them, unless a mispresentation occurs.
References
Bugaenko NN, Gorban AN, Sadovsky MG (1996) Towards the information content of nucleotide sequences. Mol Biol Mosc 30:529
Bugaenko NN, Gorban AN, Sadovsky MG (1998) Maximum entropy method in analysis of genetic text and measurement of its information content. Open Syst Inf Dyn 5:265
Carbone A, Zinovyev A, Kepes F (2003) Codon Adaptation Index as a measure of dominating codon bias. Bioinformatics 19:2005
Durand B, Zvonkin A (2004) L’héritage de Kolmogorov en Mathématiques, Berlin, pp 269–287
Gorban AN, Popova TG, Sadovsky MG (1994) Redundancy of genetic texts and mosaic structure of genomes. Mol Biology (Mosc) 28:313
Gorban AN, Karlin IV (2005) Invariant manifolds for physical and chemical kinetics. Lect. Notes Phys, 660. Springer, Berlin
Nakamura PM (2000) Codon usage: mutational bias, translational selection and mutational biases. Nucleic Acids Res 19:8023
Popova TG, Sadovsky MG (1995) Introns differ from exons in their redundancy. Russ J Genet 31:1365
Rui H, Bin W (2001) Statistically significant strings are related to regulatory elements in the promoter regions of Saccharomyces cerevisiae. Physica A 290:464
Sadovsky MG (2002a) Information capacity of symbol sequences. Open Syst Inf Dyn 9:37
Sadovsky MG (2002b) Towards the information capacity of symbol sequences. Electron Inform Control 1:82
Sadovsky MG (2002c) Towards the redundancy of viral and prokaryotic genomes. Russ J Genet 38:575
Sadovsky MG (2003) Comparison of real frequencies of strings vs. the expected ones reveals the information capacity of macromoleculae. J Biol Phys 29:23
Sadovsky MG (2005) Information capacity of biological macromoleculae reloaded ArXiv q-bio.GN 0501011 v1
Sadovsky MG (2006) Information capacity of nucleotide sequences and its applications. Bull Math Biol 68:156
Sadovsky MG, Putintzeva YA (2007) Codon usage bias measured through entropy approach, arXiv:0706.2077v1, 14 June 2007
Shannon CE, Weaver W (1949) The mathematical theory of communication. University of Illinois Press, Urbana
Sharp PM, Stenico M, Peden JF, Lloyd AT (1993) Codon usage: mutational bias, translational selection and mutational biases. Nucleic Acids Res 15:8023
Zubkov AM, Mikhailov VG (1974) Limit distributions of random variables associated with long duplications in a sequence of independent trials. Probab Theory Appl 19:173
Zvonkin AK, Levin L (1970) The complexity of finite objects and development of the concepts of information and randomness by means of the theory of algorithms. Russ Math Surv 25(6):83
Acknowledgments
We are thankful to Prof. Alexander N. Gorban from Leicester University, for valuable discussions and inspiring ideas, and to Dr. Tatyana G. Popova from the Institute of Computational Modelling of RAS for stimulating interest in this work.
Author information
Authors and Affiliations
Corresponding author
Additional information
The results present here were partially obtained due to the support from Krasnoyarsk Science Foundation.
Rights and permissions
About this article
Cite this article
Sadovsky, M.G., Putintseva, J.A. & Shchepanovsky, A.S. Genes, information and sense: complexity and knowledge retrieval. Theory Biosci. 127, 69–78 (2008). https://doi.org/10.1007/s12064-008-0032-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12064-008-0032-1