Abstract
Huntington disease (HD) is the most common neurogenetic disorder caused by expansion of the CAG repeat in the HTT gene; nevertheless, the molecular bases of the disease are not fully understood. Non-coding RNAs have demonstrated to be involved in the physiopathology of HD. However, the role of circRNAs has not been investigated. The aim of this study was to identify the circRNAs with differential expression in a murine cell line model of HD and to identify the biological pathways regulated by the differentially expressed circRNAs. CircRNA expression was analyzed through a microarray, which specifically detects circular species of RNA. The expression patterns between a murine cell line expressing mutant Huntingtin and cells expressing wild-type Huntingtin were compared. We predicted the miRNAs with binding sites for the differentially expressed circRNAs and the corresponding target genes for those miRNAs. Using the target genes, we performed a function enrichment analysis. We identified 23 circRNAs differentially expressed, 19 downregulated and four upregulated. Most of the downregulated circRNAs derive from the Rere gene. The dopaminergic synapse, MAPK, and long-term depression pathways were significantly enriched. The three identified pathways have been previously associated with the physiopathology of HD. The understanding of the circRNA-miRNA-mRNA network involved in the molecular mechanisms driving HD can lead us to identify novel biomarkers and potential therapeutic targets. To the best of our knowledge, this is the first study analyzing circRNAs in a model of Huntington disease.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
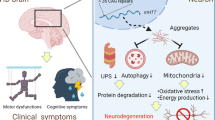
Huntington’s disease (HD) is one of the most common hereditary neurodegenerative disorders, reported prevalence per 100,000 habitants varies largely across populations, from 0.04 in Asia to 7.33 in North America [1]. Clinical manifestations of HD include motor, cognitive, and psychiatric signs and symptoms. The motor manifestations are the most evident, with chorea being the most characteristic. However, the cognitive and psychiatric manifestations, acquire great importance, since they involve important aspects of morbidity-mortality [2, 3].
HD is caused by mutation in the HTT gene, which encodes for the protein huntingtin. The molecular defect is an expansion of the CAG repeat, which encodes for glutamine (Q), in exon 1 of HTT [4]. Alleles possessing 26 or less repeats are considered normal, from 27–35 repeats, there is a risk of transmitting the disease secondary to expansion through parental meiosis, from 36–39 repeats, incomplete penetrance has been observed and above 40 repeats, alleles are considered fully penetrant for causing HD. There is an inverse correlation between the number of repeats and the age of onset of the symptoms [5].
Polyglutamine (polyQ) tracts are proposed to affect cell function through different mechanisms. Dysfunction of the proteasome system, mitochondrial metabolism, RNA processing, transport of vesicles, and neurotransmitters through the cytoskeleton are among the altered pathways [6]. Although the CAG expansion is the cause of HD, it is believed that there are other genetic and environmental factors contributing to the phenotypic variation between affected individuals [5]. Recent evidence has highlighted that not only the presence of determined genetic variants can modify the HD phenotype but also the mechanisms controlling gene expression can contribute [7].
Non-coding RNAs (ncRNAs) have emerged as key regulators of gene expression in virtually all biological processes [8]. Circular RNAs (circRNAs) are a relatively new class of ncRNA which play an important role in the regulation of gene expression through different mechanisms, one of the best characterized is by acting as microRNA (miRNA) “sponges” [9]. CircRNAs are covalently closed molecules; they are originated from a precursor mRNA (pre-mRNA) through a back-splicing mechanism [10]. This structure gives them the advantage for resisting degradation by most RNAses, allowing them to be present in the cell for long-time periods [11].
Although the presence of circRNA has been demonstrated in different types of tissue, recent studies have characterized the importance of these molecules in the central nervous system (CNS) [12]. Furthermore, studies in mammalians’ brains have observed distinctive expression of circRNAs according to the area and cell type involved, suggesting a specific area and cell-dependent patterns of regulation [13].
CircRNAs have been studied in some neurodegenerative diseases, even when they are not the direct cause of the disease; their differential expression suggests involvement in the phenotype [14]. Parsing the different mechanisms in which circRNAs are involved can lead to the identification of new therapeutic targets [15]. In addition, circRNAs can be secreted within microvesicles or exosomes into the bloodstream and they are more stable than linear RNA; therefore, they have been proposed as potential minimally invasive biomarkers of neuronal diseases [16].
To the best of our knowledge, these molecules have not been fully characterized in HD. The purpose of this study was to investigate the presence of circRNAs with differential expression in a validated HD murine cell model and identify the miRNAs, target genes, and biological pathways under the regulation of the differentially expressed circRNAs.
Methods
Cell Lines and Culture
PC12 cell line (RRID:CVCL_0481), derived from rat phaeochromocytoma, expressing Green Fluorescence Protein (GFP)-tagged exon 1 of the Huntingtin gene driven by a doxycycline-dependent Tet-On promoter, was a kind gift from David Rubinsztein from the Cambridge Institute of Medical Research. Modified cells express either a 23 or 74 polyglutamine repeats (PC12 HD-Q23 or PC12 HD-Q74) [17]. Cells were plated in 100-mm Petri dishes with high glucose Dulbecco’s modified Eagle’s medium containing L-glutamine (DMEM 12100, Gibco Life Technologies). Medium was supplemented with 2 nM L-glutamine, hygromycin (Sigma-Aldrich) 75 g/mL, 100 U/mL penicillin/streptomycin (Gibco Life Technologies), G418 100 g/Ml (Sigma-Aldrich), 10% heat-inactivated horse serum (HS) (Biowest), and 5% Tet-approved fetal bovine serum (FBS) (Clontech), and cells were incubated at 37 °C and 5% CO2. Expression of the fusion protein was induced by adding doxycycline 1 g/mL to the culture medium. For subcultures, cells were harvested when they were 80% confluent and counted using a Neubauer chamber. Subcultures were prepared using fresh medium supplemented with or without doxycycline. Expression of the fusion protein was corroborated by fluorescence microscopy using a ZEISS Axio Observer microscope 4 days after induction.
Growth Curves
Growth curves were generated to investigate how the presence of the protein containing the Q74 tract could affect cells’ growth. Cells were plated in 100-mm Petri dishes with 10 mL of DMEM media supplemented as described above. When the cells were confluent, they were harvested and transferred to a new Petri dish with 10 mL of medium until confluence was reached again. To harvest the cells, the media was removed by aspiration, the cells were washed with 2 mL of 1× phosphate-buffered saline (PBS) and then incubated in 2ml of 1× Trypsin (Gibco Life Tecnologies) for 5 min at 37°C. After incubation in trypsin solution, cells were detached by agitation. The suspension of cells was transferred to a 15-mL tube with 8ml of fresh media and mixed for counting using a Neubauer chamber. Population doublings were calculated using the formula PDL= (log10 cell count at harvest – log10 cell count at inoculation)/0.301). The value 0.301 is a constant for the conversion factor from the log10 to the natural log (log2).
Total RNA Isolation and Microarray Analysis
A proportion of the harvested cells was not plated and after trypsin detachment they were transferred to a 15-mL tube with 8ml of fresh media, mixed and centrifuged at 1100rpm for 8 min. The media was removed and the pellet washed with 10 mL of 1× PBS and centrifuged 8 min at 1100 rpm. The PBS was removed and the pellets frozen in dry ice for storage at −80°C. Total RNA from frozen cell pellets was isolated using TRIzol® Reagent (Thermo Fisher Scientific), according to manufacturer’s recommendations. Total RNA from each sample was quantified using the NanoDrop ND-1000 (Thermo Fisher Scientific). In addition, RNA Integrity and gDNA contamination were tested by Denaturing Agarose Gel Electrophoresis.
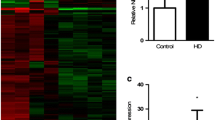
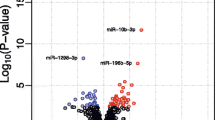
The sample preparation and microarray hybridization were performed based on the Arraystar’s standard protocols. Briefly, total RNA was digested with Rnase R (Epicentre, Inc.) to remove linear RNA and enrich circular RNA. Enriched circular RNAs were amplified and transcribed into fluorescent cRNA utilizing a random priming method (Arraystar Super RNA Labeling Kit; Arraystar). The labeled cRNAs were hybridized onto the Arraystar Rat circRNA Array (8×15K, Arraystar). After having washed the slides, the arrays were scanned by the Agilent Scanner G2505C. Agilent Feature Extraction software (version 11.0.1.1) was used to analyze acquired array images. Quantile normalization and subsequent data processing were performed using the R software limma package. Differentially expressed circRNAs with statistical significance between the two groups were identified through Volcano Plot filtering. Differentially expressed circRNAs between two samples were identified through fold change filtering. Hierarchical clustering was performed to show the distinguishable circRNAs expression pattern among samples.
Annotation for circRNA-miRNA-mRNA Interaction
One of the known functions of circRNAs is their ability to act as miRNA sponges, making them important factors for controlling gene expression. The circRNA-miRNA interaction was predicted with Arraystar’s home-made miRNA target prediction software based on TargetScan and miRanda. The best five interacting miRNAs, according to 8mer and 7mer base pairing between the miRNA seed and the circRNA sequence, were considered for further analysis. The target genes of the selected miRNAs were predicted using the R software miRNAtap package. miRNAtap retrieves predicted targets from the five most commonly cited prediction algorithms: DIANA, miRanda, PicTar, TargetScan, and miRDB. We considered only those target genes predicted by at least four algorithms.
In Silico Enrichment and Interaction Analysis
The target genes, regulated by the miRNAs able to interact with the differentially expressed circRNAs, were used as input for enrichment analysis using WebGestalt (WEB-based Gene SeT AnaLysis Toolkit) [18]. For WebGestalt analysis, we selected Rattus novergicus as the organism of interest, over-representation analysis (ORA) as the method of interest and the Kyoto Encyclopedia of Genes and Genomes (KEGG) as the functional database. Interaction networks between circRNA-miRNAs and miRNAs-mRNAs from the relevant enriched pathways identified with WebGestalt were built with the Cytoscape software [19].
Results
Expression of Huntingtin Fragment Containing Q74 Affects Cell Growth
Frozen cells were cultured in 100-mm Petri dishes with supplemented DMEM media, when cultures were 80% confluent cells were counted and subcultured at a density of 5×105 cells per Petri dish. After 10 days in culture, doxycycline (1 g/mL) was added to the PC12 HD-Q23 and PC12 HD-Q74 cells. Doxycycline was continuously added to every subculture to maintain the expression of the fusion protein containing the polyQ tract. Expression of the protein was corroborated by observation of GFP foci in the presence of doxycycline under fluorescence microscopy (data not shown). Cells were grown for 62 days in the presence of doxycycline and population doublings were calculated after every subculture as stated in the methods section. After 23 days in culture, which corresponds to 13 days under the induction of the fusion protein by doxycycline, there was an important impairment of the growth of the PC12 HD-Q74 cells (Fig. 1). This result suggests that the presence of the protein harboring a tract of 74 glutamine residues impairs cell growth.
Differential Expression of circRNAs
To identify if there were circRNAs with differential expression between the PC12 HD-Q23 and PC12 HD-Q74 cells, we used the Arraystar Rat circRNA array. This microarray is the only one with validated probes for detection of the back-splicing junction present only in circular transcripts derived from pre-mRNA processing. Expression analysis was carried out in three biological replicates from each group of cells. We used RNA treated with RNAse R for enrichment of circRNA, derived from cells exposed to doxycycline during 4 weeks. At this time point, the reduction of the growth velocity of the PC12 HD-Q74 cells started to be evident. Differential expression analysis identified four upregulated and 19 downregulated circRNAs in the PC12 HD-Q74 cells (Table 1 and Supp. Figure 1, for a more detailed description of chromosome position see Supp. table 1). The four upregulated circRNAs were originated from different chromosomal regions, while 16 of the 19 downregulated circRNAs came from the same chromosome region where the Rere gene is located. The remaining three were derived from different chromosome positions.
Unsupervised hierarchical clustering considering the 23 circRNAs with differential expression grouped together the triplicates from each condition (Supp. Figure 2). The downregulated transcripts derived from the Rere gene allow a clear distinction between both groups of cells.
circRNA-miRNA-mRNA Interaction Networks
One of the best studied functions of circRNAs is their role as miRNA “sponges” through base pairing with the seed sequence of miRNAs. We attempt to identify the five most relevant miRNAs interacting with the differentially expressed circRNAs. The miRNAs were selected based on the degree of base pairing with the seed sequence according to Arraystar’s home-made miRNA target prediction software based on TargetScan and miRanda. Table 2 displays the top five miRNAs for each circRNA, the first four rows (in italics), represent the four upregulated circRNAs. The predicted miRNAs interacting with the upregulated circRNAs were unique for each circRNA, i.e., none of the 20 recognized miRNAs is predicted to interact with more than one circRNA. On the other hand, from the 90 miRNAs interacting with the downregulated circRNAs 30 were predicted to interact with more than one circRNA; therefore, the number of different miRNAs is 60. Only the miRNA rno-miR-3568 was found to interact with one upregulated and one downregulated circRNA.
Changes in the expression of circRNA will ultimately have an impact on mRNA half-life through miRNA sequestering. Therefore, the potential target genes for each miRNA were predicted using the R software miRNAtap package. miRNAtap collects information from five of the most used algorithms: DIANA, miRanda, PicTar, TargetScan, and miRDB. Only those target genes predicted by at least four algorithms were considered for further analysis. For some of the miRNAs, no target genes were retrieved. The predicted target genes were used as input for function enrichment analysis using the WebGestalt (WEB-based Gene SeT AnaLysis Toolkit) database. Table 3 depicts the top ten enriched pathways according to KEGG through the ORA carried out in WebGestalt. Furthermore, we investigated the distribution of the target genes among Gene Ontology categories (Supp. Figure 3).
The two more statistically significant functions were the MAPK signaling pathway which is involved in cell proliferation and the dopaminergic synapse pathway, which is involved in neurological processes. Therefore, we decided to analyze these two pathways in more detail. From the 23 differentially expressed circRNAs, 15 were related to the MAPK signaling pathway, while 16 to the dopaminergic synapse pathway (Supp. table 2). All the circRNAs related to the MAPK signaling pathway were also present in the dopaminergic synapse pathway, except for the rno_circRNA_013032 which was only present in the dopaminergic synapse pathway. To visualize the interactions between molecules, we built networks with the circRNAs involved in the regulation of the miRNAs and their respective target genes belonging to both pathways. The networks display the circRNA-miRNA and the circRNA-mRNA interactions within each pathway (Supplementary figure 4). In addition, we also built a network including both, the MAPK signaling pathway and dopaminergic synapse pathway (Fig. 2).
The networks allowed us to recognize that the circRNAs with more interactions are those derived from the Rere gene. In addition, it can be noticed that the miRNAs, rno-miR-322-5p, rno-miR-16-5p, and rno-miR-15b-5p have the higher number of interactions. Twenty target genes are represented in Fig. 2b; from these, 11 are exclusive of the MAPK signaling pathway, six to the dopaminergic synapse pathway, and three are common to both pathways.
It is worth to mention that the tenth enriched function was the long-term depression pathway, which involves the response to glutamate. Supplementary table 2 shows the differentially expressed circRNAs related to the long-term depression pathway. Within the set of target genes employed for the enrichment analysis, five belong to this pathway. These five target genes are Itpr1, Gria3, Map2k1, Ppp2r1a, and Gnai3, which are already represented in Fig. 2 because they are also involved in either the MAPK signaling pathway, or the dopaminergic synapse pathway.
Discussion
Huntington disease is one of the most devastating and incapacitating neurogenetic disorders; despite the characterization of the molecular cause, the underlying molecular mechanism leading to neuronal death of specific regions of the brain needs to be unraveled. Herein, we analyze the expression of circRNAs in a murine cell model expressing mutant huntingtin. Before the differential expression analysis, we observed a significant cell growth compromise on the cells expressing the mutant huntingtin (mHtt). Afterwards, we found 23 circRNAs with differential expression between cells expressing normal versus mHtt; from them, 19 were downregulated and four upregulated. Interestingly, most of the downregulated circRNAs derived from the gene Rere (arginine (R) glutamic acid (E) repeat encoding), which belongs to the family of atrophins along with atrophin 1. The human gene RERE shares 92% homology with the rat gene. RERE protein contains two RE domains; previous experiments have demonstrated that it can localize at the cell nucleus [20]. RERE seems to act as a transcriptional co-repressor, probably through interaction with the histone deacetylase 1 (HDAC1) and, in conjunction with atrophin 1, also plays a key role in neuronal development and function [21]. Mutations in RERE can cause a disease affecting brain development (neurodevelopmental disorder with or without anomalies of the brain, eye, or heart, MIM 616975) [22]. In addition, RERE can physically interact with proteins carrying an expanded polyglutamine tract [20] through the proximal RE repeats. Dentatorubral-pallidoluysian atrophy is caused by an expansion of a CAG microsatellite with full penetrance beyond 48 repeats in atrophin 1 gene [23]. It has been demonstrated that RERE interacts with the expanded polyglutamine tract of atrophin 1, contributing to the formation of protein aggregates. This finding opens the possibility of a physical interaction between RERE and mHTT with consequent participation in aggregates formation or sequestration of RERE.
We identified the best five target-miRNAs with the capacity to interact with the differentially expressed circRNAs. Afterwards, the target genes under the regulation of those miRNAs were recognized. Enrichment analysis, using these target genes as input, showed the MAPK signaling pathway and the dopaminergic synapse as the main enriched pathways.
The role of dopamine in the physiopathology of HD has been extensively investigated. Striatal levels of dopamine and its metabolites have shown a biphasic behavior in HD patients and mouse models. The general trend is elevation of dopamine levels during initial stages with further reduction at advanced stages, whereas the expression of its receptors is reduced [24], especially subtypes D1 and D2 [25]. D1 receptor is encoded by the Drd1 gene; according to our analysis its transcript can be targeted by miR-15a-5p and miR-15b-5p, we found four downregulated circRNAs possessing MRE for these two miRNAs. Based on the ceRNA theory, the final consequence of circRNAs downregulation would be a reduced expression of the mRNA of the target gene. On this scenario, this circRNA-miRNA-mRNA axis could contribute to the reduction of the D1 receptor observed in HD. In addition, it has been demonstrated that D1 receptor, but not D2 receptor, is involved in the selectively cell death at the striatal in HD [26]. To explain how the interaction between the circRNAs and the components of the dopaminergic synapse pathway influence these findings further investigation is required.
Interestingly, besides their role on D1 receptor regulation, miR-15a-5p and miR-15b-5p have previously been involved in the physiopathology of neurodegenerative diseases through different mechanisms. Overexpression of miR-15a-5p has been observed in the cerebrospinal fluid (CSF) of Alzheimer’s disease patients [27]. Besides, experimental data support that overexpression of miR-15b-5p promotes apoptosis through regulation of Akt3 in Parkinson’s disease patients [28].
MAPK signaling pathway also showed significant enrichment. The mitogen-activated protein kinases (MAPKs) consist of a group of highly conserved proteins which respond to extracellular factors and coordinate cellular processes such as proliferation, response to stress, differentiation, and apoptosis. There are four main signaling cascades involved in the response, c-Jun N-terminal kinase (JNK), extracellular signal-regulated kinase 1 and 2 (ERK1/2), p38, and EKR5 [29]. On the other hand, dual-specific phosphatases (DUSPs, also known as mammalian dual-specificity MAPK phosphatases (MKPs)) carry out MAPKs inactivation [30]. Previous experiments have demonstrated downregulation of MKP1 in animal models and patients with HD. Overexpression of MKP1 efficiently prevents activation of JNKs and p38 having a neuroprotective effect [31]. These experiments revealed that the JNK and p38 are the main MAPK pathways involved in HD pathogenesis. A previous study, also using the PC12 HD cell model, observed increased cell dysfunction and death after activation of the JNK pathway. The same study showed that activation of the ERK pathway promoted cell survival, acting as a protective factor. The authors propose that mHtt can trigger both pathways and the cell’s fate depend on the imbalance between them [32]. The target genes under the regulation of the circRNAs with differential expression observed in our study, participate preferentially in these two signaling cascades. For example, Dusp3, which is also targeted by miR-15a-5p and miR-15b-5p, can inhibit the JNK signaling cascade [33]. Based on the ceRNA theory, the under-expression of circRNAs with MRE for the above-mentioned miRNAs will result in downregulation of Dusp3. A recent review summarizes the use of selective inhibitors of different MAPK signaling pathway components on neurodegenerative diseases [34]. Interestingly, one of the promising resources is a drug containing the JDB motif from the JNK-interacting protein 1 (JIP1). JIP1 is a scaffold protein in the JNK signaling cascade that acts as a strong inhibitor over JNKs. Our analysis predicted an interaction between circRNA_32718-miR-24-3p-JIP1 resulting in downregulation of JIP1. Even when these results need further validation, our findings open a new level for designing therapeutic strategies within this pathway.
Among the enriched pathways we also found the long-term depression (LTD), LTD represents one of the mechanisms involved in neuronal plasticity along with long-term potentiation (LTP). The synergistic interaction between LTD and LTP allows changes at the dendrite spines synapses related to learning and memory [35]. Postmortem analyses on brains of HD patients have demonstrated changes in the number and morphology of dendritic spines [36]. These changes are closely related to the cognitive symptoms observed in HD patients [37]. Even when this pathway was the tenth most relevant, it has the highest enrichment ratio. It is worth to mention that most of the genes from this pathway also play a role either in the dopaminergic synapse or MAPK pathways. These findings suggest a partially shared control of these pathways through circRNA-miRNA networks.
In conclusion, we present the first analysis of expression of circRNA in a validated murine cellular model of HD. Our data support participation on the regulation of the dopaminergic synapse, MAPK and LTD pathways, which have been previously involved in HD physiopathology. This study has some limitations. Our expression results need to be validated in other models of HD (cell lines or mouse models), experiments are required to prove the circRNA-miRNA interaction and the expression of target genes needs to be analyzed. Nevertheless, the main strength is the exploration of a different level of gene regulation in a HD model. This new knowledge could contribute for unraveling the molecular mechanisms underlying HD and designing novel therapeutic strategies.
Data availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Rawlins MD, Wexler NS, Wexler AR, Tabrizi SJ, Douglas I, Evans SJ, Smeeth L (2016) The prevalence of Huntington’s disease. Neuroepidemiology. 46:144–153
Tabrizi SJ, Leavitt BR, Landwehrmeyer GB, Wild EJ, Saft C, Barker RA, Blair NF, Craufurd D, Priller J, Rickards H, Rosser A, Kordasiewicz HB, Czech C, Swayze EE, Norris DA, Baumann T, Gerlach I, Schobel SA, Paz E et al (2019) Targeting Huntingtin expression in patients with Huntington’s disease. N Engl J Med 380:2307–2316
Ross CA, Aylward EH, Wild EJ, Langbehn DR, Long JD, Warner JH, Scahill RI, Leavitt BR, Stout JC, Paulsen JS, Reilmann R, Unschuld PG, Wexler A, Margolis RL, Tabrizi SJ (2014) Huntington disease: natural history, biomarkers and prospects for therapeutics. Nat Rev Neurol 10:204–216
Saudou F, Humbert S (2016) The biology of huntingtin. Neuron. 89:910–926
Langbehn DR, Brinkman RR, Falush D, Paulsen JS, Hayden MR, International Huntington’s Disease Collaborative Group (2004) A new model for prediction of the age of onset and penetrance for Huntington’s disease based on CAG length. Clin Genet 65:267–277
Hatters DM (2008) Protein misfolding inside cells: the case of huntingtin and Huntington’s disease. IUBMB Life 60:724–728
Gusella JF, MacDonald ME (2009) Huntington’s disease: the case for genetic modifiers. Genome Med 1:80
Sharp PA (2009) The centrality of RNA. Cell. 136:577–580
Hsiao KY, Sun HS, Tsai SJ (2017) Circular RNA - new member of noncoding RNA with novel functions. Exp Biol Med (Maywood) 242:1136–1141
Zhang XO, Wang HB, Zhang Y, Lu X, Chen LL, Yang L (2014) Complementary sequence-mediated exon circularization. Cell. 159:134–147
Li Y, Zheng Q, Bao C, Li S, Guo W, Zhao J, Chen D, Gu J, He X, Huang S (2015) Circular RNA is enriched and stable in exosomes: a promising biomarker for cancer diagnosis. Cell Res 25:981–984
Maass PG, Glažar P, Memczak S, Dittmar G, Hollfinger I, Schreyer L, Sauer AV, Toka O, Aiuti A, Luft FC, Rajewsky N (2017) A map of human circular RNAs in clinically relevant tissues. J Mol Med (Berl) 95:1179–1189
Rybak-Wolf A, Stottmeister C, Glažar P, Jens M, Pino N, Giusti S, Hanan M, Behm M, Bartok O, Ashwal-Fluss R, Herzog M, Schreyer L, Papavasileiou P, Ivanov A, Öhman M, Refojo D, Kadener S, Rajewsky N (2015) Circular RNAs in the Mammalian brain are highly abundant, conserved, and dynamically expressed. Mol Cell 58:870–885
D'Ambra E, Capauto D, Morlando M (2019) Exploring the regulatory role of circular RNAs in neurodegenerative disorders. Int J Mol Sci 20:5477. https://doi.org/10.3390/ijms20215477
Holdt LM, Kohlmaier A, Teupser D (2018) Circular RNAs as therapeutic agents and targets. Front Physiol 9:1262
Scahill RI, Wild EJ, Tabrizi SJ (2012) Biomarkers for Huntington’s disease: an update. Expert Opin Med Diagn 6:371–375
Wyttenbach A, Swartz J, Kita H, Thykjaer T, Carmichael J, Bradley J, Brown R, Maxwell M, Schapira A, Orntoft TF, Kato K, Rubinsztein DC (2001) Polyglutamine expansions cause decreased CRE-mediated transcription and early gene expression changes prior to cell death in an inducible cell model of Huntington’s disease. Hum Mol Genet 10:1829–1845
Zhang B, Kirov S, Snoddy J (2005) WebGestalt: an integrated system for exploring gene sets in various biological contexts. Nucleic Acids Res 33:W741–W748
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13:2498–2504
Yanagisawa H, Bundo M, Miyashita T, Okamura-Oho Y, Tadokoro K, Tokunaga K, Yamada M (2000) Protein binding of a DRPLA family through arginine-glutamic acid dipeptide repeats is enhanced by extended polyglutamine. Hum Mol Genet 9:1433–1442
Zoltewicz JS, Stewart NJ, Leung R, Peterson AS (2004) Atrophin 2 recruits histone deacetylase and is required for the function of multiple signaling centers during mouse embryogenesis. Development. 131:3–14
Fregeau B, Kim BJ, Hernández-García A, Jordan VK, Cho MT, Schnur RE, Monaghan KG, Juusola J, Rosenfeld JA, Bhoj E, Zackai EH, Sacharow S, Barañano K, Bosch DGM, de Vries BBA, Lindstrom K, Schroeder A, James P, Kulch P et al (2016) De novo mutations of RERE Cause a genetic syndrome with features that overlap those associated with proximal 1p36 deletions. Am J Hum Genet 98:963–970
Nagafuchi S, Yanagisawa H, Sato K, Shirayama T, Ohsaki E, Bundo M, Takeda T, Tadokoro K, Kondo I, Murayama N (1994) Dentatorubral and pallidoluysian atrophy expansion of an unstable CAG trinucleotide on chromosome 12p. Nat Genet 6:14–18
Koch ET, Raymond LA (2019) Dysfunctional striatal dopamine signaling in Huntington’s disease. J Neurosci Res 97:1636–1654
Gamez J, Lorenzo-Bosquet C, Cuberas-Borrós G, Carmona F, Hernández-Vara J, Castilló J, Castell-Conesa J (2010) Does reduced [(123)I]-FP-CIT binding in Huntington’s disease suggest pre-synaptic dopaminergic involvement? Clin Neurol Neurosurg 112:870–875
Paoletti P, Vila I, Rifé M, Lizcano JM, Alberch J, Ginés S (2008) Dopaminergic and glutamatergic signaling crosstalk in Huntington’s disease neurodegeneration: the role of p25/cyclin-dependent kinase 5. J Neurosci 28:10090–10101
Sørensen SS, Nygaard AB, Christensen T (2016) miRNA expression profiles in cerebrospinal fluid and blood of patients with Alzheimer’s disease and other types of dementia - an exploratory study. Transl Neurodegener 5:6-016-0053-5 eCollection 2016
Zhu J, Xu X, Liang Y, Zhu R (2021) Downregulation of microRNA-15b-5p Targeting the Akt3-Mediated GSK-3β/β-catenin signaling pathway inhibits cell apoptosis in Parkinson’s disease. Biomed Res Int 2021:8814862
Keshet Y, Seger R (2010) The MAP kinase signaling cascades: a system of hundreds of components regulates a diverse array of physiological functions. Methods Mol Biol 661:3–38
Seternes OM, Kidger AM, Keyse SM (2019) Dual-specificity MAP kinase phosphatases in health and disease. Biochim Biophys Acta, Mol Cell Res 1866:124–143
Taylor DM, Moser R, Régulier E, Breuillaud L, Dixon M, Beesen AA, Elliston L, Silva Santos Mde F, Kim J, Jones L, Goldstein DR, Ferrante RJ, Luthi-Carter R (2013) MAP kinase phosphatase 1 (MKP-1/DUSP1) is neuroprotective in Huntington’s disease via additive effects of JNK and p38 inhibition. J Neurosci 33:2313–2325
Apostol BL, Illes K, Pallos J, Bodai L, Wu J, Strand A, Schweitzer ES, Olson JM, Kazantsev A, Marsh JL, Thompson LM (2006) Mutant huntingtin alters MAPK signaling pathways in PC12 and striatal cells: ERK1/2 protects against mutant huntingtin-associated toxicity. Hum Mol Genet 15:273–285
Ha J, Kang E, Seo J, Cho S (2019) Phosphorylation dynamics of JNK Signaling: effects of dual-specificity phosphatases (DUSPs) on the JNK pathway. Int J Mol Sci 20:6157. https://doi.org/10.3390/ijms20246157
Ahmed T, Zulfiqar A, Arguelles S, Rasekhian M, Nabavi SF, Silva AS, Nabavi SM (2020) Map kinase signaling as therapeutic target for neurodegeneration. Pharmacol Res 160:105090
Nithianantharajah J, Hannan AJ (2013) Dysregulation of synaptic proteins, dendritic spine abnormalities and pathological plasticity of synapses as experience-dependent mediators of cognitive and psychiatric symptoms in Huntington’s disease. Neuroscience. 251:66–74
Ferrante RJ, Kowall NW, Richardson EP Jr (1991) Proliferative and degenerative changes in striatal spiny neurons in Huntington’s disease: a combined study using the section-Golgi method and calbindin D28k immunocytochemistry. J Neurosci 11:3877–3887
Raymond LA, André VM, Cepeda C, Gladding CM, Milnerwood AJ, Levine MS (2011) Pathophysiology of Huntington’s disease: time-dependent alterations in synaptic and receptor function. Neuroscience. 198:252–273
Acknowledgments
The authors acknowledge to the Consejo Nacional de Ciencia y Tecnologúa (CONACYT) and to David Rubinsztein from the Cambridge Institute of Medical Research for kindly donate the cell line.
Funding
This research was funded by Consejo Nacional de Ciencia y Tecnologúa (CONACYT) grant number FOSISS-273213
Author information
Authors and Affiliations
Contributions
Research project conception: Alejandra Camacho-Molina, Alberto Hidalgo-Bravo. Execution of experimental procedures: Ernesto Marfil-Marin, Mónica Santamaría-Olmedo, Adriana PerezGrovas-Saltijeral. Data analysis: Margarita Valdes-Flores, Adriana Ochoa-Morales, Aurelio Jara-Prado, Rosalba Sevilla-Montoya, Alejandra Camacho-Molina, Alberto Hidalgo-Bravo. Interpretation of results: Adriana Ochoa-Morales, Aurelio Jara-Prado, Santamaría-Olmedo, Adriana PerezGrovas-Saltijeral, Margarita Valdes-Flores. Draft of the manuscript: Ernesto Marfil-Marin, Rosalba Sevilla-Montoya, Alejandra Camacho-Molina, Alberto Hidalgo-Bravo. All authors reviewed and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Conflicts of interest
The authors declare no competing interests.
Ethics approval
This article employed a modified cell line derived from the organism Rattus norvegicus obtained from the University of Cambridge. No human material was used in this study. This work does not qualify for ethics approval according to the Institutional Ethics Committees from the participant institutions.
Cell lines
A cell line derived from the organism Rattus norvegicus (RRID:CVCL_0481) was obtained from David Rubinsztein from the Cambridge Institute of Medical Research.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
About this article
Cite this article
Marfil-Marin, E., Santamaría-Olmedo, M., PerezGrovas-Saltijeral, A. et al. circRNA Regulates Dopaminergic Synapse, MAPK, and Long-term Depression Pathways in Huntington Disease. Mol Neurobiol 58, 6222–6231 (2021). https://doi.org/10.1007/s12035-021-02536-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-021-02536-1