Abstract
In the development of diabetic retinopathy, retinal mitochondria are dysfunctional, and mitochondrial DNA (mtDNA) is damaged with increased base mismatches and hypermethylated cytosines. DNA methylation is also a potential source of mutation, and in diabetes, the noncoding region, the displacement loop (D-loop), experiences more methylation and base mismatches than other regions of the mtDNA. Our aim was to investigate a possible crosstalk between mtDNA methylation and base mismatches in the development of diabetic retinopathy. The effect of inhibition of Dnmts (by 5-aza-2′-deoxycytidine or Dnmt1-siRNA) on glucose-induced mtDNA base mismatches was investigated in human retinal endothelial cells by surveyor endonuclease digestion and validated by Sanger sequencing. The role of deamination factors on increased base mismatches was determined in the cells genetically modulated for mitochondrial superoxide dismutase (Sod2) or cytidine-deaminase (APOBEC3A). The results were confirmed in an in vivo model using retinal microvasculature from diabetic mice overexpressing Sod2. Inhibition of DNA methylation, or regulation of cytosine deamination, significantly inhibited an increase in base mismatches at the D-loop and prevented mitochondrial dysfunction. Overexpression of Sod2 in mice also prevented diabetes-induced D-loop hypermethylation and increase in base mismatches. The crosstalk between DNA methylation and base mismatches continued even after termination of hyperglycemia, suggesting its role in the metabolic memory phenomenon associated with the progression of diabetic retinopathy. Inhibition of DNA methylation limits the availability of methylated cytosine for deamination, suggesting a crosstalk between DNA methylation and base mismatches. Thus, regulation of DNA methylation, or its deamination, should impede the development of diabetic retinopathy by preventing formation of base mismatches and mitochondrial dysfunction.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The pathogenesis of diabetic retinopathy is highly complex and involves multiple interlinked metabolic pathways [1, 2]. Retinal mitochondria become dysfunctional leading to capillary cell apoptosis, a phenomenon which precedes the development of histopathology characteristic of diabetic retinopathy [3,4,5]. Mitochondrion is equipped with its own circular DNA (mtDNA), which encodes 13 essential proteins required for proper functioning of the electron transport chain (ETC) system. Mitochondrial DNA also contains a noncoding regulatory region, the displacement loop (D-loop), which is essential for its transcription and replication [6]. In diabetes, retinal mtDNA is damaged, and the damage is more extensive in the noncoding D-loop compared to the other coding regions of the mtDNA [7].
Recent cutting-edge research has demonstrated that diabetes induces many epigenetic modifications, including histone modifications and DNA methylation [8, 9]. The enzymes responsible for cytosine methylation, DNA methyltransferases (Dnmts), are activated in the retina, and the D-loop is hypermethylated with elevated levels of methylated cytosine (5mC). Due to damaged mtDNA, impaired transcription of mtDNA encoded NADH dehydrogenase 1 and 6 (complex I) and cytochrome b (complex III) dysfunctions the ETC system, further fueling into a self-perpetuating cycle of superoxide accumulation [10, 11].
Cytosine, however, can also undergo spontaneous deamination to form uracil [12], and 5mC can further be converted to thymine, generating base mismatch or mutations in the DNA [13]; several mtDNA mutations have been reported in many ocular diseases including pathologies associated with optic nerve, retina, and eyelids [14]. Moreover, compared to cytosine, mutation rate of 5mC is 10–50 times higher [15, 16]. Deamination of 5mC is modulated by oxidative/nitrative stress; for example, esophageal adenocarcinoma patients with higher levels of nitrotyrosine have more frequent tumors with p53 mutations at CpG dinucleotides compared to the patients with non-CpG p53 mutations [17]. Cytidine deaminases, AID/APOBECs, also catalyze 5mC deamination to thymine [18, 19], and thymine-guanine mismatch-specific glycosylases remove them. Our previous work has shown that in diabetes, retinal mtDNA has increased the number of base mismatches, especially in its D-loop, decreased mitochondrial localization of DNA glycosylase, and elevated levels of nitrotyrosine [20, 21]. Furthermore, inhibition of nitrotyrosine accumulation is shown to protect mitochondrial dysfunction and capillary cell apoptosis [22]. However, whether diabetes facilitates any crosstalk between mtDNA methylation and base mismatch formation is not clear.
Persistent high glucose is detrimental in disease pathogenesis, and the molecular events established during the early days of hyperglycemia could not be erased even after the establishment of normal glycemia, suggesting an important role of molecular signatures in metabolic memory phenomenon associated with the continued progression of diabetic complications including retinopathy [21, 23, 24]. We have shown that in diabetes, maintenance of normal glycemia, after a period of hyperglycemic insult, does not restore increase in mtDNA base mismatches [21], and mtDNA continues to be hypermethylated with sustained activation of the DNA methylation machinery [25]. Does the continued crosstalk between DNA methylation and base mismatch play any role in the metabolic memory, however, remains to be investigated.
This study aims to understand any crosstalk between mtDNA methylation and formation of base mismatches in the development, and in the continued progression, of diabetic retinopathy. Using human retinal endothelial cells (HRECs), we investigated the effect of inhibition of hyperglycemia-induced mtDNA methylation on base mismatch formation. The role of deaminases, and of mitochondrial oxidative stress, in increased formation of base mismatches was confirmed in the cells genetically modulated for APOBEC or mitochondrial superoxide dismutase (Sod2). The results were established in an in vivo model using retinal microvasculature from diabetic mice overexpressing Sod2.
Methods
Retinal Endothelial Cells
HRECs (Cell System, Kirkland, WA) were cultured in Dulbecco’s modified Eagle’s medium-F12 (DMEM-F12 medium) containing 10% heat-inactivated fetal calf serum (Sigma-Aldrich, St. Louis, MO), heparin (50 μg/mL, Sigma-Aldrich), endothelial cell growth supplement (15 μg/mL), insulin transferrin selenium (1%; Sigma-Aldrich), GlutaMAX (1%; ThermoFisher Scientific, Waltham, MA), and antibiotic/antimycotic mixture (Sigma-Aldrich) as described previously [25, 26]. Cells from the fourth to sixth passage were transfected with either Dnmt1-siRNA or APOBEC3A-siRNA (sc-35204 and sc-72514, respectively, Santa Cruz Biotechnology, Santa Cruz, CA) using transfection reagent (sc-29528; Santa Cruz Biotechnology). Parallel transfection was performed using nontargeting scrambled RNA (SC). For Sod2 overexpression, the cells were incubated with 3 μg of Sod2 plasmids (RC202330, OriGene, Rockville, MD) and TurboFectin 8.0 transfection reagent (TF81001, OriGene), as previously described [21]. The cells were then rinsed with phosphate buffer saline (PBS) and incubated in 5 or 20 mM glucose media containing 1% fetal calf serum, 9% Nu-serum, and 1 μg/mL endothelial growth supplement for 4 days [26]. As reported previously, cells incubated under such conditions maintain good endothelial cell morphology [27]. For osmotic control, cells incubated in 20 mM mannitol, instead of 20 mM glucose, were used. To determine the effect of regulation of Dnmt, or of nitric oxide synthase (the enzyme responsible for nitric oxide, NO, formation), on mtDNA base mismatches, cells incubated in the presence of a Dnmt inhibitor, 5-aza-2′-deoxycytidine (Aza; 1 μM), or NO synthase inhibitor L-NG-monomethyl arginine citrate (L-NMMA, 1 μM) [28] were also analyzed.
The role of mtDNA methylation in the continued formation of base mismatch in the progression of diabetic retinopathy was determined in cells incubated in 20 mM glucose for 4 days, followed by 5 mM glucose (with or without Aza) for four additional days.
Mice
C57BL/6J mice (Jackson Laboratory, Bar Harbor, ME) or Sod2 overexpressing transgenic mice (Tg) [29] were made diabetic by streptozotocin injection (55 mg/kg body weight) for four consecutive days. Animals with blood glucose > 250 mg/dL 3 days after the last injection were considered diabetic and were maintained for 6 months [26]. The animals were sacrificed by CO2 asphyxiation and their retina was processed. Age-matched normal mice were used as controls. The treatment of the animals conformed to the Association for Research in Vision and Ophthalmology Resolution on the Use of Animals in Research.
Microvessels were prepared by incubating freshly isolated retina (4–5 retina/group) in distilled water for 1 h at 37 °C [26]. Nonvascular tissue was then gently removed under a microscope, and the microvessels were used for the analyses.
Base mismatches in mtDNA were analyzed using Surveyor Mutation Detection kit (IDT Inc. Coralville, IA). Total DNA isolated by the phenol/chloroform was quantified by Quant-iT DNA detection kit (Invitrogen, Carlsbad, CA). Five overlapping regions of the D-loop (Fig. 2a; Table 1) and, for comparison, the gene encoding cytochrome oxidase 1 (CO1), were amplified using 50 ng DNA, region/gene-specific primers, and a high-fidelity Elongase enzyme mix (Invitrogen). The samples were digested using a mismatch-specific surveyor endonuclease [21]. Briefly, in a 60 μL digestion mixture, 6 μL of nuclease reaction buffer, 1 μL each of the enhancer and endonuclease, and 15 μL each of the PCR-amplified products were incubated at 42 °C for 40 min, and the reaction was stopped by 6 μL stop solution. The digested products were electrophoresed on a 2% agarose gel and analyzed for DNA fragmentation by visualizing under a UV transilluminator. The parent band amplicon intensity was quantified using ImageJ software.
Quantification of 5mC
Sonicated total genomic DNA was immunoprecipitated for 5mC using MethyLamp Methylated DNA capture (MeDIP) Kit (EPIGENTEK, Farmingdale, NY) [30, 31]. Enriched 5mC fractions were analyzed by qPCR for mtDNA methylation using primers specific for its D-loop (Table 1). 5mC levels were normalized against the threshold cycle (Ct) values obtained from total sonicated input DNA.
Gene Expression
Total RNA was extracted by TRIzol reagent (Invitrogen), and cDNA was synthesized using a High Capacity cDNA Reverse Transcription kit (Applied Biosystems, Foster City, CA). Real-time quantitative PCR (qPCR) was performed using SYBR Green assay in ABI 7500 Cycler detection system (Applied Biosystems). The Ct was determined, and the relative gene expression was calculated following delta delta Ct method [30, 31]. Table 1 includes the list of specific primers used.
Western blotting was performed using 40 μg protein sample using the antibodies against the proteins of interests, e.g., Dnmt1 (ab13537; Abcam, Cambridge, MA), Sod2 (sc-18504; Santa Cruz Biotechnology), or APOBEC3A (A58106; EPIGENTEK). β-Actin (Sigma-Aldrich) was employed as the loading control, and the band intensities were quantified using ImageJ software.
Sanger sequencing was performed using the Wayne State University’s DNA sequencing core to identify position of base mismatch in the D-loop. PCR products from five different overlapping regions of the D-loop (Table 1) were amplified by semiquantitative PCR and purified by QIAquick PCR Purification Kit (Qiagen, Valencia, CA). Sequencing was performed on Applied Biosystems ABI Prism 3730 DNA Analyzer using the ABI BigDye® Terminator v3.1 Cycle Sequencing Kit (Applied Biosystems). Efficiency and base call accuracy of the sequenced data were confirmed by analyzing the quality scores for all of the bases (a quality value score > 30 represents 99.9% base call accuracy with 0.001 probability of incorrect base incorporation). The efficacy of the sequencing data was further confirmed by analyzing the interference of the background signals with average noise for the four bases within the target sequence. The average signal intensity, which represents the signal strength of the nucleotide, was strong (between 436 and 1562 for all of the four nucleotides), indicating that the sequencing data was highly reliable. Sequences were then aligned against standard revised Cambridge reference sequence (rCRS) of the human mtDNA (NC_012920) for analysis [32] and examined by Sequence Scanner 2 (Applied Biosystems) and NCBI BLAST (Basic Local Alignment Search Tool) [33] to identify the mismatch base pairs.
Mitochondria were isolated using mitochondria isolation kit from ThermoFisher (Wilmington, DE) [31]. After digesting the homogenate with the kit reagents, mitochondrial rich fraction was obtained by differential centrifugation at 700×g for 10 min, followed by centrifugation at 3000×g for 15 min. The pellet was washed and suspended in PBS.
Complex III activity was assayed in a 50-μL assay volume containing 0.5 μg mitochondrial protein, 40 μM reduced decylubiquinone, and 2 mM KCN. The assay was initiated by 50 μM cytochrome c, and the reduction of cytochrome c was measured at 550 nm [10].
Nitrotyrosine levels were quantified by enzyme immunoassay using a Nitrotyrosine-EIA kit (Oxis Research, Portland, OR) by incubating nitrotyrosine standard or mitochondrial protein (25 μg protein) with nitrotyrosine antibody [22].
Mitochondrial localization of APOBEC3A was performed in HRECs by immunofluorescence technique using primary antibodies against APOBEC3A (A58106, EPIGENTEK) and CoxIV (mitochondrial marker; sc-376731; Santa Cruz Biotechnology) and DyLight green- or Texas red-conjugated secondary antibodies. The coverslips were mounted by 4′,6-diamidino-2-phenylindole (DAPI) containing mounting media (blue) (Vector Laboratories, Burlingame, CA) and were examined under a Zeiss ApoTome fluorescence microscope at ×40 magnification (Carl Zeiss, Inc., Chicago, IL) [31].
Statistical analysis was performed using Sigma Stat software (San Jose, CA). The results are expressed as mean ± standard deviation. Significance of variance among experimental groups was analyzed using one-way ANOVA. Data with normal distribution, or that did not qualify the normality distribution, was assessed by Bonferroni’s post hoc or Dunn’s tests. A p value < 0.05 was considered as statistically significant.
Results
Retinal Endothelial Cells
Regulation of DNA Methylation and Base Mismatch
Since 5mC is a frequent site of mutation [16], the effect of inhibition of cytosine methylation was evaluated on base mismatches in the D-loop. As expected [21], high glucose increased base mismatches, and Dnmt1-siRNA ameliorated glucose-induced increase in such base mismatch, as evident by reduced D-loop fragmentation (Fig. 1a). Figure 1b shows a significant decrease in the parent band amplicon intensity in Dnmt1-siRNA transfected cells, compared to that from the untransfected cells, in high glucose, confirming increase in base mismatches. Similarly, Aza also significantly reduced glucose-induced fragmentation, and the parent band intensity was not different from that observed in the cells incubated in normal glucose (Fig. 1a, b). Cells transfected with scrambled siRNA, or incubated with the transfection reagent alone, and exposed to high glucose, had no difference in the base mismatches and the parent band intensities compared to the untransfected cells incubated in high glucose. Figure 1c shows the knockdown efficiency of Dnmt1-siRNA, ~ 40% decrease in Dnmt1 mRNA or protein expressions in Dnmt1-siRNA-transfected cells.
Effect of mtDNA methylation on base mismatches. a DNA was amplified using semiquantitative PCR for the complete D-loop and digested with mismatch-specific surveyor endonuclease, and the fragmentation was analyzed on a 2% agarose gel. b The parent amplicon band intensity was quantified by densitometry analysis; the intensity of the band in 5 mM glucose was considered as 100%. c Dnmt1-siRNA knockdown efficiency was evaluated by quantifying Dnmt1 expression (mRNA and protein) by qPCR and western blot, respectively. d Amplification for the CO1 gene region of the mtDNA was analyzed for base mismatches by surveyor endonuclease digestion method, and e represents parent amplicon band intensity. Data are represented as mean ± SD, and each measurement was made in duplicate. The number of samples ranged from four each in the 5 mM, 20-5, D-si, Aza, and R groups; five each in the 20 mM and 20-5+Aza groups; and three each in the SC and Mann groups. 5 mM and 20 mM = 5 mM and 20 mM glucose, respectively; 20-5, 20 mM glucose for 4 days followed by 5 mM glucose for 4 days; Aza, 20 mM glucose + Aza; 20-5+Aza, 20 mM glucose for 4 days followed by 5 mM glucose + Aza for 4 days; D-si, SC and R, cells transfected with Dnmt1-siRNA or scrambled RNA or transfection regent alone, and incubated in 20 mM glucose for 4 days; Mann, 20 mM mannitol. *p < 0.05 vs 5 mM glucose and #p < 0.05 vs 20 mM glucose
In contrast to the D-loop region, surveyor digestion of CO1 amplicon showed undetectable fragmentation in high-glucose conditions, and the parent band amplicon intensity was similar to the cells incubated in normal glucose (Fig. 1c, d). Incubation of cells with mannitol, instead of glucose, had no effect on mtDNA fragmentation.
To understand the role of DNA methylation in the continued progression of base mismatches even after cessation of hyperglycemic insult, base mismatches were quantified in the HRECs exposed to high glucose followed by normal glucose, with this normal glucose phase being supplemented with Aza. As shown in Fig. 1a, b, even after removal of high glucose, base mismatches remained high, and the parent band amplicon intensity was low. The values obtained from this group of cells were similar to those obtained from cells incubated in continuous high glucose for the entire duration. However, when the normal glucose phase, which had followed the high-glucose insult, was supplemented with Aza, D-loop fragmentation was attenuated, and the values were comparable to those obtained from cells in continuous normal glucose.
To identify the positions of the base mismatches, complete D-loop was sequenced using five different overlapping primers (Table 1) by Sanger sequencing (Fig. 2a). Comparison of the sequence pattern, using BLAST against reference standard rCRS human mtDNA, identified a single base mismatch from cytosine to thymine at position 16,223 in region 2 (15,978–16,411 bp) (Fig. 2b, c). In contrast, cells incubated in normal glucose, or in high-glucose medium supplemented with Aza, presented no base mismatch. The quality scores for the bases ranged from 54 to 62, indicating a very high probability (> 99%) of the assigned bases. In these three groups of samples (5 mM, 20 mM, and 20+Aza), the background interference was also very marginal at the peak baselines with average noise less than 10 for all the four bases (7 for A, 6 for C, 6–7 for G, and 7–8 for T). (Please note that although couple samples had multiple base calls for a single base position, the image in Fig. 2b is representative of samples with relatively low background noise). As expected, digestion products of the region 2 amplicons presented a clear fragmentation product of ~ 200 bp in high glucose, which was absent in the Aza-treated cells (Fig. 2d).
D-loop sequencing and identification of base mismatches. a Five different overlapping D-loop amplicons were subjected to sequencing using Sanger sequencing method. b Representative electropherograms show DNA sequence scans, and the blue box on top of each scan represents the quality value score for each base call peak obtained from Sequence scanner software. c The data generated by aligning the D-loop sequence (Read) from region 2 using NCBI-BLAST online alignment tool and revised Cambridge reference sequence (rCRF; ref [32]) for human mtDNA as standard. Base mismatch at position 16,223 is highlighted in the red rectangular box and (d) was validated by digesting region-2 amplicon (d) using surveyor endonuclease followed by electrophoresis on a 2% agarose gel. Undigested region-2 amplicon (U) was used as a reference standard for comparison; arrow indicates the fragmented DNA (~200 bp) in the region-2 amplicons. Sequencing was performed in five samples each in the 5- and 20-mM groups and six samples in the Aza group
To examine any crosstalk between DNA methylation and base mismatches, 5mC levels were quantified targeting 16,099–16,277 bp of the D-loop, which is a part of the region overlapping 15,978–16,411 bp (D-loop region 2), used for the base mismatch analysis. As expected, glucose significantly increased the levels of 5mC, and this increase in 5mC was attenuated by Dnmt1-siRNA or Aza (Fig. 3a).
Regulation of cytosine methylation and base mismatches. a 5mC levels were quantified in the D-loop (16,099–16,277 bp region), and the total genomic DNA from each group of samples served as their input DNA controls. b Complex III activity was determined by measuring reduction of cytochrome c at 550 nm. Each measurement was made in duplicate in three samples each in the 5 mM, D-si, 20-5+Aza, Aza, 5+D-si, SC, and Mann groups and four samples each in the 20 mM, Aza, Sod2, NMMA, and R groups. The data are presented as mean ± SD. 5 mM and 20 mM = 5 mM and 20 mM glucose, respectively; 20–5, 20 mM glucose for 4 days followed by 5 mM glucose for 4 days; Aza and NMMA, 20 mM glucose supplemented with Aza or L-NMMA; 20-5+Aza, 20 mM glucose for 4 days followed by 5 mM glucose + Aza for 4 days; Sod2 and R, cells transfected with Sod2 plasmids or transfection reagent alone, respectively, and incubated in 20 mM glucose; Mann, 20 mM mannitol. * and # p < 0.05 vs 5 mM glucose and 20 mM glucose, respectively
Consistent with the failure of reversal of high glucose to attenuate base mismatches, the increase in 5mC levels in the D-loop was also not affected by the normal glucose that had followed high glucose (Fig. 3a). However, supplementation of Aza during reversal phase prevented an increase in 5mC, and the values were similar to those obtained from the cells incubated in continuous normal glucose (Fig. 3a).
Increase in D-loop base mismatches can compromise mitochondrial respiration [21]; the functional consequence of the inhibition of DNA methylation was investigated on mitochondrial complex III activity. As shown in Fig. 3b, Aza ameliorated glucose-induced decrease in complex III activity.
Regulation of Peroxynitrite Formation and Base Mismatch
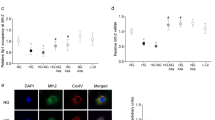
Peroxynitrite, formed by the reaction between superoxide and NO, is an important deamination factor [34]. The role of peroxynitrite in glucose-induced increase in base mismatches was confirmed by regulating the levels of both superoxide radicals and NO. Sod2 overexpression ameliorated glucose-induced increase of 5mC levels at the D-loop (Fig. 3a) and also decreased base mismatches (Fig. 4a). The parent amplicon band intensities obtained from Sod2 overexpressing cells and untransfected cells, incubated in high glucose, were significantly different from each other (Fig. 4b). Since peroxynitrite is highly reactive and its reaction with tyrosine forms a stable nitrotyrosine [35], Sod2 overexpression also ameliorated glucose-induced increase in nitrotyrosine levels in the mitochondria (Fig. 4c). Figure 4d shows ~ 50–60% increase in Sod2 expression (mRNA and protein) in the cells transfected with Sod2 plasmids compared to the untransfected cells.
Effect of Sod2 regulation on high-glucose-induced mtDNA base mismatches and nitrotyrosine levels. a Complete D-loop was analyzed for base mismatches by mismatch-specific surveyor endonuclease digestion assay. b The parent amplicon band intensity was quantified. c Nitrotyrosine levels were measured by ELISA. d Effect on Sod2 plasmid transfection on Sod2 levels were quantified by its mRNA and protein levels using β-actin as a housekeeping gene/protein. Data are represented as mean ± SD, with each measurement made in duplicate. The number of samples in each group was 4, except 3 each in the Mann and 5+Sod2 groups. *p < 0.05 vs 5 mM glucose and #p < 0.05 vs 20 mM glucose
In agreement with the persistent increase in the levels of base mismatches, reversal of hyperglycemia also failed to ameliorate glucose-induced increase in nitrotyrosine levels in the mitochondria (Fig. 4c).
To further confirm the role of peroxynitrite in base mismatches, NO synthase inhibitor L-NMMA was used; although L-NMMA failed to restore high-glucose-induced D-loop methylation (Fig. 3a), it significantly inhibited glucose-induced increase in base mismatches in the D-loop (Fig. 5a). Quantification of the amplicon band showed a higher parent band intensity in the cells incubated in high glucose in the presence of L-NMMA, compared to the cells without L-NMMA (Fig. 5b).
Effect of NO inhibition on mtDNA base mismatches. a DNA was amplified using semiquantitative PCR for complete D-loop, digested with mismatch-specific surveyor endonuclease, and analyzed on a 2% agarose gel. b The amplicon intensity was quantified, and the intensity of the band in 5 mM glucose was considered as 100%. Values are mean ± SD from four samples in each group, with each measurement made in duplicate. * and # p < 0.05 vs 5 mM and 20 mM glucose, respectively
Effect of APOBEC Deaminase on Base Mismatch
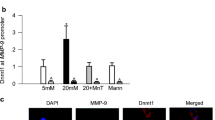
Since APOBECs are important in 5mC deamination [36], the effect of high glucose on the gene transcripts of APOBEC3A and APOBEC3G was quantified. Glucose increased APOBEC3A mRNA by ~ 50% but produced no change in APOBEC3G mRNA (Fig. 6a). Supplementation of Aza in high-glucose medium, however, had no effect on APOBEC3A transcription, and the values were similar to those obtained from the cells incubated with high glucose alone. Consistent with increased APOBEC3A transcription, glucose also increased its mitochondrial localization (Fig. 6b).
Regulation of APOBEC3A and base mismatches in mtDNA. a APOBEC3A and APOBEC3G gene transcripts were quantified by SYBR green-based qPCR using β-ACTIN as the housekeeping gene. b Mitochondrial localization of APOBEC3A was determined by immunofluorescence microscopy using Texas red-conjugated secondary antibody for APOBEC3A and DyLight green-conjugated secondary antibody for CoxIV. The cells were examined at ×40 magnification. c Base mismatches in the D-loop were analyzed by mismatch-specific surveyor endonuclease digestion assay and d the parent amplicon band intensity was quantified. e Levels of 5mC were quantified in the 16,099–16,277-bp region by methylated DNA Immunoprecipitation Kit. f Knockdown efficiency of APOBEC3A-siRNA was evaluated by quantifying APOBEC3A protein level by western blot technique using β-actin as a loading control. Data are represented as mean ± SD, with each measurement made in duplicate in four samples each in the 5 mM, 20-5, D-si, Aza, and R groups; five samples each in 20 mM and 20–5+Aza groups; and three samples each in the SC and Mann groups. 5 mM and 20 mM = 5 mM and 20 mM glucose, respectively; A-si and SC, cells transfected with APOBEC3A-siRNA or scrambled RNA, respectively, and incubated in 20 mM glucose; Mann, 20 mM mannitol. *p < 0.05 vs 5 mM glucose and #p < 0.05 vs 20 mM glucose
The expression of APOBEC3A and its mitochondrial localization also remained elevated in the cells exposed to normal glucose for 4 days, which had followed 4 days of high glucose. The addition of Aza during the normal glucose phase, which had followed high glucose, also failed to produce any beneficial effects on APOBEC3A. The values obtained from cells in the reversal group were not different from the cells in continuous high glucose alone for the entire duration (Fig. 6a, b).
Regulation of APOBEC3A by its siRNA prevented glucose-induced increase in base mismatches, and similar intensities of the parent band amplicons were observed in the untransfected cells incubated in normal glucose or mannitol. Transfection of cells with scrambled siRNA, however, had no effect on glucose-induced increase in base mismatches (Fig. 6c, d). In the same APOBEC3A-siRNA-transfected cells, glucose-induced DNA methylation was also not affected, and 5mC levels remained high (Fig. 6e). Figure 6f shows ~ 60% knockdown efficiency of APOBEC3A-siRNA in the transfected HRECs.
Diabetic Mice
Consistent with the results from HRECs, retinal microvessels from diabetic mice also presented increased base mismatches in the D-loop, which was confirmed by over 2-fold decrease in the parent amplicon band intensity (Fig. 7a, b). In the same diabetic mice, APOBEC3A expression and nitrotyrosine levels were increased by ~ 3-fold (Fig. 7c, d). Overexpression of Sod2 prevented diabetes-induced increase in base mismatches and nitrotyrosine levels (Fig. 7a–c) and had a partial, but significant, beneficial effect on the APOBEC3A expression (Fig. 7d). Consistent with its effect on APOBEC3A, Sod2 overexpression also prevented an increase of 5mC levels at the D-loop; the values obtained from Tg diabetic mice were not significantly different from those obtained from WT or Tg normal mice (Fig. 7e).
Effect of diabetes on base mismatches in D-loop and their regulation by Sod2 DNA isolated from retinal microvessels from Sod2 overexpressing mice was analyzed for a base mismatches in the D-loop using mismatch-specific surveyor endonuclease digestion kit and b represents the parent amplicon band intensity. c Nitrotyrosine levels were measured in the retinal mitochondria by ELISA. d APOBEC3A gene transcripts were quantified using 18S rRNA as the housekeeping gene, and e 5mC levels in the D-loop (15,385–15,505 bp) were quantified in the gDNA obtained from retinal microvessels using input DNA as control. WT-N, normal C57BL/6J mice; WT-D, diabetic mice; Tg-N and Tg-D, Sod2 overexpressing normal and diabetic mice. The values are represented as mean ± SD, with each measurement made for five WT-N mice, six each of WT-D and Tg-N mice, and seven Tg-D mice. *p < 0.05 compared with wild-type normal (WT-N) and #p < 0.05 diabetic (WT-D)
Discussion
Retinal mtDNA is damaged in diabetes, base mismatches are increased, and mtDNA transcription is decreased, resulting in a compromised ETC system [7, 31]. Dysfunctional ETC subsequently initiates a vicious cycle of free radicals, which continues to self-perpetuate [10, 11]. Although other regions of mtDNA show some damage, the damage is relatively higher in the D-loop region [7]. In addition to an increase in base mismatch, the D-loop is also hypermethylated with increased 5mC levels [31]. Here, our results suggest a potential crosstalk between mtDNA methylation and base mismatch; regulation of DNA methylation significantly inhibits base mismatches and prevents mitochondrial dysfunction. In addition, regulation of deamination factors, peroxynitrite and APOBEC3A, also ameliorates an increase in base mismatches. This crosstalk between DNA methylation and base mismatches continues even after the termination of hyperglycemia, suggesting its role in the metabolic memory phenomenon.
Conversion of cytosine to 5mC by Dnmts is one of the epigenetic mechanisms that cells use to lock genes in the ‘off’ position and control gene expression [37]. The mammalian mtDNA has ~ 450 CpG sites and ~ 4500 cytosines at non-CpG sites, and in diabetes, translocation of Dnmts inside the mitochondria facilitates hypermethylation of mtDNA, especially its D-loop [31]. Although CpG methylation in the mammalian genome is a natural phenomenon, it is also the major site of mutation, and > 90% of mutations are due to conversion of cytosine to thymine, or guanosine to adenine. However, 5mC can also be deaminated, and the deamination rate of 5mC is several fold higher than that of cytosine [38]. In addition, the transition rate of 5mC to thymine-guanosine is 10–50-fold higher than other transitional changes [39, 40]. Here, we provide data showing a possible crosstalk between DNA methylation and base mismatch in the development of diabetic retinopathy. Our sequencing results showing replacement of cytosine at position 16,223 by thymine at non-CpG region confirm base mismatch in the D-loop. Furthermore, this transition is prevented by Aza, suggesting a significant contribution of DNA methylation in the base mismatch formation. In support, a correlation between methylation and CpG substitution rate has been documented in a CpG mutability in the intronic regions of the human genome [41]. Gene sequence analysis has also shown a significant correlation between the extent of germ line methylation and the mutation rate [42]. Although CpG methylation is an important regulator in gene expression, we cannot rule out the possibility of hyperglycemia-induced base mismatch/DNA methylation at non-CpG sites in the D-loop region; methylation of non-CpG sites in the mtDNA has been reported in human peripheral blood [43, 44].
Increased NO is considered as one of the causative factors in cytosine deamination [34]; in esophageal adenocarcinoma, inducible NO synthase-mediated increase in nitrotyrosine accumulation has been implicated in endogenous p53 mutations at the CpG dinucleotides [17]. Reaction of NO with superoxide forms peroxynitrite, which is a potent oxidizing and genotoxic agent [45]. Our results clearly show that, although inhibition of nitric oxide synthase prevents hyperglycemia-induced increase in mitochondrial nitrotyrosine and mismatches in the D-loop, it fails to prevent an increase in 5mC levels, suggesting that the regulation of peroxynitrite might be involved only in the second step base conversion (5mC to thymine) during oxidative deamination. In addition, failure of inhibition of nitrative stress to produce any significant effect on DNA methylation further suggests that oxidative DNA damage is not the sole initiator of base mismatches, and deamination also has a critical role in their formation. The role of peroxynitrite in cytosine deamination is further supported by decreased base mismatches and DNA methylation in the cells overexpressing Sod2. Consistent with the in vitro results, D-loop in the retinal microvessels from mice overexpressing Sod2 is also protected from diabetes-induced increase in base mismatch and DNA methylation.
Deamination of 5mC to thymine is also regulated by AID/APOBEC enzymes, a family of enzymes which converts cytosine to uracil [19]. In addition to repairing DNA damage, AID/APOBECs are also implicated in the demethylation of DNA. Among many well-characterized cytidine deaminases, APOBEC3 has an effective cytosine deaminase activity. But, in contrast, because of the poor fitting of 5-methyl group in the DNA-binding pocket, deamination of 5mC by AID is not strong [36, 46]. Here, we show that the expression of APOBEC3A is significantly increased in hyperglycemia, suggesting its role in increased base mismatches in diabetes. This is further confirmed by amelioration of D-loop fragmentation by APOBEC3A-siRNA. Consistent with this, others have shown the involvement of APOBEC3A in somatic hypermutation of human mtDNA [47]. Our results also demonstrate regulation of APOBEC3A by Sod2 overexpression, raising the possibility of an indirect regulation of mtDNA deamination via mitochondrial oxidative stress. Furthermore, Aza administration, which inhibits mtDNA methylation and base mismatches, fails to have any effect on APOBEC3A expression, suggesting that the availability of deaminase substrate, i.e., 5mC, is critical for base mismatches. These results clearly imply that DNA methylation could be preceding the deamination process formation of base mismatch. Although such base mismatch can be repaired, limited mitochondrial repair capacity could lead to mtDNA mutations; many such mutations are seen in ocular disease [14]. Mismatches/mutations in mtDNA have also been reported in other vascular disease including atherosclerosis [48] and in brain microvessels [49], suggesting that mtDNA mismatch/mutation in the vascular walls can contribute to mitochondrial dysfunction and vascular pathology.
Several studies have documented that epigenetic modifications confer memory phenomenon which persists even when the external stimulus is removed, and DNA methylation is one such modification in long-term memory function [8, 25, 50, 51]. Retinal mtDNA, once methylated by prior hyperglycemia, does not reverse even after reinstitution of normal glycemia, and the regulatory region of polymerase gamma 1 remains hypermethylated [25, 50]. Furthermore, due to continued dysfunctional repair mechanism, base mismatch continues to accumulate in the mtDNA [21]. Normal glucose alone, that has followed high glucose, does not reverse D-loop methylation, and this raises the possibility that the continued DNA methylation, observed in the present study, could be one of the potential causes for the persistent high levels of base mismatches and mitochondrial dysfunction. Consistent with amelioration of mtDNA damage and progression of diabetic retinopathy by a combination of good control with a therapy targeted to quench free radicals [24], and attenuation of mtDNA methylation by direct targeting Dnmts during the reversal phase [25], here we show that the addition of Aza during the normal glucose phase attenuates mismatch formation. These results further support a crosstalk between DNA methylation and base mismatch and its role in the metabolic memory phenomenon.
In summary, we have provided a first line of evidence suggesting a positive correlation between DNA methylation and base mismatch, subsequently leading to mitochondrial dysfunction, and inhibition of DNA methylation limits the availability of 5mC for deamination to form base mismatch. Thus, regulation of DNA methylation, or its deamination, should impede the development of diabetic retinopathy by regulating base mismatch and mitochondrial dysfunction.
References
Brownlee M (2001) Biochemistry and molecular cell biology of diabetic complications. Nature 414:813–820. https://doi.org/10.1038/414813a
Frank RN (2004) Diabetic retinopathy. N Engl J Med 350:48–58. https://doi.org/10.1056/NEJMra1005073
Kern TS, Tang J, Mizutani M, Kowluru R, Nagraj R, Lorenzi M (2000) Response of capillary cell death to aminoguanidine predicts the development of retinopathy: comparison of diabetes and galactosemia. Invest Ophthalmol Vis Sci 41:3972–3978
Kowluru RA (2013) Mitochondria damage in the pathogenesis of diabetic retinopathy and in the metabolic memory associated with its continued progression. Curr Med Chem 20:3226–3233
Kowluru RA, Chan PS (2010) Metabolic memory in diabetes—from in vitro oddity to in vivo problem: role of apoptosis. Brain Res Bull 81:297–302. https://doi.org/10.1016/j.brainresbull.2009.05.006
Falkenberg M, Larsson NG, Gustafsson CM (2007) DNA replication and transcription in mammalian mitochondria. Annu Rev Biochem 76:679–699. https://doi.org/10.1146/annurev.biochem.76.060305.152028
Tewari S, Santos JM, Kowluru RA (2012) Damaged mitochondrial DNA replication system and the development of diabetic retinopathy. Antioxid Redox Signal 17:492–504. https://doi.org/10.1089/ars.2011.4333
Kowluru RA, Kowluru A, Mishra M, Kumar B (2015) Oxidative stress and epigenetic modifications in the pathogenesis of diabetic retinopathy. Prog Retin Eye Res 48:40–61. https://doi.org/10.1016/j.preteyeres.2015.05.001
Reddy MA, Zhang E, Natarajan R (2015) Epigenetic mechanisms in diabetic complications and metabolic memory. Diabetologia 58:443–455. https://doi.org/10.1007/s00125-014-3462-y
Kanwar M, Chan PS, Kern TS, Kowluru RA (2007) Oxidative damage in the retinal mitochondria of diabetic mice: possible protection by superoxide dismutase. Invest Ophthalmol Vis Sci 48:3805–3811. https://doi.org/10.1167/iovs.06-1280
Santos JM, Tewari S, Goldberg AFX, Kowluru RA (2011) Mitochondria biogenesis and the development of diabetic retinopathy. Free Radic Biol Med 51:1849–1860. https://doi.org/10.1016/j.freeradbiomed.2011.08.017
Alberts B, Johnson A, Lewis J (2002) DNA repair. Molecular biology of the cell. In: 4 edn. Garland Science, New York
Moore LD, Le T, Fan G (2013) DNA methylation and its basic function. Neuropsychopharmacology 38:23–38. https://doi.org/10.1038/npp.2012.112
Al-Enezi M, Al-Saleh H, Nasser M (2008) Mitochondrial disorders with significant ophthalmic manifestations. Middle East Afr J Ophthalmol 15:81–86. https://doi.org/10.4103/0974-9233.51998
Holliday R, Grigg GW (1993) DNA methylation and mutation. Mutat Res 285:61–67
Poulos RC, Olivier J, Wong JWH (2017) The interaction between cytosine methylation and processes of DNA replication and repair shape the mutational landscape of cancer genomes. Nucleic Acids Res 45:7786–7795. https://doi.org/10.1093/nar/gkx463
Vaninetti NM, Geldenhuys L, Porter GA, Risch H, Hainaut P, Guernsey DL, Casson AG (2008) Inducible nitric oxide synthase, nitrotyrosine and p53 mutations in the molecular pathogenesis of Barrett’s esophagus and esophageal adenocarcinoma. Mol Carcinog 47:275–285. https://doi.org/10.1002/mc.20382
Morey M, Serras F, Baguna J, Hafen E, Corominas M (2002) Modulation of the Ras/MAPK signalling pathway by the redox function of selenoproteins in Drosophila melanogaster. Dev Biol 238:145–156. https://doi.org/10.1006/dbio.2001.0389
Seplyarskiy VB, Soldatov RA, Popadin KY, Antonarakis SE, Bazykin GA, Nikolaev SI (2016) APOBEC-induced mutations in human cancers are strongly enriched on the lagging DNA strand during replication. Genome Res 26:174–182. https://doi.org/10.1101/gr.197046.115
Madsen-Bouterse SA, Mohammad G, Kanwar M, Kowluru RA (2010) Role of mitochondrial DNA damage in the development of diabetic retinopathy, and the metabolic memory phenomenon associated with its progression. Antioxid Redox Signal 13:797–805. https://doi.org/10.1089/ars.2009.2932
Mishra M, Kowluru RA (2014) Retinal mitochondrial DNA mismatch repair in the development of diabetic retinopathy, and its continued progression after termination of hyperglycemia. Invest Ophthalmol Vis Sci 55:6960–6967. https://doi.org/10.1167/iovs.14-15020
Kowluru RA, Kanwar M, Chan PS, Zhang JP (2008) Inhibition of retinopathy and retinal metabolic abnormalities in diabetic rats with AREDS-based micronutrients. Arch Ophthalmol 126:1266–1272. https://doi.org/10.1001/archopht.126.9.1266
DCCT (1993) The effect of intensive treatment of diabetes on the development and progression of long-term complications in insulin-dependent diabetes mellitus. The Diabetes Control and Complications Trial Research Group. N Engl J Med 329:977–986. https://doi.org/10.1056/NEJM199309303291401
Santos JM, Kowluru RA (2011) Role of mitochondria biogenesis in the metabolic memory associated with the continued progression of diabetic retinopathy and its regulation by lipoic acid. Invest Ophthalmol Vis Sci 52:8791–8798. https://doi.org/10.1167/iovs.11-8203
Mishra M, Kowluru RA (2016) The role of DNA methylation in the metabolic memory phenomenon associated with the continued progression of diabetic retinopathy. Invest Ophthalmol Vis Sci 57:5748–5757. https://doi.org/10.1167/iovs.16-19759
Mishra M, Kowluru RA (2017) Role of PARP-1 as a novel transcriptional regulator of MMP-9 in diabetic retinopathy. Biochim Biophys Acta 1863:1761–1769. https://doi.org/10.1016/j.bbadis.2017.04.024
Duraisamy AJ, Mishra M, Kowluru RA (2017) Crosstalk between histone and DNA methylation in regulation of retinal matrix metalloproteinase-9 in diabetes. Invest Ophthalmol Vis Sci 58:6440–6448. https://doi.org/10.1167/iovs.17-22706
Leikert JF, Rathel TR, Muller C, Vollmar AM, Dirsch VM (2001) Reliable in vitro measurement of nitric oxide released from endothelial cells using low concentrations of the fluorescent probe 4,5-diaminofluorescein. FEBS Lett 506:131–134
Kowluru RA, Kowluru V, Xiong Y, Ho YS (2006) Overexpression of mitochondrial superoxide dismutase in mice protects the retina from diabetes-induced oxidative stress. Free Radic Biol Med 41:1191–1196. https://doi.org/10.1016/j.freeradbiomed.2006.01.012
Kowluru RA, Shan Y, Mishra M (2016) Dynamic DNA methylation of matrix metalloproteinase-9 in the development of diabetic retinopathy. Lab Investig 96:1040–1049. https://doi.org/10.1038/labinvest.2016.78
Mishra M, Kowluru RA (2015) Epigenetic modification of mitochondrial DNA in the development of diabetic retinopathy. Invest Ophthalmol Vis Sci 56:5133–5142. https://doi.org/10.1167/iovs.15-16937
Bandelt HJ, Kloss-Brandstatter A, Richards MB, Yao YG, Logan I (2014) The case for the continuing use of the revised Cambridge Reference Sequence (rCRS) and the standardization of notation in human mitochondrial DNA studies. J Hum Genet 59:66–77. https://doi.org/10.1038/jhg.2013.120
Tsiatis AC, Norris-Kirby A, Rich RG, Hafez MJ, Gocke CD, Eshleman JR, Murphy KM (2010) Comparison of Sanger sequencing, pyrosequencing, and melting curve analysis for the detection of KRAS mutations: diagnostic and clinical implications. J Mol Diagn 12:425–432. https://doi.org/10.2353/jmoldx.2010.090188
Caulfield JL, Wishnok JS, Tannenbaum SR (1998) Nitric oxide-induced deamination of cytosine and guanine in deoxynucleosides and oligonucleotides. J Biol Chem 273:12689–12695
Szabo C, Ischiropoulos H, Radi R (2007) Peroxynitrite: biochemistry, pathophysiology and development of therapeutics. Nat Rev Drug Discov 6:662–680. https://doi.org/10.1038/nrd2222
Fritz EL, Papavasiliou FN (2010) Cytidine deaminases: AIDing DNA demethylation? Genes Dev 24:2107–2114. https://doi.org/10.1101/gad.1963010
Bestor TH (2000) The DNA methyltransferases of mammals. Hum Mol Genet 9:2395–2402
Ehrlich M, Norris KF, Wang RY, Kuo KC, Gehrke CW (1986) DNA cytosine methylation and heat-induced deamination. Biosci Rep 6:387–393
Jiang C, Han L, Su B, Li WH, Zhao Z (2007) Features and trend of loss of promoter-associated CpG islands in the human and mouse genomes. Mol Biol Evol 24:1991–2000. https://doi.org/10.1093/molbev/msm128
Zhao Z, Jiang C (2007) Methylation-dependent transition rates are dependent on local sequence lengths and genomic regions. Mol Biol Evol 24:23–25. https://doi.org/10.1093/molbev/msl156
Mugal CF, Ellegren H (2011) Substitution rate variation at human CpG sites correlates with non-CpG divergence, methylation level and GC content. Genome Biol 12:R58. https://doi.org/10.1186/gb-2011-12-6-r58
Xia J, Han L, Zhao Z (2012) Investigating the relationship of DNA methylation with mutation rate and allele frequency in the human genome. BMC Genomics 13(Suppl 8):S7. https://doi.org/10.1186/1471-2164-13-S8-S7
Liu B, Du Q, Chen L, Fu G, Li S, Fu L, Zhang X, Ma C et al (2016) CpG methylation patterns of human mitochondrial DNA. Sci Rep 6:23421. https://doi.org/10.1038/srep23421
Malone CS, Miner MD, Doerr JR, Jackson JP, Jacobsen SE, Wall R, Teitell M (2001) CmC(A/T)GG DNA methylation in mature B cell lymphoma gene silencing. Proc Natl Acad Sci U S A 98:10404–10409. https://doi.org/10.1073/pnas.181206898
Kamat JP (2006) Peroxynitrite: a potent oxidizing and nitrating agent. Indian J Exp Biol 44:436–447
Nowarski R, Kotler M (2013) APOBEC3 cytidine deaminases in double-strand DNA break repair and cancer promotion. Cancer Res 73:3494–3498. https://doi.org/10.1158/0008-5472.CAN-13-0728
Suspene R, Aynaud MM, Guetard D, Henry M, Eckhoff G, Marchio A, Pineau P, Dejean A et al (2011) Somatic hypermutation of human mitochondrial and nuclear DNA by APOBEC3 cytidine deaminases, a pathway for DNA catabolism. Proc Natl Acad Sci U S A 108:4858–4863. https://doi.org/10.1073/pnas.1009687108
Sazonova M, Budnikov E, Khasanova Z, Sobenin I, Postnov A, Orekhov A (2009) Studies of the human aortic intima by a direct quantitative assay of mutant alleles in the mitochondrial genome. Atherosclerosis 204:184–190. https://doi.org/10.1016/j.atherosclerosis.2008.09.001
Betts J, Jaros E, Perry RH, Schaefer AM, Taylor RW, Abdel-All Z, Lightowlers RN, Turnbull DM (2006) Molecular neuropathology of MELAS: level of heteroplasmy in individual neurones and evidence of extensive vascular involvement. Neuropathol Appl Neurobiol 32:359–373. https://doi.org/10.1111/j.1365-2990.2006.00731.x
Tewari S, Zhong Q, Santos JM, Kowluru RA (2012) Mitochondria DNA replication and DNA methylation in the metabolic memory associated with continued progression of diabetic retinopathy. Invest Ophthalmol Vis Sci 53:4881–4888. https://doi.org/10.1167/iovs.12-9732
Chen Z, Miao F, Paterson AD, Lachin JM, Zhang L, Schones DE, Wu X, Wang J et al (2016) Epigenomic profiling reveals an association between persistence of DNA methylation and metabolic memory in the DCCT/EDIC type 1 diabetes cohort. Proc Natl Acad Sci U S A 113:E3002–E3011. https://doi.org/10.1073/pnas.1603712113
Funding
The authors acknowledge the grant support from the National Institutes of Health (R01-EY014370, R01-EY017313, and R01-EY022230) and the Thomas Foundation to RAK and from Research to Prevent Blindness to the Ophthalmology Department.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Mishra, M., Kowluru, R.A. DNA Methylation—a Potential Source of Mitochondria DNA Base Mismatch in the Development of Diabetic Retinopathy. Mol Neurobiol 56, 88–101 (2019). https://doi.org/10.1007/s12035-018-1086-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-018-1086-9