Abstract
The disparate diversity in immunoglobulin (Ig) repertoire has been a subject of fascination since the emergence of prototypic adaptive immune system in vertebrates. The carboxy terminus region of activation-induced cytidine deaminase (AID) has been well established in tetrapod lineage and is crucial for its function in class switch recombination (CSR) event of Ig diversification. The absence of CSR in the paraphyletic group of fish is probably due to changes in catalytic domain of AID and lack of cis-elements in IgH locus. Therefore, understanding the arrangement of Ig genes in IgH locus and functional facets of fish AID opens up new realms of unravelling the alternative mechanisms of isotype switching and antibody diversity. Further, the teleost AID has been recently reported to have potential of catalyzing CSR in mammalian B cells by complementing AID deficiency in them. In that context, the present review focuses on the recent advances regarding the generation of diversity in Ig repertoire in the absence of AID-regulated class switching in teleosts and the possible role of T cell-independent pathway involving B cell activating factor and a proliferation-inducing ligand in activation of CSR machinery.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Owing to the increasing unpredictable threat imposed by detrimental pathogenic infiltrates, agnathans, the most primitive chordates evolved the complex adaptive immune system regulated by variable lymphocyte receptors (VLRB and VLRA) equivalent to B and T lymphocytes of jawed vertebrates [1,2,3,4]. The increasing experimental accessibility of mammalian and non-mammalian vertebrate models has established the framework of our present knowledge regarding evolution of immunoglobulin (Ig) isotypes, their function and alternative mechanisms for Ig receptor diversification [5,6,7,8,9]. It was in the late 1957 that Burnet had put forward his conjectures regarding the clonal selection theory in an attempt to explain the complex interplay of antigen–antibody interaction and how lymphocytes are selected to impose an immune response against specific antigens [10]. Burnet’s theory provided a strong base for the preponderance of experimental endeavours that followed, in order to decode the complex mechanisms tailoring antigen receptor diversification [11]. However, a direct evidence of the combinatorial rearrangement of variable and constant regions of the IgL chain gene occurring at DNA level came in 1978 when two independent groups led by Leder and Tonegawa reported the comparative analysis of germline and rearranged V gene segments of κ and λ light chain locus, respectively [12, 13]. Several other studies in line were enough to elucidate the fact that the primary diversification of the B and T cell progenitors is brought about by an antigen-independent recombination event that entails de novo random assortment of germline variable (V), diversity (D) and joining (J) gene segments to encode functional Ig molecule [14,15,16].
This lineage-specific reaction is well known as V(D)J recombination and is mediated by recombination activating genes (RAG1 and RAG2) and is supported by DNA repair factors [17,18,19]. After cognate antigen exposure, B cells are attributed to somatic hypermutation (SHM) where variable regions of the IgH and IgL chains are subjected to point mutations and are further diversified [16]. Class switch recombination (CSR) was the last of the DNA modification reaction to have evolved in amphibians. It is the region-specific recombination event that switches the constant regions of the expressed Ig heavy chain without jeopardizing the antigen specificity [20]. Therefore, it became established that antigen receptors found in all jawed vertebrates—bony fish, amphibians, reptiles, birds and mammals, are products of any two or three of lymphocyte-specific antigen receptor diversification processes—V(D)J recombination, SHM, GC (gene conversion) and CSR [5, 21,22,23].
Activation-induced cytidine deaminase (AID) belongs to a fascinating group of mutagenic proteins of apolipoprotein B mRNA-editing catalytical component (APOBEC) family and is essential for initiating both SHM [24] and CSR [25]. Although bony fish do possess AID protein, conventional CSR has not been reported to exist in fish and has emerged only in amphibians. The differences in the catalytic domain of AID, the absence of switch regions and constant region clusters in IgH locus may be responsible for the absence of CSR in this group [26, 27]. However, several studies have reported that AID from fish (zebrafish, pufferfish, catfish) have potential to induce CSR in mouse cells, indicating that early vertebrate AID already possessed the ability of catalyzing recombination reactions pertaining to receptor diversification [27, 28]. Moreover, the presence of multiple Ig isotypes, namely—IgM, IgD and IgNAR in cartilaginous fish [29,30,31] and additionally novel IgZ/T in bony fish [32,33,34,35] strongly plugs to the possibility of alternative diversification mechanisms in fish for generation of Ig repertoire.
Extensive studies in the higher vertebrates especially in humans have successfully proved that CSR induction in B cells entails two crucial signals. First one is the conventional T cell-dependent pathway which is regulated by the indirect interaction of T cell-dependent antigens to CD40 on B cells through binding CD40L ligand, and the second one is the classical pathway involving cytokine-mediated signals [36,37,38]. B cell activating factor (BAFF) and a proliferation-inducing ligand (APRIL) are crucial members of tumour necrosis factor (TNF) superfamily and have been reported to induce CSR in B cells through T cell-independent pathway [39]. This involves antigen recognition through germline-encoded pattern recognition receptors (PRRs) like Toll-like receptors (TLRs) [40] and stimulation of NF-kB pathway [41]. BAFF and APRIL thus collectively contribute in regulating residual CSR in higher vertebrates during CD40 and CD40L deficiency. Therefore, it might be hypothesized that these two factors may have critical role in generating isotype switching in lower vertebrates like fish.
In this review, the basic functions of AID in fish are discussed in the context to the absence of class switch recombination. The focus is laid on uncovering the molecular mechanisms underlying different affinity maturation events and resolving paradigms of antigen diversification and memory formation in various classes of these radically different ectothermic vertebrates. Further, the alternative possibilities for class switch recombination in fish are explored, highlighting the T cell-independent pathway and interplay of AID and BAFF/APRIL in activation of Ig isotype switching.
Evolution of AID and Its Explicit Role in Antigen Receptor Diversification Events
The discovery of AID almost two decades ago greatly revolutionized our understanding, regarding the regulation of complex secondary diversification processes like SHM, gene conversion (GC) and CSR that have implications on vertebrate physiology and B cell memory formation [25, 42,43,44,45,46]. GC is essential for underpinning V region diversification in chickens, pigs and cattle [21, 42, 47]. In contrary, humans and mice exclusively rely on SHM, while rabbits use both the maturation processes [48, 49]. Based on the theory that there must be a specific recombinase to orchestrate CSR, AID was discovered by employing a subtractive hybridization approach after stimulating B lymphocyte cell line (CH12F3) [50]. Subsequently, AID knockout studies in mice failed to exhibit SHM and CSR processes, clearly suggesting that AID is central to both affinity maturation and isotype switching events [25]. Contemporary studies in humans demonstrated that mutation in AID molecules were responsible for complete abrogation of CSR and overexpression of IgM isotype in serum of human patients with hyper-IgM syndrome [51].
Unlike higher vertebrates, the presence of affinity maturation events in gnathostomes has been a topic of debate due to two apparent shortcomings. Although fish possess B, T cells and generate memory responses like mammalian counterparts, the absence of high-affinity Igs on immunization and prototypical germinal centres that are the distinct sites for affinity maturation events in fish greatly pointed towards the complete absence of antibody diversification processes [52]. However, the identification of AID ortholog became evident by the discovery of SHM in elasmobranchs and amphibians [49, 53, 54]. The characterization of transcript sequences of primitive shark species, Heterodontus using gene-specific oligodeoxynucleotide hybridization, revealed that SHM occurs in immunoglobulin heavy chains [55]. Subsequent studies in shark light chain and IgNAR transcripts reported distinguished hypermutations, suggesting the presence of AID in lower vertebrates [56, 57]. The first evidence of the possibility of occurrence of affinity maturation in bony fish came when Cain et al. [58] carried out the antigen antibody kinetics in rainbow trout following immunization. The study reported the higher affinity of tetrameric serum IgM antibody for antigen post-immunization which might be due to the presence of pre-existing high-affinity Ig producing cells [59]. Additionally, few studies undertaken in both catfish variable heavy chain [60] and zebrafish light chains [61] revealed the nucleotide targets of SHM, suggesting the possible contribution of both affinity maturation processes and AID in generation of Ig diversity in bony fish. However, it was also evident that affinity maturation processes can also occur at alternative sites due to the absence of germinal centres in fish [62, 63]. Macrophage-populated melanomacrophage centres might serve as analogous to germinal centres for antigen retention and antibody binding in fish [52].
Although by now it has became apparent that AID protein is explicitly expressed in germinal centre centroblasts, the exact function and the molecular mechanism of the protein are yet to be explored. In addition to the expression of AID in CH12F3, Muramatsu et al. [50] reported that amino acid sequence of AID cDNA obtained in their study was homologous to APOBEC-1, a class of apo-B mRNA-editing cytidine deaminases. Therefore, based on the above report, it was hypothesized that AID has potency to edit mRNA in sequence-specific manner during both CSR and SHM events and it was designated as—RNA-editing hypothesis [24, 50]. There were other competent groups who postulated that AID mutated at DNA level and not at RNA level [64, 65]. Numerous studies in the same line were conducted in non-lymphocyte (B cells) cell lines to speculate both the hypotheses, and it was found that ectopic expression of AID could induce hypermutation and class switching in fibroblasts [66, 67] and hybridomas [68] using sophisticated tools of recombinant DNA technology. Similar results were reported in Escherichia coli [69] and yeast [70]. The best model to elucidate AID-mediated Ig gene conversion mechanism was bursal B cells (DT40) in which by using two AID knockout constructs, it was evident that AID gene disruption led to a complete termination of GC in chickens, which was reversible when AID deficiency was complemented [42]. However, similar experimental trials to understand a comparative role of AID homologs in V(D)J recombination process revealed that AID mutation did not alter the germline V gene assortment process [71]. Collectively, these findings in mice and humans clearly implied that AID is a key regulator in all three known Ig diversification processes (SHM, CSR and GC) [25, 72]. However, the site-specific V(D)J recombination process that occurs during the early development of naive B and T lymphocytes was rarely examined.
AID belongs to a class of potential nucleic acid (RNA/DNA)-editing enzymes of APOBEC family that employ ssDNA as their substrates and effectively deaminates deoxy-cytidine (dC) residues to deoxyuridine (dU) [73,74,75]. Five of the APOBEC family members have been reported yet, APOBEC1, APOBEC2, APOBEC3 (A-H) sub-branch, AID and APOBEC4 [76, 77]. APOBEC1 is capable to mutate apolipoprotein B mRNA by specifically converting cytosine at position 6666, with the help of an associated protein APOBEC-1 complementation factor (ACF) [78,79,80]. While other members of the family are exclusively DNA mutators, APOBEC2 has been reported to have implications in pattern development in Xenopus and Danio rerio embryos and is regulated by TGF-β signalling [81]. The role of AID in regulation of secondary antigen receptor diversification has already been established in several studies by editing cytosine residues [76, 82, 83]. However, APOBEC3 proteins are known to be responsible for abrogation of retroviral propagation [84] and was found to have inhibitory effect on the HIV-1, HBV and HCV replication, although clinical manifestation of its role in host-viral interaction is not clear [85, 86]. Recently, several studies have reported the crucial role of APOBEC3 A3 sub-branch members in cancer initiation [86,87,88]. APOBEC4 is the newly discovered member of the family by computational prediction approach in the tetrapod lineage; however, its function is yet to be uncovered [89]. Interestingly, APOBEC2 and AID are found to be conserved in all vertebrates. The first homolog of AID in fish was reported in channel catfish [59]. APOBEC1, APOBEC2, APOBEC3C and APOBEC4 have been successfully expressed in Protopterus sp., although the presence of AICDA or AID sequence was not ascertained [90]. APOBEC4 and AID are found in all the jawed vertebrates including cartilaginous fish and mammals (Fig. 1). However, APOBEC2 in fish is restricted to teleosts and is additionally present in frogs, chickens and mammals [59, 91, 92].
Phylogenetic tree showing the co-relationship of AID among different fish species. Heterologous nucleotide sequences of AID from various fish species were downloaded from NCBI database, and phylogenetic tree was constructed using neighbour-joining method with the MEGA 6 program. Bootstrap values were calculated from 1000 repetitions. AID nucleotide sequences from all members of Actinopterygii and Sarcopterygii classes of bony fish formed a separate cluster. AID nucleotide sequences of cartilaginous fish were chosen as outgroups. The scale bar indicates the divergence time
Structurally, the most defining AID/APOBEC proteins comprise of a zinc coordination motif containing consensus sequences, (HA/VExPCxxC) motif that is crucial for coordination of a zinc atom. The highly conserved motif is arranged in the form of catalytic pocket consisting of zinc coordinating triad of two histidines and a cysteine on the surface of glutamic acid [76, 77, 93]. X-ray and NMR studies pertaining to AID structure have made it clear that the core structure comprises of a sandwich of central β sheet between 6 and 7 α-helices interconnected with loops of variable length [88]. AID proteins have been demonstrated to possess four distinct domains each having crucial role in the collective regulatory functions of AID [92]. AID protein is composed of 198 amino acids, and prototypic four domains are known to include an APOBEC type domain, anticipated to have similar functions of cytosine conversion, a catalytic domain for targeting and binding DNA substrates, a nuclear export signal (NES) and nuclear localization signal (NLS) domain [94,95,96,97] and a dimerization domain. C-terminal NES motif of AID is crucial for CSR and has been found to be in association with CSR-related cofactors [72, 98, 99]. Studies on AID mutants have elucidated that the non-catalytical NES motif is critical for export of AID from nucleus employing chromosome region maintenance 1 (CRM1) association [100]. Extensive studies involving AID fusion constructs of zebrafish, fugu, catfish have revealed that zebrafish AID could restore the ability to induce SHM and CSR activity when transfected to mammalian cells [27, 28, 101]. Additionally, disruption of C-terminus NES domain of catfish [27] or treatment of cell cultures with leptomycin B, an inhibitor of CRM1 exportin [101], led to retention of fish AID in the nucleus. Therefore, it can be assumed that the function of C-terminus of fish AID is restricted to orchestrating AID transport out of nucleus, although having potential to regulate CSR as well.
Various Mechanisms of Ig Diversification in Fish
Agnathans: Lampreys and Hagfishes
Ever since the “Immunological Big Bang” took place, many questions were raised regarding the sudden emergence of clonally diverse antigen receptors, invasion of the ancestral RAG1 and RAG2 genes into the host genome and initiation of different affinity maturation processes [1, 102, 103]. The enigma of such critical questions began to resolve in the late 1960s when the work of eminent scientists laid the foundations of comparative immunology. The evolutionary precursors of the vertebrate antigen-specific receptors of the anticipatory immune system could be traced back to the ancestors of the jawed vertebrates [104, 105]. Urochordate and cephalochordate ancestors of chordate phylum did not exhibit the presence of any identifiable key elements of this pivotal immune system according to the experimental trials conducted on Ciona intestinalis [106] and Amphioxus [107]. In C. intestinalis, no orthologs of T cell receptors (TCRs), Ig, MHC and RAG genes were apparent, while in amphioxus, a protochordate, radically different type of secretory proteins encoded by multigene families comprising of two diverse Ig-like variable domains and a chitin-binding domain (VCBP) were reported [106,107,108]. However, none of the molecules could be compared with that of prototypic AIS (anticipatory immune system) genes. Based on these findings, it began to be anticipated that agnathans lack primordial features of adaptive immune system until the histological evidence of lymphocyte like molecules in two extant members of agnathans-lampreys and hagfish [109, 110].
Extensive experimental trails were then extended to include early jawless vertebrate group-lampreys where transcriptional analysis revealed the presence of single copies of TCR-like and CD4-like genes [111]. Concomitant non-trivial discovery of variable lymphocyte receptors (VLRs) composed of leucine-rich repeat (LRR) sequences and the fact that their diversification occurs in somatic cells reveal that vertebrates have been gifted with radically different classes of antigen receptors to combat microbial threats [2, 112, 113]. The presence of canonical T cell activity confirmed by allograft rejection [114], delayed antigen sensitivity reaction [115], production of circulating agglutinins in response to stimulation [115, 116] and the expression of transcription factors like ikaros, PU.1 and Spi-B pertaining to lymphocyte differentiation [117] in lampreys, convincingly attribute to the presence of newly encountered clonally diverse VLRs in agnathan clade.
Therefore, in contrary to jawed vertebrates who possess highly diverse TCRs and IgH locus of B cell Receptors (BCRs) comprised of complex V(D)J segments, ancestral jawless vertebrates assemble their atypical VLRs, through variable modular LRR segments which are bound to the lymphocyte surface through an invariant flexible stalk region in association with a glycosyl-phosphatidyl inositol (GPI) anchor [2, 118]. It also became appreciated that both lampreys and hagfish possess two disparate VLRs—VLRA and VLRB, that are reciprocally expressed and morphologically equivalent to TCRs and BCRs of mammalian counterparts along with a recently discovered third lymphocyte lineage—VLRC exhibiting resemblance with γδT cells [4, 119, 120]. These lymphocyte lineages are known to be diversified by somatic rearrangement of LRR cassettes that flank naive VLR gene; thus, a diverse repertoire of VLRs exhibiting unique sequences comparable to mammalian antibody repertoire (~ 1014) is generated [117, 118].
The amino acid sequence analysis and crystallographic studies undertaken to explore structural prediction and protein interaction surfaces of VLRs indicate that its hypervariable concave region is critical for the antigen recognition [121]. Pancer et al. [2] have initially revealed that the basic structure of the germline VLRs is a concave solenoid flanked by amino-terminal and carboxyl-terminal LRRs at both ends resembling to that of the Toll-like receptor ectodomains. Thus, it became evident that the germline population of VLRs is incomplete and do not express functional proteins encoding only the amino-terminal and carboxyl-terminal region of the mature VLR proteins—(LRRNT) and (LRRCT), respectively, and is intervened by non-coding sequences [119]. However, fully functional VLR proteins are encoded in lymphocytes, where germline VLRs are subjected to somatic recombination that occurs by random selection of multiple flanking LRR cassettes [118].
Unlike jawed vertebrates, the multistep LRR reassortment process in germline VLRs do not require any recombination signal sequences flanking the segments meant to be assorted which is otherwise requisite for Ig-based antibody diversification mechanism—V(D)J recombination in gnathostomes [122]. A prototypic VLR protein is composed of a (1) signal peptide (SP) followed by (2) N-terminal (LRRNT) region containing 30–38 residues; (3) a variable (LLR1) region of 18 residues followed by 7–8 (LRRV) regions of 24 residues; (4) a terminal LRRVe region containing a signature sequence; (5) a truncated LRR connecting peptide (CP) and C-terminal (LRRCT) region, to which a threonine/proline-rich stalk, GPI anchor site and a hydrophobic tail are attached [2, 119]. The structure of VLRA is conserved between both lampreys and hagfish. In addition to the above structure, lamprey VLRB has a glycosylphosphatidylinositol (GPI) membrane anchor cleavage site. VLRA and VLRC are exclusively transmembrane proteins, while VLRB can be secreted when VLRB+ lymphocytes are triggered with antigen stimulation and allowed to proliferate and differentiate to form plasma cells secreting VLRB antibodies [104, 123]. VLRA is known to have similarity with mammalian T cell lineages expressing genes unique to mammalian TCRs involved in differentiation and induction of pro-inflammatory cascade generation [4]. It can be additionally noted that VLRA+ and VLRB+ lymphocytes lineages are reported to interact with each, suggesting functional resemblance with anticipatory B and T cells of jawed vertebrates [104].
It has been predicted that an array of 1500–2400 LRR modules in the genomes of agnatha is anticipated to be responsible for huge VLR diversity [118]. This multistep assembly of the VLR genes occurs through a gene conversion mechanism and is known to be regulated by AID/APOBEC family members [59, 122]. It is important to note that no AID or AID/APOBEC family member counterparts were reported in the evolutionary primitive urochordates and cephalochordates AID/APOBEC orthologs, namely cytidine deaminase 1 (CDA1) and cytidine deaminase 2 (CDA2) are known to differentially orchestrate VLRA and VLRB assembly during gene conversion process. As these set of deaminases were first discovered in Petromyzon marinus, they were designated as PmCDA1 and PmCDA2. The two lamprey deaminases are detected in blood lymphocytes and hematopoietic tissues and are found to be evolutionary related to the AID/APOBEC family of cytidine deaminases existent in gnathostomes. A complete polypeptide of CDA1 comprises of 208 residues and is encoded by single exon, while CDA2 (311 residues) is encoded by four exons [119]. Both sets of enzymes exhibit distinguishing zinc coordination motif sequence, “HxE-PCxxC” where (x = any amino acid) and an additional AT-hook motif is present in CDA2. Extensive studies were carried out in lampreys to speculate the functions of secretory VLRB proteins, and they were found to provide humoral immunity by binding to the soluble and particulate antigens [118, 124]. When immunized with anthrax, the concentration of soluble VLR protein titers was found to increase in serum of lampreys. Further, the increased capability of VLR protein in recognizing exosporium-specific BclA protein clearly implicates its role in imparting humoral immunity.
Cartilaginous Fish
Cartilaginous fish (Class—Chondricthyes) are the earliest vertebrate group to possess prototypic immunoglobulin (Ig) molecules, TCRs and MHC genes and are capable of imparting antigen-specific immunity based on RAG-mediated V(D)J recombination against a myriad of pathogens [30, 125]. Several distinct patterns of antigen receptor diversification processes were concurrent with evolution of the primordial key genes in jawed vertebrates. All the contemporary members of jawed vertebrates have been known to possess rearranging antigen binding receptors—four types of TCRs (α, β, γ and δ), and immunoglobulin IgH and IgL chains [126]. Three types of heavy chain isotypes are demonstrated in this group of fish, namely IgM, IgD/IgW and IgNAR [31]. Sharks and skates have two divergent types of light chains that more or less resemble λ and κ IgNAR which is unique Ig isotype lacking light chain and is basically a homodimer of two heavy chains bound by peculiar disulphide bonds [54]. The most interesting features of cartilaginous fish regarding arrangement and organization of the Ig genes are presence of various segments of IgH and IgL chains in the form of clusters, comparatively decreased set of inter-family variations in the sequences of variable light and heavy chains and lack of combinatorial rearrangement of the clusters [127, 128]. Thus, it is clear that the IgH and IgL chain exists in cluster arrangement and each cluster consists of one variable (V), two or three diversity (D), one joining and a set of constant (C) region exons [119]. Common type of IgH clusters present throughout this fish group is (VH-DH1-DH2-JH-CH)n. Although it is quite impressive that lower vertebrates like agnathans (as described above) somatically diversify their so-called VLRs, the cartilaginous fish being member of jawed vertebrates still retain the capacity to somatically diversifying their antigen receptors as described below.
Primary lymphoid organs that are central for lymphocyte development in higher vertebrates, like bone marrow and well-developed lymphatic system are lacking in cartilaginous fish. Instead, Leydig cells and epigonal organs unique to this group are present that act as equivalents of bone marrow and express high levels of RAG1 and terminal deoxynucleotide transferases (TdT) [130]. RAG1 and RAG2 genes have been exclusively responsible for targeting and recognizing recombination signal sequences (RSS) adjoining the V(D)J gene segments introducing double-stranded breaks (DSBs) and facilitate in rearrangement process popularly known as V(D)J recombination, by recruiting several DNA repair enzymes [17, 131]. The presence of RSS sequences of 22 nucleotides long, downstream of V regions and 12 or 22/23 nucleotides long sequences flanking three D regions present in NAR of sharks further confirms the occurrence of deletional V(D)J recombination event [54]. Terminal transferases are specialized group of DNA polymerases that enable addition of N-nucleotides at the junction of V, D and J segments during Ig gene recombination and provide junctional diversity. Thus, it is clear that both IgH and IgL chains in sharks are brought about by RAG-mediated rearrangement process which occurs early in the germline [132].
After the discovery of the novel IgNAR in cartilaginous fish, many extensive studies have been conducted to explore the evolution of hypermutation process tailoring antigen receptor diversification. IgNAR is a unique form of Ig found in cartilaginous fish which greatly differs from other Igs and TCRs present in the group [49, 133]. The secretory immunoglobulins IgM and IgNAR are generated by an array of 100–200 clusters, comprising of few rearranging V(D)J segments and a set of constant region [126]. All evidences in cartilaginous fish obtained so far reveal that both V(D)J rearrangements in IgH and VJ rearrangements in IgL chains occurs within clusters and is restricted to occur between clusters [129] (Fig. 2a). Additionally, 22 RGYW and its complementary WRCY hotspot motifs are found in the V and J segments of nurse sharks where rate of hypermutations was high. It is also reported that the tandem substitutional changes in nurse shark Igs are regulated by error prone DNA repair mechanisms involving an AID homolog [8, 134].
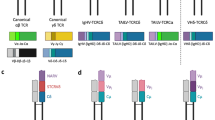
Schematic representation of IgH locus and V(D)J recombination events in Gnathostomes. a Cartilaginous fish IgH locus. The IgH locus is consisting of cluster arrangement. Combinatorial rearrangement is not possible, and both V(D)J and VJ arrangements occur within clusters, and b bony fish IgH locus. The IgH locus of the bony fish is consisting of prototypical translocon arrangement. IgZ/IgT is expressed on rearrangement of Vζ region exons with Dζ-Jζ and Cζ. Alternative splicing occurs for the expression of either IgM or IgD as Cµ and Cδ region exons in the IgH locus lie close to each other without intervening enhancers
Therefore, it is apparent that somatic hypermutation mechanism in NAR is very similar to that in mammals, where AGY hotspots are central targets for mutational events, greater concentration of mutations in CDR3 regions and mechanism of point mutations [49]. However, characterization of IgL chain assembly revealed that the pattern of hypermutation occurring in nurse shark light chains also contains tandem alterations of 2–5 bp substitutions along with the point mutations [57]. A similar study in horn shark cDNA does not show any trace of substitutions in the Cµ regions. Nurse shark Cµ regions exhibit 85–92% identity between groups and are almost 100% identical within group [134]. Additionally, V, J and C regions show many co-relations, indicating that the interlocus rearrangement hardly occurs. All these results clearly suggested that combinatorial arrangement of V(D)J and C regions, central in tetrapod lineage is not prominent in cartilaginous fish, and they undergo somatic arrangement of their Igs [134]. The point mutations generated in the IgL locus of the sharks are bias towards transitions just like the higher vertebrates [108, 129, 135]. The huge diversity in the antigen receptor repertoire is largely imparted by the junctional diversity and rearrangement of partially germline-arranged V(D)J segments, where different combinations of either individual diversity (D) segments or in combination with others and inverted combinations are joined [8, 132]. Therefore, it is clear that Igs of cartilaginous fish diversify through somatic hypermutation and tandem changes. It is hypothesized that Ig isotypes are defined based on generation of B cell lineages, similar to the expression of αβ and γδ TCRs [31, 136]. However, recently sharks have been reported to have alternative form of isotype switching which involves switching of VDJ exons of one locus with exons of constant region from different locus [52, 137].
Bony Fish (Class: Osteichthyes)
As bony fish share common ancestor with that of mammals, the IgH loci in them unlike that present in cartilaginous fish is consistent with diverse multiple families of rearranging variable heavy and light chain segments that are highly diverse and exhibit translocon arrangement. However, there are few exceptions where IgL locus of some teleosts exhibits cluster arrangement and resembles that of cartilaginous fish [138]. It is interesting to note that bony fish in addition to IgM and IgD possesses a novel Ig isotype IgZ/IgT first discovered in Danio rerio and Oncorhynchus mykiss concomitantly [32, 139]. Increasing evidences suggest that subsequent to the divergence of bony fish from mammals, whole genome duplication occurred in Teleostei as revealed by comparative analysis of pufferfish and mammalian genomes [140]. This may be the probable reason for the subtle differences found in the IgH locus of Ictalurus punctadus, O. mykiss and D. rerio studied so far in terms of the pseudogenes and the duplicated genes [141], translocon arrangement of IgH locus and cluster arrangement of IgL locus [138]. The IgH locus consists of the D, J and C regions of the newly identified isotype IgZ/IgT lying between several V segments and D, J and Cµ and Cδ regions. The IgM or IgD isotype is expressed by alternative splicing of the Cδ and Cµ region exons, respectively, while the novel IgZ/IgT isotype is expressed after alternative rearrangement of the V segments. It should be noted that the rearrangement that entails the expression of IgM locus is basically a deletional process in which (DJC)ζ is removed and (DJC)µ are joined, similar to the way TCRαδ are expressed (Fig. 2b) [142].
During both heavy and light chain rearrangements, N and P nucleotide additions in the V, J and D region occur in a very canonical way [142]. The V(D)J rearrangement takes place in a similar fashion as in cartilaginous fish and employs RAG 1 and RAG 2 proteins. RAG 1/2-mediated dsDNA breaks in RSS regions flanking Ig genes is perpetuated by hairpin formation and resolution by involving non-homologous end-joining (NHEJ) repair pathway [143]. It was determined that combinatorial rearrangements of the V and J segments can occur in the IgL clusters of the catfish by tailoring inversions and avoiding deletions. TdT is predicted to be absent in catfish; however, identification in zebrafish strengthens our notion of its explicit role in providing junctional diversity in bony fish as described previously. The location of the enhancers differs in fish in comparison with higher vertebrates like human and mice [144]. The Eµ3 enhancer is located in between the Cµ and Cδ regions and is comprised of eight octamer motifs with six variable sequences established so far [142, 144]. The enhancers have been additionally shown to have potential to bind to the Octamer-binding transcription factors (Oct1 and Oct2).
Bony fish are evolutionary first vertebrate group, in which extensive functions of AID with respect to protein structure have been established. It is apparent that basic structure of AID with certain germline B cell functions has already been developed early in the evolution; however, not all the aspects of the functions were exploited [128]. Therefore, it is likely that AID may be involved in the recruitment of sequences during somatic hypermutation events. However, switch (S) sequences are identified in the junction intervening the Cµ and Cδ region exons, suggesting that class switching is not present in teleosts similar to cartilaginous fish [142, 144]. Thus, it is clearly evident that during the course of metazoan phylogeny, the anticipatory immune system has sophisticatedly evolved and so as prototypic AID proteins regulating the affinity maturation events (Table 1).
Regulatory Mechanism of AID: Anticipated Targets and Cofactors
It was quite fascinating for many eminent immunologists to speculate the molecular mechanism of AID/AICDA in orchestration of SHM, CSR and GC. CSR and SHM are most salient mechanisms to have evolved in the vertebrates for antigen receptor diversification and enable them to recognize huge range of pathogens through the generated diverse repertoire. Through the analysis of actively transcribing Ig genes by transgenic engineering studies, it was explained that transcription might be prerequisite to generation of Ig diversity based on two crucial observations. First is the fact that ssDNA is predominant targets of AID, and second is that the reports suggest the linear correlation between transcription and hypermutation rates [73, 145]. Upon encountering antigen, naive B cells get activated followed by AID upregulation and its recruitment to active transcription sites. On reaching target location, AID directly converts cytosine to uracil generating U:G mismatches that are subjected to point mutations [73, 146]. As per its name, AID preferentially targets cytidine residues on single-stranded DNA (ssDNA) with consensus sequence WRCH (W = A/T, R = A/G, H = T/C/A) [74, 147]. Second step after deamination of uracil is recruitment of DNA repair machinery involving uracil-DNA glycosylase (UNG) responsible for the removal of uracils from the U:G mismatches introduced by AID [73]. The archetypal DNA repair pathways employing base excision repair (BER) and mismatch repair (MMR) are then initiated. It is during this point that the SHM and CSR take their respective mutational phase differently as SHM generates diversity in the V regions by introducing point mutations, while CSR regulates switching of the CH regions by generating double-stranded breaks in the switch regions [25, 147, 148].
In case of SHM, U:G lesions produced by AID are subjected to replication during which transitional or transversional mutations with a bias on transitions are introduced initiating C:G base pair formation [82, 149]. Following uracil addition, abasic (AP) sites are formed which are subsequent targets of endonucleases. AP endonuclease 1 (APE1) introduces nicks at the AP site on the opposite DNA strands adjacently creating double-stranded breaks (DSBs) in the proximity of AGCT motifs. SHM exploits MMR pathways to process the DSBs created comprising of C:G-based mutations and spreading them to form A:T-based mutations, while CSR employs activities of both BER and MMR repair pathways [43, 82, 150]. During class switch recombination (CSR), AID introduces mutations into large repetitive S regions that flank the upstream portion of all CH genes. S regions harbour multiple guanosine residues and are AGCT hotspots [25]. The AID-initiated lesions are subsequently converted into DSBs by co-opted BER and MMR activities. Deletional end-joining of the DSBs introduced is resolved by employing classical NHEJ pathway preferentially. CSR is promoted when the donor Sμ region located in the upstream end of a DSB harness downstream S region in the downstream end of a DSB of the adjacent strand. This arrangement juxtaposes the V(D)J exon and downstream CH gene that initiate isotype switching of the CH gene located in proximity to the donor switch regions [151].
Large amount of information regarding the AID structure, domain analysis, functional aspects and regulatory mechanisms pertaining to affinity maturation events have been distinct due to extensive studies which revealed the putative proteins and cardinal elements interacting with the functional motifs of AID [152,153,154,155]. It is noteworthy that the regulation of AID on SHM and CSR share several features in common but differ in their selection of DNA repair factors. CSR entirely depends on H2AX, Ku70/Ku80 containing heterodimer required for initial lesion formation and DNA-dependent protein kinases (DNA-PKcs) for processing the recognized lesions [156,157,158], whereas SHM and GC do not employ these proteins. Recent studies have indicated that the expression of AID in B cells is regulated by four conserved regions of DNA. The region I contains a HomeoBox C4/Octamer and NF-kB/SP1/SP2 binding region which regulates AID activation and repression, region II contains silencer elements which substantially regulate the activity of transcriptional factors by binding to repressor proteins like cMyb, region III is responsible for the physiological functions of AID by binding to B cell activating transcriptional factor while the region IV is known to bind to NF-kB and other factors associated with B cell activation [159]. Interestingly, C-terminus region of AID has been indicated to be negatively regulated by MDM2 and its putative target p53 in B cell avian cell line. Although MDM2 is well established as an ubiquitin ligase and is known to shuttle between nucleus and cytoplasm, it is reported to have no role in nucleo-cytoplasmic trafficking of AID [76, 160]. Replication protein A, a regulatory protein, central to replication and repair mechanism is critical for the capability of AID protein to predominantly target ssDNA and not dsDNA in a very mysterious way [161, 162]. Increased evidences suggest that AID needs to be phosphorylated and is subjected to post-translational modifications in order to get activated and is therefore associated with protein kinase A (PKA) [163,164,165]. These findings were further supported by the AID mutational events concerning phosphorylation defects which led to a decrease in somatic hypermutation. It has been shown that AID and replication protein A (RPA) interaction depends on the phosphorylation event entailing PKA at Serine-38 and Threonine-27 position necessary for placing AID in the vicinity of its ssDNA targets which is critical during CSR [161, 163]. Recently, AID has been reported to regulate gene conversion by Bach2 association in DT40 cell line [166]. Hu et al. [167] have elucidated that several heterogeneous nuclear ribonucleoprotein (hnRNPs) cofactors are associated with AID and regulate its important functions like DNA cleavage and recombination. Similar findings of RNA-dependent associations of AID monomers with hnRNP K and dimers with hnRNP U, hnRNP L were reported by Mondal et al. [72].
In contrary to mammals, analysis of AID hotspot motifs in catfish revealed that although the AID targets share common motifs in both mammals and fish, their number in fish is comparatively lesser [60]. Further, unlike mammals, AID access to its targets does not involve PKA phosphorylation at S-38 site in Takifugu, zebrafish and catfish [164]. Zebrafish AID was found to catalyze CSR in mouse cells mutated with S-38 PKA site utilizing aspartate 44 (D44) phosphorylation both in vivo and in vitro. The study showed that the introduction of PKA site in zebrafish AID mutant of aspartate 44 restored the ability to orchestrate CSR. Thus, the presence of classical PKA site in bony fish and AID EST in fish suggest that AID in tetrapods might have shared a common phylogenetic ancestor and have gradually sophisticated the regulatory features and switch elements to harness CSR in higher vertebrates [165]. Human and bony fish AID have been reported to have different deamination efficiencies [168]. Zebrafish AID was found to be more catalytically robust as compared to human and catfish AID. The similar study also elucidated that the differences in the catalytical rates in humans and bony fish AID are associated with the ssDNA binding pattern regulated by a C-terminal residue. Further, in silico study carried out by Villota-Herdoiza et al. [169] to determine the potential transcriptional genes regulating AID function in fish revealed the importance of Myb and E2f binding sites for either all or few introns of the regulatory regions in catfish and zebrafish AID locus. The study additionally predicted that coupling of zebrafish upstream region with intron 1 region would possibly show equivalent function as transcriptional enhancer complex as observed in AID locus of mouse or other higher vertebrates.
Alternative Mechanisms Involved in Regulation of Isotype Switching
In the absence of lymph node, pronephros or head kidney plays an equivalent role in the B cell lymphogenesis [170]. It has been established earlier that the AID expression is limited to germinal centres and the induction of its expression in them is regulated in a T cell-dependent and T cell-independent pathway. Similar to activated B cells, AID gets predominantly induced by any signals that are important to activate CSR in naïve B cells-like interleukin 4 (IL-4), anti-CD40 signals, lipopolysaccharide (LPS) and transforming-growth factor β (TGF-β), to name a few. These signals concomitantly regulate activation of AID expression through association of signal transducer and activator of transcription 6 (STAT6) and canonical nuclear factor-κB (NF-κB) [162, 171].
Recent evidences suggest that cytokine receptor signalling in B cells efficiently drives IgH germline transcription of (CH) regions and activates respective switch elements to initiate CSR. It is well determined that a CH gene cluster comprises of three components present in juxtaposition: the IH promoters crucial for germline transcript initiation, the S regions that are putative site of DSB breaks introduction and the CH genes [172]. In the presence of CD40 or LPS stimuli, IL-4 induces the NF-κB and STAT-6 activation, which bind to their respective promoters, initiate germline Iγ1-Cγ1 and Iε-Cε transcription for IgG1 and IgE, respectively [173]. Additionally, Basu et al. [40] have recently elucidated the crucial role of B cell activating factor (BAFF) induced by TLR and NLR ligands in modulating IgM synthesis. These evidences clearly points towards critical role of PRR stimuli and subsequent signal transduction in inducing CSR through aberrant T cell-independent pathway.
Role of PRRs: TLR-Dependent Pathway
Toll-like receptors are the pivotal components of the innate immune arm of the complex gnathostome immune system, evolved long before in invertebrate and is conserved throughout the tetrapod lineage [174, 175]. These are basically group of TYPE-I integral transmembrane glycoproteins that are equipped to recognize huge range of pathogens and thus provide non-specific immunity. The pathogen detection by TLRs is manifested by the cognate receptor–ligand binding interaction that is regulated by the unique molecular patterns associated with different pathogens recognized by corresponding pattern encoded in the receptors [174]. The molecular patterns pertaining to the pathogens include lipopolysaccharides, flagellin, peptidoglycans, zymosan, dsRNA and are popularly known as pathogen-associated molecular patterns (PAMPs). TLRs and other similar receptors in the group like NOD-like receptors, C-type lectins, RIG-like receptors are therefore known as pattern recognition receptors (PRRs) [176]. The prototypical TLR structure consists of a N-terminal region of primordial LRR repeats (LRR domain), a membrane spanning transmembrane domain and C-terminal Toll/IL-1 receptor family domain (TIR) [175].
The cognate binding of membrane bound TLRs-TLR1, TLR2, TLR4, TLR5 and endosomal TLRs—TLR3, TLR7, TLR8 with respective PAMPs or unique signature molecules located on the surface of microbes results in induction of the highly conserved TLR signalling events [171]. The canonical TLR signalling pathway comprises of MyD88 (myeloid differentiation factor 88)-dependent and MyD88-independent pathways of pro-inflammatory cytokine production and Ig diversification [174]. MyD88-independent pathway is known to harbour TRIF (TIR domain containing adaptor inducing interferon-β) that eventually leads to secretion of TYPE-1 interferons. The MyD88-dependent pathway activates molecules like IL-1RI-associated protein kinase (IRAK), TGF-β-associated kinase (TAK1), TAK1 binding protein 1 (TAB1), TAB2 and TRAF6. TRAF6 activation promotes Iκκ complex ubiquitinylation leading to the activation of NF-κB, followed by pro-inflammatory cytokine production leading to antibody production [170, 177].
LPS recognition by TLR4 in murine B cells along with co-stimulation by IL-4 has earlier been reported to induce B cell proliferation and subsequent cytokine production leading to isotype switching [178]. There is also increasing body of evidence of CSR induction in murine cells followed by co-stimulation of LPS and IFN-γ leading to isotype switching of IgG2a and LPS in combination with TGF-β imparts CSR to IgG2b and IgA. Likewise, Pone et al. [172] have earlier reported that induction of TLR1/6 by cognate ligand, Pam3CSK4 initiated cytokine-mediated CSR. These evidences pointed towards the important conclusions that TLRs not only aid in cytokine production but also contribute to modulation of B cell-regulated antibody responses [40, 177]. BCR crosslinking with O-saccharide component of LPS has elucidated the potential of B cells in regulating antibody response and specificity in an aberrant TLR ligand-independent way. This crosslinking is known to amplify B cell differentiation process by enhancing the cytokine stimuli required for isotype switching of immunoglobulins [173].
NLR-Dependent Pathway
The nucleotide oligomerization domain-like receptors (NLRs) are another set of soluble cytoplasmic PRR family members as described above. NLRs exhibit complex tripartite structure and comprises of three distinct domains that include a C-terminal leucine-rich repeat (LRR domain), a middle nucleotide oligomerization domain (NACHT) and a N-terminal protein–protein interaction domain that can be of three types—caspase activation and recruitment domain (CARD), pyrin domain (PYD) and a baculovirus inhibitor of apoptosis repeat domain (BIR) [179]. The LRR domain is meant for recognizing PAMPs, activating other NLRs in the vicinity leading to the self-oligomerization regulated by the NACHT domain, and the signal is finally processed to the downstream signalling molecules through CARD-PYD-BIR domain [171]. On antigen encounter, NLRs have been known to get activated, self-oligomerize, thus aggravating the combinatorial binding potential and recruiting a downstream signalling recruitment interacting proteins RIP2 (RICK, CARDIAK). This recruitment and signalling mediated by homotypic CARD–CARD domain lead to interaction with NF-κB complex subunits, triggering phosphorylation of IK-κB and activating NF-κB [171]. The NF-κB stimulation and MAPK (mitogen-activated protein kinase) activation collectively results in the activation of specific transcriptional factors like activator protein-1 (AP-1) and induction of pro-inflammatory cytokines and chemokines—interleukin (IL-1β), IL-6, IL8, tumour necrosis factor (TNF) and interferon (IFN-γ). Production of these cytokines and chemokines plays a significant role in the production of antibodies [180].
Role of BAFF/APRIL
Tumour-necrosis factor superfamily 13B (TNFSF 13B) of proteins or cytokines is established in all gnathostomes for a huge range of functions in inflammation, germinal centre development and apoptosis. Two pivotal cytokines include B cell activating factor (BAFF) and its closely related homolog; a proliferation-inducing ligand (APRIL) is exclusively expressed in the myeloid cells [181]. Both these cytokines share two receptors, a transmembrane activator, calcium modulator and cyclophilin ligand interactor (TACI) and B cell maturation antigen (BCMA). The third receptor is restricted to BAFF and is known as BAFF receptor (BAFFR). The role of BAFF has been established in B cell development, T cell-dependent and T cell-independent antibody response, isotype switching [182, 183] (Fig. 3).
A hypothetical diagram of T cell-independent antibody switching mechanism involving TLR. Toll-like receptors on activation by their cognate ligands like lipopolysaccharides, flagellin, dsRNA gets stimulated and activates its downstream signalling cascade through myeloid differentiation factor (MyD88) finally leading to phosphorylation of nuclear factor-kB (NF-kB). This is turn releases p65 subunit of NF-kB complex and enhances transcription of pro-inflammatory cytokines like TNF-α, IL-6, IFN-γ which in turn activates B cell activating factor (BAFF) and a proliferation-inducing ligand (APRIL). BAFF and APRIL are soluble proteins which then bind to their cognate TACI and BCMA receptors present in B cells thus activating it, leading to its proliferation and switching in terms of alternative rearrangement and splicing mechanisms in the absence of AID generating different isotype expressing plasma cells
Research on systematic mutagenesis of various portions of TACI, BCMA and BAFFR has demonstrated different essential determinants of ligand binding in terms of their selectivity and affinity. These studies have exemplified that BAFFR might not be a cognate receptor of APRIL, whereas BAFF and BCMA exhibit very weak interactions [184]. In addition to TACI and BCMA, APRIL also interacts with the polysaccharide side chains of sulfated glycosaminoglycans like heparan sulphate proteoglycans (HSPGs) [180]. This interaction is mediated by basic residues of APRIL and does not interfere with binding to either TACI or BCMA. It is noteworthy that HSPG-bound APRIL is biologically active. TACI is known to consist of two splice variants. One contains a high-affinity ligand binding site which may or may not be preceded by the other low-affinity binding site. Similar to APRIL, TACI interacts with proteoglycans such as syndecans, which are expressed on the surface of multipotent stromal cells, stimulated B cells and macrophages. This interaction results in multimerization of APRIL in the extracellular matrix. Therefore, syndecans may play crucial role as local activators of APRIL, either as ligands that activate signalling through TACI or by promoting appropriate signals to antibody-secreting cells [175].
Whether weak interactions existing between BAFF and BCMA have any physiological relevance was not clear. BAFF and APRIL disruption in BCMA-dependent plasma cells of bone marrow eventually ameliorated the number of plasma cells. However, on selective removal of either BAFF alone or APRIL alone resulted in plasma cell survival, suggesting that in vivo signal recognition by BCMA receptor can occur interruptedly by binding to both the cognate ligands. It was thus postulated that the weak interaction between BAFF and BCMA is probably stabilized by avidity effects [181]. Bombardieri et al. [177] has indicated earlier that human rheumatoid arthritis synovial fibroblasts express high levels of BAFF and APRIL in response to TLR3 induction. This TLR-dependent B cell stimulation led to the enhancement of AID expression and promoted CSR in unswitched IgD + B cells. Additionally, Castigli et al. [183] have demonstrated the role of BAFF and APRIL receptors in inducing class switching in mouse B cells by examining their mice deficient in TACI, BCMA and BAFFR receptors. All these results clearly indicate that BAFF and APRIL-may have a critical role in inducing class switching of Ig isotypes through TLR and NLR signalling pathways in the absence of conventional switching mechanism in lower vertebrates like fish.
Concluding Remarks
It is clear from the above that since the introduction of hybridoma technology, the discovery of AID and elucidation of detailed molecular mechanisms of AID-mediated affinity maturation events have revolutionized our paradigms regarding complex diversification mechanisms pertaining to both variable and constant regions of immunoglobulin molecule both in fish and in tetrapod lineage [185, 186]. Several lines of recent evidences indicate that role of AID is not restricted to orchestration of SHM and CSR events, rather it is involved in epigenetic reprogramming, disrupting retroviral replication and possess wide range of targets in B cells [161, 187]. Further, genetic manipulation and biochemical studies need to be conducted in order to gain a clear insight into the functional aspects of AID/APOBEC proteins in antigen receptor diversification mechanisms. AID regulation in the germinal B cells in a TLR-dependent and TLR-independent fashion [97], its potency in DNA mutation reactions and its probable non-Ig targets have already been reported to have huge implications in tumorigenesis [88, 188,189,190,191]. Collectively, it can now be envisioned the possible alternative class switching mechanisms regulated by AID entailing TLR-dependent Ig diversification mechanisms might be responsible for the presence of multiple Ig isotypes in earlier representatives of vertebrates—cartilaginous and bony fish.
The enigma of how VLR-based adaptive immune system emerged in lampreys in the evolutionary dichotomy between protochordates and earlier representatives of chordates [2] still remains uncovered and needs to be determined. Several extensive studies in this line needs to be carried out to address the unanswered issues such as direct role of non-functional AID of fish in tailoring Ig class switching. Furthermore, such studies would shed light in the complicated affinity maturation event and CSR evolution throughout the metazoan phylogeny. Comparative studies on AID proteins from protochordates to mammalian vertebrates and deeper information regarding their biological properties both in vivo and in vitro would probably aid in resolving this paradigm.
Abbreviations
- VLRs:
-
Variable lymphocyte receptors
- Ig:
-
Immunoglobulin
- SHM:
-
Somatic hypermutation
- CSR:
-
Class-switch recombination
- GC:
-
Gene conversion
- AID:
-
Activation-induced cytidine deaminase
- APOBEC:
-
Apolipoprotein B mRNA-editing catalytical component
- BAFF:
-
B cell activating factor
- APRIL:
-
A proliferation-inducing ligand
- TLR:
-
Toll-like receptors
- TNF:
-
Tumour necrosis factor
- NF-κB:
-
Nuclear factor-κB
- MHC:
-
Major histocompatibility complex
- MDM2:
-
Mouse double minute 2 homolog
- LPS:
-
Lipopolysaccharide
References
Laird, D. J., De Tomaso, A. W., Cooper, M. D., & Weissman, I. L. (2000). 50 million years of chordate evolution: Seeking the origins of adaptive immunity. Proceedings of the National Academy of Sciences, 97, 6924–6926.
Pancer, Z., Amemiya, C. T., Ehrhardt, G. R., Ceitlin, J., Gartland, G. L., & Cooper, M. D. (2004). Somatic diversification of variable lymphocyte receptors in the agnathan sea lamprey. Nature, 430, 174–180.
Cooper, M. D., & Alder, M. N. (2006). The evolution of adaptive immune systems. Cell, 124, 815–822.
Guo, P., Hirano, M., Herrin, B. R., Li, J., Yu, C., Sadlonova, A., et al. (2009). Dual nature of the adaptive immune system in lampreys. Nature, 459, 796–801.
Flajnik, M. F. (2002). Comparative analyses of immunoglobulin genes: Surprises and portents. Nature Reviews Immunology, 2, 688–698.
Sunyer, J. O. (2013). Fishing for mammalian paradigms in the teleost immune system. Nature Immunology, 14, 320–326.
Senger, K., Hackney, J., Payandeh, J., & Zarrin, A. A. (2015). Antibody isotype switching in vertebrates. In E. Hsu & L. D. Pasquier (Eds.), Pathogen–host interactions: antigenic variation v. somatic adaptations (pp. 295–324). Berlin: Springer.
Hsu, E. (2016). Assembly and expression of shark Ig genes. The Journal of Immunology, 196, 3517–3523.
Poole, J. R. M., Paganini, J., & Pontarotti, P. (2017). Convergent evolution of the adaptive immune response in jawed vertebrates and cyclostomes: An evolutionary biology approach based study. Developmental & Comparative Immunology. https://doi.org/10.1016/j.dci.2017.02.011.
Burnet, F. M. (1957). A modification of Jerne’s theory of antibody production using the concept of clonal selection. The Australian Journal of Science, 20, 67–69.
Hodgkin, P. D., Heath, W. R., & Baxter, A. G. (2007). The clonal selection theory: 50 years since the revolution. Nature Immunology, 8, 1019–1026.
Brack, C., Hirama, M., Lenhard-Schuller, R., & Tonegawa, S. (1978). A complete immunoglobulin gene is created by somatic recombination. Cell, 15, 1–14.
Seidman, J. G., Leder, A., Edgell, M. H., Polsky, F., Tilghman, S. M., Tiemeier, D. C., et al. (1978). Multiple related immunoglobulin variable-region genes identified by cloning and sequence analysis. Proceedings of the National Academy of Sciences of the United States of America, 75, 3881–3885.
Weigert, M. G., Cesari, I. M., Yonkovich, S. J., & Cohn, M. (1970). Variability in the lambda light chain sequences of mouse antibody. Nature, 228, 1045–1047.
Schilling, J., Clevinger, B., Davie, J. M., & Hood, L. (1980). Amino acid sequence of homogeneous antibodies to dextran and DNA rearrangments in heavy chain V-region gene segments. Nature, 283, 35–40.
Tonegawa, S. (1983). Somatic generation of antibody diversity. Nature, 302, 575–581.
Oettinger, M. A., Schatz, D. G., Gorka, C., & Baltimore, D. (1990). RAG-1 and RAG-2, adjacent genes that synergistically activate V (D) J recombination. Science, 248, 1517–1523.
Bassing, C. H., Swat, W., & Alt, F. W. (2002). The mechanism and regulation of chromosomal V (D) J recombination. Cell, 109, S45–S55.
Gellert, M. (2002). V(D)J recombination: RAG proteins, repair factors, and regulation. Annual Review of Biochemistry, 71, 101–132.
Kataoka, T., Kawakami, T., Takahashi, N., & Honjo, T. (1980). Rearrangement of immunoglobulin gamma 1-chain gene and mechanism for heavy-chain class switch. Proceedings of the National Academy of Sciences of the United States of America, 77, 919–923.
Reynaud, C. A., Anquez, V., Grimal, H., & Weill, J. C. (1987). A hyperconversion mechanism generates the chicken light chain preimmune repertoire. Cell, 48, 379–388.
Reynaud, C. A., Mackay, C. R., Müller, R. G., & Weill, J. C. (1991). Somatic generation of diversity in a mammalian primary lymphoid organ: The sheep ileal Peyer’s patches. Cell, 64, 995–1005.
Litman, G. W., Rast, J. P., & Fugmann, S. D. (2010). The origins of vertebrate adaptive immunity. Nature Reviews Immunology, 10, 543–553.
Arakawa, H., Saribasak, H., & Buerstedde, J. M. (2004). Activation-induced cytidine deaminase initiates immunoglobulin gene conversion and hypermutation by a common intermediate. PLoS Biology, 2, 0967–0971.
Muramatsu, M., Kinoshita, K., Fagarasan, S., Yamada, S., Shinkai, Y., & Honjo, T. (2000). Class switch recombination and hypermutation require activation-induced cytidine deaminase (AID), a potential RNA editing enzyme. Cell, 102, 553–563.
Stavnezer, J., & Amemiya, C. T. (2004). Evolution of isotype switching. Seminars in Immunology, 16(4), 257–275.
Wakae, K., Magor, B. G., Saunders, H., Nagaoka, H., Kawamura, A., Kinoshita, K., et al. (2006). Evolution of class switch recombination function in fish activation-induced cytidine deaminase, AID. International Immunology, 18, 41–47.
Barreto, V. M., Pan-Hammarstrom, Q., Zhao, Y., Hammarstrom, L., Misulovin, Z., & Nussenzweig, M. C. (2005). AID from bony fish catalyzes class switch recombination. The Journal of Experimental Medicine, 202, 733–738.
Wilson, M., Bengtén, E., Miller, N. W., Clem, L. W., Du Pasquier, L., & Warr, G. W. (1997). A novel chimeric Ig heavy chain from a teleost fish shares similarities to IgD. Proceedings of the National Academy of Sciences of the United States of America, 94, 4593–4597.
Flajnik, M. F., & Rumfelt, L. L. (2000). The immune system of cartilaginous fish. In L. Du Pasquier & G. W. Litman (Eds.), Origin and evolution of the vertebrate immune system (pp. 249–270). Berlin: Springer.
Dooley, H., & Flajnik, M. F. (2006). Antibody repertoire development in cartilaginous fish. Developmental & Comparative Immunology, 30, 43–56.
Danilova, N., Bussmann, J., Jekosch, K., & Steiner, L. A. (2005). The immunoglobulin heavy-chain locus in zebrafish: Identification and expression of a previously unknown isotype, immunoglobulin Z. Nature Immunology, 6, 295–302.
Zhang, Y. A., Salinas, I., Li, J., Parra, D., Bjork, S., Xu, Z., et al. (2010). IgT, a primitive immunoglobulin class specialized in mucosal immunity. Nature Immunology, 11, 827–835.
Patel, B., Banerjee, R., Basu, M., Lenka, S., Samanta, M., & Das, S. (2016). Molecular cloning of IgZ heavy chain isotype in Catla catla and comparative expression profile of IgZ and IgM following pathogenic infection. Microbiology and Immunology, 60, 561–567.
Basu, M., Lenka, S. S., Paichha, M., Swain, B., Patel, B., Banerjee, R., et al. (2016). Immunoglobulin (Ig) D in Labeo rohita is widely expressed and differentially modulated in viral, bacterial and parasitic challenges. Veterinary Immunology and Immunopathology, 179, 77–84.
Castigli, E., Fuleihan, R., Ramesh, N., Tsitsikov, E., Tsytsykova, A., & Geha, R. S. (1995). CD40 ligand/CD40 deficiency. International Archives of Allergy and Immunology, 107, 37–39.
Stavnezer, J. (1996). Immunoglobulin class switching. Current Opinion in Immunology, 8, 199–205.
Geha, R. S., Jabara, H. H., & Brodeur, S. R. (2003). The regulation of immunoglobulin E class-switch recombination. Nature Reviews Immunology, 3, 721–732.
Litinskiy, M. B., Nardelli, B., Hilbert, D. M., He, B., Schaffer, A., Casali, P., et al. (2002). DCs induce CD40-independent immunoglobulin class switching through BLyS and APRIL. Nature Immunology, 3, 822–829.
Basu, M., Lenka, S. S., Paichha, M., Patel, B., Banerjee, R., Das, S., et al. (2016). B cell activating factor is induced by toll-like receptor and NOD-like receptor ligands and plays critical role in IgM synthesis in Labeo rohita. Molecular Immunology, 78, 9–26.
Ramakrishnan, P., Wang, W., & Wallach, D. (2004). Receptor-specific signaling for both the alternative and the canonical NF-κB activation pathways by NF-κB-inducing kinase. Immunity, 21, 477–489.
Arakawa, H., Hauschild, J., & Buerstedde, J. M. (2002). Requirement of the activation-induced deaminase (AID) gene for immunoglobulin gene conversion. Science, 295, 1301–1306.
Honjo, T., Kinoshita, K., & Muramatsu, M. (2002). Molecular mechanism of class switch recombination: Linkage with somatic hypermutation. Annual Review of Immunology, 20, 165–196.
Harris, R. S., Sale, J. E., Petersen-Mahrt, S. K., & Neuberger, M. S. (2002). AID is essential for immunoglobulin V gene conversion in a cultured B cell line. Current Biology, 12, 435–438.
Delker, R. K., Fugmann, S. D., & Papavasiliou, F. N. (2009). A coming-of-age story: Activation-induced cytidine deaminase turns 10. Nature Immunology, 10, 1147–1153.
Gitlin, A. D., von Boehmer, L., Gazumyan, A., Shulman, Z., Oliveira, T. Y., & Nussenzweig, M. C. (2016). Independent roles of switching and hypermutation in the development and persistence of B lymphocyte memory. Immunity, 44, 769–781.
Parng, C. L., Hansal, S., Goldsby, R. A., & Osborne, B. A. (1996). Gene conversion contributes to Ig light chain diversity in cattle. The Journal of Immunology, 157, 5478–5486.
Weinstein, P. D., Anderson, A. O., & Maget, R. G. (1994). Rabbit IgH sequences in appendix germinal centers: VH diversification by gene conversion-like and hypermutation mechanisms. Immunity, 1, 647–659.
Diaz, M., & Flajnik, M. E. (1998). Evolution of somatic hypermutation and gene conversion in adaptive immunity. Immunological Reviews, 162, 13–24.
Muramatsu, M., Sankaranand, V. S., Anant, S., Sugai, M., Kinoshita, K., Davidson, N. O., et al. (1999). Specific expression of activation-induced cytidine deaminase (AID), a novel member of the RNA-editing deaminase family in germinal center B cells. The Journal of Biological Chemistry, 274, 18470–18476.
Revy, P., Muto, T., Levy, Y., Geissmann, F., Plebani, A., Sanal, O., et al. (2000). Activation-induced cytidine deaminase (AID) deficiency causes the autosomal recessive form of the Hyper-IgM syndrome (HIGM2). Cell, 102, 565–575.
Magor, B. G. (2015). Antibody affinity maturation in fishes—Our current understanding. Biology, 4, 512–524.
Wilson, M., Hsu, E., Marcuz, A., Courtet, M., Du Pasquier, L., & Steinberg, C. (1992). What limits affinity maturation of antibodies in Xenopus—The rate of somatic mutation or the ability to select mutants? The EMBO Journal, 11, 4337–4347.
Greenberg, A. S., Avila, D., Hughes, M., Hughes, A., McKinney, E. C., & Flajnik, M. F. (1995). A new antigen receptor gene family that undergoes rearrangement and extensive somatic diversification in sharks. Nature, 374, 168–173.
Hinds-Frey, K. R., Nishikata, H., Litman, R. T., & Litman, G. W. (1993). Somatic variation precedes extensive diversification of germline sequences and combinatorial joining in the evolution of immunoglobulin heavy chain diversity. Journal of Experimental Medicine, 178(3), 815–824.
Diaz, M., Greenberg, A. S., & Flajnik, M. F. (1998). Somatic hypermutation of the new antigen receptor gene (NAR) in the nurse shark does not generate the repertoire: Possible role in antigen-driven reactions in the absence of germinal centers. Proceedings of the National Academy of Sciences, 95(24), 14343–14348.
Lee, S. S., Tranchina, D., Ohta, Y., Flajnik, M. F., & Hsu, E. (2002). Hypermutation in shark immunoglobulin light chain genes results in contiguous substitutions. Immunity, 16, 571–582.
Cain, K. D., Jones, D. R., & Raison, R. L. (2002). Antibody–antigen kinetics following immunization of rainbow trout (Oncorhynchus mykiss) with a T-cell dependent antigen. Developmental & Comparative Immunology., 26(2), 181–190.
Saunders, H. L., & Magor, B. G. (2004). Cloning and expression of the AID gene in the channel catfish. Developmental & Comparative Immunology, 28, 657–663.
Yang, F., Waldbieser, G. C., & Lobb, C. J. (2006). The nucleotide targets of somatic mutation and the role of selection in immunoglobulin heavy chains of a teleost fish. The Journal of Immunology, 176(3), 1655–1667.
Marianes, A. E., & Zimmerman, A. M. (2011). Targets of somatic hypermutation within immunoglobulin light chain genes in zebrafish. Immunology, 132(2), 240–255.
Matsumoto, M., Lo, S. F., Carruthers, C. J., Min, J., Mariathasan, S., Huang, G., et al. (1996). Affinity maturation without germinal centres in lymphotoxin-α-deficient mice. Nature, 382, 462.
William, J., Euler, C., Christensen, S., & Shlomchik, M. J. (2002). Evolution of autoantibody responses via somatic hypermutation outside of germinal centers. Science, 297(5589), 2066–2070.
Peters, A., & Storb, U. (1996). Somatic hypermutation of immunoglobulin genes is linked to transcription initiation. Immunity, 4, 57–65.
Wiesendanger, M., Scharff, M. D., & Edelmann, W. (1998). Somatic hypermutation, transcription, and DNA mismatch repair. Cell, 94, 415–418.
Okazaki, I. M., Kinoshita, K., Muramatsu, M., Yoshikawa, K., & Honjo, T. (2002). The AID enzyme induces class switch recombination in fibroblasts. Nature, 416, 340–345.
Yoshikawa, K., Okazaki, I. M., Eto, T., Kinoshita, K., Muramatsu, M., Nagaoka, H., et al. (2002). AID enzyme-induced hypermutation in an actively transcribed gene in fibroblasts. Science, 296, 2033–2036.
Martin, A., Bardwell, P. D., Woo, C. J., Fan, M., Shulman, M. J., & Scharff, M. D. (2002). Activation-induced cytidine deaminase turns on somatic hypermutation in hybridomas. Nature, 415, 802–806.
Petersen-Mahrt, S. K., Harris, R. S., & Neuberger, M. S. (2002). AID mutates E. coli suggesting a DNA deamination mechanism for antibody diversification. Nature, 418, 99–104.
Mayorov, V. I., Rogozin, I. B., Adkison, L. R., Frahm, C., Kunkel, T. A., & Pavlov, Y. I. (2005). Expression of human AID in yeast induces mutations in context similar to the context of somatic hypermutation at GC pairs in immunoglobulin genes. BMC Immunology, 6, 10.
Dudley, D. D., Chaudhuri, J., Bassing, C. H., & Alt, F. W. (2005). Mechanism and control of V (D) J recombination versus class switch recombination: Similarities and differences. Advances in Immunology, 86, 43–112.
Mondal, S., Begum, N. A., Hu, W., & Honjo, T. (2016). Functional requirements of AID’s higher order structures and their interaction with RNA-binding proteins. Proceedings of the National Academy of Sciences of the United States of America, 113, E1545–E1554.
Di Noia, J., & Neuberger, M. S. (2002). Altering the pathway of immunoglobulin hypermutation by inhibiting uracil-DNA glycosylase. Nature, 419, 43–48.
Bransteitter, R., Pham, P., Scharff, M. D., & Goodman, M. F. (2003). Activation-induced cytidine deaminase deaminates deoxycytidine on single-stranded DNA but requires the action of RNase. Proceedings of the National Academy of Sciences of the United States of America, 100, 4102–4107.
Dickerson, S. K., Market, E., Besmer, E., & Papavasiliou, F. N. (2003). AID mediates hypermutation by deaminating single stranded DNA. The Journal of Experimental Medicine, 197, 1291–1296.
Conticello, S. G. (2008). The AID/APOBEC family of nucleic acid mutators. Genome Biology, 9, 229.
Larijani, M., & Martin, A. (2012). The biochemistry of activation-induced deaminase and its physiological functions. Seminars in Immunology, 24(4), 255–263.
Chester, A., Scott, J., Anant, S., & Navaratnam, N. (2000). RNA editing: Cytidine to uridine conversion in apolipoprotein B mRNA. Biochimica et Biophysica Acta (BBA)-Gene Structure and Expression, 1494, 1–13.
Kinoshita, K., & Honjo, T. (2001). Linking class-switch recombination with somatic hypermutation. Nature Reviews Molecular Cell Biology, 2, 493–503.
Chester, A., Weinreb, V., Carter, C. W., & Navaratnam, N. (2004). Optimization of apolipoprotein B mRNA editing by APOBEC1 apoenzyme and the role of its auxiliary factor, ACF. Rna, 10, 1399–1411.
Vonica, A., Rosa, A., Arduini, B. L., & Brivanlou, A. H. (2011). APOBEC2, a selective inhibitor of TGFβ signaling, regulates left–right axis specification during early embryogenesis. Developmental Biology, 350, 13–23.
Honjo, T., Muramatsu, M., & Fagarasan, S. (2004). AID: How does it aid antibody diversity? Immunity, 20, 659–668.
Conticello, S. G., Thomas, C. J., Petersen-Mahrt, S. K., & Neuberger, M. S. (2005). Evolution of the AID/APOBEC family of polynucleotide (deoxy) cytidine deaminases. Molecular Biology and Evolution, 22, 367–377.
Cullen, B. R. (2006). Role and mechanism of action of the APOBEC3 family of antiretroviral resistance factors. Journal of Virology, 80, 1067–1076.
Gandhi, S. K., Siliciano, J. D., Bailey, J. R., Siliciano, R. F., & Blankson, J. N. (2008). Role of APOBEC3G/F-mediated hypermutation in the control of human immunodeficiency virus type 1 in elite suppressors. Journal of Virology, 82, 3125–3130.
Siriwardena, S., Chen, K., & Bhagwat, A. S. (2016). The functions and malfunctions of AID/APOBEC family deaminases: The known knowns and the known unknowns. Chemical Reviews, 116, 12688–12710.
Starrett, G. J., Luengas, E. M., McCann, J. L., Ebrahimi, D., Temiz, N. A., Love, R. P., et al. (2016). The DNA cytosine deaminase APOBEC3H haplotype I likely contributes to breast and lung cancer mutagenesis. Nature Communications, 7, 12918.
King, J. J., & Larijani, M. (2017). A novel regulator of activation-induced cytidine deaminase/APOBeCs in immunity and cancer: Schrödinger’s CATalytic Pocket. Frontiers in Immunology, 8, 351. https://doi.org/10.3389/fimmu.2017.00351.
Rogozin, I. B., Basu, M. K., Jordan, I. K., Pavlov, Y. I., & Koonin, E. V. (2005). APOBEC4, a new member of the AID/APOBEC family of polynucleotide (deoxy) cytidine deaminases predicted by computational analysis. Cell Cycle, 4, 1281–1285.
Tacchi, L., Larragoite, E. T., Muñoz, P., Amemiya, C. T., & Salinas, I. (2015). African lungfish reveal the evolutionary origins of organized mucosal lymphoid tissue in vertebrates. Current Biology, 25(18), 2417–2424.
Saunders, H. L., Oko, A. L., Scott, A. N., Fan, C. W., & Magor, B. G. (2010). The cellular context of AID expressing cells in fish lymphoid tissues. Developmental & Comparative Immunology, 34, 669–676.
Barreto, V. M., & Magor, B. G. (2011). Activation-induced cytidine deaminase structure and functions: A species comparative view. Developmental & Comparative Immunology, 35, 991–1007.
King, J. J., Manuel, C. A., Barrett, C. V., Raber, S., Lucas, H., Sutter, P., et al. (2015). Catalytic pocket inaccessibility of activation-induced cytidine deaminase is a safeguard against excessive mutagenic activity. Structure, 23, 615–627.
Shinkura, R., Ito, S., Begum, N. A., Nagaoka, H., Muramatsu, M., Kinoshita, K., et al. (2004). Separate domains of AID are required for somatic hypermutation and class-switch recombination. Nature Immunology, 5, 707–712.
Geisberger, R., Rada, C., & Neuberger, M. S. (2009). The stability of AID and its function in class-switching are critically sensitive to the identity of its nuclear-export sequence. Proceedings of the National Academy of Sciences of the United States of America, 106, 6736–6741.
Xu, Z., Zan, H., Pone, E. J., Mai, T., & Casali, P. (2012). Immunoglobulin class-switch DNA recombination: Induction, targeting and beyond. Nature Reviews Immunology, 12, 517–531.
Zan, H., & Casali, P. (2013). Regulation of Aicda expression and AID activity. Autoimmunity, 46, 83–101.
Barreto, V., Reina-San-Martin, B., Ramiro, A. R., McBride, K. M., & Nussenzweig, M. C. (2003). C-terminal deletion of AID uncouples class switch recombination from somatic hypermutation and gene conversion. Molecular Cell, 12, 501–508.
Ta, V. T., Nagaoka, H., Catalan, N., Durandy, A., Fischer, A., Imai, K., et al. (2003). AID mutant analyses indicate requirement for class-switch-specific cofactors. Nature Immunology, 4, 843–848.
McBride, K. M., Barreto, V., Ramiro, A. R., Stavropoulos, P., & Nussenzweig, M. C. (2004). Somatic hypermutation is limited by CRM1-dependent nuclear export of activation-induced deaminase. The Journal of Experimental Medicine, 199, 1235–1244.
Ichikawa, H. T., Sowden, M. P., Torelli, A. T., Bachl, J., Huang, P., Dance, G. S., et al. (2006). Structural phylogenetic analysis of activation-induced deaminase function. The Journal of Immunology, 177, 355–361.
Flajnik, M. F., & Kasahara, M. (2010). Origin and evolution of the adaptive immune system: Genetic events and selective pressures. Nature Reviews Genetics, 11, 47–59.
Ramsden, D. A., Weed, B. D., & Reddy, Y. V. (2010). V (D) J recombination: Born to be wild. Seminars in Cancer Biology, 20(4), 254–260.
Cooper, M. D., & Herrin, B. R. (2010). How did our complex immune system evolve? Nature Reviews Immunology, 10, 2–3.
Hirano, M., Das, S., Guo, P., & Cooper, M. D. (2011). The evolution of adaptive immunity in vertebrates. Advances in Immunology, 109, 125–157.
Dehal, P., Satou, Y., Campbell, R. K., Chapman, J., Degnan, B., De Tomaso, A., et al. (2002). The draft genome of Ciona intestinalis: Insights into chordate and vertebrate origins. Science, 298, 2157–2167.
Cannon, J. P., Haire, R. N., & Litman, G. W. (2002). Identification of diversified genes that contain immunoglobulin-like variable regions in a protochordate. Nature Immunology, 3, 1200–1207.
Kasahara, M., Suzuki, T., & Du Pasquier, L. (2004). On the origins of the adaptive immune system: Novel insights from invertebrates and cold-blooded vertebrates. Trends in Immunology, 25, 105–111.
Klein, J., Sato, A., & Mayer, W. E. (1999). The major histocompatibility complex: Evolution, structure and function (pp. 3–26). Tokyo: Springer.
Herrin, B. R., & Cooper, M. D. (2010). Alternative adaptive immunity in jawless vertebrates. The Journal of Immunology, 185, 1367–1374.
Pancer, Z., Mayer, W. E., Klein, J., & Cooper, M. D. (2004). Prototypic T cell receptor and CD4-like coreceptor are expressed by lymphocytes in the agnathan sea lamprey. Proceedings of the National Academy of Sciences of the United States of America, 101, 13273–13278.
Pancer, Z., Saha, N. R., Kasamatsu, J., Suzuki, T., Amemiya, C. T., Kasahara, M., et al. (2005). Variable lymphocyte receptors in hagfish. Proceedings of the National Academy of Sciences of the United States of America, 102, 9224–9229.
Schatz, D. G. (2007). DNA deaminases converge on adaptive immunity. Nature Immunology, 8, 551–553.
Perey, D. Y., Finstad, J., Pollara, B., & Good, R. A. (1968). Evolution of the immune response. VI. First and second set skin homograft rejections in primitive fishes. Laboratory Investigation, 19, 591–597.
Finstad, J., & Good, R. A. (1964). The evolution of the immune response: III. Immunologic responses in the lamprey. The Journal of Experimental Medicine, 120, 1151.
Marchalonis, J. J., & Edelman, G. M. (1968). Phylogenetic origins of antibody structure: III. Antibodies in the primary immune response of the sea lamprey, Petromyzon marinus. The Journal of Experimental Medicine, 127, 891–914.
Shintani, S., Terzic, J., Sato, A., Saraga-Babic, M., O’hUigin, C., Tichy, H., et al. (2000). Do lampreys have lymphocytes? The Spi evidence. Proceedings of the National Academy of Sciences of the United States of America, 97, 7417–7422.
Alder, M. N., Rogozin, I. B., Iyer, L. M., Glazko, G. V., Cooper, M. D., & Pancer, Z. (2005). Diversity and function of adaptive immune receptors in a jawless vertebrate. Science, 310, 1970–1973.
Rogozin, I. B., Iyer, L. M., Liang, L., Glazko, G. V., Liston, V. G., Pavlov, Y. I., et al. (2007). Evolution and diversification of lamprey antigen receptors: Evidence for involvement of an AID-APOBEC family cytosine deaminase. Nature Immunology, 8, 647–656.
Kishishita, N., & Nagawa, F. (2014). Evolution of adaptive immunity: Implications of a third lymphocyte lineage in lampreys. Bioessays, 36, 244–250.
Kim, H. M., Oh, S. C., Lim, K. J., Kasamatsu, J., Heo, J. Y., Park, B. S., et al. (2007). Structural diversity of the hagfish variable lymphocyte receptors. The Journal of Biological Chemistry, 282, 6726–6732.
Nagawa, F., Kishishita, N., Shimizu, K., Hirose, S., Miyoshi, M., Nezu, J., et al. (2007). Antigen-receptor genes of the agnathan lamprey are assembled by a process involving copy choice. Nature Immunology, 8, 206–213.
Hirano, M., Guo, P., McCurley, N., Schorpp, M., Das, S., Boehm, T., et al. (2013). Evolutionary implications of a third lymphocyte lineage in lampreys. Nature, 501, 435–438.
Boehm, T., McCurley, N., Sutoh, Y., Schorpp, M., Kasahara, M., & Cooper, M. D. (2012). VLR-based adaptive immunity. Annual Review of Immunology, 30, 203–220.
Zhu, C., Lee, V., Finn, A., Senger, K., Zarrin, A. A., Du Pasquier, L., et al. (2012). Origin of immunoglobulin isotype switching. Current Biology, 22, 872–880.
Rast, J. P., & Litman, G. W. (1998). Towards understanding the evolutionary origins and early diversification of rearranging antigen receptors. Immunological Reviews, 166, 79–86.
Lee, S. S., Fitch, D., Flajnik, M. F., & Hsu, E. (2000). Rearrangement of immunoglobulin genes in shark germ cells. The Journal of Experimental Medicine, 191, 1637–1648.
Eason, D. D., Cannon, J. P., Haire, R. N., Rast, J. P., Ostrov, D. A., & Litman, G. W. (2004). Mechanisms of antigen receptor evolution. Seminars in Immunology, 16(4), 215–226.
Hinds, K. R., & Litman, G. W. (1986). Major reorganization of immunoglobulin VH segmental elements during vertebrate evolution. Nature, 320, 546–549.
Rumfelt, L. L., McKinney, E. C., Taylor, E., & Flajnik, M. F. (2002). The development of primary and secondary lymphoid tissues in the nurse shark Ginglymostoma cirratum: B-cell zones precede dendritic cell immigration and T-cell zone formation during ontogeny of the spleen. Scandinavian Journal of Immunology, 56, 130–148.
Van Gent, D. C., Ramsden, D. A., & Gellert, M. (1996). The RAG1 and RAG2 proteins establish the 12/23 rule in V (D) J recombination. Cell, 85, 107–113.
Rumfelt, L. L., Avila, D., Diaz, M., Bartl, S., McKinney, E. C., & Flajnik, M. F. (2001). A shark antibody heavy chain encoded by a nonsomatically rearranged VDJ is preferentially expressed in early development and is convergent with mammalian IgG. Proceedings of the National Academy of Sciences of the United States of America, 98, 1775–1780.
Scapigliati, G. (2013). Functional aspects of fish lymphocytes. Developmental & Comparative Immunology, 41, 200–208.
Malecek, K., Brandman, J., Brodsky, J. E., Ohta, Y., Flajnik, M. F., & Hsu, E. (2005). Somatic hypermutation and junctional diversification at Ig heavy chain loci in the nurse shark. The Journal of Immunology, 175, 8105–8115.
Kato, L., Stanlie, A., Begum, N. A., Kobayashi, M., Aida, M., & Honjo, T. (2012). An evolutionary view of the mechanism for immune and genome diversity. The Journal of Immunology, 188, 3559–3566.
Eason, D. D., Litman, R. T., Luer, C. A., Kerr, W., & Litman, G. W. (2004). Expression of individual immunoglobulin genes occurs in an unusual system consisting of multiple independent loci. European Journal of Immunology, 34, 2551–2558.
Zhang, C., Du Pasquier, L., & Hsu, E. (2013). Shark IgW C region diversification through RNA processing and isotype switching. The Journal of Immunology, 191, 3410–3418.
Ghaffari, S. H., & Lobb, C. J. (1997). Structure and genomic organization of a second class of immunoglobulin light chain genes in the channel catfish. The Journal of Immunology, 159, 250–258.
Hansen, J. D., Landis, E. D., & Phillips, R. B. (2005). Discovery of a unique Ig heavy-chain isotype (IgT) in rainbow trout: Implications for a distinctive B cell developmental pathway in teleost fish. Proceedings of the National Academy of Sciences of the United States of America, 102, 6919–6924.
Jaillon, O., Aury, J. M., Brunet, F., Petit, J. L., Stange-Thomann, N., Mauceli, E., et al. (2004). Genome duplication in the teleost fish Tetraodon nigroviridis reveals the early vertebrate proto-karyotype. Nature, 431, 946–957.
Hsu, E., Pulham, N., Rumfelt, L. L., & Flajnik, M. F. (2006). The plasticity of immunoglobulin gene systems in evolution. Immunological Reviews, 210, 8–26.
Bengtén, E., Clem, L. W., Miller, N. W., Warr, G. W., & Wilson, M. (2006). Channel catfish immunoglobulins: Repertoire and expression. Developmental & Comparative Immunology, 30, 77–92.
Fillatreau, S., Six, A., Magadan, S., Castro, R., Sunyer, J. O., & Boudinot, P. (2013). The astonishing diversity of Ig classes and B cell repertoires in teleost fish. Frontiers in immunology, 4, 28.
Hikima, J. I., Jung, T. S., & Aoki, T. (2011). Immunoglobulin genes and their transcriptional control in teleosts. Developmental & Comparative Immunology, 35, 924–936.
Vallur, A. C., Yabuki, M., Larson, E. D., & Maizels, N. (2007). AID in antibody perfection. Cellular and Molecular Life Sciences, 64, 555–565.
Sohail, A., Klapacz, J., Samaranayake, M., Ullah, A., & Bhagwat, A. S. (2003). Human activation-induced cytidine deaminase causes transcription-dependent, strand-biased C to U deaminations. Nucleic Acids Research, 31, 2990–2994.
Pham, P., Bransteitter, R., Petruska, J., & Goodman, M. F. (2003). Processive AID-catalysed cytosine deamination on single-stranded DNA simulates somatic hypermutation. Nature, 424, 103–107.
DiMenna, L. J., & Chaudhuri, J. (2016). Regulating infidelity: RNA-mediated recruitment of AID to DNA during class switch recombination. European Journal of Immunology, 46, 523–530.
Hwang, J. K., Alt, F. W., & Yeap, L. S. (2015). Related mechanisms of antibody somatic hypermutation and class switch recombination. Microbiology Spectrum, 3, 1–35.
Chatterji, M., Unniraman, S., McBride, K. M., & Schatz, D. G. (2007). Role of activation-induced deaminase protein kinase A phosphorylation sites in Ig gene conversion and somatic hypermutation. The Journal of Immunology, 179, 5274–5280.
Matthews, A. J., Zheng, S., DiMenna, L. J., & Chaudhuri, J. (2014). Regulation of immunoglobulin class-switch recombination: Choreography of noncoding transcription, targeted DNA deamination, and long-range DNA repair. Advances in Immunology, 122, 1–57.
Patenaude, A. M., Orthwein, A., Hu, Y., Campo, V. A., Kavli, B., Buschiazzo, A., et al. (2009). Active nuclear import and cytoplasmic retention of activation-induced deaminase. Nature Structural & Molecular Biology, 16, 517–527.
Pavri, R., Gazumyan, A., Jankovic, M., Di Virgilio, M., Klein, I., Ansarah-Sobrinho, C., et al. (2010). Activation-induced cytidine deaminase targets DNA at sites of RNA polymerase II stalling by interaction with Spt5. Cell, 143, 122–133.
Stavnezer, J., & Schrader, C. E. (2014). IgH chain class switch recombination: Mechanism and regulation. The Journal of Immunology, 193, 5370–5378.
Pham, P., Afif, S. A., Shimoda, M., Maeda, K., Sakaguchi, N., Pedersen, L. C., et al. (2016). Structural analysis of the activation-induced deoxycytidine deaminase required in immunoglobulin diversification. DNA Repair, 43, 48–56.
Casellas, R., Nussenzweig, A., Wuerffel, R., Pelanda, R., Reichlin, A., Suh, H., et al. (1998). Ku80 is required for immunoglobulin isotype switching. The EMBO Journal, 17, 2404–2411.
Manis, J. P., Gu, Y., Lansford, R., Sonoda, E., Ferrini, R., Davidson, L., et al. (1998). Ku70 is required for late B cell development and immunoglobulin heavy chain class switching. The Journal of Experimental Medicine, 187, 2081–2089.
Reina-San-Martin, B., Difilippantonio, S., Hanitsch, L., Masilamani, R. F., Nussenzweig, A., & Nussenzweig, M. C. (2003). H2AX is required for recombination between immunoglobulin switch regions but not for intra-switch region recombination or somatic hypermutation. The Journal of Experimental Medicine, 197, 1767–1778.
Chandra, V., Bortnick, A., & Murre, C. (2015). AID targeting: Old mysteries and new challenges. Trends in Immunology, 36, 527–535.
MacDuff, D. A., Neuberger, M. S., & Harris, R. S. (2006). MDM2 can interact with the C-terminus of AID but it is inessential for antibody diversification in DT40 B cells. Molecular Immunology, 43, 1099–1108.
Chaudhuri, J., Khuong, C., & Alt, F. W. (2004). Replication protein A interacts with AID to promote deamination of somatic hypermutation targets. Nature, 430, 992–998.
de Yébenes, V. G., & Ramiro, A. R. (2006). Activation-induced deaminase: Light and dark sides. Trends in Molecular Medicine, 12, 432–439.
Basu, U., Chaudhuri, J., Alpert, C., Dutt, S., Ranganath, S., Li, G., et al. (2005). The AID antibody diversification enzyme is regulated by protein kinase A phosphorylation. Nature, 438, 508–511.
Basu, U., Wang, Y., & Alt, F. W. (2008). Evolution of phosphorylation-dependent regulation of activation-induced cytidine deaminase. Molecular Cell, 32, 285–291.
Basu, U., Franklin, A., & Alt, F. W. (2009). Post-translational regulation of activation-induced cytidine deaminase. Philosophical Transactions of the Royal Society of London B: Biological Sciences, 364, 667–673.
Budzyńska, P. M., Kyläniemi, M. K., Kallonen, T., Soikkeli, A. I., Nera, K. P., Lassila, O., et al. (2017). Bach2 regulates AID-mediated immunoglobulin gene conversion and somatic hypermutation in DT40 B cells. European Journal of Immunology. https://doi.org/10.1002/eji.201646895.
Hu, W., Begum, N. A., Mondal, S., Stanlie, A., & Honjo, T. (2015). Identification of DNA cleavage-and recombination-specific hnRNP cofactors for activation-induced cytidine deaminase. Proceedings of the National Academy of Sciences of the United States of America, 112, 5791–5796.
Dancyger, A. M., King, J. J., Quinlan, M. J., Fifield, H., Tucker, S., Saunders, H. L., et al. (2012). Differences in the enzymatic efficiency of human and bony fish AID are mediated by a single residue in the C terminus modulating single-stranded DNA binding. The FASEB Journal, 26(4), 1517–1525.
Villota-Herdoiza, D., Pila, E. A., Quiniou, S., Waldbieser, G. C., & Magor, B. G. (2013). Transcriptional regulation of teleost Aicda genes. Part 1—Suppressors of promiscuous promoters. Fish & Shellfish Immunology, 35(6), 1981–1987.
Cerutti, A. (2008). The regulation of IgA class switching. Nature Reviews Immunology, 8, 421–434.
Bekeredjian-Ding, I., & Jego, G. (2009). Toll-like receptors–sentries in the B-cell response. Immunology, 128, 311–323.
Pone, E. J., Zan, H., Zhang, J. S., Al-Qahtani, A., Xu, Z., & Casali, P. (2010). Toll-like receptors and B-cell receptors synergize to induce immunoglobulin class-switch DNA recombination: Relevance to microbial antibody responses. Critical Reviews in Immunology, 30, 1–29.
Xu, W., Santini, P. A., Matthews, A. J., Chiu, A., Plebani, A., He, B., et al. (2008). Viral double-stranded RNA triggers Ig class switching by activating upper respiratory mucosa B cells through an innate TLR3 pathway involving BAFF. The Journal of Immunology, 181, 276–287.
Medzhitov, R., Preston-Hurlburt, P., & Janeway, C. A. (1997). A human homologue of the Drosophila Toll protein signals activation of adaptive immunity. Nature, 388, 394–397.
Rauta, P. R., Samanta, M., Dash, H. R., Nayak, B., & Das, S. (2014). Toll-like receptors (TLRs) in aquatic animals: Signaling pathways, expressions and immune responses. Immunology Letters, 158, 14–24.
Akira, S., Uematsu, S., & Takeuchi, O. (2006). Pattern recognition and innate immunity. Cell, 124, 783–801.
Bombardieri, M., Kam, N. W., Brentano, F., Choi, K., Filer, A., Kyburz, D., et al. (2011). A BAFF/APRIL-dependent TLR3-stimulated pathway enhances the capacity of rheumatoid synovial fibroblasts to induce AID expression and Ig class-switching in B cells. Annals of the Rheumatic Diseases, 70, 1857–1865.
Mandler, R., Finkelman, F. D., Levine, A. D., & Snapper, C. M. (1993). IL-4 induction of IgE class switching by lipopolysaccharide-activated murine B cells occurs predominantly through sequential switching. The Journal of Immunology, 150, 407–418.
Castigli, E., Wilson, S. A., Scott, S., Dedeoglu, F., Xu, S., Lam, K. P., et al. (2005). TACI and BAFF-R mediate isotype switching in B cells. The Journal of Experimental Medicine, 201, 35–39.
He, B., Qiao, X., & Cerutti, A. (2004). CpG DNA induces IgG class switch DNA recombination by activating human B cells through an innate pathway that requires TLR9 and cooperates with IL-10. The Journal of Immunology, 173, 4479–4491.
Schneider, P. (2005). The role of APRIL and BAFF in lymphocyte activation. Current Opinion in Immunology, 17, 282–289.
Thompson, J. S., Schneider, P., Kalled, S. L., Wang, L., Lefevre, E. A., Cachero, T. G., et al. (2000). BAFF binds to the tumor necrosis factor receptor-like molecule B cell maturation antigen and is important for maintaining the peripheral B cell population. The Journal of Experimental Medicine, 192, 129–136.
Castigli, E., Scott, S., Dedeoglu, F., Bryce, P., Jabara, H., Bhan, A. K., et al. (2004). Impaired IgA class switching in APRIL-deficient mice. Proceedings of the National Academy of Sciences of the United States of America, 101, 3903–3908.
Day, E. S., Cachero, T. G., Qian, F., Sun, Y., Wen, D., Pelletier, M., et al. (2005). Selectivity of BAFF/BLyS and APRIL for binding to the TNF family receptors BAFFR/BR3 and BCMA. Biochemistry, 44, 1919–1931.
Barreto, V. M., Ramiro, A. R., & Nussenzweig, M. C. (2005). Activation-induced deaminase: Controversies and open questions. Trends in Immunology, 26, 90–96.
Moris, A., Murray, S., & Cardinaud, S. (2014). AID and APOBECs span the gap between innate and adaptive immunity. Frontiers in Microbiology, 5, 1–13.
Abdouni, H., King, J. J., Suliman, M., Quinlan, M., Fifield, H., & Larijani, M. (2013). Zebrafish AID is capable of deaminating methylated deoxycytidines. Nucleic Acids Research, 41, 5457–5468.
Harris, R. S., Sheehy, A. M., Craig, H. M., Malim, M. H., & Neuberger, M. S. (2003). DNA deamination: Not just a trigger for antibody diversification but also a mechanism for defense against retroviruses. Nature Immunology, 4, 641–643.
Okazaki, I. M., Kotani, A., & Honjo, T. (2007). Role of AID in tumorigenesis. Advances in Immunology, 94, 245–273.
Liu, M., & Schatz, D. G. (2009). Balancing AID and DNA repair during somatic hypermutation. Trends in Immunology, 30, 173–181.
Robbiani, D. F., Bunting, S., Feldhahn, N., Bothmer, A., Camps, J., Deroubaix, S., et al. (2009). AID produces DNA double-strand breaks in non-Ig genes and mature B cell lymphomas with reciprocal chromosome translocations. Molecular Cell, 36, 631–641.
Guselnikov, S. V., Ramanayake, T., Erilova, A. Y., Mechetina, L. V., Najakshin, A. M., Robert, J., et al. (2008). The Xenopus FcR family demonstrates continually high diversification of paired receptors in vertebrate evolution. BMC Evolutionary Biology, 8, 148.
Acknowledgements
Authors wish to thank the National Agricultural Science Fund (NASF/BS-4003) of the Indian Council of Agricultural Research (ICAR), Government of India, for providing the research grant.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Patel, B., Banerjee, R., Samanta, M. et al. Diversity of Immunoglobulin (Ig) Isotypes and the Role of Activation-Induced Cytidine Deaminase (AID) in Fish. Mol Biotechnol 60, 435–453 (2018). https://doi.org/10.1007/s12033-018-0081-8
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-018-0081-8