Abstract
Wheat stripe (yellow) rust, caused by Puccinia striiformis West. f. sp. tritici (Pst), is one of the most destructive diseases in many wheat-growing countries, especially in China, the largest stripe rust epidemic area in the world. Growing the resistant cultivars is an effective, economic and environmentally friendly way to control this disease. Wheat cultivar Zhengmai 7698 has shown a high-level resistance to wheat stripe rust. To elucidate its genetic characteristics and location of the resistance gene, Zhengmai 7698 was crossed with susceptible variety Taichung 29 to produce \(\hbox {F}_{{1}}\), \(\hbox {F}_{{2}}\) and \(\hbox {BC}_{{1}}\) progeny generations. The genetic analysis showed that the stripe rust resistance in Zhengmai 7698 to Pst predominant race CYR32 was controlled by a single-dominant gene, namedYrZM. Bulked segregant analysis and simple sequence repeat (SSR) markers were used to map the gene. Four SSR markers, Xbarc198, Xwmc179, Xwmc786 and Xwmc398 on chromosome 6BL were polymorphic between the parents and resistance, and susceptible bulks. A linkage genetic map was constructed using 212 \(\hbox {F}_{{2}}\) plants in the sequential order of Xwmc398, Xwmc179, YrZM, Xbarc198, Xwmc786. As this gene is effective against predominant race CYR32, it is useful in combination with other resistance genes for developing new wheat cultivars with resistance to stripe rust.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Wheat stripe (yellow) rust is a typical air-borne disease. This disease is a serious threat to wheat production in Asia, North America, Europe and other wheat-growing areas. China is the largest epidemic region of wheat stripe rust in the world and its occurrence, and damage are especially frequent and severe (Wan et al. 2004). Stripe rust reduces wheat yields by damaging plants and shrinking the grain. Many severe epidemics of wheat stripe rust have occurred since 1950, including the nation-wide extremely severe epidemics in 1950, 1964, 1990 and 2002 which caused multimillion tons of yield losses in these epidemic years. Highly virulent races and widely grown susceptible cultivars were attributed to the epidemics.
Due to the appearance of new races CYR30, CYR31, CYR32 and CYR33 since 1995, many wheat cultivars of the Fan 6 derivatives, Mianyang series and Shuiyuan-derived cultivars have become susceptible. In recent years, a new group of Guinong 22-virulent races that infect wheat cultivars with Yr24 (=Yr26) has been rapidly increasing in frequency and distribution. The predominant races CYR32, CYR33 and new races have accelerated the turnover frequency of wheat cultivars. This has put forward higher requirement of developing wheat cultivars with effective resistance to stripe rust in China.
The use of resistance genes against stripe rust is essential in wheat breeding programmes. Stripe rust resistance genes in wheat have been designed with the symbol ‘Yr’ (Lupton and Macer 1962). Numerous genes for stripe rust resistance have been identified, including 78 Yr permanently named genes (McIntosh et al. 2017) and hundreds of temporarily named genes. Many resistance genes were identified in common wheat, while some came from wheat-related species, such as Yr5 from Triticum spela album (Macer 1966);Yr7, Yr53, Yr64 and Yr65 from T. durum (Macer 1966; Xu et al. 2013; Cheng et al. 2014); Yr9 from Secale cereale (Macer 1975); Yr15, Yr35 and Yr36 from T. dicoccoides (Gerechter-Amitai et al. 1989; Marais et al. 2005a; Uauy et al. 2005);Yr24 (=Yr26) from T. turgidum L. (McIntosh and Lagudah 2000; Ma et al. 2001); Yr8, Yr17, Yr28, Yr37, Yr38, Yr40 and Yr42 from different Aegilops species (Riley et al. 1968; Robert et al. 1999; Singh et al. 2000; Marais et al. 2005b, 2006; Kuraparthy et al. 2007; Marais et al. 2009) and Yr50 from Thinopyrum intermedium (Liu et al. 2013). Most of the genes with resistance to stripe rust were on B group chromosomes. Due to quick development of new races of Puccinia striiformis f. sp. tritici (Pst), wheat cultivars with race-specific resistance usually become susceptible about 3–5 years after release. Currently, most reported seedling resistance genes are ineffective against race CYR32 in China (Yang et al. 2003). Although, a lot of efforts have been made to identify genes for resistance and develop stripe rust-resistant cultivars, it is still a great challenge to have enough wheat cultivars with effective and long-lasting resistance. Therefore, the best strategy for controlling wheat stripe rust is to introduce new resistance genes into wheat-breeding programmes, broaden the genetic basis, and develop cultivars with durable resistance and many other excellent traits.
Zhengmai 7698 with high quality and yield potential was developed by the Molecular Breeding Laboratory, Wheat Research Center, Henan Academy of Agricultural Sciences. It has been grown in more than 13 million acres since its release in 2011. The cultivar has resistance to stripe rust. The objective of this study was to identify the resistance gene(s) through genetic analysis and molecular mapping.
Materials and methods
Plant materials and Pst inoculum
Stripe rust resistant cultivar Zhengmai 7698 was developed from a cross between Zhengmai 9405 and 4B269, and then crossed with Zhoumai 16. Spring wheat Taichung 29 was susceptible to all known Chinese races of Pst. An \(\hbox {F}_{{2}}\) population derived from crossing Zhengmai 7698 with the susceptible cultivar Taichung 29 was used for genetic analysis and molecular mapping. In addition, \(\hbox {F}_{{1}}\) plants were backcrossed with Taichung 29 to develop a \(\hbox {BC}_{{1}}\) population to confirm the \(\hbox {F}_{{2}}\) results. A highly susceptible wheat cultivar, Mingxian 169, was used as a disease spreader in the field tests.
Single urediniospore isolates of eight Pst races (CYR17, CYR26, CYR27, CYR29, CYR31, CYR32, CYR33 and V26) maintained at the Institute of Plant Protection, Chinese Academy of Agricultural Sciences, China, were used for evaluating wheat cultivars. The isolate of race CYR32, which is prevalent throughout China, and virulent on all Chinese differential cultivars except for Zhong 4 and all world differential cultivars except for Moro, was used to phenotype the segregating populations for genetic analysis and molecular mapping. Fresh urediniospores of an isolate were produced on susceptible wheat cultivar Mingxian 169.
Resistance evaluation to wheat stripe rust
Seedlings of Zhengmai 7698 were grown in a greenhouse under controlled conditions of \(20^{\circ }\hbox {C}\), humidity 80% and 14 h daily light. Plants at the fully expanded one-leaf stage were inoculated with fresh urediniospores suspended in aqueous solution containing 0.05% v/v Tween 20. The inoculated seedlings were placed in a dew chamber at \(10^{\circ }\hbox {C}\) for 24 h and then transferred into a controlled greenhouse with a diurnal cycle of 14 h of light at \(18^{\circ }\hbox {C}\) and 10 h of dark at \(12^{\circ }\hbox {C}\). \(\hbox {F}_{{1}}\), \(\hbox {BC}_{{1}}\) and \(\hbox {F}_{{2}}\) plants were inoculated with CYR32. After 14–15 days, when the susceptible cultivar Mingxian 169 was fully sporulating, infection type (IT) data were recorded according to the scale of 0–4 (McIntosh et al. 1995). ITs 0–2 were considered resistant whereas ITs 3–4 were susceptible.
Field tests were conducted in the Langfang Experimental Station in the Hebei province. \(\hbox {F}_{{2}}\) plants were grown in rows with 15–20 seeds planted in a 1-m row. Mingxian 169 was planted as a susceptible control around the field and inoculated at the jointing stage with urediniospores of CYR32 suspended in 0.05% v/v Tween 20. The inoculated plants were covered with plastics overnight to facilitate dew formation for Pst infection. The plastics were removed in the following morning. ITs were scored when Mingxian 169 had more than 80% severity. The IT data were classified as resistant (R, IT 0–2) and susceptible (S, IT 3–4).
DNA extraction, PCR amplification, electrophoresis and gel visualization
Genomic DNA was extracted from leaf tissues of each \(\hbox {F}_{{2}}\) plant and the parents using a modified method of cetyltrimethylammonium bromide (CTAB) (Rogers and Bendich 1985) and dissolved in \(100\,\mu \hbox {L}\) TE-buffer (10 mM Tris, pH 8.0, 1 mM EDTA). The DNA was quantified with a NanoDrop 1000 Spectrophotometer (Thermo Fisher Scientific, USA) and assessed in 1% agarose gel. DNA samples were diluted to a concentration of 50 ng/\(\mu \hbox {L}\) before storing at \(-20^{\circ }\hbox {C}\).
To identify the DNA markers that are associated with stripe rust resistance, the bulked segregant analysis (BSA) method was used. DNA samples from 10 R (IT 0 0;) and 10 S (IT 4) \(\hbox {F}_{{2}}\) plants were mixed to construct the R and S bulks, respectively and then evaluated with simple sequence repeat (SSR) markers together with the parents. SSR markers in the WMC, GWM, BARC, CFA, CFD and GDM series (http://wheat.pw.usda.gov/ggpages/SSR) were chosen across the 21 wheat chromosomes according to the consensus map of Somers et al. (2004).
PCR was performed in a 10-\(\mu \hbox {L}\) solution containing 2 \(\mu \hbox {L}\) of 50 ng/\(\mu \hbox {L}\) template DNA, 1 \(\mu \hbox {L}\) of \(10\times \) PCR buffer, 0.2 \(\mu \hbox {L}\) of 5 U/\(\mu \hbox {L}\) Taq DNA polymerase, 2.5 mM of each dNTP, 5 \(\mu \hbox {mol/L}\) forward and reverse primer solutions and sterilized double distilled \(\hbox {H}_{{2}}\)O. Amplification was performed in a GeneAmp PCR System 9700 programmed as follows: 5 min at \(94^{\circ }\hbox {C}\) for initial denaturation; 35 cycles each consisting of 1 min at \(94^{\circ }\hbox {C}\) for denaturation, 1 min at \(50^{\circ }\hbox {C}\), \(55^{\circ }\hbox {C}\) or \(60^{\circ }\hbox {C}\) for annealing depending on individual primers, 1 min at \(72^{\circ }\hbox {C}\) for extension and finally a 10 min extension step at \(72^{\circ }\hbox {C}\). Six microlitre formamide loading buffer (98% formamide, 10 mM EDTA (pH 8.0), 0.5% (w/v) xylene cyanol and 0.5% (w/v) bromophenol blue) was added to each PCR product. After 5 min denaturation at \(94^{\circ }\hbox {C}\), 5 \(\mu \hbox {L}\) of the PCR product and loading buffer mixture for each sample was loaded for electrophoresis in a 6% polyacrylamide gel in 1\(\times \) TBE buffer (89 mM Tris, 89 mM boric acid, 2 mM EDTA, pH 8.3) (Bassam et al. 1991). Electrophoresis was carried out at 1400 V for 1.0–1.5 h. Gel staining and visualization was done as previously described (Chen et al. 1998). Polymorphic markers were used to genotype the \(\hbox {F}_{{2}}\) population. Genotype data were used to construct a genetic map and locate the resistance gene.
Mapping and data analysis
A linkage map was established using software JoinMap ver. 3.0 (van Ooijen and Voorrips 2001). A regression mapping algorithm and Kosambi’s function were utilized to convert recombination fractions into map distances and order the markers (Kosambi 1943).
Chi-square \((\chi ^{2})\) tests were applied to evaluate the \(\hbox {F}_{{2}}\) segregation ratios of R/S plants to determine the goodness of a theoretical ratio.
Results
Response of Zhengmai 7698 to Pst races
When tested in the seedling stage in the greenhouse, Zhengmai 7698 was resistant (IT 0;) with necrotic spots without any uredinia to all tested races except for V26, while Taichung 29 was susceptible (IT 4) to all races (table 1).
Reactions of progenies to CYR32
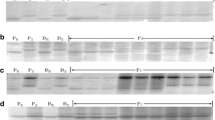
In the progeny test with race CYR32, Taichung 29 was susceptible (4) and Zhengmai 7698 was resistant as described above. All \(\hbox {F}_{{1}}\) plants were resistant and \(\hbox {F}_{{2}}\) plants segregated into resistant and susceptible reactions (figure 1), indicating dominant resistance of Zhengmai 7698. The \(\hbox {F}_{{2}}\) segregation fits a 3 resistant:1 susceptible ratio \((\chi ^{2}\,{=}\,1.73, P\,{=}\,0.19)\), indicating one gene for resistance (table 2). The 32 tested \(\hbox {BC}_{1}\) plants were divided into 15 resistant and 17 susceptible plants. The observed segregation ratio was consistent with the expected ratio of 1:1 \((\chi ^{2}=0.13, P\,{=}\,0.72)\). These results showed that Zhengmai 7698 has a dominant gene conferring resistance to race CYR32.
In the field evaluation, Taichung 29 was susceptible (IT 4), Zhengmai 7698 was resistant (IT 0;) and \(\hbox {F}_{{1}}\) plants were also resistant (IT 0;). The \(212 \hbox { F}_{{2}}\) plants segregated into 158 resistant and 54 susceptible fitting the ratio of 3:1 \((\chi ^{2}=0.03, P\,{=}\,0.86)\), indicating that resistance under field conditions was also controlled by a single-dominant gene (table 2). The segregation of resistant and susceptible \(\hbox {BC}_{1}\) plants fit the 1:1 ratio \((\chi ^{2}=0.04, P\,{=}\,0.84)\). The field results confirmed a single-dominant gene for resistance to CYR32 and the gene was tentatively designated as YrZM.
Screening of polymorphic markers
From 240 tested SSR markers, which were selected to cover the 21 common wheat chromosome 5 markers, Xwmc786, Xwmc398, Xwmc179, Xbarc198 and Xwmc597, were polymorphic between the parents, and between the resistant and susceptible bulks (figure 2), indicating that these markers were linked to YrZM. The five markers were all from chromosome 6B, suggesting that YrZM was likely located on chromosome 6B.
Linkage analysis and mapping resistance gene
The associations of the five polymorphic SSR markers with the resistance gene were tested with 212 \(\hbox {F}_{{2}}\) plants. Because the dominant marker Xwmc597 was present in Taichung 29 and susceptible bulk, it was not used to construct the linkage map. Other four markers were present in Zhengmai 7698 and the resistant bulk. Figure 3 shows the amplification results of marker Xbarc198 for the two parents and some \(\hbox {F}_{{2}}\) plants. Three of the markers were codominant and one was dominant (table 3). Codominant markers Xwmc398, Xwmc179 and Xbarc198 segregated at the expected 1:2:1 ratio for homozygous resistant, heterozygous and homozygous susceptible in the \(\hbox {F}_{{2}}\) population \((\chi ^{2}=0.03\,{-}\,0.46, P\,{=}\,0.50\,{-}\,0.86)\). The segregation of marker Xwmc786 fit the expected ratio 3:1 \((\chi ^{2}=0.91, P\,{=}\,0.34)\) for presence and absence in the \(\hbox {F}_{{2}}\) population (table 3).
Amplification of \(\hbox {F}_{{2}}\) plants of cross Taichung 29/Zhengmai 7698 using the primers of SSR marker Xbarc198. \(\hbox {P}_{{1}}\), Zhengmai 7698; \(\hbox {P}_{{2}}\), Taichung 29. A, homozygous resistant; B, homozygous susceptible; H, heterozygote. The band associated with Yr is indicated by the arrow.
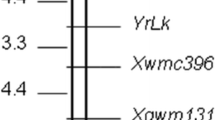
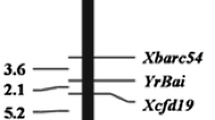
A linkage map was constructed with the four segregating markers (figure 4). YrZM was flanked by markers Xwmc179 and Xbarc198 at genetic distances of 2.7 and 2.4 cM, respectively.
Discussion
The first requirement for breeding programmes is to have resistance sources which can be used for developing cultivars with resistance to major diseases and pests. Many genetic studies of wheat stripe rust have identified one, two or more genes with additive effects on disease resistance. Wheat cultivar Zhengmai 7698 was derived from a complex cross involving three parental varieties (Zhengmai 9405/ Zhengmai 4B269/Zhoumai 16). The cultivar is resistant to many Pst races, including predominant races CYR17, CYR26 and CYR32. CYR32, virulent to Yr1, Yr2, Yr3, Yr4, Yr6, Yr7, Yr9, Yr27, YrA, YrAlba, YrCle, YrCV, YrGaby, YrRes, YrSD, YrSP and YrSu circumvents resistance in many wheat cultivars and has caused a severe epidemic in recent decades in China (Kang et al. 2010). In addition to CYR32, CYR33 has become predominant, which has made more cultivars susceptible (Kang et al. 2010). Only a few genes for race-specific all stage resistance, such asYr5 and Yr15 are effective against Chinese Pst races. In this study, Zhengmai 7698 was found to be resistant to Pst races CYR32 and CYR33, indicating that its resistance is effective against the major predominant races. Genetic analysis indicated that resistance of Zhengmai 7698 against CYR32 was conferred by a single dominant gene, tentatively designated as YrZM. The closely linked SSR primers mapped YrZM to the long arm of chromosome wheat 6B. The two flanking markers, Xbarc198 and Xwmc179, should be useful for identifying YrZM in breeding programmes.
Currently, four formally named Yr gene loci (Yr4, Yr35, Yr36 and Yr78) have been identified on chromosome 6B (Wang and Chen 2017). The typical carriers of these genes are Cappelle-Desprez (Yr4a), Hybrid 46 (Yr4b), 98M71 (Yr35), Glupro (Yr36) and PI 519805 (Yr78). Cappelle-Desprez was first released in France in 1946 and widely cultivated in Western Europe up to 1970s (Lupton and Macer 1962; Worland and Law 1986; Bonjean et al. 2001). It is known to possess the seedling resistance genes Yr4a (de Vallavieille-Pope et al. 1990) on chromosome 6B and Chen et al. (1996) also reported Yr4b in Hybrid 46 on chromosome 6B. However, their specific chromosomal location is unknown. Based on the seedling tests with different Chinese Pst races, Cappelle-Desprez and Hybrid 46 were susceptible to CYR32, indicating that Yr4a and Yr4b were different from YrZM. Yr35 on the short arm of chromosome 6B was transferred from T. turgidum spp. dicoccoides to hexaploid wheat by Marais et al. (2005a). Yr36, also located on chromosome 6BS and from T.turgidum ssp. dicoccoides, confers high-temperature and adult-plant resistance (Uauy et al. 2005). Through pedigree analysis, it was concluded that YrZM was different from Yr35 and Yr36. Yr78 for adult-plant resistance to stripe rust was identified in PI 519805 and several other common wheat cultivars and mapped to chromosome 6BS (Dong et al. 2017). As YrZM is located on chromosome 6BL and resistant to CYR32 and CYR33, it is different from all permanently named Yr genes on chromosome 6B.
There were three regions of importance in chromosome 6B for stripe rust resistance (Rosewarne et al. 2013): QRYr6B.1 referred to the short arm; QRYr6B.2 mainly contained Yr36 and the final region was identified in Pastor (Rosewarne et al. 2012) and Pavon (William et al. 2006). Due to the differences in marker platforms and associated linkage maps, it was difficult to accurately compare the chromosomal location of YrZM with previously identified quantitative trait loci (QTL). By comparing chromosome 6B with other published SSR maps based on shared markers, the YrZM locus is different from previously reported QTL for stripe rust resistance (figure 5). As marker platforms will continue to be developed and improved, and more linkage maps will be published with combinations of SSR and SNP markers, we will be able to compare the YrZM linkage map with maps containing similar chromosomal regions.
To improve the durability of resistance in wheat cultivars and to achieve sustainability of stripe rust management, it is important to diversify the resistance gene in breeding programmes and deploy diverse effective resistance genes in wheat cultivars. As a widely grown cultivar for high yield and stripe rust resistance, Zhengmai 7698 is also a valuable cultivar to be used in breeding programmes. As its stripe rust resistance gene YrZM is race-specific and ineffective against the recently emerged race groups, e.g. V26, caution needs to be taken to use the gene in combinations with other effective genes and especially for nonrace specific adult-plant resistance genes. The flanking markers Xbarc198 and Xwmc179 can be used for marker-assisted selection in pyramiding YrZM with other resistance genes.
References
Bassam B. J., Caetano-Anollés G. and Gresshoff P. M. 1991 Fast and sensitive silver staining of DNA in polyacrylamide gels. Anal. Biochem. 196, 80–83.
Bonjean A. P., Doussinault G. and Stragliati J. 2001 French wheat pool. In The world wheat book. A history of wheat breeding (ed. A. P. Bonjean and W. J. Angus), pp. 128–165. Lavoisier Publishing, Paris.
Chen X. M., Jones S. S. and Line R. F. 1996 Chromosomal location of genes for resistance to Puccinia striiformis in seven wheat cultivars with resistance genes at the Yr3 and Yr4 loci. Phytopathology 86, 1228–1233.
Chen X. M., Line R. F. and Leung H. 1998 Genome scanning for resistance-gene analogs in rice, barley, and wheat by high-resolution electrophoresis. Theor. Appl. Genet. 97, 345–355.
Cheng P., Xu L. S., Wang M. N., See D. R. and Chen X. M. 2014 Molecular mapping of genes Yr64 and Yr65 for stripe rust resistance in hexaploid derivatives of durum wheat accessions PI 331260 and PI 480016. Theor. Appl. Genet. 127, 2267–2277.
de Vallavieille-Pope C., Picard-Formery H., Radulovic S. and Johnson R. 1990 Specific resistance factors to yellow rust in seedlings of some French wheat varieties and races of Puccinia striiformis west end in France. Agronomie 2, 103–113.
Dong Z. Z., Hegarty J. W., Zhang J. L., Zhang W. J., Chao S. M., Chen X. M. et al. 2017 Validation and characterization of a QTL for adult plant resistance to stripe rust on wheat chromosome arm 6BS (Yr78). Theor. Appl. Genet. 130, 2127–2137.
Gerechter-Amitai Z. K., Van Silfhout C. H., Grama A. and Kleitman F. 1989 Yr15 – a new gene for resistance to Puccinia striiformis in Triticum dicoccoides sel. G-25. Euphytica 43, 187–190.
Kang Z. S., Zhao J., Han D. J., Zhang H. C., Wang X. J., Wang C. F. et al. 2010 Status of wheat rust research and control in China. BGRI 2010 Technical Workshop, pp. 1–21. St Petersburg, Russia.
Kosambi D. D. 1943 The estimation of map distances from recombination values. Ann. Eugen. 12, 172–175.
Kuraparthy V., Chhuneja P., Dhaliwal H. S., Kaur S., Bowden R. L. and Gill B. S. 2007 Characterization and mapping of cryptic alien introgression from Aegilops geniculata with new leaf rust and stripe rust resistance genes Lr57 and Yr40 in wheat. Theor. Appl. Genet. 114, 1379–1389.
Liu J., Chang Z., Zhang X., Yang Z., Li X., Jia J. et al. 2013 Putative Thinopyrum intermedium-derived stripe rust resistance gene Yr50 maps on wheat chromosome arm 4BL. Theor. Appl. Genet. 126, 265–274.
Lupton F. C. H. and Macer R. C. F. 1962 Inheritance of resistance to yellow rust (Puccinia glumarum Erikss. and Henn) in seven varieties of wheat. Trans. Br. Mycol. Soc. 45, 21–45.
Ma J. X., Zhou R. G., Dong Y. S., Wang L. F., Wang X. M. and Jia J. Z. 2001 Molecular mapping and detection of the yellow rust resistance gene Yr26 in wheat transferred from Triticum turgidum L. using microsatellite markers. Euphytica 120, 219–226.
Macer R. C. F. 1966 The formal and monosomic genetic analysis of stripe rust (Puccinia striiformis) resistance in wheat. Hereditas (Suppl. 2), 127–142.
Marais G. F., McCallum B., Snyman J. E., Pretorius Z. A. and Marais A. S. 2005a Leaf rust and stripe rust resistance genes Lr54 and Yr37 transferred to wheat from Aegilops kotschyi. Plant Breed. 124, 538–541.
Marais G. F., Pretorius Z. A., Wellings C. R., McCallum B. and Marais A. S. 2005b Leaf rust and stripe rust resistance genes transferred to common wheat from Triticum dicoccoides. Euphytica 143, 115–123.
Marais G. F., McCallum B. and Marais A. S. 2006 Leaf rust and stripe rust resistance genes derived from Aegilops sharonensis. Euphytica 149, 373–380.
Marais F., Marais A., McCallum B. and Pretorius Z. 2009 Transfer of leaf rust and stripe rust resistance genes Lr62 and Yr42 from Aegilops neglecta Req. ex Bertol. to common wheat. Crop Sci. 49, 871–879.
McIntosh R. A. and Lagudah E. S. 2000 Cytogenetic studies in wheat XVIII. Gene Yr24 for resistance to stripe rust. Plant Breed. 119, 81–93.
McIntosh R. A., Wellings C. R. and Park R. F. 1995 Wheat rust: an atlas of resistance genes. CSIRO, Sydney.
McIntosh R. A., Dubcovsky J., Rogers W. J., Morris C. and Xia X. C. 2017 Catalogue of gene symbols for wheat: 2017 supplement (https://shigen.nig.ac.jp/wheat/komugi/genes/symbolClassList.jsp).
Riley R., Chapman V. and Johnson R. 1968 Introduction of yellow rust resistance of Aegilops comosa into wheat by genetically induced homoeologous recombination. Nature 217, 383–384.
Robert O., Abelard C. and Dedryver F. 1999 Identification of molecular markers of the detection of the yellow rust resistance gene YrI7 in wheat. Mol. Breed. 5, 167–175.
Rogers S. O. and Bendich A. J. 1985 Extraction of DNA from milligram amounts of fresh, herbarium and mummified plant tissues. Plant Mol. Biol. 5, 69–76.
Rosewarne G. M., Singh R. P., Huerta-Espino J., Herrera-Foessel S. A., Forrest K. L., Hayden M. J. et al. 2012 Analysis of leaf and stripe rust severities reveals pathotype changes and multiple minor QTLs associated with resistance in an Avocet 3 Pastor wheat population. Theor. Appl. Genet. 124, 1283–1294.
Rosewarne G. M., Herrera-Foessel S. A., Singh R. P., Huerta-Espino J., Lan C. X. and He Z. H. 2013 QTL trait loci of stripe rust resistance in wheat. Theor. Appl. Genet. 126, 2427–2449.
Singh R. P., Nelson J. C. and Sorrells M. E. 2000 Mapping Yr28 and other genes for resistance to stripe rust in wheat. Crop Sci. 40, 1148–1155.
Somers D. J., Isaac P. and Edwards K. 2004 A high-density microsatellite consensus map for bread wheat (Triticum aestivum L.). Theor. Appl. Genet. 109, 1105–1114.
Uauy C., Brevis J., Chen X., Khan I., Jackson L., Chicaiza O. et al. 2005 High-temperature adult-plant (HTAP) stripe rust resistance gene Yr36 from Triticum turgidum ssp. dicoccoides is closely linked to the grain protein content locus Gpc-B1. Theor. Appl. Genet. 112, 97–105.
van Ooijen J. W. and Voorrips R. E. 2001 JoinMap 3.0, Software for the calculation of genetic linkage maps. Plant Research International, Wageningen, The Netherlands.
Wan A., Zhao Z., Chen X., He Z., Jin S., Jia Q. et al. 2004 Wheat stripe rust epidemic and virulence of Puccinia striiformis f. sp. tritici in China in 2002. Plant Dis. 88, 896–904.
Wang M. N. and Chen X. M. 2017 Stripe rust resistance. In Stripe rust (ed. X. M. Chen and Z. S. Kang), chapter 5, pp. 353–558. Springer, Dordrecht.
William H. M., Singh R. P., Huertaespino J., Palacios G. and Suenaga K. 2006 Characterization of genetic loci conferring adult plant resistance to leaf rust and stripe rust in spring wheat. Genome 49, 977–990.
Worland A. J. and Law C. N. 1986 Genetic analysis of chromosome 2D of wheat. I. The location of genes affecting height, day-length insensitivity, hybrid dwarfism and yellow rust resistance. \(Z\). Pflanzenzücht. 96, 331–345.
Xu L. S., Wang M. N., Cheng P., Kang Z. S., Hulbert S. H. and Chen X. M. 2013 Molecular mapping of Yr53, a new gene for stripe rust resistance in durum wheat accession PI 480148 and its transfer to common wheat. Theor. Appl. Genet. 126, 523–533.
Yang Z. M., Xie C. J. and Sun Q. X. 2003 Situation of the sources of stripe rust resistance of wheat in the post-CY32 era in China. Acta Agron. Sin. 29, 161–168.
Acknowledgements
This study is supported by the National Key Basic Research Programme of China (2013CB127700) and the National Natural Science Foundation of China (nos. 31272033 and 31261140370). We thank Dr. Chen Xianming (Department of Plant Pathology, Washington State University) for editing the language.
Author information
Authors and Affiliations
Corresponding author
Additional information
Corresponding editor: Manoj Prasad
Rights and permissions
About this article
Cite this article
Li, H., Feng, J., Xu, X. et al. Genetic analysis and location of a resistance gene for Puccinia striiformis f. sp. tritici in wheat cultivar Zhengmai 7698. J Genet 97, 931–937 (2018). https://doi.org/10.1007/s12041-018-0986-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12041-018-0986-9