Abstract
WD repeat containing protein 5 (WDR5), Retinoblastoma Binding Protein 5 (RbBP5), Absent-Small-Homeotic-2-Like protein (ASH2L), and Dumpy-30 (Dpy30) have been reported to be the integral and shared components of all the SET1 family of histone 3 lysine 4 histone methyltransferase (HMT) complexes. Collectively called the WRAD complex, these proteins are pivotal to the HMT activity of the SET1 complexes. Recent reports highlight the novel non-canonical functions of WRAD in cellular processes other than its well-studied role in histone methylation and gene expression. In this review, we examine the diversity in emerging transcription-independent functions of WRAD.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Post-translational histone modifications add another layer of regulation to the ever changing chromatin landscape to activate or repress transcription. N- or C-terminal tails of histones can be methylated, phosphorylated, acetylated, ubiquitinated and, even, sumoylated or ribosylated to impact gene expression. All these histone modifications recruit chromatin remodelling protein complexes or alter the structure of chromatin (Bannister and Kouzarides 2011). In particular, lysine methylation regulates the activation (H3K4, H3K36, and H3K79) as well as repression (H3K9, H3K27, and H4K20) of transcription by methylation of specific residues in the histones (reviewed in Martin and Zhang 2005). Histone 3 Lysine 4 methylation (H3K4me) is closely associated with the activated state of the chromatin (Martin and Zhang 2005; Eissenberg and Shilatifard 2010). In mammals, the SET1 family of histone methyltransferases (HMT) is responsible for depositing the H3K4 methylation mark on promoters of active genes (Ernst and Vakoc 2012; Piunti and Shilatifard 2016).
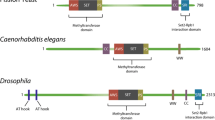
The SET1 family include at least six multi-protein complexes namely Mixed Lineage Leukemia (MLL1 or MLL), MLL2, MLL3, MLL4, Set1A, and Set1B proteins (Cao 2012; Ernst and Vakoc 2012). All these enzymes have unique non-redundant roles (Ansari and Mandal 2010; Shilatifard 2012; Shinsky and Cosgrove 2015). While Set1A and Set1B impart H3K4me3 mark to majority of gene promoters in mammals, MLL1 has been shown to regulate a subset of genes important during development, including the Hox loci (Wu et al. 2008). In contrast, MLL3 and MLL4 are involved in depositing the H3K4 mono-methylation mark at the enhancers (Herz et al. 2012; Hu et al. 2013). Like other methyltransferases, all members from this family act as multi-protein complexes with common subunits namely, WD repeat containing protein 5 (WDR5), Retinoblastoma Binding Protein 5 (RbBP5), Absent-Small-Homeotic-2-Like protein (ASH2L) and Dumpy-30 protein (DPY30) (Dou et al. 2006; Steward et al. 2006; Takahashi et al. 2011; Ernst and Vakoc 2012; van Nuland et al. 2013). Additional context-specific subunits also associate to impart unique properties to the complex (Yokoyama et al. 2004; Cho et al. 2007; Wu et al. 2008; Patel et al. 2007, 2014).
2 The conventional functions of WRAD
The WDR5, RbBP5, ASH2L, and DPY30 collectively called WRAD, along with the Set1/MLL Su(var)3-9, Enhancer-of-zeste, Trithorax (SET) domain form a minimal ‘core complex’ capable of optimal enzymatic activity (Dou et al. 2006; Steward et al. 2006; Patel et al. 2011; Ernst and Vakoc 2012; Dharmarajan et al. 2012; Shinsky and Cosgrove 2015; Zhang et al. 2015). In vitro reconstitution studies with the purified WRAD proteins and the catalytic SET domain of MLL have provided deeper insights into the role of WRAD in the functions and the assembly of the MLL complex (Steward et al. 2006; Odho et al. 2010; Dharmarajan et al. 2012; Senisterra et al. 2013). In vitro, MLL by itself is a mono-methyltransferase (Patel et al. 2008b; reviewed in Cosgrove and Patel 2010). However in association with WRAD, it is capable of di- and tri-methylation activities (Zhang et al. 2015). While WDR5, RbBP5, and ASH2L form the minimal WRA complex required for the methylation activity observed with the WRAD complex, DPY-30 increases the catalytic efficiency and specificity of WRAD for histone H3 peptide (Cosgrove and Patel 2010; Patel et al. 2011).
Interestingly, in vitro studies also suggest that WRAD, which lacks SET domain, can methylate histone 3 peptide (Cosgrove and Patel 2010; Patel et al. 2009, 2011). WRAD can mono-methylate histone 3 peptide by itself but requires the presence of MLL SET domain (even when catalytically inactive) for nucleosome mono-methylation (Patel et al. 2009, 2011; Dharmarajan et al. 2012).
Two independent studies showed that WDR5 subunit bridges the interaction between MLL and the WRAD complex. WDR5 engages in direct interactions with the conserved arginine-containing WDR5 interacting (Win) motif localized in the C-terminus of MLL protein (Patel et al. 2008a, 2008b; Dharmarajan et al. 2012). This motif is conserved in SET1 family members and shares sequence homology with histone H3 N-terminus. Correspondingly, Phe133 and Phe263 amino acid residues of WDR5 sandwich the guanidinium of central arginine of the Win motif of MLL (Arg3765). Both in vitro and in vivo studies suggest that point mutation of either Phe133 or Phe263 in WDR5 or Arg3765 in MLL nearly abolishes the interaction between MLL and WRA sub-complex (Patel et al. 2008a; Song and Kingston 2008). However, more recently, it has been suggested that RbBP5-ASH2L heterodimer can bind to MLL directly albeit with weak affinity (Shinsky et al. 2014; Li et al. 2016).
In addition to the catalytic activity, the components of the WRAD complex are also implicated in the maintenance of the global levels of H3K4 methylation and, stability, product specificity and recruitment of SET1 complexes on the chromatin (Briggs et al. 2001; Nagy et al. 2002; Dou et al. 2006; Steward et al. 2006; Smith et al. 2011; Ernst and Vakoc 2012; Zhang et al. 2015).
3 Non-conventional functions of WRAD
Apart from participation of WRAD in the transcriptional activation of the genes by histone methylation, recent findings suggest transcriptional-independent roles of WRAD in the regulation of the cell cycle progression (figure 1). WRAD along with MLL has recently been shown to regulate the mitotic progression in a transcriptional independent manner (Ali et al. 2014). Loss of MLL by RNA interference (RNAi) gave rise to defects in S-phase and M-phase progression. While, after extensive mutational analysis, the regulation of the S-phase progression could be attributed to the activity of Transcriptional activation domain (TAD) of MLL (which acts independently of the SET domain), the domain responsible for influencing mitotic progression was not obvious.
Diverse functional roles of WRAD. The figure shows the model of WDR5, RbBP5, ASH2L, and DPY30 (WRAD) components participating in conventional (1) and non-conventional (2) functions. (1) The WRAD complex associates with MLL and methylates the promoters to regulate the transcription of genes during development. (2a-i, ii) Members of WRAD complex participate in cell-cycle regulation controlling S- and M-phase progression. (2a-iii) WDR5 subunit of WRAD associates with diverse interaction partners such as KIF4, MKLP10, and PRC1 at the midbody to regulate Cytokinesis; (2b) DPY30 component of WRAD interacts with BIG1 to modulate endosomal transport. WDR5, W: Red; RbBP5, R: Green; ASH2L, A: Blue, and DPY30, D: Orange.
Out of the several mitotic defects observed upon knockdown of MLL, two were quantified: (i) binuclei, arising from defective cytokinesis and (ii) micronuclei, marker of error-prone mitotic segregation (Ali et al. 2014). These mitotic defects were phenocopied when each subunit of WRAD was knocked down using RNAi individually, suggesting that the whole MLL ‘core complex’ was required. Further, we could demonstrate that the WDR5-MLL interaction was crucial for this function of MLL, as a recombinant MLL protein with point mutation in the Win motif was unable to rescue mitotic defects, observed upon the knockdown of endogenous MLL protein. However, further analysis highlighted differences from conventional MLL-WRAD activity, most notable being the non-requirement of MLL SET domain for this function. These observations led us to speculate if WRAD complex also enhances the transcriptional activity of MLL (in addition to its histone methyltransferase activity) using its transcriptional activation domain. We put this hypothesis to test by assessing the Win-motif mutant of MLL for its cell proliferative (S-phase) functions, which is dependent on the TAD domain. Even though, we observed that the mutation in Win motif had no effect on the cell proliferation functions of MLL, later experiments revealed that MLL may not require WRAD for its cell proliferation functions. Hence, we cannot say with certainty if WRAD will affect the TAD activity of MLL or not. But what is clear from our studies is that just the MLL-WRAD interaction is enough for mitotic progression, and neither the transcriptional activity (via the TAD) nor the methyltransferase activity (via the SET domain) of MLL plays any role here (Ali et al. 2014). Hence WRAD has a novel function with MLL.
Although this study highlights the emerging non-transcriptional, SET domain- and TAD-independent functions of MLL-WRAD, the underlying molecular mechanism is yet to be investigated. It is likely that WRAD may expand the interactome of MLL, fetching in new specific interaction partners for the proper regulation of mitosis. Alternatively, it may act as an HMT by itself, performing methylation of non-histone proteins required in mitosis. In any case, it will be exciting to find out the sub-cellular localizations, interactions, and most importantly, functions of MLL-WRAD complex during mitotic progression to discern the underlying mechanism of mitotic regulation by this complex.
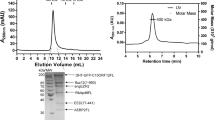
Recently, another study demonstrated that WDR5 localizes to the midbody and interacts with the midbody-localized proteins such kinesins (KIF4, and MKLP10) and PRC1 proteins, thereby participating in the proper progression of cytokinesis (Bailey et al. 2015). Although this finding could show that central arginine-binding cavity of WDR5 is crucial for its midbody localization, yet the proteins involved in the recruitment of WDR5 to the midbody were not explored further (Bailey et al. 2015). In addition, DPY-30 subunit of WRAD has been shown to localize to the trans-Golgi compartments and regulate the recycling of the endosomes (Xu et al. 2009). The same study demonstrated that the depletion of RAD (RbBP5, ASH2L, and DPY30) perturbed the recycling of the endosomes (Xu et al. 2009). Even though these studies have not tested all subunits of WRAD complex; it is very likely that all of them are involved. Supporting this hypothesis, our studies observed cytokinesis defects with all WRAD subunits, even though only WDR5 has been shown to localize to the midbody (Ali et al. 2014; Bailey et al. 2015). However, whether MLL is involved in all the above mentioned functions of WRAD is difficult to predict. Interestingly, in size exclusion chromatography of mammalian nuclear extracts, WRA complex in addition to eluting with MLL, also elutes as a distinct ∼150 kDa complex lacking MLL, indicating that it may exist as an independent entity in the cell (Steward et al. 2006). Therefore, there is a possibility that WRAD may function far beyond the previously studied nuclear HMT function.
4 Future directions
An ever-increasing body of literature has correlated the activity of WRAD with the SET1 family for the histone methyltransferase function. While all these functions of WRAD well connect with its propensity to bind to the gene promoters within the nucleus, recently WRAD complex subunits have been suggested to play roles beyond its nuclear functions. It will be interesting to observe if contribution of WRAD supersedes its thoroughly studied partners to regulate diverse cellular functions.
References
Ali A, Veeranki SN and Tyagi S 2014 A SET-domain-independent role of WRAD complex in cell-cycle regulatory function of mixed lineage leukemia. Nucleic Acids Res. 42 7611–7624
Ansari KI and Mandal SS 2010 Mixed lineage leukemia: roles in gene expression, hormone signaling and mRNA processing. FEBS J. 277 1790–1804
Bailey JK, Fields AT, Cheng K, Lee A, Wagenaar E, Lagrois R, Schmidt B, Xia B, et al. 2015 WD repeat-containing protein 5 (WDR5) localizes to the midbody and regulates abscission. J. Biol. Chem. 290 8987–9001
Bannister AJ and Kouzarides T 2011 Regulation of chromatin by histone modifications. Cell Res. 21 381–395
Briggs SD, Bryk M, Strahl BD, Cheung WL, Davie JK, Dent SYR, Winston F and David Allis C 2001 Histone H3 lysine 4 methylation is mediated by Set1 and required for cell growth and rDNA silencing in Saccharomyces cerevisiae. Genes Dev. 15 3286–3295
Cao F 2012 MLL/SET1 complex: from yeast to human. Fungal Genomics Biol. 2 1–2
Cho YW, Hong T, Hong SH, Guo H, Yu H, Kim D, Guszczynski T, Dressler GR, et al. 2007 PTIP associates with MLL3- and MLL4-containing histone H3 lysine 4 methyltransferase complex. J. Biol. Chem. 282 20395–20406
Cosgrove MS and Patel A 2010 Mixed lineage leukemia: a structure-function perspective of the MLL1 protein. FEBS J. 277 1832–1842
Dharmarajan V, Lee JH, Patel A, Skalnik DG and Cosgrove MS 2012 Structural basis for WDR5 interaction (Win) motif recognition in human SET1 family histone methyltransferases. J. Biol. Chem. 287 27275–27289
Dou Y, Milne TA, Ruthenburg AJ, Lee S, Lee JW, Verdine GL, Allis CD and Roeder RG 2006 Regulation of MLL1 H3K4 methyltransferase activity by its core components. Nat. Struct. Mol. Biol. 13 713–719
Eissenberg JC and Shilatifard A 2010 Histone H3 lysine 4 (H3K4) methylation in development and differentiation. Dev. Biol. 339 240–249
Ernst P and Vakoc CR 2012 WRAD: enabler of the SET1-family of H3K4 methyltransferases. Brief. Funct. Genomics. 11 217–226
Herz HM, Mohan M, Garruss AS, Liang K, Takahashi Y-h, Mickey K, Voets O, Verrijzer CP, et al. 2012 Enhancer-associated H3K4 monomethylation by trithorax-related, the drosophila homolog of mammalian MLL3/MLL4. Genes Dev. 26 2604–2620
Hu D, Gao X, Morgan MA, Herz H-M, Smith ER and Shilatifard A 2013 The MLL3/MLL4 branches of the COMPASS family function as major histone H3K4 monomethylases at enhancers. Mol. Cell. Biol. 33 4745–4754
Li Y, Han J, Zhang Y, Cao F, Liu Z, Li S, Wu J, Hu C, et al. 2016 Structural basis for activity regulation of MLL family methyltransferases. Nature 530 447–452
Martin C and Zhang Y 2005 The diverse functions of histone lysine methylation. Nat. Rev. Mol. Cell Biol. 6 838–849
Nagy PL, Griesenbeck J, Kornberg RD and Cleary ML 2002 A trithorax-group complex purified from Saccharomyces cerevisiae is required for methylation of histone H3. Proc. Natl. Acad. Sci. USA 99 90–94
Odho Z, Southall SM and Wilson JR 2010 Characterization of a novel WDR5-binding site that recruits RbBP5 through a conserved motif to enhance methylation of histone H3 lysine 4 by mixed lineage leukemia protein-1. J. Biol. Chem. 285 32967–32976
Patel SR, Kim D, Levitan I and Dressler GR 2007 The BRCT-domain containing protein PTIP links PAX2 to a histone H3, lysine 4 methyltransferase complex. Dev. Cell 13 580–592
Patel A, Dharmarajan V and Cosgrove MS 2008a Structure of WDR5 bound to mixed lineage leukemia protein-1 peptide. J. Biol. Chem. 283 32158–32161
Patel A, Vought VE, Dharmarajan V and Cosgrove MS 2008b A conserved arginine-containing motif crucial for the assembly and enzymatic activity of the mixed lineage leukemia protein-1 core complex. J. Biol. Chem. 283 32162–32175
Patel A, Dharmarajan V, Vought VE and Cosgrove MS 2009 On the mechanism of multiple lysine methylation by the human mixed lineage leukemia protein-1 (MLL1) core complex. J. Biol. Chem. 284 24242–24256
Patel A, Vought VE, Dharmarajan V and Cosgrove MS 2011 A novel non-SET domain multi-subunit methyltransferase required for sequential nucleosomal histone H3 methylation by the mixed lineage leukemia protein-1 (MLL1) core complex. J. Biol. Chem. 286 3359–3369
Patel A, Vought VE, Swatkoski S, Viggiano S, Howard B, Dharmarajan V, Monteith KE, Kupakuwana G, et al. 2014 Automethylation activities within the mixed lineage leukemia-1 (MLL1) core complex reveal evidence supporting a “two-active site” model for multiple histone H3 lysine 4 methylation. J. Biol. Chem. 289 868–884
Piunti A and Shilatifard A 2016 Epigenetic balance of gene expression by Polycomb and COMPASS families. Science 352 aad9780
Senisterra G, Wu H, Allali-Hassani A, Wasney GA, Barsyte-Lovejoy D, Dombrovski L, Dong A, Nguyen KT, et al. 2013 Small-molecule inhibition of MLL activity by disruption of its interaction with WDR5. Biochem. J. 449 151–159
Shilatifard A 2012 The COMPASS family of histone H3K4 methylases: mechanisms of regulation in development and disease pathogenesis. Annu. Rev. Biochem. 81 65–95
Shinsky SA and Cosgrove MS 2015 Unique role of the WD-40 repeat protein 5 (WDR5) subunit within the mixed lineage leukemia 3 (MLL3) histone methyltransferase complex. J. Biol. Chem. 290 25819–25833
Shinsky SA, Hu M, Vought VE, Ng SB, Bamshad MJ, Shendure J and Cosgrove MS 2014 A non-active-site SET domain surface crucial for the interaction of MLL1 and the RbBP5/Ash2L heterodimer within MLL family core complexes. J. Mol. Biol. 426 2283–2299
Smith E, Lin C and Shilatifard A 2011 The super elongation complex (SEC) and MLL in development and disease. Genes Dev. 25 661–672
Song JJ and Kingston RE 2008 WDR5 interacts with mixed lineage leukemia (MLL) protein via the histone H3-binding pocket. J. Biol. Chem. 283 35258–35264
Steward MM, Lee J, O’Donovan A, Wyatt M, Bernstein BE, Shilatifard A, Donovan AO, Wyatt M, et al. 2006 Molecular regulation of H3K4 trimethylation by ASH2L, a shared subunit of MLL complexes. Nat. Struct. Mol. Biol. 13 852–854
Takahashi Y-h, Westfield GH, Oleskie AN, Trievel RC, Shilatifard A and Skiniotis G 2011 Structural analysis of the core COMPASS family of histone H3K4 methylases from yeast to human. Proc. Natl. Acad. Sci. 108 20526–20531
van Nuland R, Smits AH, Pallaki P, Jansen PWTC, Vermeulen M and Timmers HTM 2013 Quantitative dissection and stoichiometry determination of the human SET1/MLL histone methyltransferase complexes. Mol. Cell. Biol. 33 2067–2077
Wu M, Wang PF, Lee JS, Martin-Brown S, Florens L, Washburn M and Shilatifard A 2008 Molecular regulation of H3K4 trimethylation by Wdr82, a component of human Set1/COMPASS. Mol. Cell. Biol. 28 7337–7344
Xu Z, Gong Q, Xia B, Groves B, Zimmermann M, Mugler C, Mu D, Matsumoto B, et al. 2009 A role of histone H3 lysine 4 methyltransferase components in endosomal trafficking. J. Cell Biol. 186 343–353
Yokoyama A, Wang Z, Wysocka J, Sanyal M, Aufiero DJ, Kitabayashi I, Herr W and Cleary ML 2004 Leukemia proto-oncoprotein MLL forms a SET1-like histone methyltransferase complex with menin to regulate Hox gene expression. Mol. Cell. Biol. 24 5639–5649
Zhang Y, Mittal A, Reid J, Reich S, Gamblin SJ and Wilson JR 2015 Evolving catalytic properties of the MLL family SET domain. Structure 23 1921–1933
Acknowledgements
We thank members of the Laboratory of Cell Cycle Regulation for valuable feedback. AA is the recipient of Junior and Senior Research Fellowships of the Council of Scientific and Industrial Research (CSIR), India, towards the pursuit of a Ph.D. degree of Manipal University. This work was supported in part by a grant from DBT (to ST; BT/BR15453/BRB/10/927/2011), DST (to ST; SB/SO/BB-069/2013) and CDFD core funds.
Author information
Authors and Affiliations
Corresponding author
Additional information
[Ali A and Tyagi S 2017 Diverse roles of WDR5-RbBP5-ASH2L-DPY30 (WRAD) complex in the functions of the SET1 histone methyltransferase family. J. Biosci.]
Graduate Studies, Manipal University, Manipal, India
Rights and permissions
About this article
Cite this article
Ali, A., Tyagi, S. Diverse roles of WDR5-RbBP5-ASH2L-DPY30 (WRAD) complex in the functions of the SET1 histone methyltransferase family. J Biosci 42, 155–159 (2017). https://doi.org/10.1007/s12038-017-9666-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12038-017-9666-9