Abstract
Parkinson’s disease (PD) is the most common movement disorder disease, and its pathological feature is the degenerative loss of dopaminergic neurons in the substantia nigra compacta (SNc). In this study, we investigated whether distinct stress conditions target F-box protein Fbw7β via converging mechanisms. Our results showed that the 6-hyroxydopamine (6-OHDA), which causes PD in animals’ models, led to decreased stability of Fbw7β in DA neuronal SN4741 cells. Further experiments suggested that oxidized Fbw7β bound to heat-shock cognate protein 70 kDa, the key regulator for chaperone-mediated autophagy (CMA), at a higher affinity. Oxidative stress also increased the level of lysosomal-associated membrane protein 2A (LAMP2A), the rate-limiting receptor for CMA substrate flux, and stimulated CMA activity. These changes resulted in accelerated degradation of Fbw7β. 6-OHDA induced Fbw7β oxidation and increased LAMP2A in the SNc region of the mouse models. Consistently, the levels of oxidized Fbw7β were higher in postmortem PD brains compared with the controls. These findings for the first time revealed the specific mechanism of ubiquitin ligases, oxidative stress, and CMA-mediated protein degradation, to provide a new theoretical basis for further clarifying the mechanism of PD.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Parkinson’s disease (PD) is the most common movement disorder disease [1]. The incidence of PD rises progressively with the increase of age, mainly characterized by limb movement disorder such as resting tremor, postural dysfunction, and bradykinesia [2]. Some patients are also accompanied with cognitive impairment, which seriously affects their quality of life and social work [3]. Histopathologically, the disease is characterized by relatively selective loss of dopaminergic neurons in the substantia nigra compacta (SNc) in the midbrain and their terminals in the striatum [2]. The cause of continuous loss of dopamine neurons in PD is still unclear until now. Due to the increased population aging, neurodegenerative diseases, typically represented by PD, have become the major cause of death and have brought heavy burden to patients [1].
Autophagy refers to the process that cells disassemble cellular components by lysosome, which includes macroautophagy, microautophagy, and chaperone-mediated autophagy (CMA) according to the delivery mode of the substrate to the lysosome [4]. It is very important to maintain the normal and stable autophagy function for the steady state of intracellular environment. The imbalance of autophagic pathway plays an important role in pathogenic mechanisms of different diseases, including cancer [5], aging diseases [6], infectious diseases [7], cardiovascular diseases [8], and neurological diseases [9]. Neurons are particularly vulnerable to the autophagy [10]. Existing studies have shown that autophagy disorder may cause chronic neurodegenerative diseases such as Alzheimer’s disease [11], PD [12], Huntington’s disease [13], amyotrophic lateral sclerosis [14], and neuron injuries by ischemia [15] and trauma [16]. The deletion of macroautophagy-related genes in the central nervous system can lead to neurodegenerative damage [17, 18]. Two major regulatory factors to adjust CMA are heat-shock cognate protein 70 (Hsc70) and lysosome-associated membrane protein 2A (LAMP2A) [19]. Several PD-related proteins, such as mutated α-synuclein and ubiquitin C-terminal hydrolase L1 (UCH-L1), disrupt the CMA process [20, 21], suggesting that CMA may underlie PD pathogenesis [22].
Neurotoxins mainly induce neuronal death of DA by oxidative stress [23], and 6-OHDA induces oxidative stress by damaging mitochondria [24]. Fbw7β is a member of F-box proteins that provide substrate specificity for SCF (Skp, Cullin, F-box containing complex) ubiquitin ligases. Fbw7β has been reported to collaborate with parkin for proteasomal degradation and promotes neuronal survival in response to oxidative stress [25]. In order to observe the influences of 6-OHDA on Fbw7β degradation and whether oxidative stress participates in the regulation of function and metabolism of Fbw7β by oxidative modification of Fbw7β, we used 6-OHDA to treat SN4741 cells and detected the changes of expression in the nucleus and cytoplasm and degree of Fbw7β oxidation and its correlation with 6-OHDA. Our results showed that the oxidative stress caused oxidation modifications of Fbw7β in vitro and in the SNc areas of mice and postmortem PD brain tissues. Oxidized Fbw7β is preferentially removed by CMA, and failure to remove damaged Fbw7β by CMA is correlated with an increase injury to oxidative stress. Research has also shown that dysfunction of the CMA is closely related to the onset of Parkinson’s disease [26]. This research for the first time elaborated the pathogenesis of Parkinson’s from the angle of oxidative stress, CMA, and metabolism of key factor of ubiquitin ligase.
Methods and Materials
Animals
All animal studies were performed according to the protocols and guidelines of the institutional care and use committee of Huazhong University of Science and Technology. Adult male C57BL/6 mice were purchased and housed in standard enriched environment cages in a temperature-controlled room with a 12-h light/dark cycle and free access to food. The animals were given at least a week acclimation period to the facility before the administration of 6-OHDA (Sigma, St. Louis, MO).
The 6-OHDA mouse model of PD was created as previously described [27, 28]. Mice were randomly assigned to two groups. The group of PD model mice (n = 9) received injections of 6-OHDA (3 μg in 2 μl) into the left SNc of midbrain at the following coordinates: AP, −3.16; ML, −3.85; and DV, −2.64 mm. The group of control animals (n = 9) received only saline at the same coordinates.
Patient Cases
Brain tissue was collected from Tongji Medical College, Huazhong University of Science and Technology. A clinical diagnosis was established according to the Chinese Neurology Society clinical diagnostic criteria for PD [29], which was drawn up referring to the United Kingdom Brain Bank criteria. The diagnostic criteria for PD include a slow disease onset with at least two of the following manifestations: (1) resting tremor or bradykinesia responsive to levodopa, (2) rigidity, or (3) abnormal gait/posture [29]. Healthy controls were selected based on age and no history of neuropathological diagnosis of neurologic or psychiatric disease. The study population consisted of ten control subjects (five male and five female; mean age 80.2 ± 2.13 years, range 74–89 years) and ten patients with PD (six male and four female; mean age 81.2 ± 2.44 years, range 71–90 years). The Ethics Committee of Tongji Medical College approved all procedures, and all subjects provided written informed consents. The postmortem samples were snap-frozen and stored in liquid nitrogen until further processing. The delay in tissue processing was between 2.25 and 5.5 h (control group 3.60 ± 1.0 h; PD group 3.21 ± 1.2 h). The storage duration in liquid nitrogen was between 7 and 26 weeks (control group 17.6 ± 12.4 weeks; PD group 19.1 ± 10.6 weeks).
Antibodies and Chemicals
Antibodies were purchased from the following vendors: anti-Fbw7β from Bethyl Laboratories (Montgomery, TX); anti-Hsc70, anti-LAMP2A, and anti-HMGB1 from Abcam (Cambridge, MA); and horseradish peroxidase (HRP)-streptavidin from Amersham Bioscience (Arlington Heights, IL). The OxyBlot protein oxidation detection kit was purchased from Chemicon International (Tamecula, CA), 6-OHDA, iodoacetamide (IAM), and α-tocopherol from Sigma, and biotin-iodoacetamide from Invitrogen (Carlsbad, CA).
Cell Culture and Transfection
Mouse embryonic substantia nigra-derived cell line SN4741 was purchased from American Type Culture Collection (Manassas, VA, USA). Cells were cultured with DMEM medium containing 5% fetal bovine serum (GIBCO, Carlsbad, CA) in the incubator (95% air and 5% CO2) at 37 °C. SN4741 cells were inoculated into 6-well plate with 3 × 105 cells per well to culture till 80% fusion and the medium was changed by serum-free DMEM for further 6-h cultivation, and then the cells were stimulated with 6-OHDA (5 ng/ml) according to the experimental requirement and then collected for detection. For cell transfection, the pGCSIL vector encoding the mouse Fbw-7β was purchased from GenePharma (Shanghai, China). Cell transfection was introduced by Lipofectamin 2000 (Invitrogen) in accordance with the manufacturer’s protocol. The overexpression was confirmed by western blot.
Immunofluorescence Staining
Cells were seeded onto coverslips in 6-well culture plates. Cells were washed with PBS and fixed by 4% paraformaldehyde for 15 min at room temperature. Cells were then washed three times with PBS and permeabilized with 1% BSA in PBS solution with 0.1% Triton X-100 at 4 °C for 5 min. Cells were blocked with 1% BSA in PBS for 30 min. Thereafter, incubated with anti-LAMP2A antibody (1:200) overnight, washed thrice with PBS, incubated with anti-rabbit antibody at room temperature, and washed thrice with PBS. The nuclei were stained with DAPI. Cells were mounted with anti-fade solution and visualized using confocal microscopy with LAS AF software (Leica, Wetzlar, Germany).
Immunoprecipitation of BIAM-Labeled Fbw7β
The cells were treated with 6-OHDA as indicated and lysed in oxygen-free lysis buffer (50-mM Bis-Tris-HCl (pH 6.5), 0.5% Triton X-100, 0.5% deoxycholate, 0.1% sodium dodecyl sulfate (SDS), 150-mM NaCl, 1-mM EDTA, protease inhibitor, and 20-μM BIAM). After incubation at 37 °C in darkness for 30 min, the labeling reaction was stopped by adding IAM to 5 mM. Fbw7β was precipitated and immunoblotted with anti-streptavidin HRP.
Protein Oxidation Detection
The SN4741 cells were allowed to reach 70–80% confluence. After treatment with 6-OHDA for 18 h, the cells were lysed in a buffer containing 50-mM DTT. A small amount of lysate (30 μg) was applied for western blot assay to determine the relative expression of Fbw7β (quantified by ImageJ software). The volume of each sample was calculated according to the Fbw7β protein expression level. Precleared lysates were incubated with the Fbw7β primary antibody (Bethyl) and protein A/G agarose at 4 °C overnight. The complexes were washed three times with lysis buffer and then twice with low-salt buffer (0.1% SDS, 1% Triton, 2-mM Tris-HCl pH 7.8, 150-mM NaCl) and twice with high-salt buffer (0.1% SDS, 1% Triton, 2-mM Tris-HCl pH 7.8, 500-mM NaCl). The levels of oxidized Fbw7β in purified protein were detected using the OxyBlot protein oxidation detection kit. The DNPH derivatization was carried out on 6 μg of protein for 15 min according to the manufacturer’s instructions. One-dimensional electrophoresis was carried out on 12% SDS/polyacrylamide gel after DNPH derivatization. Proteins were transferred to nitrocellulose membranes which were then incubated in the primary antibody solution (anti-DNP 1:150) for 2 h, followed by incubation in second antibody solution (1:300) for 1 h at room temperature. The washing procedure was repeated eight times within 40 min. Immunoreactive bands were visualized by enhanced chemiluminescence (ECL; Amersham Biosciences) which was then exposed to Biomax L film (Kodak, Rochester, NY). For quantification, ECL signals were digitized using ImageJ software.
GST Pull-down Assay
The SN4741 cells were washed in PBS, lysed in a lysis buffer (20-mM Tris pH 7.4, 40-mM NaCl, 1-mM EDTA, 1-mM EGTA, 1-mM Na3VO4, 10-mM NaF, 10-mM sodium pyrophosphate, 10-mM sodium β-glycerophosphate, and 1% Triton X-100), and centrifuged. The supernatant was incubated with purified GST fusion proteins on Sepharose beads (GE, Boston, MA) for 2 h at 4 °C.
Western Blot Assay
The cells were resuspended in the Laemmli sample buffer (pH 6.8, 62.5-mM Tris-Cl, 0.01% bromphenol blue, 2% SDS, 2% β-mercaptoethanol, and 10% glycerol), separated by SDS-PAGE, and transferred to a nitrocellulose membrane (Amersham Biosciences). The membranes were blocked with 10% low-fat milk in TBS and probed with primary antibodies. Followed by chemiluminescence (ECL; Amersham Biosciences), the HRP-conjugated secondary antibodies (Amersham Bioscience) were utilized to visualize the bands of protein. The ImageJ was used to quantify the intensities of band. More than three independent experiments were carried out.
Statistics
Results are expressed as the means ± standard error of the mean (SEM). Differences between multiple groups were compared using one-way ANOVA with post hoc Bonferroni correction. Differences between two groups were evaluated using a two-tailed unpaired Student’s t test. The significance level was set at P < 0.05 which indicated a strongly significant difference. All statistical analyses were performed using SPSS 10.0 software (SPSS Inc., Chicago, IL).
Results
Oxidative Modification by Neurotoxin 6-OHDA on Fbw7β of SN4741 Cells
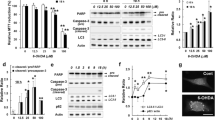
Results showed that after 6-OHDA exposure, expression of Fbw7β in SN4741 cells was reduced with increasing dose (Fig. 1a, P < 0.05). We further detected the changes of oxidative modification level in SN4741 cells by different methods. After treatment with 6-OHDA, Fbw7β was immunoprecipitated from lysates of SN4741 cells and analyzed for carbonyl oxidation using the OxyBlot protocol [30]. Exposure to 6-OHDA caused a marked increase in the level of carbonyl oxidation of Fbw7β in a time-dependent manner in SN4741 cells (Fig. 1b, P < 0.05). In order to confirm the oxidative modification of oxidative stress on Fbw7β, we used BIAM to further detect and determine the above research results [30]. Specifically, SN4741 cells treated with 6-OHDA were lysed in a buffer containing BIAM. Fbw7β was then isolated from the cell lysate by immunoprecipitation and was subsequently labeled with BIAM. The result indicated that 6-OHDA decreased the BIAM labeling of Fbw7β protein to a great extent, revealing increased oxidation of cysteine residues (Fig. 1c, P < 0.05). Furthermore, we tested the anti-oxidant vitamin E and showed that it protected Fbw7β from 6-OHDA-induced reduction (Fig. 1d, P < 0.05). Together, these findings clearly demonstrate that Fbw7β is readily oxidized under 6-OHDA exposure.
6-OHDA induced Fbw7β oxidation increasing. a Altered expression of Fbw7β protein after 6-OHDA treatment (20 or 40 μM, 24 h) by western blot. b 6-OHDA induced altered levels of oxidized Fbw7βdetected by DNP immunoblotting. Cell lysates treated with 20- or 40-μM 6-OHDA for 24 h were immunoprecipitated with anti-Fbw7β antibody. The precipitates were immunoblotted for DNP and Fbw7β. c 6-OHDA induced altered levels of BIAM-labeled Fbw7β. Cell lysates treated with 20- or 40-μM 6-OHDA for 24 h were immunoprecipitated with anti-Fbw7β antibody. The precipitates were immunoblotted for streptavidin-HRP and Fbw7β. d Anti-oxidants vitamin E blocked 6-OHDA induced Fbw7β oxidation. Cells were treated with vitamin E and 6-OHDA for 24 h. Fbw7β expression was analyzed by western blot. The photographs are representative of three independent experiments. The bar graphs show the quantification of protein expression (mean ± SEM, *P < 0.05)
Influences of Neurotoxin 6-OHDA on CMA Degradation Pathway in SN4741 Cells
Our studies in Fig. 1 showed that levels of Fbw7β are oxidatively modified after 6-OHDA treatment. We further investigated whether the neurotoxin-induced loss of Fbw7β proteins was mediated by a lysosomal pathway. We used lysosomal inhibitor NH4Cl and then exposed SN4741 cells to 6-OHDA with or without NH4Cl. The results showed that 6-OHDA significantly reduced Fbw7β levels in SN4741 cells, and the inhibition of lysosomal function with NH4Cl significantly blocked the 6-OHDA-induced loss of Fbw7β (Fig. 2a, P < 0.05). Also, the reduction of Fbw7β expression by 6-OHDA was not significantly inhibited when using macroautophagy inhibitor 3-MA (Fig. 2b, P > 0.05). Inhibition of the proteasome through MG132 also failed to attenuate 6-OHDA-induced loss of Fbw7β (Fig. 2c, P > 0.05). These findings suggest that proteasomes do not play a major role in 6-OHDA-induced degradation of Fbw7β. Furthermore, the inhibition of caspase-3 activity by z-DEVD-fmk did not reverse the 6-OHDA-induced Fbw7β reduction (Fig. 2d, P > 0.05). Together, these results support CMA as the primary mechanism by which 6-OHDA promotes Fbw7β degradation.
6-OHDA induced Fbw7β degradation through lysosome-mediated pathway. a NH4Cl blocked 6-OHDA-induced Fbw7β degradation. Cells were exposed to 6-OHDA with or without NH4Cl for 18 h and Fbw7β protein expression was analyzed by western blot. b 3-MA did not block 6-OHDA-induced Fbw7β degradation. Cells were exposed to 6-OHDA with or without 3-MA for 12 h and analyzed Fbw7β protein expression by western blot. c Proteasome inhibitor MG132 did not block 6-OHDA-induced Fbw7β degradation. Cells were exposed to 6-OHDA with or without MG132 for 12 h and analyzed Fbw7β protein expression. d Caspase-3 inhibitor z-DEVD-fmk did not block 6-OHDA-induced Fbw7β degradation. Cells were exposed to 6-OHDA with or without z-DEVD-fmk for 12 h and Fbw7β protein expression was analyzed. The photographs are representative of three independent experiments. The bar graphs show the quantification of protein expression (mean ± SEM, *P < 0.05)
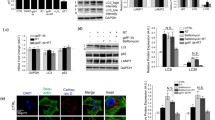
Previous study showed that neurotoxin 6-OHDA can activate CMA of dopaminergic SN4741 cell [30]. We further observed the regulation of 6-OHDA on expression level of LAMP2A—the key rate-limiting factor of CMA. Changes of LAMP2A were detected through immunofluorescence and western blot assays. Results showed that after treated with 6-OHDA, LAMP2A expression in SN4741 cells was obviously increased (Fig. 3a, P < 0.05). Western blot results were consistent with the fluorescence results (Fig. 3b, P < 0.05). Through transfection with anti-sense-LAMP2A to interfere with LAMP2A expression, Fbw7β loss induced by 6-OHDA was prevented (Fig. 3c, P < 0.05). However, western blot assay at the same time showed that SN cells treated by 6-OHDA did not obviously change the expression of another key CMA regulator HSC70 (Fig. 3d, P > 0.05). These findings suggest that 6-OHDA may significantly enhance CMA activity via LAMP2A in the SN4741 cells and the degradation of Fbw7β induced by 6-OHDA was mainly through CMA pathway.
6-OHDA-induced Fbw7β degradation through LAMP2A-mediated pathway. a 6-OHDA induced LAMP2A expression. Cells were treated with 6-OHDA for 6 h and LAMP2A expression was analyzed by immunofluorescent staining (scale bar 100 μm). The bar graph shows the quantification results (*P < 0.05). b LAMP2A expression after 6-OHDA treatment was determined by western blot. c LAMP2A knockdown blocked 6-OHDA induced Fbw7β degradation. Cells were transfected with LAMP2A siRNA and treated with 6-OHDA (40 μM) for 12 h. The protein levels were determined by western blot. d Hsc70 expression after 6-OHDA treatment did not change. The photographs are representative of three independent experiments. The bar graphs show the quantification of protein expression (mean ± SEM, * P < 0.05)
Oxidative Stress Produced by Neurotoxin 6-OHDA Can Enhance the Combination of Fbw7β and Hsc70 and Promote Its Degradation in Lysosome
To confirm the influences of oxidative modification effect of neurotoxin on CMA and metabolic degradation of Fbw7β, we first tested whether 6-OHDA affects the interaction between Fbw7β and Hsc70. We transfected overexpressed Fbw7β to SN4741 cells and then treated the cells with 6-OHDA for 12 h. The overexpression was confirmed by western blot in Supplementary Fig. S1. After that, cellular lysate and glutathione-S-transferase (GST)-Hsc70 were co-incubated to detect the combination ability of Fbw7β and GST-Hsc70. Results showed that exposure to 6-OHDA enhanced the interaction between Fbw7β and GST-Hsc70 in a pull-down assay (Fig. 4a, P < 0.05). In addition, 6-OHDA significantly increased the interaction between endogenous Fbw7β and Hsc70 in a co-immunoprecipitation assay (Fig. 4b, P < 0.05). These results strongly support the fact that 6-OHDAinduced oxidization of Fbw7β significantly enhances its interaction with Hsc70 and promotes its degradation by CMA.
6-OHDA induced increased binding of oxidized Fbw7β to Hsc70. a 6-OHDA induced increased binding of Fbw7β to endogenous Hsc70. After 18 h of 6-OHDA treatment, Fbw7β co-immunoprecipitation with Hsc70 was conducted. b 6-OHDA induced increased binding of Fbw7β to GST-Hsc70. Cells were transfected with Flag- Fbw7β and treated with 6-OHDA for 18 h. The binding level of Fbw7β was blotted after GST pull-down assay. The photographs are representative of three independent experiments. The bar graphs show the quantification of protein expression (mean ± SEM, *P < 0.05)
Fbw7β Oxidation Level and CMA Changes in 6-OHDA Parkinson’s Disease in Mice as well as in Postmortem PD Patient Brain Samples
To corroborate the findings made in a DA neuronal cell line, we determined Fbw7β oxidation in brain tissues in vivo. We employed 6-OHDA injection in mice through a well-established rodent PD model [27, 28], and the impaired motor activity confirmed that the PD phenotype was successfully induced (Supplementary Fig. S2). Brains of mice were taken 3 days after stereotactic positioned injection of 6-OHDA to mice’s SNc. DNP assay found oxidation level of Fbw7β was obviously increased compared with the control group (injected with saline) (Fig. 5a, P < 0.05). Consistent with our cellular studies, 6-OHDA also caused a decrease in the level of Fbw7β in the mouse SNc (Supplementary Fig. S3A, P < 0.05).
6-OHDA induced oxidation of Fbw7β in mice and oxidation level of Fbw7β in PD patients. a 6-OHDA induced Fbw7β oxidation in the SNc region of mice. The lysates which were prepared from SNc region of 6-OHDA model mice were immunoprecipitated with Fbw7β antibody and blotted for oxidation of Fbw7β. The bar graph was quantitative analysis of density of bands (mean ± SEM, n = 4, *P < 0.05). b Oxidation of Fbw7β increased in the striata of human PD patients. The lysates were prepared from striata of postmortem PD patients and controls. The bar graph was quantitative analysis of the density of bands (mean ± SEM, n = 5 patients and 4 controls, *P < 0.05)
To strengthen these findings, we tested the level of Fbw7β carbonyl oxidation in postmortem PD brain tissues. We immunoprecipitated Fbw7β from control and PD postmortem brain lysates and blotted for carbonyl formation. This analysis showed that the level of Fbw7β carbonyl formation in PD patients was obviously increased compared with the control group (Fig. 5b, P < 0.05). To confirm that the oxidation of Fbw7β was not the result of handling process of the postmortem samples, we test the oxidation level of HMGB1 in the PD and control group. By searching PubMed database, there were no studies showing that the oxidation of HMGB1 changed in PD. The result showed that there was not difference of oxidation level of HMGB1 between these two groups (Supplemental Fig. S4). However, the total level of Fbw7β protein did not change in postmortem PD brains compared with the controls (Supplementary Fig. S3B, P > 0.05). This result indicated that additional mediators were involved in the regulation of Fbw7β expression in PD status and the mechanisms need further investigation. Overall, these studies indicate that Fbw7β is oxidized in vivo in the condition of PD.
Discussions
Oxidative stress is considered a major cause of death and missing of DA neurons, but researchers mainly focused on regulation on damage of mitochondrial function and lacked of studies on regulation mechanism of other organelles [31, 32]. Our study found that the oxidative modification and regulation of neurotoxin-induced oxidative modification on ubiquitin ligase Fbw7β reveal the new mechanism of oxidative stress in regulating cell function.
F-box protein Fbw7β has been shown to regulate the survival of neuron in the nerve cells [25]. Previous study has shown that Fbw7β would produce different forms of changes under a variety of environmental conditions, including oxidative stress [25]. The bulk accumulation of deactivated Fbw7β could induce cell death, and the cells were more sensitive to toxin when mitochondrion Fbw7β level or function was decreased [25]. Our experimental results showed that the 6-OHDA was involved in regulating Fbw7β protein stability. 6-OHDA treatment can lead to decreased Fbw7β expression in SN cells. Further experiments suggested that Fbw7β with oxidative modification induced by 6-OHDA was increased. Our study demonstrated that the neurotoxins-induced oxidative stress led to degradation and dysfunction of Fbw7β.
Previous research found that in the non-neuron cells, a certain degree of oxidative stress can stimulate the production of CMA [33]. Oxidative stress can damage and interfere with many cellular and molecular processes and affect the CMA pathway. In this study, we established cell model of PD with neurotoxin 6-OHDA in dopaminergic neuron SN4741 cells as previously described [34, 35]. Experimental results showed that neurotoxin would also upregulate CMA function in the neurons and increase the LAMP2A expressions—key regulatory factor of CMA. Therefore, 6-OHDA may enhance the function of CMA via up-regulating LAMP2A. The latest study showed that Fbw7β played an important role in neuron survival [25]. In addition to increasing the expression level of LAMP2A to improve the overall activity of CMA, our data showed that 6-OHDA also enhances the combination of Fbw7β with Hsc70, another key regulatory factor of CMA, enabling the delivery of Fbw7β to LAMP2A. On the whole, 6-OHDA improves the CMA-mediated degradation of Fbw7β through two ways: increasing the combination with Hsc70 and increasing the expression of LAMP2A.
We also found that oxidative modification of Fbw7β was increased in brain tissues of PD mice model and PD patients. PD mouse model was successfully established by unilateral stereotactic targeted injection of 6-OHDA to mouse SNc. 6-OHDA induced Fbw7β oxidation in the SNc region of the mouse brain. We also obtained the brain striatum tissue samples from postmortem PD patients and normal control. The levels of oxidized Fbw7β were higher in postmortem PD brains compared with the controls. These results suggest that oxidation-mediated inhibition of Fbw7β may underlie the pathogenic process in PD.
In conclusion, our present study shows that Fbw7β is oxidized and degraded through CMA special pathway. These findings for the first time reveal the specific mechanism of F-box protein ubiquitin ligases, oxidative stress, and CMA-mediated protein degradation, which provides a new theoretical basis for further clarifying the mechanism of PD diseases. Thus, it provides a new thought for the prevention and treatment of PD disease to promote the research on treatment of PD and other neurodegenerative diseases.
6-OHDA, 6-hydroxydopamine; ANOVA, analysis of variance; BIAM, biotinylated iodoacetamide; CMA, chaperone-mediated autophagy; DA, dopamine; DAPI, 4′,6-diamidino-2-phenylindole; DMEM, Dulbecco’s modified Eagle’s medium; DNP, dinitrophenol; Fbw7β, F-box/WD repeat-containing protein 7β; GST, glutathione-S-transferase; HRP, horseradish peroxidase; Hsc70, heat-shock cognate protein 70 kDa; LAMP2A, lysosomal-associated membrane protein 2A; PBS, phosphate-buffered saline; PD, Parkinson’s disease; SCF, Skp, Cullin, F-box containing complex; SDS, sodium dodecyl sulfate; SNc, substantia nigra compacta
References
Kalia LV, Lang AE (2016) Parkinson disease in 2015: evolving basic, pathological and clinical concepts in PD. Nat Rev Neurol 12(2):65–66. doi:10.1038/nrneurol.2015.249
Rizek P, Kumar N, Jog MS (2016) An update on the diagnosis and treatment of Parkinson disease. CMAJ De l'Association Medicale Canadienne 188(16):1157–1165. doi:10.1503/cmaj.151179
Gratwicke J, Jahanshahi M, Foltynie T (2015) Parkinson’s disease dementia: a neural networks perspective. Brain 138(Pt 6):1454–1476. doi:10.1093/brain/awv104
Wang B, Abraham N, Gao G, Yang Q (2016) Dysregulation of autophagy and mitochondrial function in Parkinson’s disease. Transl Neurodegener 5:19. doi:10.1186/s40035-016-0065-1
Kenific CM, Debnath J (2015) Cellular and metabolic functions for autophagy in cancer cells. Trends Cell Biol 25(1):37–45. doi:10.1016/j.tcb.2014.09.001
Lapierre LR, Kumsta C, Sandri M, Ballabio A, Hansen M (2015) Transcriptional and epigenetic regulation of autophagy in aging. Autophagy 11(6):867–880. doi:10.1080/15548627.2015.1034410
Joven J, Guirro M, Marine-Casado R, Rodriguez-Gallego E, Menendez JA (2014) Autophagy is an inflammation-related defensive mechanism against disease. Adv Exp Med Biol 824:43–59. doi:10.1007/978-3-319-07320-0_6
Orogo AM, Gustafsson AB (2015) Therapeutic targeting of autophagy: potential and concerns in treating cardiovascular disease. Circ Res 116(3):489–503. doi:10.1161/CIRCRESAHA.116.303791
Xilouri M, Stefanis L (2010) Autophagy in the central nervous system: implications for neurodegenerative disorders. CNS Neurol Disord Drug Targets 9(6):701–719
Hu Z, Yang B, Mo X, Xiao H (2015) Mechanism and regulation of autophagy and its role in neuronal diseases. Mol Neurobiol 52(3):1190–1209. doi:10.1007/s12035-014-8921-4
Zare-Shahabadi A, Masliah E, Johnson GV, Rezaei N (2015) Autophagy in Alzheimer’s disease. Rev Neurosci 26(4):385–395. doi:10.1515/revneuro-2014-0076
Trinh J, Farrer M (2013) Advances in the genetics of Parkinson disease. Nat Rev Neurol 9(8):445–454. doi:10.1038/nrneurol.2013.132
Martin DD, Ladha S, Ehrnhoefer DE, Hayden MR (2015) Autophagy in Huntington disease and huntingtin in autophagy. Trends Neurosci 38(1):26–35. doi:10.1016/j.tins.2014.09.003
Lee JK, Shin JH, Lee JE, Choi EJ (2015) Role of autophagy in the pathogenesis of amyotrophic lateral sclerosis. Biochim Biophys Acta 1852(11):2517–2524. doi:10.1016/j.bbadis.2015.08.005
Wei K, Wang P, Miao CY (2012) A double-edged sword with therapeutic potential: an updated role of autophagy in ischemic cerebral injury. CNS Neurosci Ther 18(11):879–886. doi:10.1111/cns.12005
Lipinski MM, Wu J, Faden AI, Sarkar C (2015) Function and mechanisms of autophagy in brain and spinal cord trauma. Antioxid Redox Signal 23(6):565–577. doi:10.1089/ars.2015.6306
Hara T, Nakamura K, Matsui M, Yamamoto A, Nakahara Y, Suzuki-Migishima R, Yokoyama M, Mishima K et al (2006) Suppression of basal autophagy in neural cells causes neurodegenerative disease in mice. Nature 441(7095):885–889. doi:10.1038/nature04724
Komatsu M, Waguri S, Chiba T, Murata S, Iwata J, Tanida I, Ueno T, Koike M et al (2006) Loss of autophagy in the central nervous system causes neurodegeneration in mice. Nature 441(7095):880–884. doi:10.1038/nature04723
Kaushik S, Cuervo AM (2012) Chaperone-mediated autophagy: a unique way to enter the lysosome world. Trends Cell Biol 22(8):407–417. doi:10.1016/j.tcb.2012.05.006
Kabuta T, Furuta A, Aoki S, Furuta K, Wada K (2008) Aberrant interaction between Parkinson disease-associated mutant UCH-L1 and the lysosomal receptor for chaperone-mediated autophagy. J Biol Chem 283(35):23731–23738. doi:10.1074/jbc.M801918200
Orenstein SJ, Kuo SH, Tasset I, Arias E, Koga H, Fernandez-Carasa I, Cortes E, Honig LS et al (2013) Interplay of LRRK2 with chaperone-mediated autophagy. Nat Neurosci 16(4):394–406. doi:10.1038/nn.3350
Bejarano E, Cuervo AM (2010) Chaperone-mediated autophagy. Proc Am Thorac Soc 7(1):29–39. doi:10.1513/pats.200909-102JS
Blesa J, Trigo-Damas I, Quiroga-Varela A, Jackson-Lewis VR (2015) Oxidative stress and Parkinson’s disease. Front Neuroanat 9:91. doi:10.3389/fnana.2015.00091
Chaturvedi RK, Flint Beal M (2013) Mitochondrial diseases of the brain. Free Radic Biol Med 63:1–29. doi:10.1016/j.freeradbiomed.2013.03.018
Ekholm-Reed S, Goldberg MS, Schlossmacher MG, Reed SI (2013) Parkin-dependent degradation of the F-box protein Fbw7beta promotes neuronal survival in response to oxidative stress by stabilizing Mcl-1. Mol Cell Biol 33(18):3627–3643. doi:10.1128/MCB.00535-13
Cai Z, Zeng W, Tao K, E Zhen, Wang B, Yang Q (2015) Chaperone-mediated autophagy: roles in neuroprotection. Neurosci Bull 31 (4):452–458. doi:10.1007/s12264-015-1540-x
Johansson S, Lee IH, Olson L, Spenger C (2005) Olfactory ensheathing glial co-grafts improve functional recovery in rats with 6-OHDA lesions. Brain 128(Pt 12):2961–2976. doi:10.1093/brain/awh644
Magraoui FE, Reidick C, Meyer HE, Platta HW (2015) Autophagy-related deubiquitinating enzymes involved in health and disease. Cell 4(4):596–621. doi:10.3390/cells4040596
Chen S (2014) Chinese guidelines for the diagnosis criteria and treatment of Parkinson’s disease (3rd edition). Chin J Neurol 47(6):428–433. doi:10.3760/cma.j.issn.1006-7876.2014.06
Gao L, She H, Li W, Zeng J, Zhu J, Jones DP, Mao Z, Gao G et al (2014) Oxidation of survival factor MEF2D in neuronal death and Parkinson’s disease. Antioxid Redox Signal 20(18):2936–2948. doi:10.1089/ars.2013.5399
Song J, Kim J (2016) Degeneration of dopaminergic neurons due to metabolic alterations and Parkinson’s disease. Front Aging Neurosci 8:65. doi:10.3389/fnagi.2016.00065
Subramaniam SR, Chesselet MF (2013) Mitochondrial dysfunction and oxidative stress in Parkinson’s disease. Prog Neurobiol 106-107:17–32. doi:10.1016/j.pneurobio.2013.04.004
Saha T (2012) LAMP2A overexpression in breast tumors promotes cancer cell survival via chaperone-mediated autophagy. Autophagy 8(11):1643–1656. doi:10.4161/auto.21654
Zhang S, Gui XH, Huang LP, Deng MZ, Fang RM, Ke XH, He YP, Li L et al (2016) Neuroprotective effects of beta-asarone against 6-hydroxy dopamine-induced parkinsonism via JNK/Bcl-2/Beclin-1 pathway. Mol Neurobiol 53(1):83–94. doi:10.1007/s12035-014-8950-z
Huang L, Xue Y, Feng D, Yang R, Nie T, Zhu G, Tao K, Gao G et al (2017) Blockade of RyRs in the ER attenuates 6-OHDA-induced calcium overload, cellular hypo-excitability and apoptosis in dopaminergic neurons. Front Cell Neurosci 11:52. doi:10.3389/fncel.2017.00052
Acknowledgements
The authors thank Ms. Lin Zhang for her English proof reading. This project was supported by grant from the National Natural Science Foundation of China (81471377).
Author information
Authors and Affiliations
Contributions
Xiufeng Wang, acquisition of data, literature research, and manuscript preparation; Heng Zhai, acquisition of data; Fang Wang, study concept and design, critical revision of the manuscript for important intellectual content, and study supervision.
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Electronic Supplementary Material
Supplementary Fig. S1
The overexpression of Fbw7β was confirmed by western blot. SN4741 cells were inoculated into six-well plate to culture till 80% fusion and the pGCSIL vector encoding the mouse Fbw-7β and control was transfected into SN4741 cells (JPEG 62 kb)
Supplementary Fig. S2
Motor activity examination after 6-OHDA lesioning. Amphetamine (2.5 mg/kg)-induced rotations were counted after 6-OHDA lesioning, as described in the supplementary methods. a Time course of rotational behavior. b Total number of rotations in 60 min. Values shown are means ± SD of three experiments (total nine mice in each group). * P < 0.05 (JPEG 181 kb)
Supplementary Fig. S3
Levels of Fbw7β in the SNc region of 6-OHDA-treated mice and PD patients. a Lysates prepared from 6-OHDA-treated mice brains were analyzed for Fbw7β by western blot (* P < 0.05). b Fbw7β levels in the brain of human PD patients analyzed by western blot (JPEG 216 kb)
Supplementary Fig. S4
The oxidization level of HMGB1 in PD patients and control. The lysates were prepared from the striata of postmortem PD patients and controls (JPEG 56 kb)
ESM 1
(DOCX 15 kb)
Rights and permissions
About this article
Cite this article
Wang, X., Zhai, H. & Wang, F. 6-OHDA Induces Oxidation of F-box Protein Fbw7β by Chaperone-Mediated Autophagy in Parkinson’s Model. Mol Neurobiol 55, 4825–4833 (2018). https://doi.org/10.1007/s12035-017-0686-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-017-0686-0