Abstract
One of the shared hallmarks of neurodegenerative diseases is the accumulation of misfolded proteins. Therefore, it is suspected that normal proteostasis is crucial for neuronal survival in the brain and that the malfunction of this mechanism may be the underlying cause of neurodegenerative diseases. The accumulation of amyloid plaques (APs) composed of amyloid-beta peptide (Aβ) aggregates and neurofibrillary tangles (NFTs) composed of misfolded Tau proteins are the defining pathological markers of Alzheimer’s disease (AD). The accumulation of these proteins indicates a faulty protein quality control in the AD brain. An impaired ubiquitin-proteasome system (UPS) could lead to negative consequences for protein regulation, including loss of function. Another pivotal mechanism for the prevention of misfolded protein accumulation is the utilization of molecular chaperones. Molecular chaperones, such as heat shock proteins (HSPs) and FK506-binding proteins (FKBPs), are highly involved in protein regulation to ensure proper folding and normal function. In this review, we elaborate on the molecular basis of AD pathophysiology using recent data, with a particular focus on the role of the UPS and molecular chaperones as the defensive mechanism against misfolded proteins that have prion-like properties. In addition, we propose a rational therapy approach based on this mechanism.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Alzheimer’s disease (AD) is an incurable, progressive brain degeneration that results in a loss of higher cognitive function, memory impairment, language deterioration, depression, and other manifestations. This disease is named after a German neuropathologist, Alois Alzheimer, who first presented the case of his patient Auguste Deter in 1906 [1, 2]. An increase in human life expectancy has coincided with a rapid increase in neurodegenerative disease incidence in recent years. A recent statistical study projected that in 2050 approximately 13.8 million people in the USA will suffer from AD unless preventive measures are taken. AD has gained widespread attention because of its high economic costs, which have reached $400 billion per year in the USA alone, as well as the social costs, which are more difficult to quantify [3, 4].
AD-related pathological markers include a progressive death of neurons in specific areas with an accumulation of intracellular neurofibrillary tangles (NFTs) and extracellular depositions of amyloid plaques (APs). NFTs are composed of the misfolded hyperphosphorylated microtubule-associated protein Tau (MAPT or Tau), whereas APs are extracellular deposits of misfolded and aggregated amyloid-beta peptides (Aβ). Because both NFTs and APs are persistently found in areas with severe neuronal death, these proteins were considered to be the main cause of neuronal loss and the emergence of dementia, which is a crucial symptom of AD; however, numerous drug trials based on these proteins have failed to provide a useful AD therapy [5, 6]. A postmortem study demonstrated that the misfolded protein accumulation is a shared pattern in many neurodegenerative diseases, including AD [7, 8]. Therefore, it was concluded that an accumulation of misfolded proteins is the potential cause of neurodegeneration in AD.
Here, we review recent global gene expression and proteomic studies of AD brains versus normal brains and in AD-affected regions versus unaffected regions. The obtained data underline the crucial role of protein regulation systems, especially the ubiquitin-proteasome system (UPS), as well as molecular chaperones and co-chaperones. These interconnected systems reveal how AD pathology commences at the molecular level, long before the development of any symptoms. This review also discusses potential areas for AD research that focus on protein regulatory mechanisms as the basis for novel AD therapeutics.
AD Development in the Brain
The development of AD pathology is not isolated to one specific brain region; rather, it spreads throughout nearly the entire brain in a predictable pattern [9]. The earliest area affected by NFTs in AD is the medial temporal lobe, beginning at the transentorhinal region [9–12]. Subsequently, the limbic system, which is composed of the hippocampus and amygdala, followed by the parahippocampal area and frontal lobe exhibit NFT accumulation [12]. AD lesions are found in the most susceptible regions even at the presymptomatic stage. This fact was further confirmed by magnetic resonance imaging (MRI) scans that indicate the expansion of cerebrospinal fluid (CSF), which correlates with brain volume loss in the temporal lobe area [13]. Moreover, presymptomatic individuals and mildly affected patients show a significant rate of volume loss in the hippocampal area. As the disease progresses, the inferior and lateral temporal lobes begin to show accelerated atrophy rates. This observation suggests that atrophy begins from the medial temporal lobe area and spreads to the adjacent inferior and temporal lobes [14]. The mechanism of how misfolded proteins spread to other parts of the brain remains elusive; however, we address the hypothesis that this mechanism involves a prion-like transmission [15].
Genetics of AD
For a small proportion of cases (~5 %), AD is caused by a genetic mutation in the genes involved in processing AD hallmark proteins, including the Aβ precursor protein (APP) and presenilin-1 and presenilin-2 (PS1 and PS2, respectively) [16, 17]. The genetic form of AD is referred to as familial AD or early-onset AD (EOAD), whereas the sporadic form of AD is referred to as late-onset AD (LOAD) [18]. LOAD is the most prevalent type of AD and is likely caused by normal aging [19, 20] and its associated consequences, such as increased protein misfolding, oxidative stress, and perturbed cell regulation [21, 22]. Familial AD or EOAD is often diagnosed at ages younger than 65 years, whereas LOAD is commonly diagnosed in patients older than 65 years [2, 17, 20].
A possible link between APP and AD emerged in the late 1980s [23–26]. The evidence for a pivotal genetic role of APP in the progression of AD pathology stemmed from the shared APs with Down’s syndrome [27, 28]. Amino acid (aa) sequencing of Down’s syndrome-derived Aβ indicated results similar to AD, which suggests that genetics plays a role in the formation of APs [28, 29]. Down’s syndrome is characterized by trisomy of chromosome (chr) 21; as a consequence, the extra dose of the APP gene contributes to the lifelong overexpression of Aβ [30, 31]. This accumulation of Aβ in patients with Down’s syndrome often results in the early appearance of APs, which are an AD hallmark [32, 33]. These data underline the importance of AP formation in the development of AD-like pathology.
Mutant APP770 (V717I) was the first genetic mutation reported in AD [34, 35]. To date, various mutations in the APP gene have been identified in EOAD cases [36–45]. The exact mechanism by which APP mutations promote AD remains unknown. However, it has been demonstrated that mutations in close proximity to secretase cleavage sites can alter the cleavage and processing of APP, which may favor the production of the highly amyloidogenic Aβ42 fragment [46, 47].
Even though the APP mutation is the first AD-related mutation that was discovered, mutations in the PS1/2-encoding genes are the most prevalent genetic cause of EOAD, with the majority of mutations in PS1 [48]. This issue was raised after a reported poor correlation between EOAD and a specific genetic marker in the APP region on chr 21 [49, 50]. It was subsequently discovered that missense mutations in the PS-encoding genes shift the production of Aβ toward the AP-promoting toxic Aβ species [51, 52]. The role of the PS1/2 in the production of Aβ will be discussed later in this review. Extensive studies on PS mutants and gene knock-out models have provided a clear understanding of its role in the development of APs, and various mutations of the PS1/2 genes have been identified and reviewed elsewhere [53]. Another gene linked to AD is apolipoprotein E4 (APOE4). Mutations in the APOE4 gene, while not directly involved in the development of AD, are a genetic factor that increases the risk of AD up to 15-fold [54, 55]. The APOE4 mechanism that affects AD is not in the scope of this review; thus, readers are directed to other review papers [56, 57]. A frame-shifted mutant ubiquitin B (UBB) gene, UBB+, has also been documented to produce an AD phenotype [58, 59]. However, UBB is not directly involved in the current major hypothesis for AD pathogenesis, including the AP or NFT hypotheses. A more comprehensive assessment of UBB+ and its pathophysiological significance in AD is discussed later in this review.
APs
APs, one of the two major hallmarks of AD, predominantly consist of 40 or 42 aa Aβ (Aβ40–42). These Aβ species are commonly found in postmortem AD brains and are neurotoxic [28, 29, 60, 61]. The Aβ peptide is a product of regulated intramembrane proteolysis (RIP) of the APP protein, which is ubiquitously and constitutively expressed in many human cells [62]. APs are also often associated with dystrophic neurites, fibrillar paired helical filaments (PHFs) of hyperphosphorylated Tau, and activated glial cells [63]. The frequency of APs in AD brains led to the development of the “amyloid cascade hypothesis.” This idea has been the de facto foundation of the majority of AD research for the previous 20 years and has been previously reviewed by Selkoe [61, 64]. Diffuse plaques are a subtype of APs that are more granular and less compacted than the APs commonly found in a normal aging brain; furthermore, these plaques do not induce neuronal death or microglia activation [65, 66]. Accordingly, diffuse plaques are considered a presymptomatic sign of AD and can be observed in human brain biopsies and animal models long before the development of a full AD phenotype. To understand how Aβ is formed, we must first briefly discuss APP, the precursor protein of Aβ.
APP
APP is a type 1 transmembrane glycoprotein with hypothesized functions as a cell surface receptor [23, 67]. It has heterogeneous molecular weights between 110 and 135 kDa because of various posttranslational modifications (e.g., N and O glycosylation, phosphorylation, and sulfation) [2, 68]. APP is also a target for alternative splicing, which results in six isoforms (mainly 695, 751, and 770 aa residues) [2, 62, 69–71]. Neurons specifically express a high level of the APP695 isoform, which is not ubiquitously expressed in other cell types [72]. The difference between the APP695 and APP751 isoforms is the presence of a 56 aa Kunitz-type serine protease inhibitor (KPI) motif [70, 73, 74].
A structural analysis of APP has identified several domains; however, the fundamental physiological role of this protein remains unclear. As a transmembrane protein, APP has extracellular, transmembrane, and intracellular regions. Several domains have been identified within these regions, such as the heparin-binding domain (HBD), the Cu(II)-binding domain (CUBD), and the RERMS (Arg-Glu-Arg-Met-Ser) sequence [75–79]. These findings suggest that the physiological function of APP and its derived domains are related to neurotrophic cell signaling, synaptic plasticity, and oxidative defense [80–84]. For example, a recent study demonstrated that the HBD of sAPPα has neuroprotective properties in traumatic brain injury [85].
Regulated Intramembrane Proteolysis of APP
RIP is the processing of APP involving serial cleavages at different sites, which results in the production of several possible fragments, including non-amyloidogenic and soluble APP fragments (sAPPα and sAPPβ), an APP intracellular C-terminal domain (AICD), and amyloidogenic Aβ, as determined by the specific site of APP cleavage. This cleavage occurs at three different sites according to the activities of three different enzymes: α-, β-, and γ-secretases (Fig. 1). RIP primarily occurs intracellularly within the trans-Golgi network (TGN) and the endoplasmic reticulum (ER) compartment and partially on the cell surface [86–89].
RIP of APP. APP has α-, β-, and γ-secretase cleaving sites and a KPI domain (left). APP can be cleaved in a non-amyloidogenic pathway by α-secretase, thus producing sAPPα (top right). Amyloidogenic processing of APP by sequential cleaving with β- and γ-secretases generates sAPPβ, Aβ, and the nuclear signaling molecule AICD (bottom right)
The first possible product of RIP is sAPPα [90], which is produced when APP is cleaved by α-secretase. The proposed biological functions of sAPPα are related to neuritogenesis/axonal outgrowth, cell proliferation, glial differentiation, and memory and learning-related processes, such as long-term potentiation (LTP), metal homeostasis, and neuroprotection [81, 91–97]. The detailed physiological role of sAPPα has been extensively reviewed by Chasseigneaux and Allinquant in 2012 [91].
α-Secretase is a zinc metalloproteinase from the A disintegrin and metalloproteinase (ADAM) family that cleaves native APP within the transmembrane region and mediates the shedding of sAPPα from the cell membrane. Because α-secretase cleaves APP within the Aβ region (i.e., K16 of Aβ or aa residue number 687 of APP770), the α-cleavage of APP does not produce an amyloidogenic product [98–100].
ADAM family proteins play an important role in the ectodomain shedding events of diverse biological molecules (e.g., cytokines, growth factors, receptors, and other signaling molecules) and have recently been suggested to be involved in defense mechanisms against cellular prion proteins (PrPs) [101–103]. A comprehensive list of ADAM substrates has been reviewed by Huovila et al. [104]. ADAM family proteins are crucial mediators during the development of the central nervous system (CNS), with effects that largely mediate Notch signaling [105, 106].
The knock-out of ADAM10 in transgenic mice results in multiple defects, including the disruption of Notch signaling, which ultimately induce prenatal death; however, fibroblast cells retain normal APP processing, which suggests ADAM activity is tissue-specific and ADAMs may compensate for one another [105–107]. The overexpression of ADAM10 in transgenic mice results in the competitive cleavage of APP and release of sAPPα, thus reducing the formation of Aβ and its downstream pathology [108]. Furthermore, the in vivo administration of an ADAM10 inhibitor peptide in mice successfully mimicked sporadic AD-like pathology [109].
ADAM17, also known as tumor necrosis factor-α converting enzyme (TACE), cleaves APP and Notch [110]. ADAM17 is also responsible for the shedding of cell adhesion proteins, cytokine and growth factor receptors, and many ligands of the epidermal growth factor receptor (EGFR) [111, 112]. In human primary neuron cultures, an ADAM17 inhibitor prevents the α-secretase activity of ADAM17 for APP shedding [113]. Another ADAM protein, ADAM9, has also been reported to have α-secretase activity for APP [114].
sAPPβ is the second possible product of APP proteolysis, which occurs inside the TGN and plasma membrane [89, 115]. It is produced when native APP is cleaved by β-secretase. The sAPPβ and sAPPα isoforms have identical sequences with the exception that sAPPβ is missing the last 16 aa residues at its C terminus. The functions of sAPPα and sAPPβ are generally similar because their common domains are identical. However, sAPPβ has significantly lower neuroprotective properties and is not involved in LTP because of the missing aa residues at the C terminus [116]. In neurons, the most significant β-secretase is the aspartyl protease beta-site APP cleaving enzyme 1 (BACE1). The cleavage site of BACE1 is in the N terminus of the resulting Aβ, between residues 671 and 672 of APP770. The β-secretase activity is regulated through binding to other proteins, such as the reticulon family proteins (RTNs). A co-localization study demonstrated that RTN3 is a BACE1-binding partner in neurons and negatively modulates BACE1 activity [117], and a gene expression study showed that RTN3 is under-expressed in AD brains [118]. Thus, the downregulation of RTN3 may result in increased Aβ production in neurons [119].
Amyloidogenic Aβ is produced when APP undergoes serial cleavages by β- and γ-secretases. γ-Secretase is a multi-subunit protease complex that consists of PS1 or PS2, nicastrin, anterior pharynx-defective 1 (APH1), and presenilin enhancer 2 (PEN2) [120, 121]. Upon β-/γ-cleavage, the γ-secretase cleaves APP within the transmembrane domain, thus producing an Aβ fragment that ranges from 38 to 43 aa residues in length. It has been suggested that the GxGD motif of PS1 is the active site for APP cleavage specificity [122]. A subsequent study indicated that a missense mutation in L383 (i.e., x residue of the GxGD motif) of PS1 shifted APP cleavage to produce Aβ42 [123]. This mechanism may explain why the modulation of PS1 activity or a mutation in the PS1 gene favors the production of Aβ42.
The majority of Aβs found in APs are in the Aβ40 form [124]; however, it is the Aβ42 fragment that has been highly studied in AD research because of its potency to induce aggregation and neurotoxicity [125]. Aβ42 is considered more hydrophobic than Aβ40 due to its two extra stretches of hydrophobic residues; thus, it more readily forms a β conformation [126, 127]. The elevated hydrophobicity increases the potential of Aβ42 to form insoluble aggregates and consequently promotes its toxicity [128–130]. Under normal physiological conditions, the Aβ42/Aβ40 ratio is maintained at approximately 1:9; however, the level of Aβ42 increases in AD pathology. This perturbed Aβ42/Aβ40 ratio is thought to contribute to the formation of neurotoxic aggregates, which eventually cause cell death [36, 51, 131, 132].
AICD
AICD is another product of γ-secretase cleavage of APP. AICD has properties similar to the Notch intracellular domain (NICD), which is also mediated by γ-secretase [105]. Therefore, it has been hypothesized that AICD may play a pivotal role in neuron signaling pathways. For example, AICD was reported to be transported into the nucleus by an FE65 adaptor protein (otherwise known as APBB1) and subsequently connected to TIP60 (official gene symbol KAT5) to form the AICD-FE65-TIP60 (AFT) complex, which regulates gene transcription [133–137]. The AFT complex is transcriptionally active via the histone acetylation activity of TIP60 [138] with vast downstream target genes, such as lipoprotein receptor-related protein 1 (LRP1) [139], APP [140], glycogen synthase kinase 3β (GSK3β) [141, 142], p53 [143], and EGFR (Fig. 2) [144].
AICD signaling from the membrane to the nucleus. AICD is released through RIP by γ-secretase. One known trigger for this process is TAG1 binding to APP. Released AICD binds to the adaptor protein FE65; it subsequently translocates to the nucleus, where these proteins bind to TIP60 and form the AFT complex to regulate gene expression. GSK3β, APP, p53, and DAB1 are overexpressed by aberrant AICD signaling, whereas the expression of LRP1, EGFR, and neurogenesis genes are repressed
The activation of AICD signaling can produce changes in gene expression and cell proteome metabolism, such as the upregulation of DAB1, an AICD interacting protein, in AD frontal cortex brain samples [145]. Moreover, it has been suggested that AICD can mediate neuron-specific apoptosis [146]. AICD also binds to Numb, a Notch inhibitor, and competes with the Notch signaling pathway [147]. Ma et al. showed that AICD release increased after the binding of the transient axonal glycoprotein 1 (TAG1), an extracellular ligand to the extracellular part of APP [67]. This study also demonstrated that released AICD binds to FE65 and negatively modulates neurogenesis. TAG1 is a member of the F3 protein family and is a functional ligand for the Notch receptor [148]. The overexpression of AICD in mice has been reported to induce AD-like characteristics because of the overactivation of GSK3β, which induces the hyperphosphorylation of Tau [149]. This study also found that AICD levels are increased in the brains of AD patients. Thus, aberrant AICD signaling may contribute to the development of AD pathophysiology [150].
Physiological Function of Aβ
Despite its strong link to neurodegeneration in AD, Aβ has exhibited various neuroprotective and neurotrophic effects at suboligomer-promoting concentrations [91, 151, 152]. A study reported that the Aβ1–28 fragment has a neurotrophic effect on hippocampal pyramidal neurons in vitro [152]. Soluble monomeric Aβ40 can induce neurogenesis in neural progenitor cells, whereas Aβ42 favors astrogenesis [153]. Monomeric Aβ40 has been demonstrated to be involved in insulin/insulin-like growth factor (IGF-1) receptor signaling and displays neuroprotective properties [154]. Additional interesting results have been discussed by Calafiore et al. [155]. This group showed that the Aβ25–35 fragment and neurotoxic Aβ42 are not harmful to neural precursor cells (NPC). Furthermore, this study indicated that Aβ increased neurogenesis. A similar result was achieved in another study which suggested that the neurogenic effect of the Aβ42 oligomer is mediated by the mitogen-activated protein kinase (MAPK) pathway [156].
Aβ Pathophysiology
Many hypotheses speculate how APs and Aβ induce cell death, including mechanisms related to oxidative stress, calcium imbalance, mitochondrial damage, the disruption of energy production, the phosphorylation of Tau, and UPS impairment [157–163]. The potential role of oxidative damage as a downstream process in AD pathogenesis has garnered a great deal of interest [164, 165]. This interest is mainly because the brain is the most highly oxygen-metabolizing organ in the body; thus, it is highly susceptible to oxidative damage [166–170]. Moreover, the brain has a high lipid content that is prone to oxidation [165, 167]. Aβ-induced free radical production promotes cell-damaging species that cause oxidized proteins, oxidized DNA, and alterations in brain phospholipids. A relationship between Aβ and free radical-mediated oxidative stress has been proven in vitro and in vivo [158, 159, 171–173]. However, despite the plethora of data that support this hypothesis, some studies have reached a contrasting conclusion [174–177]. Moreover, the role of antioxidants in AD pathology remains unclear [178]. These data suggest that the direct and specific role of Aβ is rather marginal; thus, the correlation between Aβ and neurodegeneration is not as direct as previously suggested [179–181] and has been described with an “airbag” analogy [182]. A more convincing hypothesis suggests that Aβ induces Tau hyperphosphorylation, which enhances Tau aggregation and leads to neurodegeneration [163, 183–189].
AD and NFTs
Another significant AD pathophysiology hallmark is the NFTs, which are filamentous aggregates of the intracellular insoluble hyperphosphorylated Tau protein [190, 191]. Apart from AD, NFTs have been linked to other tauopathies, including frontotemporal dementia and Pick’s disease [192–194]. NFTs were first described in the 1980s and consist of smaller units referred to as PHFs [195]. PHF is a fibril-like aggregation of proteins, which was subsequently discovered to be a wound form of the Tau protein [196, 197].
Tau and Its Physiological Functions
Tau was first characterized in the 1970s from a porcine brain as a protein that binds to tubulin and is essential for microtubule formation [198–200]. In addition to its significance in AD, Tau mutations are responsible for the frontotemporal dementia related to chr 17 (FTDP-17; human Tau is located on chr17q21). Tau is crucial for the formation of a functional microtubule system and is almost exclusively expressed in neurons, particularly in axons [199, 201–203]. Tau is highly soluble and rich in phosphorylation-prone aa residues. Based on its native function, Tau is involved in the stabilization of the axonal microtubule architecture for neurite extension, the establishment of cell polarity, and neuronal axonal transport [204–209]. The role of axonal transport in neurons is critical for the maintenance of axons, synapses, and synaptic activities [210]. Because neuronal function is sensitive to any disturbance of the microtubule-based system, errors in the axonal transport system have been clearly linked to neurodegenerative conditions [211, 212].
The Tau protein has a few notable regions [213, 214]. One region is the tubulin-binding repeat domain, which is highly conserved and spans the C-terminal half of the protein. Some isoforms have three of this repeat, whereas other isoforms have four; thus, these isoforms are often referred to as 3R or 4R Tau. Adjacent to this domain are basic proline-rich repeats, which promote Tau protein binding using tubulin-binding repeats that bind to specific pockets of positively charged prolines attached to the negatively charged microtubule surface. This second domain is in the N-terminal region of Tau, which has either one or two highly acidic sets of 29 aa residues depending on the isoform. This domain is also presumed to protrude because its negatively charged region repels it from the microtubule surface [213]. These various domain combinations generate the six known Tau isoforms that are generated by alternative splicing during mRNA maturation [215]. These isoforms, which range from 352 to 451 aa in length, were characterized from PHFs [215], and all six isoforms have been demonstrated to be aberrantly phosphorylated [216].
AD and the Pathophysiology of Tau Misfolding
Another striking characteristic of the Tau protein is its hyperphosphorylation. As previously described, hyperphosphorylated misfolded Tau causes fibrillization and subsequent aggregation in AD pathophysiology. In the 441 aa residues of each Tau isoform, 79 aa residues of serine or threonine can be potentially phosphorylated [215]. Numerous kinases are linked to Tau phosphorylation, both from proline-directed kinases and non-proline-directed kinases. The major known Tau kinases are GSK3, cyclin-dependent kinase 5 (CDK5), MAPKs, protein kinase A (PKA), protein kinase C (PKC), calcium/calmodulin-dependent protein kinase II (CaMKII), casein kinase II (CKII), MAP/microtubule-affinity-regulating kinases (MARKs), dual-specificity tyrosine-regulated kinase 1A (Dyrk1A), and the AMP-activated kinase (AMPK) [186, 187, 217–222]. As a result of the diverse potential of phosphorylating enzymes, Tau phosphorylation can be a downstream of a myriad of events, including Ca2+-mediated toxicity [223, 224], impaired insulin signaling [225, 226], and brain metabolic deficiency [227]. The hyperphosphorylation of Tau alters its binding to microtubules and promotes its disassociation [162, 228, 229]. Phosphorylated Tau, however, is also subject to various protein phosphatases, such as PP1, PP2A, PP2B, and PP5 [230]. In vitro experiments have demonstrated that Tau dephosphorylation can restore its ability to bind to microtubules [162]. Other than phosphorylation, Tau is a known target for acetylation, deamidation, and ubiquitination [231–234].
Furthermore, the Tau protein is also classified as an intrinsically disordered protein (IDP). Similar to other IDPs, Tau is highly hydrophilic, which easily prevents appropriate folding and a lack of a well-defined three-dimensional structure under physiological conditions [235]. IDPs have been reported to play key roles in complex diseases, including cancer, cardiovascular disease, amyloidosis, neurodegenerative diseases, and diabetes [236]. Interestingly, IDPs are involved in most of the other major neurodegenerative diseases, for example, α-synuclein in Parkinson’s disease (PD), the PrP in transmissible spongiform encephalopathies, polyglutamine in Huntington’s disease (HD), mutant superoxide dismutase 1 (SOD1), and TAR DNA-binding protein of 43 kDa (TDP-43), as well as several others in amyotrophic lateral sclerosis (ALS) [237–241].
Transmission of Aβ and Tau
Studies suggest that misfolded proteins can be transmitted to adjacent cells and ultimately to other brain regions. However, the exact mechanism of this phenomenon is still unclear. One hypothesis is that the misfolded protein acts as a prion-like particle that can replicate and transmit itself to an adjacent cell, where it initiates the nucleation process of protein aggregation in the receiving cell [15, 242–248]. It has been proposed that a prion-like transmission of misfolded proteins (Aβ or Tau) from neuron to neuron could be responsible for the progressive neurodegeneration in AD, which also appears in the absence of Tau aggregation in the transentorhinal region and cortical Aβ pathology. This prion-like transmission may subsequently trigger misfolded Tau accumulation even during the asymptomatic phase of the disease [246, 249].
The misfolded protein transmission hypothesis is supported by data which indicate that the injection of brain extracts, which contain clumps of misfolded amyloidogenic Aβ into a normal mouse brain, triggers the spread of APs [250–252]. Similar experimental results have demonstrated that injected Tau protein triggered extracellular protein aggregates that were internalized by the cells, thereby inducing further Tau aggregation [253–255]. Firm evidence has indicated that mutant Tau can induce the aggregation of normal Tau in the human Tau-expressing mouse brain. More importantly, injected Tau was able to transmit the aggregation to different brain regions [253]. Furthermore, the newly aggregated human Tau can induce Tau aggregation when serially injected in healthy wild-type mouse brains [256].
The prion-like propagation of properties of misfolded Tau is also shared with IDPs in other neurodegenerative diseases, for example, α-synuclein in PD [257–259], polyglutamine in HD [260], and mutant SOD1 in ALS [261]. TDP-43, an RNA binding protein that is involved in ALS and frontotemporal lobar degeneration with ubiquitin (FTLD-U or FTLD-TDP), also exhibits prion-like transmission [240].
The propagation mechanism of misfolded proteins from one cell to another cell is unclear. Currently, two mechanisms of misfolded protein propagation have been proposed. The first mechanism involves the release and uptake of misfolded proteins [244, 262] that were potentially released through membrane damage [260], cell lysis [260], or exocytosis [263, 264]. These first two release options likely contribute to AD pathology via trauma and aging, whereas the third release option may be relevant in cases of neuronal synaptic transmission [264]. The evidence for the hypothesis of this mechanism has been demonstrated for Aβ [251] and Tau [253, 254]. The second mechanism uses a direct cell-to-cell propagation via tunneling nanotubes [244, 265], and it has been observed using a fluorescent-labeled PrP related to Creutzfeldt-Jakob disease (CJD); however, this second mechanism has not been shown in AD to date [246, 265].
Taken together, these data suggest that there is a strong correlation between proteinopathies, prion-like transmission, and AD; these findings imply that the impairment of the UPS, which is responsible for the removal of misfolded proteins, such as prion-like aggregates, occurs at an early stage of AD [15, 246, 266]. In addition, other pivotal proteins of the chaperone/proteasomal pathways, such as heat shock proteins (HSPs), FK506-binding proteins (FKBPs), and tripartite motif (TRIM) family proteins, were also altered in AD [118, 267].
The UPS
Protein degradation was initially thought to be an unregulated, non-specific, lysosome-dependent process. However, the discovery of the UPS has altered the paradigm of protein degradation because this alternative pathway provides a system with high specificity and involves a complex control mechanism. The UPS was first described in 1979 with the discovery of the 26S proteasome, an active proteolytic protein complex [268, 269].
The UPS is responsible for degrading the majority of cellular proteins and maintaining protein homeostasis [270]. It has been conserved during evolution because of its importance in cell regulation; thus, it is not surprising that it has approximately 1,000 components (which compose approximately 5 % of the genome), and the aberration of its function can lead to many diseases, including neurodegeneration [271]. The UPS is distinct from the other proteasome systems (e.g., lysosomal proteases) because it is ATP- and ubiquitin-dependent. The (poly)ubiquitination process initiates this cascade by covalently linking an ubiquitin molecule to a target protein. Subsequently, additional ubiquitin molecules are linked to form a polyubiquitin chain.
Ubiquitin is an 8.5 kDa protein composed of 76 aa. The ubiquitin protein is transcribed from UBB, ubiquitin C (UBC), RPS27a, and UBCEP2 genes. However, only the first two genes encode for a polyubiquitin precursor that is significant in the UPS signaling cascade. The other genes encode an ubiquitin that fuses to ribosomal proteins [272, 273]. The regulation of the attached protein depends on which ubiquitin lysine residues are linked. In the context of protein degradation, K-48 is the most significant residue because it signals for anchored protein degradation via the UPS. Other polyubiquitin functions may involve DNA repair, cell cycle regulation, lysosomal degradation, kinase modification, and endocytosis [273].
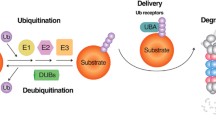
The 26S proteasome is the proteolytic complex of the UPS system, which is composed of a single 20S core catalytic protein and capped with 19S regulatory proteins. The ubiquitination of proteins occurs posttranslationally. The cascade is initiated with the activation of an ubiquitin molecule catalyzed by an E1 ligase, the activating enzyme that forms an ubiquitin-adenylate intermediate. This process is followed by a reaction of the intermediate bound to an E1 cysteine residue to form a high-energy thioester bond, allowing the ubiquitin molecule to be transferred to E2, a conjugating enzyme. The third enzyme, E3 ligase, acts as a substrate-recognition enzyme bound to the target protein. Finally, ubiquitin is transferred from the E2 enzyme to a lysine in the target protein (Fig. 3) [273, 274]. The E3 ligase is a pivotal enzyme in the UPS cascade and is crucial for substrate specificity. E3 plays a critical role in selecting the substrates, and the regulation of either its catalytic activity or substrate interaction properties is important for the UPS signaling pathway [273, 274].
The ubiquitin-proteasome system. The ubiquitin-proteasome degradation cascade begins when E1 activates an ubiquitin molecule at its lysine residue to create a high-energy thioester bond. The ubiquitin is then transferred to a reactive cysteine residue of the E2 enzyme. Meanwhile, the E3 enzyme (e.g., TRIM32/37) is bound to a target protein and moves it to the E2-ubiquitin, which transfers the ubiquitin molecule to the lysine residue of the target protein. This process is repeated for several rounds to obtain a polyubiquitinated target protein. This target protein is then recognized by the 26S proteasome and undergoes degradation
The repetitive addition of ubiquitin moieties to the target protein results in polyubiquitination. As a result of its pivotal role in maintaining protein homeostasis, any impairment of the UPS can cause disease; thus, the UPS plays an important role in multiple diseases, including neurodegenerative pathologies in the CNS [275].
The TRIM family proteins are E3 ligases with a RING finger motif, which have ubiquitin ligase activity and are involved in various cellular processes, including apoptosis, tumorigenesis, and the viral response [276–278]. For example, the mutation of TRIM37 is linked to Mulibrey nanism [279], whereas TRIM32 is associated with skin carcinogenesis and is highly expressed in skin-derived tumors [280]. TRIM32 is also involved in mouse neural-progenitor cell signaling during neuronal differentiation [281] and in skeletal muscle remodeling and myogenic differentiation [282]. TRIM32 plays an important role in the proper formation of neurofilaments and axons. In addition, the knock-out of TRIM32 in mice resulted in reductions in the diameter of myelinated axons and low expression of type II (fast) myosin isoforms [283]. The knock-out of TRIM32 or TRIM37 using siRNA decreased survival in SHSY5Y neuroblastoma cells [118].
The UPS and Neurodegeneration
The accumulation of toxic protein aggregates is a common characteristic of neurodegenerative diseases, including AD, PD, HD, ALS, dementia with Lewy bodies (DLB), and prion diseases [7]. It has been demonstrated that Tau, α-synuclein, and TDP-43 co-occurred in AD and DLB [284, 285]. To protect cell function, several defense mechanisms against protein aggregate formation have been suggested [244, 270]. The UPS plays a crucial role in the management of general protein turnover and the degradation of misfolded proteins [272]; therefore, it is an important defensive mechanism against neurotoxic protein aggregation and prion-like protein propagation.
Evidence has indicated that a perturbed UPS can lead to the accumulation of toxic protein aggregates; thus, a perturbed UPS is likely to contribute to the development of many neurodegenerative disorders [266]. A well-known example of UPS impairment occurs in PD, which is a neurodegenerative disease characterized by the loss of dopaminergic neurons in the substantia nigra pars compacta. A loss-of-function mutation in Parkin, an E3 ligase, is a common familial cause of PD. The impairment of Parkin can lead to impaired aggregated α-synuclein degradation and the accumulation of unwanted protein aggregates [286]. In PD models, the inhibition of the UPS leads to the death of dopaminergic neurons in rat brains and primary neurons [287–289].
Proteasome inhibition has also been linked to ALS pathology. The proteasome inhibition in primary neurons (cortical, hippocampal, and motor neurons) resulted in the accumulation of TDP-43 aggregates [290]. An in vivo experiment of the conditional knock-out of a proteasome subunit gene resulted in the induction of ALS pathogenesis in mice; however, the knock-out of an autophagy-related gene failed to develop ALS pathology [291].
A recent study by Giannini et al. assessed the activity of the 20S and 26S proteasomes in young and aging rat brains and demonstrated that in aged rat brains, the proteasome’s substrate degradation activity was significantly decreased [292]. However, the ability of the 26S proteasome-specific polyubiquitin substrate degradation system was not impaired but rather slightly increased. This finding indicates that the 26S proteasome is not impaired during the normal aging process. Taken together, these results indicate that proteasome impairment is a key hallmark of neurodegenerative brains and is not observed in normal aging brains.
Increasing UPS activity in an HD patient’s fibroblast-derived neurons has been shown to improve cell survival [293]. In this study, UPS activity was rescued by overexpressing PA28γ, a 26S proteasome activator. Following exposure to various toxins, HD-derived neurons exhibited an increased susceptibility to cell death compared with neurons derived from donor healthy cells. However, when PA28γ was activated, HD-derived neurons exhibited an increased tolerance.
Autophagy is another major mechanism for the clearance of protein aggregates [21]. It depends on the lysosomal degradation of proteins by enveloping target proteins or organelles inside specialized compartments. A targeted protein for autophagy must be conjugated by an ubiquitin-like protein in a way that is similar to E1-E2-E3 signaling in the UPS [294]. Autophagy has been reported to be involved in various neurodegenerative diseases [295] and has recently been reviewed [294, 296–298].
Chaperones and Co-chaperones
Molecular chaperones, together with co-chaperones, are specialized machinery that assist protein folding and prevent toxic protein aggregation [299, 300]. The binding of chaperones to newly synthesized proteins aids in their correct folding. Many factors can promote protein misfolding, such as oxidative stress, aging, and misfolding-prone mutations. Chaperones also maintain general proteostasis, including protein trafficking and the targeting of severely misfolded proteins for proteolytic degradation [300]. As previously discussed, aggregating proteins may be cytotoxic; thus, the prevention of protein aggregation is critical for cell survival.
The HSPs belong to a major molecular chaperone family that assists protein folding by reversibly binding to unfolded and misfolded proteins to prevent their aggregation. In mammals, HSPs are classified according to their properties into several families, i.e., HSP90, HSP70, HSP60, HSP40, and small HSPs [301].
HSP90 (a 90 kDa HSP) is a major heat shock protein class involved in the stress response and interacts with a wide array of target proteins. It contains three domains, including an ATP-binding domain at its N terminus (~25 kDa) that displays low intrinsic ATPase activity, a middle segment for interaction with target proteins, and a C-terminal domain that is essential for protein dimerization [302]. Co-chaperone binding of HSP90 can alter enzyme activity; for example, the binding of Aha1 and Cpr6 can activate ATPase, whereas Hop/Sti1, Cdc37, and p23 suppress this activity [302].
Another important HSP class is HSP70. These proteins are one of the most ubiquitous classes of chaperones and are involved in diverse cellular functions [303, 304]. HSP70s have two domains, including a nucleotide binding domain for ATP-binding and a substrate binding domain used for target protein attachment [305]. Detailed aspects of the HSP70 machinery have been reviewed by Kampinga and Craig [306]. HSP70 is also required for the ubiquitin-dependent protein degradation of short-lived and aberrant proteins by the UPS. The interplay between HSP90 and HSP70 during the protein degradation cascade is intricate and is not completely understood. HSP90 stabilizes proteins by preventing an exposed hydrophobic cleft in its target protein and HSP70-dependent ubiquitination [307].
FKBP Co-chaperones
FKBP is a major member of the immunophilin protein family and a known co-chaperone for the HSP family [308]. The physiological function of FKBP is mediated by peptidyl-prolyl cis/trans isomerase activity (PPIase) in the FK506-binding domain (FKBD). The FKBD is the binding site for the immunosuppressant molecule FK506 (tacrolimus) [309]. Somarelli et al. classified 45 FKBPs across 23 species based on a tertiary structure prediction algorithm that resulted in six distinct groups of FKBPs [310]. In addition to chaperoning protein folding, FKBP plays a role in the regulation of cytokines, histone assembly, steroid receptor signaling, calcineurin signaling, and apoptosis [310, 311]. FKBPs are highly present in neuronal tissues and exhibit neuroprotective properties [312, 313]. There are 15 known FKBPs expressed in humans. A summary of a subset of FKBPs in the UniProt database is shown in Fig. 4.
The FKBP domains based on the UniProt database. All FKBPs have a characteristic FKBD that exhibits PPIase activity. FKBP14 has two additional EF-hand domains for Ca2+ signaling regulation. FKBP38 has three TPR domains for HSP90 binding and a transmembrane retention domain. FKBP51 and FKBP52 have two similar FKBDs and three TPRs
FKBP12 (also known as FKBP1A) is the smallest and contains only a single FKBD [309]. FKBP38 (official gene symbol FKBP8) contains an unusual FKBD but functions similarly to FKBP12. It has been demonstrated that FKBP38 prevents apoptosis via the regulation of Bcl-2 to mitochondria; however, upon interaction with presenilins, FKBP38 promotes apoptosis by reducing mitochondrial Bcl-2 [314, 315]. FKBP51 and FKBP52 (also known as FKBP5 and FKBP4, respectively) have two FKBDs that are slightly different from each other and three tetratricopeptide (TPR) repeats [316, 317]. The TPR domain allows FKBP to mediate protein-protein interactions, and it has been identified as a binding site for HSP90. Based on the UniProt database, FKBP14 (other alias FKBP22) has a single FKBP domain and two EF-hand domains for Ca2+-mediated signaling. Whereas other FKBPs are commonly found in the cytoplasm, FKBP14 is located in the ER [318]. FKBP14 plays an important role in Notch signaling in Drosophila and may control γ-secretase activity via the regulation of presenilin, which thereby affects Notch cleavage [319].
Chaperones and Co-chaperones in AD
Resolving the role of chaperones and co-chaperones in AD can be an intricate task. A number of studies have identified links between chaperones and neurodegeneration. Dickey et al. demonstrated that phosphorylated Tau was selectively degraded via the HSP90-carboxyl terminus of the HSP70-interacting protein (CHIP) complex [234] (Fig. 5). CHIP is an ubiquitin ligase that selectively targets phosphorylated Tau and promotes Tau degradation. This same research group reported that the deletion of CHIP increases the phosphorylation of Tau in mouse brains [320]. Although the significance of the HSP and co-chaperone interplay in AD pathology remains unclear, increasing evidence suggests that HSP-UPS-mediated aggregate clearance is the pivotal mechanism of AD pathology and further emphasizes the role of HSP90 as a mediator of Tau protein regulation and degradation. However, the role of HSP90 in Tau degradation is paradoxical: evidence suggests that HSP90 both promotes and prevents the formation of abnormal Tau, depending on its interaction with different target proteins or chaperones [321] (Fig. 5). An insight regarding the interaction between HSP90 activity and Tau folding has been reported in a recent finding [322]. The authors used state-of-the-art nuclear magnetic resonance (NMR) technology to determine the specific binding sites of the HSP90-Tau interaction.
Predicted interactions between Tau protein, FKBP and CHIP in AD. FKBP12 catalyzes the hyperphosphorylated Tau assembly into PHF. The fate of HSP90-bound hyperphosphorylated Tau is determined by the binding of the HSP90 co-chaperone molecule. The binding of an FKBP51 co-chaperone will drive the accumulation of Tau, which is antagonized by FKBP52. In contrast, the binding of HSP90 to the CHIP co-chaperone will lead to hyperphosphorylated Tau proteasomal degradation
The earliest study on the role of FKBPs in AD was conducted by Kraemer et al. who screened 46 genes that induced Tau pathology using Caenorhabditis elegans as a model [323]. A subset of the genes identified belonged to the chaperone or co-chaperone families. An in vivo knock-down experiment of FKBP51 by siRNA resulted in drastically reduced Tau and phosphorylated Tau levels, whereas the knock-down of FKBP52 slightly increased total Tau, which suggests that FKBP51 and FKBP52 have opposing actions [324]. Furthermore, the overexpression of FKBP51 increased Tau and phosphorylated Tau levels. To further confirm the involvement of FKBP51 in AD, a co-localization study in human AD brains indicated that FKBP51 was strongly co-localized with Tau. An F130A mutant of FKBP51, which abolished PPIase activity, did not prevent Tau degradation by chymotrypsin, indicating that FKBP51 prevents Tau degradation via its PPIase activity. A similar result was achieved by Blair et al. [325]. Furthermore, Blair et al. revealed that FKBP51 accumulates in AD brains and promotes the formation of Tau oligomers (Fig. 5).
Moreover, FKBP52 interacts with tubulin proteins and negatively modulates microtubule formation. The knock-down of FKBP52 results in neurite outgrowth and the induction of neuronal differentiation [326], whereas the overexpression of FKBP52 reduced Aβ-mediated toxicity in Drosophila; furthermore, the ablation of FKBP52 has opposing effects [327].
FKBP12 is observed in the area of neurodegeneration and is strongly associated with the protein aggregates in AD, PD, and DLB [328]. For example, an immunoblot analysis showed that FKBP12 expression is lower in AD total brain homogenates compared with controls and FKBP12 was identified in the intracellular NFTs of the hippocampus in AD brains [329], which suggest that FKBP12 is tangled with aberrant Tau protein. FKBP12 also has been reported to play a role in APP localization and processing. FKBP12 overexpression shifts the APP processing pathway towards the production of amyloidogenic products by changing the subcellular localization of APP [330]. It has also been reported that FKBP12 can bind AICD; however, the implication of this phenomenon in AD remains to be discovered [331].
FKBPs also play a role in other neurodegenerative diseases, including PD. The involvement of FKBP12 and FKBP52 in α-synuclein aggregation has been previously discussed by Gerard et al. [332, 333]. The overexpression of FKBP12 and FKBP52 in SHSY5Y cells induced α-synuclein aggregation, whereas inhibition, using FK506 and siRNA, reduced α-synuclein aggregation. Furthermore, an in vivo analysis indicated that the injection of FK506 (FKBP inhibitor) into PD mouse brains resulted in reduced α-synuclein aggregation.
Another distinct chaperone involved in AD pathogenesis is the peptidyl-prolyl cis/trans isomerase NIMA-interacting 1 (PIN1). PIN1 has been demonstrated to restore Tau binding to the microtubules via the promotion of Tau dephosphorylation [334, 335]. PIN1 has also been demonstrated to alter APP processing; its overexpression reduced Aβ production in vitro, whereas a PIN1 knock-out had the opposite outcome [336].
Clinical Studies Indicate UPS and Chaperone Deregulation in AD
As previously described, the pathophysiology of AD is initiated by an accumulation of aggregated Aβ and hyperphosphorylated Tau. The failure to clear these misfolded proteins may induce AD pathophysiology. The first evidence that underlines the importance of the UPS in AD is the cross-reactivity of ubiquitin in NFTs [337]. It has been observed that ubiquitination of the cerebral protein is impaired in human AD brains, which is accompanied by decreased proteolytic activity. Salon et al. reported that E1 and E2 ligase activities are lower in AD brains [338]. In agreement with this finding, an earlier postmortem study by Keller et al. identified a decrease in proteasome function in human AD brains compared with age- and gender-matched controls [339]. Moreover, the brain regions that demonstrated the most significant decrease in proteasome function are the regions most susceptible to AD, including the hippocampus, parahippocampal gyrus, superior and middle temporal gyri, and inferior parietal lobule. On the other hand, no significant decreases in proteasome function were observed in the occipital lobe or cerebellum, areas comparatively unaffected in AD pathology. Furthermore, this study showed that while the expression of the proteasome subunit(s) was uninterrupted, the function was inhibited. In agreement with previous studies, Keck et al.’s findings indicate that proteasome activity is decreased in AD brains [340]. Moreover, this study revealed that the proteasome activity was inhibited by PHFs and polymerized Tau, but not by soluble Tau. Isolated PHFs from AD brains and polymerized recombinant Tau protein significantly inhibited proteasome activity; however, unpolymerized recombinant Tau did not yield proteasome inhibition.
This theory is further supported by previous results that indicated proteasome activity impairment in AD-related neurodegeneration [266, 267, 341–344]. Furthermore, a human in vivo study that used the metabolic marker 13C6-leucine to trace the production and clearance rates of Aβ40 and Aβ42 in the CSF indicated that Aβ was produced at the same rate in both the AD (which ranged from very mild to mild AD) and control groups; however, the Aβ clearance in the AD group was significantly (~35 %) lower than the control group. [345]. The impaired proteolysis by the UPS may contribute to this observed phenomenon. The impeded proteolysis observed in very mild stages of AD suggests that proteolysis impairment precedes any symptoms from the development of AD pathology. Additional supporting evidence originated from a correlation study that identified a strong correlation between AD severity and Tau accumulation in AD brains [346]. This study also demonstrated a significant correlation between the level and activity of proteasome subunits and Tau accumulation in non-AD brains, which indicates that the UPS is crucial to prevent Tau aggregation.
Various groups have reported that the UPS can also be inhibited by Aβ [161, 347–351]. The use of scanning transmission electron microscopy has demonstrated that gold-labeled Aβ binds to the 20S proteasome, the catalytic core of the UPS, and inhibits its proteolytic activity [349]. This event, together with the activation of kinases, may explain why EOAD patients have observable symptoms at an accelerated rate. EOAD patients exhibit an increased production of Aβ that causes impaired UPS action, which is, alongside with contemporaneous activation of kinases that phosphorylate Tau, followed by the accumulation of misfolded Tau; these effects ultimately induce neuronal death accompanied by early-stage cognitive dysfunctions.
A clinical study by Yokota et al. indicated that Tau mRNA expression is not necessarily upregulated in AD brains; however, a related and more recent global proteome study by Manavalan et al. has shown that Tau protein is accumulated twofold in the temporal lobes of AD brains, which suggests that an impaired UPS is one of the pivotal causative factors of AD [118, 267]. This finding is in agreement with another previous report that indicated that impaired protein clearance, not increased protein production, causes the accumulation of neurotoxic aggregates [339, 340, 345]. Another important finding by Manavalan et al. was that most of the differentially regulated proteins from their study have a strong interaction with UBC. UBC is a polyubiquitin precursor involved in protein degradation by the UPS. These results underline that a perturbed UPS is almost certainly involved in the development of AD pathology [21, 271, 345].
In a global gene expression study of human AD subjects, Yokota et al. determined the differential gene expressions of TRIM32 and TRIM37, which are E3 ligase-encoding genes [118]. These data indicate that TRIM32/37 are expressed more than twofold in the occipital lobe (the brain region less susceptible to AD) compared with normal aging controls. Therefore, TRIM32/37 overexpression in the occipital lobe may prevent neurodegeneration, and areas affected by AD may have a perturbed UPS. Additional evidence for this phenomenon is that UBB+, a frame-shifted mutant ubiquitin, causes neuronal death and accumulates in the neurons of AD patients with this mutation [344]. UBB+ is a potent inhibitor of 26S proteasome [352]. Thus, this finding further emphasizes that the UPS can be inhibited in several ways that all lead to neuronal death in AD.
The upregulation of FKBP14 in the temporal lobes of AD brains has also been observed [118]. As previously described, immunophilins with PPIase activity may play a key role in AD pathogenesis by binding to hyperphosphorylated Tau and preventing ubiquitination and degradation. This notion is supported by reports which indicate that the inhibition of FKBPs by FK506 has a neuroprotective effect [353–355]. A study reported that FK506 also inhibits prion replication in a mouse model of CJD [356]. One explanation for these effects is that FK506 inhibits the formation of misfolded protein by inhibiting the PPIase activity of FKBP. Therefore, an increased level of small immunophilins, such as FKBP14, may represent another risk factor that prevents proper Tau degradation in neurons, which may thus contribute to AD pathogenesis.
Discussion
Despite more than 20 years as the de facto basis for AD research, the amyloid cascade hypothesis has failed to provide a clear mechanism of AD pathogenesis or halt disease progress in a myriad of drug trials [5, 6, 357]. In contrast, the Tau-based hypothesis has provided a more reasonable molecular explanation. First, other tauopathies exhibit neurodegeneration, even in the absence of APs. Thus, APs may induce neurodegeneration (as in EOAD) without being crucial (as in LOAD). A therapy based on a Tau aggregation inhibitor (TAI) has demonstrated more promising results compared with Aβ drugs. In a large phase 2 study of 321 subjects, methylthioninium chloride (methylene blue) was reported to stabilize the progression of both mild and moderate AD over 50 weeks [358, 359]. Therefore, a novel approach for the prevention of misfolded Tau accumulation is critically needed.
Here, we propose a hypothesis that AD is caused by collective insults to the ability of neurons to maintain proper proteostasis, especially in Tau protein regulation mediated by the UPS (Fig. 6). In neurons, Tau protein is exceptionally important because of its role in the maintenance of axonal transport and synaptic health. A failure to maintain Tau proteostasis results in the deposition of tangled Tau, which eventually manifests as PHFs and NFTs. Unfortunately, Tau protein is an IDP that is highly susceptible to aggregation without proper regulation and is subject to numerous modifications (i.e., phosphorylation and acetylation) that can induce Tau misfolding [217, 232, 233, 237]. Furthermore, in animal studies, Tau has exhibited prion-like characteristics that can be spread as aggregated units to nearby neurons [253, 254]. These intrinsic characteristics of Tau shed light on how AD is almost asymptomatic during its early stages and then exponentially progresses during the later stages.
The proposed AD model that involves Tau phosphorylation, prion-like misfolded protein propagation, and the deregulation of UPS, HSP90, and FKBP co-chaperones. In AD, various physiological conditions are altered, including kinase overexpression or Aβ-induction increases in cellular kinase activities. As a result, the phosphorylation-prone Tau protein becomes hyperphosphorylated and loses its ability to bind with microtubules. It is subsequently misfolded and acts as a seed for prion-like transmission to adjacent neurons where it aggregates into NFTs. In AD brains, the UPS system is impaired as a result of inhibition by Aβ or aberrant gene expression, which is possibly triggered by an aberrant AICD signaling cascade. TRIM32/37, important E3-ligases, are downregulated, which impairs the protein degradation pathway. In addition, the downregulation of HSP90 might also impair hyperphosphorylated Tau clearance. HSP90 and CHIP target hyperphosphorylated Tau for proteasomal degradation. The FKBP14 co-chaperone, which is upregulated in AD brains, prevents Tau degradation by binding to Tau and increasing its stability via interaction with the PPIase domain. Detached Tau loses its function and impairs the axonal transport pivotal for neuron survival. In this paradigm, Tau is involved in both the loss-of-function of axonal transport and gain-of-toxicity via prion-like transmission; in addition, impaired UPS and HSP90 function and FKBP14 overexpression promote pathophysiological conditions
One source of alterations for Tau proteostasis is the downregulation of E3 ligase in AD-susceptible areas. In the AD brain, HSP90 is a binding partner for CHIP and has been reported to be downregulated. Previous reports have indicated that CHIP selectively targets phosphorylated Tau, and the deletion of CHIP leads to an accumulation of phosphorylated Tau protein [234, 320]. The downregulation of HSP90 in AD-affected areas may result in the impairment of CHIP function, which thus promotes Tau accumulation. We postulate that CHIP may be the canonical E3 ligase that targets Tau. In AD-affected neurons, CHIP is either impaired or overloaded. A clinical study by Yokota et al. demonstrated that TRIM32/37 upregulation in the occipital lobe may result in a gain-of-function of the UPS; therefore, this protective mechanism enables certain brain regions to become less susceptible to AD pathogenesis [118]. Interestingly, it has been reported that the overexpression of Parkin, an E3 ligase that is associated with familial PD, ameliorates Tau and Aβ aggregates in an AD mouse model [360]. This finding implies that the activity of canonical E3 ligases for Tau or Aβ can be compensated by other E3 ligases.
Another important finding from this clinical investigation is the involvement of small immunophilins. The overexpression of FKBP14 in the temporal lobe of AD brains suggests that it might promote Tau toxicity and inhibits Tau clearance via the stabilization of aggregated Tau and the prevention of its degradation. Another previous study also indicated that another small immunophilin, FKBP12, is bound to aberrant Tau in NFTs. This finding suggests that small immunophilins have a crucial role in the development of AD pathogenesis and could become important drug targets or biomarkers.
In light of these findings, further studies are needed to investigate the role of TRIM ligases in Tau ubiquitination and to shed more light on E3 ligases other than CHIP that are involved in the Tau degradation pathway. The rationale is that E3 ligases are one of the largest protein families in humans and are encoded by more than 600 genes [361]; thus, the identification of canonical E3 ligases involved in Tau ubiquitination will provide a better understanding of AD pathogenesis at the molecular level. The discovery of non-canonical E3 ligases that target Tau can aid in the development or identification of compensatory E3 ligases, which can be used as therapeutics for the treatment of AD (Fig. 7).
E3 ligases and FKBP14 are potential drug targets for AD therapy. Increasing the activity of non-canonical E3 ligases by drugs or gene therapy could prevent Tau aggregation. CHIP has been suggested to target phosphorylated Tau for ubiquitination and subsequent degradation; therefore, CHIP is hypothesized to be a canonical E3 ligase for phosphorylated Tau. We postulate that in the AD brain, CHIP is impaired or overloaded, which causes a failure to maintain Tau turnover. In the occipital lobe, a region less impacted by AD, TRIM32/37 act as compensatory E3 ligases and help to maintain Tau protein turnover. As previously described, FKBP14 catalyzes Tau aggregation. FK506 (an FKBP inhibitor) has been reported to have neuroprotective effects; however, it also acts as an immunosuppressant. Novel drug therapies for AD should also focus on a non-immunosuppressive FKBP inhibitor that can cross the blood-brain barrier or a compound that can activate E3 ligases
The role of FKBPs has not been widely studied in AD. The inhibition of FKBPs by FK506 has been demonstrated to be neuroprotective in prion diseases. It is suggested that prions assert their toxicity via the activation of the calcineurin pathway, whereas the inhibition of calcineurin signaling by FK506 prevents neurodegeneration [362]. However, we also postulate that the neuroprotective effects of FK506 are achieved via the inhibition of misfolded protein formation, rather than calcineurin signaling. A study in neuroblastoma cells indicated that the addition of FK506 did not induce autophagy; however, it prevented PrP replication in cell culture [356]. The inhibition of FKBPs successfully attenuated protein aggregation in a mouse PD model [332]. Considering the shared properties between Tau and α-synuclein aggregates, non-immunosuppressive FKBP inhibitors could represent candidates for a therapy to prevent Tau aggregation (Fig. 7). A focus on FKBP inhibitors as drug candidates has additional advantages. FKBPs have been well studied for their pharmacological interest, several sequences have been identified, and three-dimensional structures from X-ray and NMR studies are available; therefore, screening strategies for the discovery of new drug candidates have been established [363].
Conclusions
Our review of the recent research data based on (pre)clinical studies elucidated a potential mechanism for AD pathogenesis as a result of the malfunction of the UPS and deregulation of molecular chaperones. AD neurons exhibit impaired proteostasis that ultimately leads to misfolded protein accumulation and manifests as characteristic AD lesions. We postulate that the impairment of E3 ligases, notably HSP90-CHIP and TRIM32/37, and the deregulation of molecular chaperones, notably FKBP immunophilins, are the real culprits that cause misfolded Tau protein accumulation. Thus, future AD research should not only focus on the amyloid cascade hypothesis but also on the involvement of the UPS and molecular chaperones to facilitate the understanding of the neuropathogenesis of AD and other tauopathies.
Abbreviations
- aa:
-
Amino acid
- AD:
-
Alzheimer’s disease
- ADAM:
-
A disintegrin and metalloproteinase
- AFT:
-
AICD-FE65-TIP60
- AICD:
-
APP intracellular c-terminal domain
- ALS:
-
Amyotrophic lateral sclerosis
- AMPK:
-
AMP-activated kinase
- AP:
-
Amyloid plaque
- APH1:
-
Anterior pharynx-defective 1
- APOE4:
-
Apolipoprotein E4
- APP:
-
Amyloid precursor protein
- Aβ:
-
Amyloid-beta peptide
- BACE1:
-
Beta-site APP cleaving enzyme 1
- CaMK:
-
Calcium/calmodulin-dependent protein kinase II
- CDK5:
-
Cyclin-dependent kinase 5
- CHIP:
-
Carboxyl terminus of HSP70-interacting protein
- chr:
-
Chromosome
- CJD:
-
Creutzfeldt-Jakob disease
- CKII:
-
Casein kinase II
- CNS:
-
Central nervous system
- CSF:
-
Cerebrospinal fluid
- CUBD:
-
Cu(II)-binding domain
- DLB:
-
Dementia with Lewy bodies
- Dyrk1A:
-
Dual-specificity tyrosine-regulated kinase 1A
- EGFR:
-
Epidermal growth factor receptor
- EOAD:
-
Early-onset AD
- ER:
-
Endoplasmic reticulum
- FKBD:
-
FK506-binding domain
- FKBP:
-
FK506-binding protein
- FTDP-17:
-
Frontotemporal dementia related to chr 17
- HBD:
-
Heparin-binding domain
- HD:
-
Huntington’s disease
- HSP:
-
Heat shock protein
- IDP:
-
Intrinsically disordered protein
- KPI:
-
Kunitz-type serine protease inhibitor
- LOAD:
-
Late-onset AD
- LRP1:
-
Lipoprotein receptor-related protein 1
- LTP:
-
Long-term potentiation
- MAP:
-
Microtubule-associated protein
- MAPK:
-
Mitogen-activated protein kinase
- MAPT:
-
Microtubule-associated protein Tau
- MARK:
-
MAP/microtubule-affinity-regulating kinases
- MRI:
-
Magnetic resonance imaging
- NFT:
-
Neurofibrillary tangle
- NICD:
-
Notch intracellular domain
- NPC:
-
Neural precursor cell
- PD:
-
Parkinson’s disease
- PEN2:
-
Presenilin enhancer 2
- PHF:
-
Paired helical filament
- PIN1:
-
Peptidyl-prolyl cis/trans isomerase NIMA-interacting 1
- PKA:
-
Protein kinase A
- PKC:
-
Protein kinase C
- PrP:
-
Prion protein
- PS1/2:
-
Presenilin-1/Presenilin-2
- RIP:
-
Regulated intramembrane proteolysis
- RTN:
-
Reticulon family protein
- SOD1:
-
Superoxide dismutase 1
- SVZ:
-
Subventricular zone
- TACE:
-
Tumor necrosis factor-α converting enzyme
- TAG1:
-
Transient axonal glycoprotein
- TAI:
-
Tau aggregation inhibitor
- TDP-43:
-
TAR DNA-binding protein-43
- TGN:
-
Trans-Golgi network
- TRIM:
-
Tripartite motif
- UBB:
-
Ubiquitin B
- UBC:
-
Ubiquitin C
- UPS:
-
Ubiquitin-proteasome system
References
Graeber MB, Kösel S, Egensperger R, Banati RB, Müller U, Bise K, Hoff P, Möller HJ, Fujisawa K, Mehraein P (1997) Rediscovery of the case described by Alois Alzheimer in 1911: historical, histological and molecular genetic analysis. Neurogenetics 1(1):73–80. doi:10.1007/s100480050011
Selkoe DJ (2001) Alzheimer’s disease: genes, proteins, and therapy. Physiol Rev 81(2):741–766
Hebert LE, Weuve J, Scherr PA, Evans DA (2013) Alzheimer disease in the United States (2010–2050) estimated using the 2010 census. Neurology 80(19):1778–1783. doi:10.1212/WNL.0b013e31828726f5
Thies W, Bleiler L (2013) 2013 Alzheimer’s disease facts and figures. Alzheimers Dement 9(2):208–245. doi:10.1016/j.jalz.2013.02.003
Castello MA, Soriano S (2014) On the origin of Alzheimer’s disease. Trials and tribulations of the amyloid hypothesis. Ageing Res Rev 13:10–12. doi:10.1016/j.arr.2013.10.001
Cummings J, Morstorf T, Zhong K (2014) Alzheimer’s disease drug-development pipeline: few candidates, frequent failures. Alzheimers Res Ther 6(4):37
Taylor JP, Hardy J, Fischbeck KH (2002) Toxic proteins in neurodegenerative disease. Science 296(5575):1991–1995. doi:10.1126/science.1067122
Benowitz LI, Rodriguez W, Paskevich P, Mufson EJ, Schenk D, Neve RL (1989) The amyloid precursor protein is concentrated in neuronal lysosomes in normal and Alzheimer disease subjects. Exp Neurol 106(3):237–250
Thal DR, Rüb U, Orantes M, Braak H (2002) Phases of Aβ-deposition in the human brain and its relevance for the development of AD. Neurology 58(12):1791–1800. doi:10.1212/wnl.58.12.1791
Braak H, Braak E (1991) Neuropathological stageing of Alzheimer-related changes. Acta Neuropathol 82(4):239–259
Delacourte A, David JP, Sergeant N, Buee L, Wattez A, Vermersch P, Ghozali F, Fallet-Bianco C, Pasquier F, Lebert F, Petit H, Di Menza C (1999) The biochemical pathway of neurofibrillary degeneration in aging and Alzheimer’s disease. Neurology 52(6):1158–1165
Johnson KA, Fox NC, Sperling RA, Klunk WE (2012) Brain imaging in Alzheimer disease. Cold Spring Harb Perspect Med 2(4):a006213. doi:10.1101/cshperspect.a006213
Scahill RI, Schott JM, Stevens JM, Rossor MN, Fox NC (2002) Mapping the evolution of regional atrophy in Alzheimer’s disease: unbiased analysis of fluid-registered serial MRI. Proc Natl Acad Sci U S A 99(7):4703–4707. doi:10.1073/pnas.052587399
Smith AD (2002) Imaging the progression of Alzheimer pathology through the brain. Proc Natl Acad Sci U S A 99(7):4135–4137. doi:10.1073/pnas.082107399
Frost B, Diamond MI (2010) Prion-like mechanisms in neurodegenerative diseases. Nat Rev Neurosci 11(3):155–159. doi:10.1038/nrn2786
Tanzi RE (2012) The genetics of Alzheimer disease. Cold Spring Harb Perspect Med 2(10):a006296. doi:10.1101/cshperspect.a006296
Bekris LM, Yu C-E, Bird TD, Tsuang DW (2010) Review article: genetics of Alzheimer disease. J Geriatr Psychiatry Neurol 23(4):213–227. doi:10.1177/0891988710383571
Tanzi RE, Bertram L (2001) New frontiers in Alzheimer’s disease genetics. Neuron 32(2):181–184. doi:10.1016/S0896-6273(01)00476-7
Lindsay J, Laurin D, Verreault R, Hébert R, Helliwell B, Hill GB, McDowell I (2002) Risk factors for Alzheimer’s disease: a prospective analysis from the Canadian study of health and aging. Am J Epidemiol 156(5):445–453. doi:10.1093/aje/kwf074
Mayeux R, Stern Y (2012) Epidemiology of Alzheimer disease. Cold Spring Harb Perspect Med 2(8):a006239. doi:10.1101/cshperspect.a006239
Ihara Y, Morishima-Kawashima M, Nixon R (2012) The ubiquitin–proteasome system and the autophagic–lysosomal system in Alzheimer disease. Cold Spring Harb Perspect Med 2(8):a006361. doi:10.1101/cshperspect.a006361
Troen BR (2003) The biology of aging. Mt Sinai J Med 70(1):3–22
Kang J, Lemaire H-G, Unterbeck A, Salbaum JM, Masters CL, Grzeschik K-H, Multhaup G, Beyreuther K, Muller-Hill B (1987) The precursor of Alzheimer’s disease amyloid A4 protein resembles a cell-surface receptor. Nature 325(6106):733–736
Ghiso J, Tagliavini F, Timmers WF, Frangione B (1989) Alzheimer’s disease amyloid precursor protein is present in senile plaques and cerebrospinal fluid: immunohistochemical and biochemical characterization. Biochem Biophys Res Commun 163(1):430–437
Autilio-Gambetti L, Morandi A, Tabaton M, Schaetzle B, Kovacs D, Perry G, Sharma S, Cornette J, Greenberg B, Gambetti P (1988) The amyloid percursor protein of Alzheimer disease is expressed as a 130 kDa polypeptide in various cultured cell types. FEBS Lett 241(1–2):94–98
Clark AW, Krekoski CA, Parhad IM, Liston D, Julien JP, Hoar DI (1989) Altered expression of genes for amyloid and cytoskeletal proteins in Alzheimer cortex. Ann Neurol 25(4):331–339. doi:10.1002/ana.410250404
Masters CL, Simms G, Weinman NA, Multhaup G, McDonald BL, Beyreuther K (1985) Amyloid plaque core protein in Alzheimer disease and Down syndrome. Proc Natl Acad Sci U S A 82(12):4245–4249
Glenner GG, Wong CW (1984) Alzheimer’s disease and Down’s syndrome: sharing of a unique cerebrovascular amyloid fibril protein. Biochem Biophys Res Commun 122(3):1131–1135
Glenner GG, Wong CW (1984) Alzheimer’s disease: initial report of the purification and characterization of a novel cerebrovascular amyloid protein. Biochem Biophys Res Commun 120(3):885–890. doi:10.1016/S0006-291X(84)80190-4
Tanzi R, Gusella J, Watkins P, Bruns G, St George-Hyslop P, Van Keuren M, Patterson D, Pagan S, Kurnit D, Neve R (1987) Amyloid beta protein gene: cDNA, mRNA distribution, and genetic linkage near the Alzheimer locus. Science 235(4791):880–884. doi:10.1126/science.2949367
Korenberg JR, Pulst SM, Neve RL, West R (1989) The Alzheimer amyloid precursor protein maps to human chromosome 21 bands q21.105-q21.05. Genomics 5(1):124–127
Tokuda T, Fukushima T, Ikeda S, Sekijima Y, Shoji S, Yanagisawa N, Tamaoka A (1997) Plasma levels of amyloid beta proteins Abeta1-40 and Abeta1-42(43) are elevated in Down’s syndrome. Ann Neurol 41(2):271–273. doi:10.1002/ana.410410220
Mehta PD, Capone G, Jewell A, Freedland RL (2007) Increased amyloid β protein levels in children and adolescents with Down syndrome. J Neurol Sci 254(1):22–27
Goate A, Chartier-Harlin M-C, Mullan M, Brown J, Crawford F, Fidani L, Giuffra L, Haynes A, Irving N, James L, Mant R, Newton P, Rooke K, Roques P, Talbot C, Pericak-Vance M, Roses A, Williamson R, Rossor M, Owen M, Hardy J (1991) Segregation of a missense mutation in the amyloid precursor protein gene with familial Alzheimer’s disease. Nature 349(6311):704–706
Murrell J, Farlow M, Ghetti B, Benson M (1991) A mutation in the amyloid precursor protein associated with hereditary Alzheimer’s disease. Science 254(5028):97–99. doi:10.1126/science.1925564
Suzuki N, Cheung T, Cai X, Odaka A, Otvos L, Eckman C, Golde T, Younkin S (1994) An increased percentage of long amyloid beta protein secreted by familial amyloid beta protein precursor (beta APP717) mutants. Science 264(5163):1336–1340. doi:10.1126/science.8191290
Murrell JR, Hake AM, Quaid KA, Farlow MR, Ghetti B (2000) Early-onset Alzheimer disease caused by a new mutation (V717L) in the amyloid precursor protein gene. Arch Neurol 57(6):885–887. doi:10.1001/archneur.57.6.885
Kwok JBJ, Li QX, Hallupp M, Whyte S, Ames D, Beyreuther K, Masters CL, Schofield PR (2000) Novel Leu723Pro amyloid precursor protein mutation increases amyloid beta 42(43) peptide levels and induces apoptosis. Ann Neurol 47(2):249–253. doi:10.1002/1531-8249(200002)47:2<249::aid-ana18>3.0.co;2-8
Kumar-Singh S, De Jonghe C, Cruts M, Kleinert R, Wang R, Mercken M, De Strooper B, Vanderstichele H, Löfgren A, Vanderhoeven I, Backhovens H, Vanmechelen E, Kroisel PM, Van Broeckhoven C (2000) Nonfibrillar diffuse amyloid deposition due to a γ 42‐secretase site mutation points to an essential role for N‐truncated Aβ42 in Alzheimer’s disease. Hum Mol Genet 9(18):2589–2598. doi:10.1093/hmg/9.18.2589
Ancolio K, Dumanchin C, Barelli H, Warter JM, Brice A, Campion D, Frébourg T, Checler F (1999) Unusual phenotypic alteration of β amyloid precursor protein (βAPP) maturation by a new Val-715 → Met βAPP-770 mutation responsible for probable early-onset Alzheimer’s disease. Proc Natl Acad Sci U S A 96(7):4119–4124. doi:10.1073/pnas.96.7.4119
Eckman CB, Mehta ND, Crook R, Perez-tur J, Prihar G, Pfeiffer E, Graff-Radford N, Hinder P, Yager D, Zenk B, Refolo LM, Mihail Prada C, Younkin SG, Hutton M, Hardy J (1997) A new pathogenic mutation in the APP gene (I716V) increases the relative proportion of Aβ42(43). Hum Mol Genet 6(12):2087–2089. doi:10.1093/hmg/6.12.2087
Mullan M, Crawford F, Axelman K, Houlden H, Lilius L, Winblad B, Lannfelt L (1992) A pathogenic mutation for probable Alzheimer’s disease in the APP gene at the N-terminus of beta-amyloid. Nat Genet 1(5):345–347. doi:10.1038/ng0892-345
Hendriks L, Vanduijn CM, Cras P, Cruts M, Vanhul W, Vanharskamp F, Warren A, McInnis MG, Antonarakis SE, Martin JJ, Hofman A, Vanbroeckhoven C (1992) Presenile-dementia and cerebral-hemorrhage linked to a mutation at codon-692 of the beta-amyloid precursor protein gene. Nat Genet 1(3):218–221. doi:10.1038/ng0692-218
Lan M-Y, Liu J-S, Wu Y-S, Peng C-H, Chang Y-Y (2014) A novel APP mutation (D678H) in a Taiwanese patient exhibiting dementia and cerebral microvasculopathy. J Clin Neurosci 21(3):513–515. doi:10.1016/j.jocn.2013.03.038
Kero M, Paetau A, Polvikoski T, Tanskanen M, Sulkava R, Jansson L, Myllykangas L, Tienari PJ (2013) Amyloid precursor protein (APP) A673T mutation in the elderly Finnish population. Neurobiol Aging 34(5):1518.e1–1518.e3. doi:10.1016/j.neurobiolaging.2012.09.017
Suárez-Calvet M, Belbin O, Pera M, Badiola N, Magrané J, Guardia-Laguarta C, Muñoz L, Colom-Cadena M, Clarimón J, Lleó A (2014) Autosomal-dominant Alzheimer’s disease mutations at the same codon of amyloid precursor protein differentially alter Aβ production. J Neurochem 128(2):330–339. doi:10.1111/jnc.12466
De Jonghe C, Esselens C, Kumar-Singh S, Craessaerts K, Serneels S, Checler F, Annaert W, Van Broeckhoven C, De Strooper B (2001) Pathogenic APP mutations near the γ-secretase cleavage site differentially affect Aβ secretion and APP C-terminal fragment stability. Hum Mol Genet 10(16):1665–1671. doi:10.1093/hmg/10.16.1665
Russo C, Schettini G, Saido TC, Hulette C, Lippa C, Lannfelt L, Ghetti B, Gambetti P, Tabaton M, Teller JK (2000) Neurobiology: presenilin-1 mutations in Alzheimer’s disease. Nature 405(6786):531–532. doi:10.1038/35014735
Tanzi RE, George-Hyslop PHS, Haines JL, Polinsky RJ, Nee L, Foncin J-F, Neve RL, McClatchey AI, Conneally PM, Gusella JF (1987) The genetic defect in familial Alzheimer’s disease is not tightly linked to the amyloid [beta]-protein gene. Nature 329(6135):156–157
Vitek MP, Rasool CG, de Sauvage F, Vitek SM, Bartus RT, Beer B, Ashton RA, Macq AF, Maloteaux JM, Blume AJ et al (1988) Absence of mutation in the beta-amyloid cDNAs cloned from the brains of three patients with sporadic Alzheimer’s disease. Brain Res 464(2):121–131
Borchelt DR, Thinakaran G, Eckman CB, Lee MK, Davenport F, Ratovitsky T, Prada C-M, Kim G, Seekins S, Yager D, Slunt HH, Wang R, Seeger M, Levey AI, Gandy SE, Copeland NG, Jenkins NA, Price DL, Younkin SG, Sisodia SS (1996) Familial Alzheimer’s disease–linked presenilin 1 variants elevate Aβ1–42/1–40 ratio in vitro and in vivo. Neuron 17(5):1005–1013. doi:10.1016/S0896-6273(00)80230-5
Lemere CA, Lopera F, Kosik KS, Lendon CL, Ossa J, Saido TC, Yamaguchi H, Ruiz A, Martinez A, Madrigal L (1996) The E280A presenilin 1 Alzheimer mutation produces increased Aβ42 deposition and severe cerebellar pathology. Nat Med 2(10):1146–1150
Hardy J (1997) Amyloid, the presenilins and Alzheimer’s disease. Trends Neurosci 20(4):154–159. doi:10.1016/S0166-2236(96)01030-2
Saunders AM, Strittmatter WJ, Schmechel D, George-Hyslop PH, Pericak-Vance MA, Joo SH, Rosi BL, Gusella JF, Crapper-MacLachlan DR, Alberts MJ, Hulette C, Crain B, Goldgaber D, Roses AD (1993) Association of apolipoprotein E allele ∈4 with late-onset familial and sporadic Alzheimer’s disease. Neurology 43(8):1467–1472
Corder EH, Saunders AM, Strittmatter WJ, Schmechel DE, Gaskell PC, Small GW, Roses AD, Haines JL, Pericak-Vance MA (1993) Gene dose of apolipoprotein E type 4 allele and the risk of Alzheimer’s disease in late onset families. Science 261(5123):921–923
Strittmatter WJ, Roses AD (1996) Apolipoprotein E and Alzheimer’s disease. Annu Rev Neurosci 19(1):53–77. doi:10.1146/annurev.ne.19.030196.000413
Michaelson DM (2014) ApoE4: the most prevalent yet understudied risk factor for Alzheimer’s disease. Alzheimers Dement. doi:10.1016/j.jalz.2014.06.015
Fergusson J, Landon M, Lowe J, Ward L, van Leeuwen FW, Mayer RJ (2000) Neurofibrillary tangles in progressive supranuclear palsy brains exhibit immunoreactivity to frameshift mutant ubiquitin-B protein. Neurosci Lett 279(2):69–72. doi:10.1016/S0304-3940(99)00917-9
van Leeuwen FW, de Kleijn DP, van den Hurk HH, Neubauer A, Sonnemans MA, Sluijs JA, Koycu S, Ramdjielal RD, Salehi A, Martens GJ, Grosveld FG, Peter J, Burbach H, Hol EM (1998) Frameshift mutants of beta amyloid precursor protein and ubiquitin-B in Alzheimer’s and Down patients. Science 279(5348):242–247
Yankner BA, Dawes LR, Fisher S, Villa-Komaroff L, Oster-Granite ML, Neve RL (1989) Neurotoxicity of a fragment of the amyloid precursor associated with Alzheimer’s disease. Science 245(4916):417–420
Selkoe DJ (2000) Toward a comprehensive theory for Alzheimer’s disease. Hypothesis: Alzheimer’s disease is caused by the cerebral accumulation and cytotoxicity of amyloid beta-protein. Ann N Y Acad Sci 924:17–25
Selkoe DJ (1998) The cell biology of beta-amyloid precursor protein and presenilin in Alzheimer’s disease. Trends Cell Biol 8(11):447–453. doi:10.1016/S0962-8924(98)01363-4
Wippold FJ, Cairns N, Vo K, Holtzman DM, Morris JC (2008) Neuropathology for the neuroradiologist: plaques and tangles. Am J Neuroradiol 29(1):18–22. doi:10.3174/ajnr.A0781
Masters CL, Selkoe DJ (2012) Biochemistry of amyloid β-protein and amyloid deposits in Alzheimer disease. Cold Spring Harb Perspect Med a006262. doi: 10.1101/cshperspect.a006262
Joachim CL, Morris JH, Selkoe DJ (1989) Diffuse senile plaques occur commonly in the cerebellum in Alzheimer’s disease. Am J Pathol 135(2):309–319
Lord A, Philipson O, Klingstedt T, Westermark G, Hammarstrom P, Nilsson KP, Nilsson LN (2011) Observations in APP bitransgenic mice suggest that diffuse and compact plaques form via independent processes in Alzheimer’s disease. Am J Pathol 178(5):2286–2298. doi:10.1016/j.ajpath.2011.01.052
Ma QH, Futagawa T, Yang WL, Jiang XD, Zeng L, Takeda Y, Xu RX, Bagnard D, Schachner M, Furley AJ, Karagogeos D, Watanabe K, Dawe GS, Xiao ZC (2008) A TAG1-APP signalling pathway through Fe65 negatively modulates neurogenesis. Nat Cell Biol 10(3):283–294. doi:10.1038/ncb1690
Selkoe DJ, Podlisny MB, Joachim CL, Vickers EA, Lee G, Fritz LC, Oltersdorf T (1988) Beta-amyloid precursor protein of Alzheimer disease occurs as 110- to 135-kilodalton membrane-associated proteins in neural and nonneural tissues. Proc Natl Acad Sci U S A 85(19):7341–7345
Tanaka S, Shiojiri S, Takahashi Y, Kitaguchi N, Ito H, Kameyama M, Kimura J, Nakamura S, Ueda K (1989) Tissue-specific expression of three types of beta-protein precursor mRNA: enhancement of protease inhibitor-harboring types in Alzheimer’s disease brain. Biochem Biophys Res Commun 165(3):1406–1414
Johnson SA, Pasinetti GM, May PC, Ponte PA, Cordell B, Finch CE (1988) Selective reduction of mRNA for the β-amyloid precursor protein that lacks a Kunitz-type protease inhibitor motif in cortex from Alzheimer brains. Exp Neurol 102(2):264–268. doi:10.1016/0014-4886(88)90104-5
Johnson SA, Rogers J, Finch CE (1989) APP-695 transcript prevalence is selectively reduced during Alzheimer’s disease in cortex and hippocampus but not in cerebellum. Neurobiol Aging 10(6):755–760
Haass C, Hung AY, Selkoe DJ (1991) Processing of beta-amyloid precursor protein in microglia and astrocytes favors an internal localization over constitutive secretion. J Neurosci 11(12):3783–3793
Tanzi RE, McClatchey AI, Lamperti ED, Villa-Komaroff L, Gusella JF, Neve RL (1988) Protease inhibitor domain encoded by an amyloid protein precursor mRNA associated with Alzheimer’s disease. Nature 331(6156):528–530
Kitaguchi N, Takahashi Y, Tokushima Y, Shiojiri S, Ito H (1988) Novel precursor of Alzheimer’s disease amyloid protein shows protease inhibitory activity. Nature 331(6156):530–532. doi:10.1038/331530a0
Small DH, Nurcombe V, Reed G, Clarris H, Moir R, Beyreuther K, Masters CL (1994) A heparin-binding domain in the amyloid protein precursor of Alzheimer’s disease is involved in the regulation of neurite outgrowth. J Neurosci 14(4):2117–2127
Mok SS, Sberna G, Heffernan D, Cappai R, Galatis D, Clarris HJ, Sawyer WH, Beyreuther K, Masters CL, Small DH (1997) Expression and analysis of heparin-binding regions of the amyloid precursor protein of Alzheimer’s disease. FEBS Lett 415(3):303–307. doi:10.1016/S0014-5793(97)01146-0
Rossjohn J, Cappai R, Feil SC, Henry A, McKinstry WJ, Galatis D, Hesse L, Multhaup G, Beyreuther K, Masters CL, Parker MW (1999) Crystal structure of the N-terminal, growth factor-like domain of Alzheimer amyloid precursor protein. Nat Struct Biol 6(4):327–331
Barnham KJ, McKinstry WJ, Multhaup G, Galatis D, Morton CJ, Curtain CC, Williamson NA, White AR, Hinds MG, Norton RS, Beyreuther K, Masters CL, Parker MW, Cappai R (2003) Structure of the Alzheimer’s disease amyloid precursor protein copper binding domain: a regulator of neuronal copper homeostasis. J Biol Chem 278(19):17401–17407. doi:10.1074/jbc.M300629200
Ninomiya H, Roch JM, Sundsmo MP, Otero DAC, Saitoh T (1993) Amino acid sequence RERMS represents the active domain of amyloid β/A4 protein precursor that promotes fibroblast growth. J Cell Biol 121(4):879–886
Lazarov O, Demars MP (2012) All in the family: how the APPs regulate neurogenesis. Front Neurosci 6:81. doi:10.3389/fnins.2012.00081
O’Brien RJ, Wong PC (2011) Amyloid precursor protein processing and Alzheimer’s disease. Annu Rev Neurosci 34:185–204. doi:10.1146/annurev-neuro-061010-113613
Oh ES, Savonenko AV, King JF, Fangmark Tucker SM, Rudow GL, Xu G, Borchelt DR, Troncoso JC (2009) Amyloid precursor protein increases cortical neuron size in transgenic mice. Neurobiol Aging 30(8):1238–1244. doi:10.1016/j.neurobiolaging.2007.12.024
Dawkins E, Small DH (2014) Insights into the physiological function of the β-amyloid precursor protein: beyond Alzheimer’s disease. J Neurochem 756–769. doi: 10.1111/jnc.12675
Perez RG, Zheng H, Van der Ploeg LHT, Koo EH (1997) The β-amyloid precursor protein of Alzheimer’s disease enhances neuron viability and modulates neuronal polarity. J Neurosci 17(24):9407–9414
Corrigan F, Thornton E, Roisman LC, Leonard AV, Vink R, Blumbergs PC, van den Heuvel C, Cappai R (2014) The neuroprotective activity of the amyloid precursor protein against traumatic brain injury is mediated via the heparin binding site in residues 96-110. J Neurochem 128(1):196–204. doi:10.1111/jnc.12391
Greenfield JP, Tsai J, Gouras GK, Hai B, Thinakaran G, Checler F, Sisodia SS, Greengard P, Xu H (1999) Endoplasmic reticulum and trans-Golgi network generate distinct populations of Alzheimer β-amyloid peptides. Proc Natl Acad Sci U S A 96(2):742–747. doi:10.1073/pnas.96.2.742
Selkoe DJ, Yamazaki T, Citron M, Podlisny MB, Koo EH, Teplow DB, Haass C (1996) The role of APP processing and trafficking pathways in the formation of amyloid β-protein. Ann N Y Acad Sci 777(1):57–64. doi:10.1111/j.1749-6632.1996.tb34401.x
Parvathy S, Hussain I, Karran EH, Turner AJ, Hooper NM (1999) Cleavage of Alzheimer’s amyloid precursor protein by α-secretase occurs at the surface of neuronal cells. Biochemistry (Mosc) 38(30):9728–9734. doi:10.1021/bi9906827
Skovronsky DM, Moore DB, Milla ME, Doms RW, Lee VM-Y (2000) Protein kinase C-dependent α-secretase competes with β-secretase for cleavage of amyloid-β precursor protein in the trans-Golgi network. J Biol Chem 275(4):2568–2575. doi:10.1074/jbc.275.4.2568
Palmert MR, Podlisny MB, Witker DS, Oltersdorf T, Younkin LH, Selkoe DJ, Younkin SG (1989) The beta-amyloid protein precursor of Alzheimer disease has soluble derivatives found in human brain and cerebrospinal fluid. Proc Natl Acad Sci U S A 86(16):6338–6342
Chasseigneaux S, Allinquant B (2012) Functions of Abeta, sAPPalpha and sAPPbeta: similarities and differences. J Neurochem 120(Suppl 1):99–108. doi:10.1111/j.1471-4159.2011.07584.x
Chasseigneaux S, Dinc L, Rose C, Chabret C, Coulpier F, Topilko P, Mauger G, Allinquant B (2011) Secreted amyloid precursor protein β and secreted amyloid precursor protein α induce axon outgrowth in vitro through Egr1 signaling pathway. PLoS ONE 6(1):e16301
Hartl D, Klatt S, Roch M, Konthur Z, Klose J, Willnow TE, Rohe M (2013) Soluble alpha-APP (sAPPalpha) regulates CDK5 expression and activity in neurons. PLoS ONE 8(6):e65920
Gakhar‐Koppole N, Hundeshagen P, Mandl C, Weyer SW, Allinquant B, Müller U, Ciccolini F (2008) Activity requires soluble amyloid precursor protein α to promote neurite outgrowth in neural stem cell‐derived neurons via activation of the MAPK pathway. Eur J Neurosci 28(5):871–882
Baratchi S, Evans J, Tate WP, Abraham WC, Connor B (2012) Secreted amyloid precursor proteins promote proliferation and glial differentiation of adult hippocampal neural progenitor cells. Hippocampus 22(7):1517–1527
Demars MP, Bartholomew A, Strakova Z, Lazarov O (2011) Soluble amyloid precursor protein: a novel proliferation factor of adult progenitor cells of ectodermal and mesodermal origin. Stem Cell Res Ther 2:36
Hasebe N, Fujita Y, Ueno M, Yoshimura K, Fujino Y, Yamashita T (2013) Soluble β-amyloid precursor protein alpha binds to p75 neurotrophin receptor to promote neurite outgrowth. PLoS ONE 8(12):e82321
Asai M, Hattori C, Szabó B, Sasagawa N, Maruyama K, S-i T, Ishiura S (2003) Putative function of ADAM9, ADAM10, and ADAM17 as APP α-secretase. Biochem Biophys Res Commun 301(1):231–235. doi:10.1016/S0006-291X(02)02999-6
Allinson TM, Parkin ET, Turner AJ, Hooper NM (2003) ADAMs family members as amyloid precursor protein alpha-secretases. J Neurosci Res 74(3):342–352. doi:10.1002/jnr.10737
Deuss M, Reiss K, Hartmann D (2008) Part-time-secretases: the functional biology of ADAM 9, 10 and 17. Curr Alzheimer Res 5(2):187–201
Liang J, Kong Q (2012) α-Cleavage of cellular prion protein. Prion 6(5):453–460
van der Vorst EP, Keijbeck AA, de Winther MP, Donners MM (2012) A disintegrin and metalloproteases: molecular scissors in angiogenesis, inflammation and atherosclerosis. Atherosclerosis 224(2):302–308
Yamamoto S, Higuchi Y, Yoshiyama K, Shimizu E, Kataoka M, Hijiya N, Matsuura K (1999) ADAM family proteins in the immune system. Immunol Today 20(6):278–284. doi:10.1016/S0167-5699(99)01464-4
Huovila A-PJ, Turner AJ, Pelto-Huikko M, Kärkkäinen I, Ortiz RM (2005) Shedding light on ADAM metalloproteinases. Trends Biochem Sci 30(7):413–422. doi:10.1016/j.tibs.2005.05.006
Hartmann D, Tournoy J, Saftig P, Annaert W, De Strooper B (2001) Implication of APP secretases in notch signaling. J Mol Neurosci 17(2):171–181
Hartmann D, de Strooper B, Serneels L, Craessaerts K, Herreman A, Annaert W, Umans L, Lubke T, Lena Illert A, von Figura K, Saftig P (2002) The disintegrin/metalloprotease ADAM 10 is essential for Notch signalling but not for alpha-secretase activity in fibroblasts. Hum Mol Genet 11(21):2615–2624
Yang P, Baker KA, Hagg T (2006) The ADAMs family: coordinators of nervous system development, plasticity and repair. Prog Neurobiol 79(2):73–94
Postina R, Schroeder A, Dewachter I, Bohl J, Schmitt U, Kojro E, Prinzen C, Endres K, Hiemke C, Blessing M, Flamez P, Dequenne A, Godaux E, van Leuven F, Fahrenholz F (2004) A disintegrin-metalloproteinase prevents amyloid plaque formation and hippocampal defects in an Alzheimer disease mouse model. J Clin Invest 113(10):1456–1464. doi:10.1172/JCI20864
Epis R, Marcello E, Gardoni F, Vastagh C, Malinverno M, Balducci C, Colombo A, Borroni B, Vara H, Dell’Agli M, Cattabeni F, Giustetto M, Borsello T, Forloni G, Padovani A, Di Luca M (2010) Blocking ADAM10 synaptic trafficking generates a model of sporadic Alzheimer’s disease. Brain 133(11):3323–3335. doi:10.1093/brain/awq217
Arribas J, Esselens C (2009) ADAM17 as a therapeutic target in multiple diseases. Curr Pharm Des 15(20):2319–2335
Rose-John S (2013) ADAM17, shedding, TACE as therapeutic targets. Pharmacol Res 71:19–22
Scheller J, Chalaris A, Garbers C, Rose-John S (2011) ADAM17: a molecular switch to control inflammation and tissue regeneration. Trends Immunol 32(8):380–387
Blacker M, Noe MC, Carty TJ, Goodyer CG, LeBlanc AC (2002) Effect of tumor necrosis factor-alpha converting enzyme (TACE) and metalloprotease inhibitor on amyloid precursor protein metabolism in human neurons. J Neurochem 83(6):1349–1357
Hotoda N, Koike H, Sasagawa N, Ishiura S (2002) A secreted form of human ADAM9 has an α-secretase activity for APP. Biochem Biophys Res Commun 293(2):800–805. doi:10.1016/S0006-291X(02)00302-9
Huse JT, Pijak DS, Leslie GJ, Lee VM-Y, Doms RW (2000) Maturation and endosomal targeting of β-site amyloid precursor protein-cleaving enzyme: the Alzheimer’s disease β-secretase. J Biol Chem 275(43):33729–33737. doi:10.1074/jbc.M004175200
Taylor CJ, Ireland DR, Ballagh I, Bourne K, Marechal NM, Turner PR, Bilkey DK, Tate WP, Abraham WC (2008) Endogenous secreted amyloid precursor protein-α regulates hippocampal NMDA receptor function, long-term potentiation and spatial memory. Neurobiol Dis 31(2):250–260. doi:10.1016/j.nbd.2008.04.011
He W, Lu Y, Qahwash I, Hu X-Y, Chang A, Yan R (2004) Reticulon family members modulate BACE1 activity and amyloid-[beta] peptide generation. Nat Med 10(9):959–965. doi:10.1038/nm1088
Yokota T, Mishra M, Akatsu H, Tani Y, Miyauchi T, Yamamoto T, Kosaka K, Nagai Y, Sawada T, Heese K (2006) Brain site-specific gene expression analysis in Alzheimer’s disease patients. Eur J Clin Investig 36(11):820–830. doi:10.1111/j.1365-2362.2006.01722.x
Shi Q, Ge Y, Sharoar MG, He W, Xiang R, Zhang Z, Hu X, Yan R (2014) Impact of RTN3 deficiency on expression of BACE1 and amyloid deposition. J Neurosci 34(42):13954–13962. doi:10.1523/jneurosci. 1588-14.2014
Serneels L, Van Biervliet J, Craessaerts K, Dejaegere T, Horre K, Van Houtvin T, Esselmann H, Paul S, Schafer MK, Berezovska O, Hyman BT, Sprangers B, Sciot R, Moons L, Jucker M, Yang Z, May PC, Karran E, Wiltfang J, D’Hooge R, De Strooper B (2009) Gamma-secretase heterogeneity in the Aph1 subunit: relevance for Alzheimer’s disease. Science 324(5927):639–642. doi:10.1126/science.1171176
Dries DR, Yu G (2008) Assembly, maturation, and trafficking of the gamma-secretase complex in Alzheimer’s disease. Curr Alzheimer Res 5(2):132–146
Yamasaki A, Eimer S, Okochi M, Smialowska A, Kaether C, Baumeister R, Haass C, Steiner H (2006) The GxGD motif of presenilin contributes to catalytic function and substrate identification of gamma-secretase. J Neurosci 26(14):3821–3828. doi:10.1523/jneurosci. 5354-05.2006
Kretner B, Fukumori A, Kuhn P-H, Pérez-Revuelta BI, Lichtenthaler SF, Haass C, Steiner H (2013) Important functional role of residue x of the presenilin GxGD protease active site motif for APP substrate cleavage specificity and substrate selectivity of γ-secretase. J Neurochem 125(1):144–156. doi:10.1111/jnc.12124
Mori H, Takio K, Ogawara M, Selkoe DJ (1992) Mass spectrometry of purified amyloid beta protein in Alzheimer’s disease. J Biol Chem 267(24):17082–17086
Portelius E, Mattsson N, Andreasson U, Blennow K, Zetterberg H (2011) Novel aβ isoforms in Alzheimer’s disease—their role in diagnosis and treatment. Curr Pharm Des 17(25):2594–2602
Sgourakis NG, Yan Y, McCallum SA, Wang C, Garcia AE (2007) The Alzheimer’s peptides Aβ40 and 42 adopt distinct conformations in water: a combined MD/NMR study. J Mol Biol 368(5):1448–1457. doi:10.1016/j.jmb.2007.02.093
Lim KH, Collver HH, Le YTH, Nagchowdhuri P, Kenney JM (2007) Characterizations of distinct amyloidogenic conformations of the Aβ (1–40) and (1–42) peptides. Biochem Biophys Res Commun 353(2):443–449. doi:10.1016/j.bbrc.2006.12.043
Zhang Y, McLaughlin R, Goodyer C, LeBlanc A (2002) Selective cytotoxicity of intracellular amyloid β peptide1–42 through p53 and Bax in cultured primary human neurons. J Cell Biol 156(3):519–529. doi:10.1083/jcb.200110119
Mucke L, Masliah E, Yu G-Q, Mallory M, Rockenstein EM, Tatsuno G, Hu K, Kholodenko D, Johnson-Wood K, McConlogue L (2000) High-level neuronal expression of Aβ1–42 in wild-type human amyloid protein precursor transgenic mice: synaptotoxicity without plaque formation. J Neurosci 20(11):4050–4058
Burdick D, Soreghan B, Kwon M, Kosmoski J, Knauer M, Henschen A, Yates J, Cotman C, Glabe C (1992) Assembly and aggregation properties of synthetic Alzheimer’s A4/beta amyloid peptide analogs. J Biol Chem 267(1):546–554
Duff K, Eckman C, Zehr C, Yu X, Prada CM, Pereztur J, Hutton M, Buee L, Harigaya Y, Yager D, Morgan D, Gordon MN, Holcomb L, Refolo L, Zenk B, Hardy J, Younkin S (1996) Increased amyloid-beta 42(43) in brains of mice expressing mutant presenilin 1. Nature 383(6602):710–713. doi:10.1038/383710a0
Jan A, Gokce O, Luthi-Carter R, Lashuel HA (2008) The ratio of monomeric to aggregated forms of Abeta40 and Abeta42 is an important determinant of amyloid-beta aggregation, fibrillogenesis, and toxicity. J Biol Chem 283(42):28176–28189. doi:10.1074/jbc.M803159200
Kimberly WT, Zheng JB, Guenette SY, Selkoe DJ (2001) The intracellular domain of the beta-amyloid precursor protein is stabilized by Fe65 and translocates to the nucleus in a notch-like manner. J Biol Chem 276(43):40288–40292. doi:10.1074/jbc.C100447200
Gao Y, Pimplikar SW (2001) The gamma-secretase-cleaved C-terminal fragment of amyloid precursor protein mediates signaling to the nucleus. Proc Natl Acad Sci U S A 98(26):14979–14984. doi:10.1073/pnas.261463298
Cupers P, Orlans I, Craessaerts K, Annaert W, De Strooper B (2001) The amyloid precursor protein (APP)-cytoplasmic fragment generated by gamma-secretase is rapidly degraded but distributes partially in a nuclear fraction of neurones in culture. J Neurochem 78(5):1168–1178
Slomnicki LP, Lesniak W (2008) A putative role of the amyloid precursor protein intracellular domain (AICD) in transcription. Acta Neurobiol Exp (Wars) 68(2):219–228
Beckett C, Nalivaeva NN, Belyaev ND, Turner AJ (2012) Nuclear signalling by membrane protein intracellular domains: the AICD enigma. Cell Signal 24(2):402–409. doi:10.1016/j.cellsig.2011.10.007
Cao X, Südhof TC (2001) A transcriptively active complex of APP with Fe65 and histone acetyltransferase Tip60. Science 293(5527):115–120. doi:10.1126/science.1058783
Liu Q, Zerbinatti CV, Zhang J, Hoe H-S, Wang B, Cole SL, Herz J, Muglia L, Bu G (2007) Amyloid precursor protein regulates brain apolipoprotein E and cholesterol metabolism through lipoprotein receptor LRP1. Neuron 56(1):66–78. doi:10.1016/j.neuron.2007.08.008
von Rotz RC, Kohli BM, Bosset J, Meier M, Suzuki T, Nitsch RM, Konietzko U (2004) The APP intracellular domain forms nuclear multiprotein complexes and regulates the transcription of its own precursor. J Cell Sci 117(19):4435–4448. doi:10.1242/jcs.01323
Ryan KA, Pimplikar SW (2005) Activation of GSK-3 and phosphorylation of CRMP2 in transgenic mice expressing APP intracellular domain. J Cell Biol 171(2):327–335. doi:10.1083/jcb.200505078
Kim H-S, Kim E-M, Lee J-P, Park CH, Kim S, Seo J-H, Chang K-A, Yu E, Jeong S-J, Chong YH, Suh Y-H (2003) C-terminal fragments of amyloid precursor protein exert neurotoxicity by inducing glycogen synthase kinase-3β expression. FASEB J 17(13):1951–1953. doi: 10.1096/fj.03-0106fje
Checler F, Sunyach C, Pardossi-Piquard R, Sevalle J, Vincent B, Kawarai T, Girardot N, St George-Hyslop P, da Costa CA (2007) The gamma/epsilon-secretase-derived APP intracellular domain fragments regulate p53. Curr Alzheimer Res 4(4):423–426
Y-w Z, Wang R, Liu Q, Zhang H, Liao F-F, Xu H (2007) Presenilin/γ-secretase-dependent processing of β-amyloid precursor protein regulates EGF receptor expression. Proc Natl Acad Sci U S A 104(25):10613–10618. doi:10.1073/pnas.0703903104
Muller T, Loosse C, Schrotter A, Schnabel A, Helling S, Egensperger R, Marcus K (2011) The AICD interacting protein DAB1 is up-regulated in Alzheimer frontal cortex brain samples and causes deregulation of proteins involved in gene expression changes. Curr Alzheimer Res 8(5):573–582
Nakayama K, Ohkawara T, Hiratochi M, Koh CS, Nagase H (2008) The intracellular domain of amyloid precursor protein induces neuron-specific apoptosis. Neurosci Lett 444(2):127–131. doi:10.1016/j.neulet.2008.08.034
Roncarati R, Sestan N, Scheinfeld MH, Berechid BE, Lopez PA, Meucci O, McGlade JC, Rakic P, D’Adamio L (2002) The gamma-secretase-generated intracellular domain of beta-amyloid precursor protein binds Numb and inhibits Notch signaling. Proc Natl Acad Sci U S A 99(10):7102–7107. doi:10.1073/pnas.102192599
Hu QD, Ang BT, Karsak M, Hu WP, Cui XY, Duka T, Takeda Y, Chia W, Sankar N, Ng YK, Ling EA, Maciag T, Small D, Trifonova R, Kopan R, Okano H, Nakafuku M, Chiba S, Hirai H, Aster JC, Schachner M, Pallen CJ, Watanabe K, Xiao ZC (2003) F3/contactin acts as a functional ligand for Notch during oligodendrocyte maturation. Cell 115(2):163–175
Ghosal K, Vogt DL, Liang M, Shen Y, Lamb BT, Pimplikar SW (2009) Alzheimer’s disease-like pathological features in transgenic mice expressing the APP intracellular domain. Proc Natl Acad Sci U S A 106(43):18367–18372. doi:10.1073/pnas.0907652106
Konietzko U (2012) AICD nuclear signaling and its possible contribution to Alzheimer’s disease. Curr Alzheimer Res 9(2):200–216
Yankner B, Duffy L, Kirschner D (1990) Neurotrophic and neurotoxic effects of amyloid beta protein: reversal by tachykinin neuropeptides. Science 250(4978):279–282. doi:10.1126/science.2218531
Whitson J, Selkoe D, Cotman C (1989) Amyloid beta protein enhances the survival of hippocampal neurons in vitro. Science 243(4897):1488–1490. doi:10.1126/science.2928783
Chen Y, Dong C (2008) A[beta]40 promotes neuronal cell fate in neural progenitor cells. Cell Death Differ 16(3):386–394
Giuffrida M, Tomasello F, Caraci F, Chiechio S, Nicoletti F, Copani A (2012) Beta-amyloid monomer and insulin/IGF-1 signaling in Alzheimer’s disease. Mol Neurobiol 46(3):605–613. doi:10.1007/s12035-012-8313-6
Calafiore M, Battaglia G, Zappala A, Trovato-Salinaro E, Caraci F, Caruso M, Vancheri C, Sortino MA, Nicoletti F, Copani A (2006) Progenitor cells from the adult mouse brain acquire a neuronal phenotype in response to beta-amyloid. Neurobiol Aging 27(4):606–613. doi:10.1016/j.neurobiolaging.2005.03.019
Lopez-Toledano MA, Shelanski ML (2004) Neurogenic effect of beta-amyloid peptide in the development of neural stem cells. J Neurosci 24(23):5439–5444. doi:10.1523/JNEUROSCI. 0974-04.2004
Reddy PH (2009) Amyloid beta, mitochondrial structural and functional dynamics in Alzheimer’s disease. Exp Neurol 218(2):286–292. doi:10.1016/j.expneurol.2009.03.042
Butterfield DA, Yatin SM, Link CD (1999) In vitro and in vivo protein oxidation induced by Alzheimer’s disease amyloid beta-peptide (1-42). Ann N Y Acad Sci 893:265–268
Yatin SM, Varadarajan S, Link CD, Butterfield DA (1999) In vitro and in vivo oxidative stress associated with Alzheimer’s amyloid beta-peptide (1-42). Neurobiol Aging 20(3):325–330, discussion 339–342
Yatin SM, Aksenova M, Aksenov M, Markesbery WR, Aulick T, Butterfield DA (1998) Temporal relations among amyloid beta-peptide-induced free-radical oxidative stress, neuronal toxicity, and neuronal defensive responses. J Mol Neurosci 11(3):183–197. doi:10.1385/JMN:11:3:183
Almeida CG, Takahashi RH, Gouras GK (2006) β-amyloid accumulation impairs multivesicular body sorting by inhibiting the ubiquitin-proteasome system. J Neurosci 26(16):4277–4288. doi:10.1523/jneurosci. 5078-05.2006
Busciglio J, Lorenzo A, Yeh J, Yankner BA (1995) β-Amyloid fibrils induce tau phosphorylation and loss of microtubule binding. Neuron 14(4):879–888. doi:10.1016/0896-6273(95)90232-5
Zempel H, Thies E, Mandelkow E, Mandelkow EM (2010) Abeta oligomers cause localized Ca(2+) elevation, missorting of endogenous Tau into dendrites, Tau phosphorylation, and destruction of microtubules and spines. J Neurosci 30(36):11938–11950. doi:10.1523/jneurosci. 2357-10.2010
Butterfield DA, Drake J, Pocernich C, Castegna A (2001) Evidence of oxidative damage in Alzheimer’s disease brain: central role for amyloid β-peptide. Trends Mol Med 7(12):548–554. doi:10.1016/S1471-4914(01)02173-6
Axelsen PH, Komatsu H, Murray IVJ (2011) Oxidative stress and cell membranes in the pathogenesis of Alzheimer’s disease. Physiology 26(1):54–69. doi:10.1152/physiol.00024.2010
Adibhatla RM, Hatcher JF (2010) Lipid oxidation and peroxidation in CNS health and disease: from molecular mechanisms to therapeutic opportunities. Antioxid Redox Signal 12(1):125–169. doi:10.1089/ars.2009.2668
Rauchova H, Vokurkova M, Koudelova J (2012) Hypoxia-induced lipid peroxidation in the brain during postnatal ontogenesis. Physiol Res 61(Suppl 1):S89–S101
De Georgia MA (2014) Brain tissue oxygen monitoring in neurocritical care. J Intensive Care Med. doi:10.1177/0885066614529254
Radak D, Resanovic I, Isenovic ER (2014) Link between oxidative stress and acute brain ischemia. Angiology 65(8):667–676. doi:10.1177/0003319713506516
Rodrigo R, Fernandez-Gajardo R, Gutierrez R, Matamala JM, Carrasco R, Miranda-Merchak A, Feuerhake W (2013) Oxidative stress and pathophysiology of ischemic stroke: novel therapeutic opportunities. CNS Neurol Disord Drug Targets 12(5):698–714
Smith MA, Hirai K, Hsiao K, Pappolla MA, Harris PLR, Siedlak SL, Tabaton M, Perry G (1998) Amyloid-β deposition in Alzheimer transgenic mice is associated with oxidative stress. J Neurochem 70(5):2212–2215. doi:10.1046/j.1471-4159.1998.70052212.x
Xie H, Hou S, Jiang J, Sekutowicz M, Kelly J, Bacskai BJ (2013) Rapid cell death is preceded by amyloid plaque-mediated oxidative stress. Proc Natl Acad Sci U S A 110(19):7904–7909. doi:10.1073/pnas.1217938110
Behl C, Davis JB, Lesley R, Schubert D (1994) Hydrogen peroxide mediates amyloid β protein toxicity. Cell 77(6):817–827. doi:10.1016/0092-8674(94)90131-7
Kontush A, Berndt C, Weber W, Akopyan V, Arlt S, Schippling S, Beisiegel U (2001) Amyloid-β is an antioxidant for lipoproteins in cerebrospinal fluid and plasma. Free Radic Biol Med 30(1):119–128. doi:10.1016/S0891-5849(00)00458-5
Kontush A, Donarski N, Beisiegel U (2001) Resistance of human cerebrospinal fluid to in vitro oxidation is directly related to its amyloid-β content. Free Radic Res 35(5):507–517. doi:10.1080/10715760100301521
Zou K, Gong JS, Yanagisawa K, Michikawa M (2002) A novel function of monomeric amyloid β-protein serving as an antioxidant molecule against metal-induced oxidative damage. J Neurosci 22(12):4833–4841
Smith MA, Casadesus G, Joseph JA, Perry G (2002) Amyloid-β and τ serve antioxidant functions in the aging and Alzheimer brain. Free Radic Biol Med 33(9):1194–1199. doi:10.1016/S0891-5849(02)01021-3
Markesbery WR (1997) Oxidative stress hypothesis in Alzheimer’s disease. Free Radic Biol Med 23(1):134–147. doi:10.1016/S0891-5849(96)00629-6
Skaper SD (2012) Alzheimer’s disease and amyloid: culprit or coincidence. Int Rev Neurobiol 102:277–316
Golde TE, Dickson D, Hutton M (2006) Filling the gaps in the a cascade hypothesis of Alzheimer’s disease. Curr Alzheimer Res 3(5):421–430
Drachman DA (2014) The amyloid hypothesis, time to move on: amyloid is the downstream result, not cause, of Alzheimer’s disease. Alzheimers Dement 10(3):372–380
Krstic D, Knuesel I (2013) The airbag problem-a potential culprit for bench-to-bedside translational efforts: relevance for Alzheimer’s disease. Acta Neuropathol Commun 1(1):62. doi:10.1186/2051-5960-1-62
Zempel H, Mandelkow E-M (2011) Linking amyloid-β and tau: amyloid-β induced synaptic dysfunction via local wreckage of the neuronal cytoskeleton. Neurodegener Dis 10(1–4):64–72
Zempel H, Luedtke J, Kumar Y, Biernat J, Dawson H, Mandelkow E, Mandelkow EM (2013) Amyloid-beta oligomers induce synaptic damage via Tau-dependent microtubule severing by TTLL6 and spastin. EMBO J 32(22):2920–2937. doi:10.1038/emboj.2013.207
Ferreira A, Lu Q, Orecchio L, Kosik KS (1997) Selective phosphorylation of adult tau isoforms in mature hippocampal neurons exposed to fibrillar Aβ. Mol Cell Neurosci 9(3):220–234. doi:10.1006/mcne.1997.0615
Yu W, Polepalli J, Wagh D, Rajadas J, Malenka R, Lu B (2012) A critical role for the PAR-1/MARK-tau axis in mediating the toxic effects of Aβ on synapses and dendritic spines. Hum Mol Genet 21(6):1384–1390
Mairet-Coello G, Courchet J, Pieraut S, Courchet V, Maximov A, Polleux F (2013) The CAMKK2-AMPK kinase pathway mediates the synaptotoxic effects of Aβ oligomers through tau phosphorylation. Neuron 78(1):94–108
Zheng W-H, Bastianetto S, Mennicken F, Ma W, Kar S (2002) Amyloid β peptide induces tau phosphorylation and loss of cholinergic neurons in rat primary septal cultures. Neuroscience 115(1):201–211
Lewis J, Dickson DW, Lin W-L, Chisholm L, Corral A, Jones G, Yen S-H, Sahara N, Skipper L, Yager D, Eckman C, Hardy J, Hutton M, McGowan E (2001) Enhanced neurofibrillary degeneration in transgenic mice expressing mutant tau and APP. Science 293(5534):1487–1491. doi:10.1126/science.1058189
Spillantini MG, Goedert M (1998) Tau protein pathology in neurodegenerative diseases. Trends Neurosci 21(10):428–433. doi:10.1016/S0166-2236(98)01337-X
Mandelkow E-M, Mandelkow E (1998) Tau in Alzheimer’s disease. Trends Cell Biol 8(11):425–427. doi:10.1016/S0962-8924(98)01368-3
Tolnay P (1999) Review: tau protein pathology in Alzheimer’s disease and related disorders. Neuropathol Appl Neurobiol 25(3):171–187. doi:10.1046/j.1365-2990.1999.00182.x
Brandt R, Hundelt M, Shahani N (2005) Tau alteration and neuronal degeneration in tauopathies: mechanisms and models. Biochim Biophys Acta (BBA)-Mol Basis Dis 1739(2):331–354
Ludolph A, Kassubek J, Landwehrmeyer B, Mandelkow E, Mandelkow EM, Burn D, Caparros‐Lefebvre D, Frey K, De Yebenes J, Gasser T (2009) Tauopathies with parkinsonism: clinical spectrum, neuropathologic basis, biological markers, and treatment options. Eur J Neurol 16(3):297–309
Ihara Y, Abraham C, Selkoe DJ (1983) Antibodies to paired helical filaments in Alzheimer’s disease do not recognize normal brain proteins. Nature 304(5928):727–730
Grundke-Iqbal I, Iqbal K, Quinlan M, Tung YC, Zaidi MS, Wisniewski HM (1986) Microtubule-associated protein tau. A component of Alzheimer paired helical filaments. J Biol Chem 261(13):6084–6089
Grundke-Iqbal I, Iqbal K, Tung YC, Quinlan M, Wisniewski HM, Binder LI (1986) Abnormal phosphorylation of the microtubule-associated protein tau (tau) in Alzheimer cytoskeletal pathology. Proc Natl Acad Sci U S A 83(13):4913–4917
Cleveland DW, Hwo S-Y, Kirschner MW (1977) Purification of tau, a microtubule-associated protein that induces assembly of microtubules from purified tubulin. J Mol Biol 116(2):207–225
Weingarten MD, Lockwood AH, Hwo SY, Kirschner MW (1975) A protein factor essential for microtubule assembly. Proc Natl Acad Sci U S A 72(5):1858–1862
Witman GB, Cleveland DW, Weingarten MD, Kirschner MW (1976) Tubulin requires tau for growth onto microtubule initiating sites. Proc Natl Acad Sci U S A 73(11):4070–4074
Schoenfeld TA, Obar RA (1994) Diverse distribution and function of fibrous microtubule-associated proteins in the nervous system. Int Rev Cytol 151:67–137
Buée L, Bussière T, Buée-Scherrer V, Delacourte A, Hof PR (2000) Tau protein isoforms, phosphorylation and role in neurodegenerative disorders. Brain Res Rev 33(1):95–130. doi:10.1016/S0165-0173(00)00019-9
Binder LI, Frankfurter A, Rebhun LI (1985) The distribution of tau in the mammalian central nervous system. J Cell Biol 101(4):1371–1378. doi:10.1083/jcb.101.4.1371
González-Billault C, Engelke M, Jiménez-Mateos EM, Wandosell F, Cáceres A, Avila J (2002) Participation of structural microtubule-associated proteins (MAPs) in the development of neuronal polarity. J Neurosci Res 67(6):713–719. doi:10.1002/jnr.10161
Caceres A, Mautino J, Kosik KS (1992) Suppression of MAP2 in cultured cerebellar macroneurons inhibits minor neurite formation. Neuron 9(4):607–618. doi:10.1016/0896-6273(92)90025-9
Liu C-wA, Lee G, Jay DG (1999) Tau is required for neurite outgrowth and growth cone motility of chick sensory neurons. Cell Motil Cytoskeleton 43(3):232–242. doi:10.1002/(SICI)1097-0169(1999)43:3<232::AID-CM6>3.0.CO;2-7
Kempf M, Clement A, Faissner A, Lee G, Brandt R (1996) Tau binds to the distal axon early in development of polarity in a microtubule- and microfilament-dependent manner. J Neurosci 16(18):5583–5592
Ebneth A, Godemann R, Stamer K, Illenberger S, Trinczek B, Mandelkow E-M, Mandelkow E (1998) Overexpression of Tau protein inhibits kinesin-dependent trafficking of vesicles, mitochondria, and endoplasmic reticulum: implications for Alzheimer’s disease. J Cell Biol 143(3):777–794. doi:10.1083/jcb.143.3.777
Terwel D, Dewachter I, Van Leuven F (2002) Axonal transport, tau protein, and neurodegeneration in Alzheimer’s disease. Neuromol Med 2(2):151–165. doi:10.1385/NMM:2:2:151
Kanaan NM, Pigino GF, Brady ST, Lazarov O, Binder LI, Morfini GA (2013) Axonal degeneration in Alzheimer’s disease: when signaling abnormalities meet the axonal transport system. Exp Neurol 246:44–53. doi:10.1016/j.expneurol.2012.06.003
Morfini GA, Burns M, Binder LI, Kanaan NM, LaPointe N, Bosco DA, Brown RH, Brown H, Tiwari A, Hayward L, Edgar J, Nave K-A, Garberrn J, Atagi Y, Song Y, Pigino G, Brady ST (2009) Axonal transport defects in neurodegenerative diseases. J Neurosci 29(41):12776–12786. doi:10.1523/jneurosci. 3463-09.2009
De Vos KJ, Grierson AJ, Ackerley S, Miller CCJ (2008) Role of axonal transport in neurodegenerative diseases. Annu Rev Neurosci 31(1):151–173. doi:10.1146/annurev.neuro.31.061307.090711
Hirokawa N, Shiomura Y, Okabe S (1988) Tau proteins: the molecular structure and mode of binding on microtubules. J Cell Biol 107(4):1449–1459
Avila J, Lucas JJ, Perez M, Hernandez F (2004) Role of tau protein in both physiological and pathological conditions. Physiol Rev 84(2):361–384. doi:10.1152/physrev.00024.2003
Goedert M, Spillantini MG, Jakes R, Rutherford D, Crowther RA (1989) Multiple isoforms of human microtubule-associated protein tau: sequences and localization in neurofibrillary tangles of Alzheimer’s disease. Neuron 3(4):519–526. doi:10.1016/0896-6273(89)90210-9
Goedert M, Spillantini MG, Cairns NJ, Crowther RA (1992) Tau proteins of Alzheimer paired helical filaments: abnormal phosphorylation of all six brain isoforms. Neuron 8(1):159–168. doi:10.1016/0896-6273(92)90117-V
Avila J (2006) Tau phosphorylation and aggregation in Alzheimer’s disease pathology. FEBS Lett 580(12):2922–2927. doi:10.1016/j.febslet.2006.02.067
Ferrer I, Gomez-Isla T, Puig B, Freixes M, Ribe E, Dalfo E, Avila J (2005) Current advances on different kinases involved in tau phosphorylation, and implications in Alzheimer’s disease and tauopathies. Curr Alzheimers Res 2(1):3–18
Gong CX, Iqbal K (2008) Hyperphosphorylation of microtubule-associated protein tau: a promising therapeutic target for Alzheimer disease. Curr Med Chem 15(23):2321–2328
Schwalbe M, Biernat J, Bibow S, Ozenne V, Jensen MR, Kadavath H, Blackledge M, Mandelkow E, Zweckstetter M (2013) Phosphorylation of human tau protein by microtubule affinity-regulating kinase 2. Biochemistry (Mosc) 52(50):9068–9079. doi:10.1021/bi401266n
Drewes G, Lichtenberg-Kraag B, Doring F, Mandelkow EM, Biernat J, Goris J, Doree M, Mandelkow E (1992) Mitogen activated protein (MAP) kinase transforms tau protein into an Alzheimer-like state. EMBO J 11(6):2131–2138
Mandelkow EM, Drewes G, Biernat J, Gustke N, Van Lint J, Vandenheede JR, Mandelkow E (1992) Glycogen synthase kinase-3 and the Alzheimer-like state of microtubule-associated protein tau. FEBS Lett 314(3):315–321
Demuro A, Parker I, Stutzmann GE (2010) Calcium signaling and amyloid toxicity in Alzheimer disease. J Biol Chem 285(17):12463–12468. doi:10.1074/jbc.R109.080895
Lin K-F, Chang RC-C, Suen K-C, So K-F, Hugon J (2004) Modulation of calcium/calmodulin kinase-II provides partial neuroprotection against beta-amyloid peptide toxicity. Eur J Neurosci 19(8):2047–2055. doi:10.1111/j.0953-816X.2004.03245.x
Steen E, Terry BM, Rivera EJ, Cannon JL, Neely TR, Tavares R, Xu XJ, Wands JR, de la Monte SM (2005) Impaired insulin and insulin-like growth factor expression and signaling mechanisms in Alzheimer’s disease—is this type 3 diabetes? J Alzheimers Dis 7(1):63–80
Correia SC, Santos RX, Perry G, Zhu X, Moreira PI, Smith MA (2011) Insulin-resistant brain state: the culprit in sporadic Alzheimer’s disease? Ageing Res Rev 10(2):264–273. doi:10.1016/j.arr.2011.01.001
Blass JP (2001) Brain metabolism and brain disease: is metabolic deficiency the proximate cause of Alzheimer dementia? J Neurosci Res 66(5):851–856. doi:10.1002/jnr.10087
Lindwall G, Cole RD (1984) Phosphorylation affects the ability of tau protein to promote microtubule assembly. J Biol Chem 259(8):5301–5305
Gustke N, Steiner B, Mandelkow EM, Biernat J, Meyer HE, Goedert M, Mandelkow E (1992) The Alzheimer-like phosphorylation of tau protein reduces microtubule binding and involves Ser-Pro and Thr-Pro motifs. FEBS Lett 307(2):199–205
Liu F, Grundke-Iqbal I, Iqbal K, Gong CX (2005) Contributions of protein phosphatases PP1, PP2A, PP2B and PP5 to the regulation of tau phosphorylation. Eur J Neurosci 22(8):1942–1950. doi:10.1111/j.1460-9568.2005.04391.x
Watanabe A, Takio K, Ihara Y (1999) Deamidation and isoaspartate formation in smeared tau in paired helical filaments: unusual properties of the microtubule-binding domain of tau. J Biol Chem 274(11):7368–7378. doi:10.1074/jbc.274.11.7368
Min S-W, Cho S-H, Zhou Y, Schroeder S, Haroutunian V, Seeley WW, Huang EJ, Shen Y, Masliah E, Mukherjee C, Meyers D, Cole PA, Ott M, Gan L (2010) Acetylation of tau inhibits its degradation and contributes to tauopathy. Neuron 67(6):953–966. doi:10.1016/j.neuron.2010.08.044
Cohen TJ, Guo JL, Hurtado DE, Kwong LK, Mills IP, Trojanowski JQ, Lee VMY (2011) The acetylation of tau inhibits its function and promotes pathological tau aggregation. Nat Commun 2:252. doi:10.1038/ncomms1255
Dickey CA, Kamal A, Lundgren K, Klosak N, Bailey RM, Dunmore J, Ash P, Shoraka S, Zlatkovic J, Eckman CB, Patterson C, Dickson DW, Nahman NS Jr, Hutton M, Burrows F, Petrucelli L (2007) The high-affinity HSP90-CHIP complex recognizes and selectively degrades phosphorylated tau client proteins. J Clin Invest 117(3):648–658. doi:10.1172/JCI29715
Tompa P (2002) Intrinsically unstructured proteins. Trends Biochem Sci 27(10):527–533. doi:10.1016/S0968-0004(02)02169-2
Uversky VN, Oldfield CJ, Dunker AK (2008) Intrinsically disordered proteins in human diseases: introducing the D2 concept. Annu Rev Biophys 37(1):215–246. doi:10.1146/annurev.biophys.37.032807.125924
Skrabana R, Sevcik J, Novak M (2006) Intrinsically disordered proteins in the neurodegenerative processes: formation of tau protein paired helical filaments and their analysis. Cell Mol Neurobiol 26(7–8):1083–1095. doi:10.1007/s10571-006-9083-3
Dyson HJ, Wright PE (2005) Intrinsically unstructured proteins and their functions. Nat Rev Mol Cell Biol 6(3):197–208
Uversky VN (2010) Targeting intrinsically disordered proteins in neurodegenerative and protein dysfunction diseases: another illustration of the D(2) concept. Expert Rev Proteomics 7(4):543–564. doi:10.1586/epr.10.36
Nonaka T, Masuda-Suzukake M, Arai T, Hasegawa Y, Akatsu H, Obi T, Yoshida M, Murayama S, Mann DM, Akiyama H, Hasegawa M (2013) Prion-like properties of pathological TDP-43 aggregates from diseased brains. Cell Rep 4(1):124–134. doi:10.1016/j.celrep.2013.06.007
Tsuiji H, Iguchi Y, Furuya A, Kataoka A, Hatsuta H, Atsuta N, Tanaka F, Hashizume Y, Akatsu H, Murayama S, Sobue G, Yamanaka K (2013) Spliceosome integrity is defective in the motor neuron diseases ALS and SMA. EMBO Mol Med 5(2):221–234. doi:10.1002/emmm.201202303
Eisele YS, Bolmont T, Heikenwalder M, Langer F, Jacobson LH, Yan ZX, Roth K, Aguzzi A, Staufenbiel M, Walker LC, Jucker M (2009) Induction of cerebral beta-amyloidosis: intracerebral versus systemic Abeta inoculation. Proc Natl Acad Sci U S A 106(31):12926–12931. doi:10.1073/pnas.0903200106
Langer F, Eisele YS, Fritschi SK, Staufenbiel M, Walker LC, Jucker M (2011) Soluble Abeta seeds are potent inducers of cerebral beta-amyloid deposition. J Neurosci 31(41):14488–14495. doi:10.1523/JNEUROSCI. 3088-11.2011
Brundin P, Melki R, Kopito R (2010) Prion-like transmission of protein aggregates in neurodegenerative diseases. Nat Rev Mol Cell Biol 11(4):301–307. doi:10.1038/nrm2873
Jucker M, Walker LC (2013) Self-propagation of pathogenic protein aggregates in neurodegenerative diseases. Nature 501(7465):45–51. doi:10.1038/nature12481
Braak H, Del Tredici K (2011) Alzheimer’s pathogenesis: is there neuron-to-neuron propagation? Acta Neuropathol 121(5):589–595. doi:10.1007/s00401-011-0825-z
Lee S-J, Desplats P, Sigurdson C, Tsigelny I, Masliah E (2010) Cell-to-cell transmission of non-prion protein aggregates. Nat Rev Neurol 6(12):702–706
Prusiner SB (2012) A unifying role for prions in neurodegenerative diseases. Science 336(6088):1511–1513
Goedert M, Clavaguera F, Tolnay M (2010) The propagation of prion-like protein inclusions in neurodegenerative diseases. Trends Neurosci 33(7):317–325. doi:10.1016/j.tins.2010.04.003
Eisele YS, Obermuller U, Heilbronner G, Baumann F, Kaeser SA, Wolburg H, Walker LC, Staufenbiel M, Heikenwalder M, Jucker M (2010) Peripherally applied Abeta-containing inoculates induce cerebral beta-amyloidosis. Science 330(6006):980–982. doi:10.1126/science.1194516
Meyer-Luehmann M, Coomaraswamy J, Bolmont T, Kaeser S, Schaefer C, Kilger E, Neuenschwander A, Abramowski D, Frey P, Jaton AL, Vigouret JM, Paganetti P, Walsh DM, Mathews PM, Ghiso J, Staufenbiel M, Walker LC, Jucker M (2006) Exogenous induction of cerebral beta-amyloidogenesis is governed by agent and host. Science 313(5794):1781–1784. doi:10.1126/science.1131864
Soto C, Estrada L, Castilla J (2006) Amyloids, prions and the inherent infectious nature of misfolded protein aggregates. Trends Biochem Sci 31(3):150–155. doi:10.1016/j.tibs.2006.01.002
Clavaguera F, Bolmont T, Crowther RA, Abramowski D, Frank S, Probst A, Fraser G, Stalder AK, Beibel M, Staufenbiel M, Jucker M, Goedert M, Tolnay M (2009) Transmission and spreading of tauopathy in transgenic mouse brain. Nat Cell Biol 11(7):909–913. doi:10.1038/ncb1901
Frost B, Jacks RL, Diamond MI (2009) Propagation of tau misfolding from the outside to the inside of a cell. J Biol Chem 284(19):12845–12852. doi:10.1074/jbc.M808759200
Guo JL, Lee VM-Y (2011) Seeding of normal tau by pathological tau conformers drives pathogenesis of Alzheimer-like tangles. J Biol Chem 286(17):15317–15331. doi:10.1074/jbc.M110.209296
Clavaguera F, Akatsu H, Fraser G, Crowther RA, Frank S, Hench J, Probst A, Winkler DT, Reichwald J, Staufenbiel M, Ghetti B, Goedert M, Tolnay M (2013) Brain homogenates from human tauopathies induce tau inclusions in mouse brain. Proc Natl Acad Sci U S A 110(23):9535–9540. doi:10.1073/pnas.1301175110
Desplats P, Lee HJ, Bae EJ, Patrick C, Rockenstein E, Crews L, Spencer B, Masliah E, Lee SJ (2009) Inclusion formation and neuronal cell death through neuron-to-neuron transmission of alpha-synuclein. Proc Natl Acad Sci U S A 106(31):13010–13015. doi:10.1073/pnas.0903691106
Mougenot A-L, Nicot S, Bencsik A, Morignat E, Verchère J, Lakhdar L, Legastelois S, Baron T (2012) Prion-like acceleration of a synucleinopathy in a transgenic mouse model. Neurobiol Aging 33(9):2225–2228. doi:10.1016/j.neurobiolaging.2011.06.022
Masuda-Suzukake M, Nonaka T, Hosokawa M, Oikawa T, Arai T, Akiyama H, Mann DM, Hasegawa M (2013) Prion-like spreading of pathological alpha-synuclein in brain. Brain 136(Pt 4):1128–1138. doi:10.1093/brain/awt037
Ren PH, Lauckner JE, Kachirskaia I, Heuser JE, Melki R, Kopito RR (2009) Cytoplasmic penetration and persistent infection of mammalian cells by polyglutamine aggregates. Nat Cell Biol 11(2):219–225. doi:10.1038/ncb1830
Münch C, O’Brien J, Bertolotti A (2011) Prion-like propagation of mutant superoxide dismutase-1 misfolding in neuronal cells. Proc Natl Acad Sci U S A 108(9):3548–3553. doi:10.1073/pnas.1017275108
Moreno-Gonzalez I, Soto C (2011) Misfolded protein aggregates: mechanisms, structures and potential for disease transmission. Semin Cell Dev Biol 22(5):482–487. doi:10.1016/j.semcdb.2011.04.002
Jaiswal JK, Fix M, Takano T, Nedergaard M, Simon SM (2007) Resolving vesicle fusion from lysis to monitor calcium-triggered lysosomal exocytosis in astrocytes. Proc Natl Acad Sci U S A 104(35):14151–14156. doi:10.1073/pnas.0704935104
Mohamed NV, Herrou T, Plouffe V, Piperno N, Leclerc N (2013) Spreading of tau pathology in Alzheimer’s disease by cell-to-cell transmission. Eur J Neurosci 37(12):1939–1948. doi:10.1111/ejn.12229
Gousset K, Schiff E, Langevin C, Marijanovic Z, Caputo A, Browman DT, Chenouard N, de Chaumont F, Martino A, Enninga J, Olivo-Marin JC, Mannel D, Zurzolo C (2009) Prions hijack tunnelling nanotubes for intercellular spread. Nat Cell Biol 11(3):328–336. doi:10.1038/ncb1841
Ciechanover A, Brundin P (2003) The ubiquitin proteasome system in neurodegenerative diseases: sometimes the chicken, sometimes the egg. Neuron 40(2):427–446. doi:10.1016/S0896-6273(03)00606-8
Manavalan A, Mishra M, Feng L, Sze SK, Akatsu H, Heese K (2013) Brain site-specific proteome changes in aging-related dementia. Exp Mol Med 45:e39. doi:10.1038/emm.2013.76
Hershko A, Ciechanover A, Rose IA (1979) Resolution of the ATP-dependent proteolytic system from reticulocytes: a component that interacts with ATP. Proc Natl Acad Sci U S A 76(7):3107–3110
Thompson SJ, Loftus LT, Ashley MD, Meller R (2008) Ubiquitin-proteasome system as a modulator of cell fate. Curr Opin Pharmacol 8(1):90–95. doi:10.1016/j.coph.2007.09.010
Jana NR (2012) Protein homeostasis and aging: role of ubiquitin protein ligases. Neurochem Int 60(5):443–447. doi:10.1016/j.neuint.2012.02.009
Weissman AM, Shabek N, Ciechanover A (2011) The predator becomes the prey: regulating the ubiquitin system by ubiquitylation and degradation. Nat Rev Mol Cell Biol 12(9):605–620
Dennissen FJ, Kholod N, van Leeuwen FW (2012) The ubiquitin proteasome system in neurodegenerative diseases: culprit, accomplice or victim? Prog Neurobiol 96(2):190–207. doi:10.1016/j.pneurobio.2012.01.003
Pickart CM, Eddins MJ (2004) Ubiquitin: structures, functions, mechanisms. Biochim Biophys Acta (BBA) - Mol Cell Res 1695(1–3):55–72. doi:10.1016/j.bbamcr.2004.09.019
Scheffner M, Nuber U, Huibregtse JM (1995) Protein ubiquitination involving an E1-E2-E3 enzyme ubiquitin thioester cascade. Nature 373(6509):81–83
Ciechanover A, Schwartz AL (2004) The ubiquitin system: pathogenesis of human diseases and drug targeting. Biochim Biophys Acta (BBA) - Mol Cell Res 1695(1–3):3–17. doi:10.1016/j.bbamcr.2004.09.018
Nisole S, Stoye JP, Saib A (2005) TRIM family proteins: retroviral restriction and antiviral defence. Nat Rev Microbiol 3(10):799–808. doi:10.1038/nrmicro1248
Hatakeyama S (2011) TRIM proteins and cancer. Nat Rev Cancer 11(11):792–804. doi:10.1038/nrc3139
Meroni G, Diez-Roux G (2005) TRIM/RBCC, a novel class of ‘single protein RING finger’ E3 ubiquitin ligases. Bioessays 27(11):1147–1157. doi:10.1002/bies.20304
Avela K, Lipsanen-Nyman M, Idänheimo N, Seemanová E, Rosengren S, Mäkelä TP, Perheentupa J, de la Chapelle A, Lehesjoki A-E (2000) Gene encoding a new RING-B-box-Coiled-coil protein is mutated in mulibrey nanism. Nat Genet 25(3):298–301
Horn EJ, Albor A, Liu Y, El-Hizawi S, Vanderbeek GE, Babcock M, Bowden GT, Hennings H, Lozano G, Weinberg WC, Kulesz-Martin M (2004) RING protein Trim32 associated with skin carcinogenesis has anti-apoptotic and E3-ubiquitin ligase properties. Carcinogenesis 25(2):157–167. doi:10.1093/carcin/bgh003
Schwamborn JC, Berezikov E, Knoblich JA (2009) The TRIM-NHL protein TRIM32 activates microRNAs and prevents self-renewal in mouse neural progenitors. Cell 136(5):913–925. doi:10.1016/j.cell.2008.12.024
Kudryashova E, Kudryashov D, Kramerova I, Spencer MJ (2005) Trim32 is a ubiquitin ligase mutated in limb girdle muscular dystrophy type 2H that binds to skeletal muscle myosin and ubiquitinates actin. J Mol Biol 354(2):413–424. doi:10.1016/j.jmb.2005.09.068
Kudryashova E, Wu J, Havton LA, Spencer MJ (2009) Deficiency of the E3 ubiquitin ligase TRIM32 in mice leads to a myopathy with a neurogenic component. Hum Mol Genet 18(7):1353–1367. doi:10.1093/hmg/ddp036
Higashi S, Iseki E, Yamamoto R, Minegishi M, Hino H, Fujisawa K, Togo T, Katsuse O, Uchikado H, Furukawa Y, Kosaka K, Arai H (2007) Concurrence of TDP-43, tau and α-synuclein pathology in brains of Alzheimer’s disease and dementia with Lewy bodies. Brain Res 1184:284–294. doi:10.1016/j.brainres.2007.09.048
Ishizawa T, Mattila P, Davies P, Wang D, Dickson DW (2003) Colocalization of Tau and alpha‐synuclein epitopes in lewy bodies. J Neuropathol Exp Neurol 62(4):389–397
Zhang Y, Dawson VL, Dawson TM (2001) Parkin: clinical aspects and neurobiology. Clin Neurosci Res 1(6):467–482. doi:10.1016/S1566-2772(01)00025-1
Dawson TM, Dawson VL (2003) Molecular pathways of neurodegeneration in Parkinson’s disease. Science 302(5646):819–822. doi:10.1126/science.1087753
McNaught KSP, Björklund LM, Belizaire R, Isacson O, Jenner P, Olanow CW (2002) Proteasome inhibition causes nigral degeneration with inclusion bodies in rats. Neuroreport 13(11):1437–1441
McNaught KSP, Mytilineou C, JnoBaptiste R, Yabut J, Shashidharan P, Jenner P, Olanow CW (2002) Impairment of the ubiquitin-proteasome system causes dopaminergic cell death and inclusion body formation in ventral mesencephalic cultures. J Neurochem 81(2):301–306. doi:10.1046/j.1471-4159.2002.00821.x
van Eersel J, Ke YD, Gladbach A, Bi M, Götz J, Kril JJ, Ittner LM (2011) Cytoplasmic accumulation and aggregation of TDP-43 upon proteasome inhibition in cultured neurons. PLoS ONE 6(7):e22850. doi:10.1371/journal.pone.0022850
Tashiro Y, Urushitani M, Inoue H, Koike M, Uchiyama Y, Komatsu M, Tanaka K, Yamazaki M, Abe M, Misawa H, Sakimura K, Ito H, Takahashi R (2012) Motor neuron-specific disruption of proteasomes, but not autophagy, replicates amyotrophic lateral sclerosis. J Biol Chem 287(51):42984–42994. doi:10.1074/jbc.M112.417600
Giannini C, Kloß A, Gohlke S, Mishto M, Nicholson TP, Sheppard PW, Kloetzel P-M, Dahlmann B (2013) Poly-Ub-substrate-degradative activity of 26S proteasome is not impaired in the aging rat brain. PLoS ONE 8(5):e64042. doi:10.1371/journal.pone.0064042
Seo H, Sonntag K-C, Kim W, Cattaneo E, Isacson O (2007) Proteasome activator enhances survival of Huntington’s disease neuronal model cells. PLoS ONE 2(2):e238. doi:10.1371/journal.pone.0000238
Kirkin V, McEwan DG, Novak I, Dikic I (2009) A role for ubiquitin in selective autophagy. Mol Cell 34(3):259–269. doi:10.1016/j.molcel.2009.04.026
Nixon RA, Wegiel J, Kumar A, Yu WH, Peterhoff C, Cataldo A, Cuervo AM (2005) Extensive involvement of autophagy in Alzheimer disease: an immuno-electron microscopy study. J Neuropathol Exp Neurol 64(2):113–122
Wong E, Cuervo AM (2010) Autophagy gone awry in neurodegenerative diseases. Nat Neurosci 13(7):805–811
Nedelsky NB, Todd PK, Taylor JP (2008) Autophagy and the ubiquitin-proteasome system: collaborators in neuroprotection. Biochim Biophys Acta (BBA) - Mol Basis Dis 1782(12):691–699. doi:10.1016/j.bbadis.2008.10.002
Nixon RA, Yang D-S (2011) Autophagy failure in Alzheimer’s disease—locating the primary defect. Neurobiol Dis 43(1):38–45. doi:10.1016/j.nbd.2011.01.021
Bukau B, Horwich AL (1998) The Hsp70 and Hsp60 chaperone machines. Cell 92(3):351–366
Kim YE, Hipp MS, Bracher A, Hayer-Hartl M, Ulrich Hartl F (2013) Molecular chaperone functions in protein folding and proteostasis. Annu Rev Biochem 82(1):323–355. doi:10.1146/annurev-biochem-060208-092442
Richter K, Haslbeck M, Buchner J (2010) The heat shock response: life on the verge of death. Mol Cell 40(2):253–266
Pearl LH, Prodromou C (2006) Structure and mechanism of the Hsp90 molecular chaperone machinery. Annu Rev Biochem 75:271–294. doi:10.1146/annurev.biochem.75.103004.142738
Mayer MP, Bukau B (2005) Hsp70 chaperones: cellular functions and molecular mechanism. Cell Mol Life Sci 62(6):670–684. doi:10.1007/s00018-004-4464-6
Mayer MP (2013) Hsp70 chaperone dynamics and molecular mechanism. Trends Biochem Sci 38(10):507–514. doi:10.1016/j.tibs.2013.08.001
Doyle SM, Genest O, Wickner S (2013) Protein rescue from aggregates by powerful molecular chaperone machines. Nat Rev Mol Cell Biol 14(10):617–629. doi:10.1038/nrm3660
Kampinga HH, Craig EA (2010) The HSP70 chaperone machinery: J proteins as drivers of functional specificity. Nat Rev Mol Cell Biol 11(8):579–592. doi:10.1038/nrm2941
Pratt WB, Morishima Y, Peng HM, Osawa Y (2010) Proposal for a role of the Hsp90/Hsp70-based chaperone machinery in making triage decisions when proteins undergo oxidative and toxic damage. Exp Biol Med (Maywood) 235(3):278–289. doi:10.1258/ebm.2009.009250
Cao W, Konsolaki M (2011) FKBP immunophilins and Alzheimer’s disease: a chaperoned affair. J Biosci 36(3):493–498
Van Duyne GD, Standaert RF, Karplus PA, Schreiber SL, Clardy J (1993) Atomic structures of the human immunophilin FKBP-12 complexes with FK506 and rapamycin. J Mol Biol 229(1):105–124
Somarelli JA, Lee SY, Skolnick J, Herrera RJ (2008) Structure-based classification of 45 FK506-binding proteins. Proteins Struct Funct Bioinforma 72(1):197–208. doi:10.1002/prot.21908
Liu J, Farmer JD Jr, Lane WS, Friedman J, Weissman I, Schreiber SL (1991) Calcineurin is a common target of cyclophilin-cyclosporin A and FKBP-FK506 complexes. Cell 66(4):807–815. doi:10.1016/0092-8674(91)90124-H
Kang CB, Hong Y, Dhe-Paganon S, Yoon HS (2008) FKBP family proteins: immunophilins with versatile biological functions. Neurosignals 16(4):318–325
Charters AR, Kobayashi M, Butcher SP (1994) The subcellular distribution of FK506 binding proteins in rat brain. Biochem Soc Trans 22(4):412 s
Shirane M, Nakayama KI (2003) Inherent calcineurin inhibitor FKBP38 targets Bcl-2 to mitochondria and inhibits apoptosis. Nat Cell Biol 5(1):28–37. doi:10.1038/ncb894
Wang H-Q, Nakaya Y, Du Z, Yamane T, Shirane M, Kudo T, Takeda M, Takebayashi K, Noda Y, Nakayama KI, Nishimura M (2005) Interaction of presenilins with FKBP38 promotes apoptosis by reducing mitochondrial Bcl-2. Hum Mol Genet 14(13):1889–1902. doi:10.1093/hmg/ddi195
Sinars CR, Cheung-Flynn J, Rimerman RA, Scammell JG, Smith DF, Clardy J (2003) Structure of the large FK506-binding protein FKBP51, an Hsp90-binding protein and a component of steroid receptor complexes. Proc Natl Acad Sci U S A 100(3):868–873. doi:10.1073/pnas.0231020100
Wu B, Li P, Liu Y, Lou Z, Ding Y, Shu C, Ye S, Bartlam M, Shen B, Rao Z (2004) 3D structure of human FK506-binding protein 52: implications for the assembly of the glucocorticoid receptor/Hsp90/immunophilin heterocomplex. Proc Natl Acad Sci U S A 101(22):8348–8353
Boudko SP, Ishikawa Y, Nix J, Chapman MS, Bachinger HP (2014) Structure of human peptidyl-prolyl cis-trans isomerase FKBP22 containing two EF-hand motifs. Protein Sci 23(1):67–75. doi:10.1002/pro.2391
van de Hoef DL, Bonner JM, Boulianne GL (2013) FKBP14 is an essential gene that regulates presenilin protein levels and Notch signaling in Drosophila. Development 140(4):810–819. doi:10.1242/dev.081356
Dickey CA, Yue M, Lin W-L, Dickson DW, Dunmore JH, Lee WC, Zehr C, West G, Cao S, Clark AMK, Caldwell GA, Caldwell KA, Eckman C, Patterson C, Hutton M, Petrucelli L (2006) Deletion of the ubiquitin ligase CHIP leads to the accumulation, but not the aggregation, of both endogenous phospho- and caspase-3-cleaved tau species. J Neurosci 26(26):6985–6996. doi:10.1523/jneurosci. 0746-06.2006
Jinwal UK, Koren J 3rd, Dickey CA (2013) Reconstructing the Hsp90/Tau machine. Curr Enzym Inhib 9(1):41–45. doi:10.2174/1573408011309010006
Karagoz GE, Duarte AM, Akoury E, Ippel H, Biernat J, Moran Luengo T, Radli M, Didenko T, Nordhues BA, Veprintsev DB, Dickey CA, Mandelkow E, Zweckstetter M, Boelens R, Madl T, Rudiger SG (2014) Hsp90-Tau complex reveals molecular basis for specificity in chaperone action. Cell 156(5):963–974. doi:10.1016/j.cell.2014.01.037
Kraemer BC, Burgess JK, Chen JH, Thomas JH, Schellenberg GD (2006) Molecular pathways that influence human tau-induced pathology in Caenorhabditis elegans. Hum Mol Genet 15(9):1483–1496. doi:10.1093/hmg/ddl067
Jinwal UK, Koren J 3rd, Borysov SI, Schmid AB, Abisambra JF, Blair LJ, Johnson AG, Jones JR, Shults CL, O’Leary JC 3rd, Jin Y, Buchner J, Cox MB, Dickey CA (2010) The Hsp90 cochaperone, FKBP51, increases Tau stability and polymerizes microtubules. J Neurosci 30(2):591–599. doi:10.1523/jneurosci. 4815-09.2010
Blair LJ, Nordhues BA, Hill SE, Scaglione KM, O’Leary JC 3rd, Fontaine SN, Breydo L, Zhang B, Li P, Wang L, Cotman C, Paulson HL, Muschol M, Uversky VN, Klengel T, Binder EB, Kayed R, Golde TE, Berchtold N, Dickey CA (2013) Accelerated neurodegeneration through chaperone-mediated oligomerization of tau. J Clin Invest 123(10):4158–4169. doi:10.1172/jci69003
Chambraud B, Belabes H, Fontaine-Lenoir V, Fellous A, Baulieu EE (2007) The immunophilin FKBP52 specifically binds to tubulin and prevents microtubule formation. FASEB J 21(11):2787–2797. doi:10.1096/fj.06-7667com
Sanokawa-Akakura R, Cao W, Allan K, Patel K, Ganesh A, Heiman G, Burke R, Kemp FW, Bogden JD, Camakaris J, Birge RB, Konsolaki M (2010) Control of Alzheimer’s amyloid beta toxicity by the high molecular weight immunophilin FKBP52 and copper homeostasis in Drosophila. PLoS ONE 5(1):e8626. doi:10.1371/journal.pone.0008626
Avramut M, Achim CL (2002) Immunophilins and their ligands: insights into survival and growth of human neurons. Physiol Behav 77(4–5):463–468. doi:10.1016/S0031-9384(02)00934-4
Sugata H, Matsuo K, Nakagawa T, Takahashi M, Mukai H, Ono Y, Maeda K, Akiyama H, Kawamata T (2009) A peptidyl–prolyl isomerase, FKBP12, accumulates in Alzheimer neurofibrillary tangles. Neurosci Lett 459(2):96–99
Liu FL, Liu TY, Kung FL (2014) FKBP12 regulates the localization and processing of amyloid precursor protein in human cell lines. J Biosci 39(1):85–95
Liu F-L, Liu P-H, Shao H-W, Kung F-L (2006) The intracellular domain of amyloid precursor protein interacts with FKBP12. Biochem Biophys Res Commun 350(2):472–477. doi:10.1016/j.bbrc.2006.09.073
Gerard M, Deleersnijder A, Daniels V, Schreurs S, Munck S, Reumers V, Pottel H, Engelborghs Y, Van den Haute C, Taymans JM, Debyser Z, Baekelandt V (2010) Inhibition of FK506 binding proteins reduces alpha-synuclein aggregation and Parkinson’s disease-like pathology. J Neurosci 30(7):2454–2463. doi:10.1523/JNEUROSCI. 5983-09.2010
Deleersnijder A, Van Rompuy AS, Desender L, Pottel H, Buee L, Debyser Z, Baekelandt V, Gerard M (2011) Comparative analysis of different peptidyl-prolyl isomerases reveals FK506-binding protein 12 as the most potent enhancer of alpha-synuclein aggregation. J Biol Chem 286(30):26687–26701. doi:10.1074/jbc.M110.182303
Liou YC, Sun A, Ryo A, Zhou XZ, Yu ZX, Huang HK, Uchida T, Bronson R, Bing G, Li X, Hunter T, Lu KP (2003) Role of the prolyl isomerase Pin1 in protecting against age-dependent neurodegeneration. Nature 424(6948):556–561. doi:10.1038/nature01832
Holzer M, Gartner U, Stobe A, Hartig W, Gruschka H, Bruckner MK, Arendt T (2002) Inverse association of Pin1 and tau accumulation in Alzheimer’s disease hippocampus. Acta Neuropathol 104(5):471–481. doi:10.1007/s00401-002-0581-1
Pastorino L, Sun A, Lu PJ, Zhou XZ, Balastik M, Finn G, Wulf G, Lim J, Li SH, Li X, Xia W, Nicholson LK, Lu KP (2006) The prolyl isomerase Pin1 regulates amyloid precursor protein processing and amyloid-beta production. Nature 440(7083):528–534. doi:10.1038/nature04543
Perry G, Friedman R, Shaw G, Chau V (1987) Ubiquitin is detected in neurofibrillary tangles and senile plaque neurites of Alzheimer disease brains. Proc Natl Acad Sci U S A 84(9):3033–3036
López Salon M, Morelli L, Castaño EM, Soto EF, Pasquini JM (2000) Defective ubiquitination of cerebral proteins in Alzheimer’s disease. J Neurosci Res 62(2):302–310. doi:10.1002/1097-4547(20001015)62:2<302::AID-JNR15>3.0.CO;2-L
Keller JN, Hanni KB, Markesbery WR (2000) Impaired proteasome function in Alzheimer’s disease. J Neurochem 75(1):436–439. doi:10.1046/j.1471-4159.2000.0750436.x
Keck S, Nitsch R, Grune T, Ullrich O (2003) Proteasome inhibition by paired helical filament-tau in brains of patients with Alzheimer’s disease. J Neurochem 85(1):115–122
Berke SJS, Paulson HL (2003) Protein aggregation and the ubiquitin proteasome pathway: gaining the UPPer hand on neurodegeneration. Curr Opin Genet Dev 13(3):253–261. doi:10.1016/S0959-437X(03)00053-4
Lindsten K, de Vrij FMS, Verhoef LGGC, Fischer DF, van Leeuwen FW, Hol EM, Masucci MG, Dantuma NP (2002) Mutant ubiquitin found in neurodegenerative disorders is a ubiquitin fusion degradation substrate that blocks proteasomal degradation. J Cell Biol 157(3):417–427. doi:10.1083/jcb.200111034
Layfield R, Cavey JR, Lowe J (2003) Role of ubiquitin-mediated proteolysis in the pathogenesis of neurodegenerative disorders. Ageing Res Rev 2(4):343–356. doi:10.1016/S1568-1637(03)00025-4
de Vrij FMS, Jacqueline A, Gregori LF, David F, Hermens WTJMC, Goldgaber D, Verhaagen J, Van Leeuwen FW, Hol EM (2001) Mutant ubiquitin expressed in Alzheimer’s disease causes neuronal death. FASEB J 15(14):2680–2688. doi:10.1096/fj.01-0438com
Mawuenyega KG, Sigurdson W, Ovod V, Munsell L, Kasten T, Morris JC, Yarasheski KE, Bateman RJ (2010) Decreased clearance of CNS β-amyloid in Alzheimer’s disease. Science 330(6012):1774. doi:10.1126/science.1197623
Yen SS (2011) Proteasome degradation of brain cytosolic tau in Alzheimer’s disease. Int J Clin Exp Pathol 4(4):385–402
Oh S, Hong HS, Hwang E, Sim HJ, Lee W, Shin SJ, Mook-Jung I (2005) Amyloid peptide attenuates the proteasome activity in neuronal cells. Mech Ageing Dev 126(12):1292–1299. doi:10.1016/j.mad.2005.07.006
Tseng BP, Green KN, Chan JL, Blurton-Jones M, LaFerla FM (2008) Aβ inhibits the proteasome and enhances amyloid and tau accumulation. Neurobiol Aging 29(11):1607–1618. doi:10.1016/j.neurobiolaging.2007.04.014
Gregori L, Hainfeld JF, Simon MN, Goldgaber D (1997) Binding of amyloid β protein to the 20 S proteasome. J Biol Chem 272(1):58–62. doi:10.1074/jbc.272.1.58
Shringarpure R, Grune T, Sitte N, Davies* KJA (2000) 4-Hydroxynonenal-modified amyloid-β peptide inhibits the proteasome: possible importance in Alzheimer’s disease*. CMLS, Cell Mol Life Sci 57(12):1802–1809. doi: 10.1007/PL00000660
Gregori L, Fuchs C, Figueiredo-Pereira ME, Van Nostrand WE, Goldgaber D (1995) Amyloid β-protein inhibits ubiquitin-dependent protein degradation in vitro. J Biol Chem 270(34):19702–19708. doi:10.1074/jbc.270.34.19702
Lam YA, Pickart CM, Alban A, Landon M, Jamieson C, Ramage R, Mayer RJ, Layfield R (2000) Inhibition of the ubiquitin-proteasome system in Alzheimer’s disease. Proc Natl Acad Sci U S A 97(18):9902–9906. doi:10.1073/pnas.170173897
Klettner A, Baumgrass R, Zhang Y, Fischer G, Bürger E, Herdegen T, Mielke K (2001) The neuroprotective actions of FK506 binding protein ligands: neuronal survival is triggered by de novo RNA synthesis, but is independent of inhibition of JNK and calcineurin. Mol Brain Res 97(1):21–31. doi:10.1016/S0169-328X(01)00286-8
Lee KH, Won R, Kim UJ, Kim GM, Chung M-A, Sohn J-H, Lee BH (2010) Neuroprotective effects of FK506 against excitotoxicity in organotypic hippocampal slice culture. Neurosci Lett 474(3):126–130. doi:10.1016/j.neulet.2010.03.009
Winter C, Schenkel J, Bürger E, Eickmeier C, Zimmermann M, Herdegen T (1999) The immunophilin ligand FK506, but not GPI-1046, protects against neuronal death and inhibits c-Jun expression in the substantia nigra pars compacta following transection of the rat medial forebrain bundle. Neuroscience 95(3):753–762. doi:10.1016/S0306-4522(99)00486-8
Karapetyan YE, Sferrazza GF, Zhou M, Ottenberg G, Spicer T, Chase P, Fallahi M, Hodder P, Weissmann C, Lasmézas CI (2013) Unique drug screening approach for prion diseases identifies tacrolimus and astemizole as antiprion agents. Proc Natl Acad Sci U S A 110(17):7044–7049. doi:10.1073/pnas.1303510110
Chiang K, Koo EH (2014) Emerging therapeutics for Alzheimer’s disease. Annu Rev Pharmacol Toxicol 54(1):381–405. doi:10.1146/annurev-pharmtox-011613-135932
Harrington C, Rickard JE, Horsley D, Harrington KA, Hindley KP, Riedel G, Theuring F, Seng KM, Wischik CM (2008) O1-06-04: methylthioninium chloride (MTC) acts as a Tau aggregation inhibitor (TAI) in a cellular model and reverses Tau pathology in transgenic mouse models of Alzheimer’s disease. Alzheimers Dement 4(Supplement 4):T120–T121. doi:10.1016/j.jalz.2008.05.259
Wischik CM, Bentham P, Wischik DJ, Seng KM (2008) O3-04-07: Tau aggregation inhibitor (TAI) therapy with rember™ arrests disease progression in mild and moderate Alzheimer’s disease over 50 weeks. Alzheimers Dement 4(Supplement 4):T167. doi:10.1016/j.jalz.2008.05.438
Hong X, Liu J, Zhu G, Zhuang Y, Suo H, Wang P, Huang D, Xu J, Huang Y, Yu M, Bian M, Sheng Z, Fei J, Song H, Behnisch T, Huang F (2014) Parkin overexpression ameliorates hippocampal long-term potentiation and β-amyloid load in an Alzheimer’s disease mouse model. Hum Mol Genet 23(4):1056–1072. doi:10.1093/hmg/ddt501
Deshaies RJ, Joazeiro CAP (2009) RING domain E3 ubiquitin ligases. Annu Rev Biochem 78(1):399–434. doi:10.1146/annurev.biochem.78.101807.093809
Mukherjee A, Morales-Scheihing D, Gonzalez-Romero D, Green K, Taglialatela G, Soto C (2010) Calcineurin inhibition at the clinical phase of prion disease reduces neurodegeneration, improves behavioral alterations and increases animal survival. PLoS Pathog 6(10):e1001138. doi:10.1371/journal.ppat.1001138
Blackburn EA, Walkinshaw MD (2011) Targeting FKBP isoforms with small-molecule ligands. Curr Opin Pharmacol 11(4):365–371. doi:10.1016/j.coph.2011.04.007
Acknowledgments
This work was supported by the research fund of Hanyang University.
Conflict of Interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Sulistio, Y.A., Heese, K. The Ubiquitin-Proteasome System and Molecular Chaperone Deregulation in Alzheimer’s Disease. Mol Neurobiol 53, 905–931 (2016). https://doi.org/10.1007/s12035-014-9063-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-014-9063-4