Abstract
MDM4 is an important p53-negative regulator, consequently, it is involved in cell proliferation, DNA repair, and apoptosis regulation. MDM4 overexpression and amplification are described to lead to cancer formation, metastasis, and poor disease prognosis. Several MDM4 SNPs are in non-coding regions, and some affect the MDM4 regulation by disrupting the micro RNA binding site in 3'UTR (untranslated region). Here, we gathered several association studies with different MDM4 SNPs and populations to understand the relationship between its SNPs and solid tumor risk. Many studies failed to replicate their results regarding different populations, cancer types, and risk genotypes, leading to conflicting conclusions. We suggested that distinct haplotype patterns in different populations might affect the association between MDM4 SNPs and cancer risk. Thus, we propose to investigate some linkage SNPs in specific haplotypes to provide informative MDM4 markers for association studies with cancer.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
MDM4 (mouse double minute 4), also known as MDMX and HDMX, is in 1q32 loci of the human genome [1] which encodes a protein capable of forming a heterodimer with MDM2 (mouse double minute 2) [2]. MDM2 is an E3 ubiquitin ligase that binds to the guardian of the genome p53 promoting its proteasomal degradation. This activity can be enhanced by MDM4-MDM2 complex formation [3]. MDM2 is a paralogous gene of MDM4, and both proteins present 33% identity in humans. These genes emerged after the ancestral MDM duplication in primitive vertebrates. The ancestral protein was more MDM2-like, while MDM4 evolved to lose its ubiquitin ligase activity [4]. Under normal conditions, these two proteins have autoregulatory feedback, a mechanism in which p53 can activate MDM2 transcription [5], and the MDM2-MDM4 heterodimer activity regulation is responsible for maintaining low levels of p53 when this protein is not necessary, preserving intracellular homeostasis [6]. During cellular stress, MDM2 promotes self-degradation and MDM4 degradation, leaving p53 free to activate transcription factors, promoting DNA repair, cell cycle arrest, and apoptosis. Thus, the p53-MDM2-MDM4 regulatory axis alterations are mainly known to promote cancer [7].
The central regulator of the p53 pathway is MDM2; however, MDM4 participates in the p53 level regulation in different ways (Fig. 1). The p53 inhibition occurs by direct MDM4-p53 protein binding (Fig. 1A), which abolishes the tumor suppression from its function, and blocks its transcriptional activity [8, 9]. MDM4 stabilizes MDM2, enhancing p53 ubiquitination and its proteasomal degradation by forming the MDM2-MDM4 heterodimer (Fig. 1B) [10, 11]. The MDM2-MDM4 heterodimer can promote p53 synthesis during genotoxic stress acting as IRES (Internal Ribosome Entry Site) transacting factors to activate p53 mRNA translation (Fig. 1C) [12]. Furthermore, a contrasting activity of MDM4 has been described, which requires further investigation, as a facilitator of the interaction between p53 and BCL2 (B-Cell Leukemia/Lymphoma 2) after genotoxic stress by promoting the displacement of p53 phosphorylated in Ser46 to mitochondria, releasing cytochrome c in the cytoplasm, and signaling apoptosis (Fig. 1D) [13]. In a p53-independent manner, MDM2-MDM4 heterodimer acts as a cell cycle promoter maintaining high levels of transcriptional factors E2F1, E2F3, and p73 when wild-type p53 is absent (Fig. 1E) [14]. MDM4, alone or in cooperation with MDM2, can promote the transition from G1 to the early S phase, binding directly to p21 to mediate its proteasomal degradation (Fig. 1F) [15]. These MDM4 activities can contribute to cancer development, and some MDM4 transcripts responsible for encoding proteins are related to different cancer types [16].
MDM4 interaction with p53 and other proteins of cell cycle regulation. MDM4 is involved in some steps in the p53 pathway and other cell cycle control pathways. MDM4 can act alone or in a heterodimer with MDM2. MDM4 can directly bind and inhibit p53 activity A and enhances the p53 proteasomal degradation forming an MDM4-MDM2 heterodimer B In genotoxic stress conditions, MDM4 can act as an IRES transacting factor to increase p53 mRNA C and facilitates the interaction between p53 and BCL2 in mitochondria to promote apoptosis D The MDM4-MDM2 heterodimer interacts with the E2F protein family E and MDM4, alone or in cooperation with MDM2, mediates p21 proteasomal degradation to induce cell cycle progression F Created with BioRender.com
Tumor development depends on pathways frequently deregulated in cancer which often play essential roles in promoting aberrant splicing [17]. MDM4 protein diversity is created by alternative splicing (Fig. 2), and the most described isoforms in cancer studies are MDM4-FL (full-length) and MDM4-S (short form). MDM4-FL is more stable than the latter [18], and it has the complete MDM4 protein structure with all four conserved regions: the N-terminal portion with p53 binding domain, the acid domain, zinc finger, and RING finger domain in the C-terminal portion (Fig. 2A) [2]. The skipping of exon 6 results in a frameshift and a premature stop codon that encodes a short carboxy-truncated MDM4 protein (MDM4-S) (Fig. 2B) containing the p53 binding domain and a few residues in the C-terminal portion [19]. Some authors described MDM4-S as presenting a higher affinity with p53 when compared with MDM4-FL [20], and MDM4-S presents higher expression levels than MDM4-FL in tumor cells [21,22,23]. MDM4-S expression is related to more aggressive p53 mutated cancer, and the MDM4-S/MDM4-FL ratio could be useful as a potential prognostic biomarker [22, 24, 25]. Some MDM4 isoforms are also related to cancer: MDM4-211(Fig. 2C) mRNA is highly expressed in thyroid tumor cell line (ARO) [26], and it is frequently expressed in papillary thyroid carcinoma samples, while MDM4-FL is downregulated [27]; MDM4-A (Fig. 2D) mRNA is significantly more expressed in human melanoma samples than MDM4-FL, and its expression is correlated to poor survival [28]; MDM4-A and MDM4-G (Fig. 2E) were first isolated from an ovarian cancer cell line (C33A), and they both might affect p53 activity. MDM4-G stabilizes MDM2, while MDM4-A inhibits p53 [29]. MDM4-ALT1 (Fig. 2F) and MDM4-ALT2 (Fig. 2G) expressions were described by Chandler et al. [30] in tumor breast cell lines (MCF-7), and MDM4-ALT2 was associated with high metastatic risk of rhabdomyosarcoma [31]. However, MDM4 isoforms need further investigation to evaluate their roles in carcinogenesis.
MDM4 isoforms related to cancer. The isoforms transcripts and proteins (represented by letters A to G) are named by Ensembl codes (ENST and ENSP, respectively). Each conserved protein domain and its respective transcript exons have the same color, i.e., p53 binding domain (orange), acid domain (red), zinc finger (Zn) (blue), RING finger domain (purple) and missing domains (gray). The MDM4 UTR portions are in white color. The MDM4 isoform sizes is shown by amino acids (aa) number. The MDM4-FL is the full-length isoform with 490 aa with all exons and conserved domains A MDM4-S isoform has only the p53 binding domain, and its transcript skips exon 6, creating a frameshift and a premature stop codon in exon 7 B MDM4-211 isoform presents only the RING finger domain after its exons 3 to 10 are removed C MDM4-A isoform has an incomplete acid domain since its transcript skips exon 9 D MDM4-G isoform has no p53 binding domain, and its transcript skips exons 3 to 5 E MDM4-Alt1 isoform has only the p53 binding domain since its transcript skips exons 6 to 9, with a premature stop codon in exon 10 F MDM4-Alt2 isoform has only the zinc finger and RING finger domains after its exons 4 to 9 are removed G
Alternative splicing is also a fundamental process to regulate gene expression. However, it is also known that MDM4 is amplified or overexpressed in many human cancers, such as glioma, lung, colon, breast, retinoblastoma, prostate, squamous cell carcinoma of head and neck, gastric, and acute myeloid leukemia (AML) [32,33,34,35,36,37,38,39,40]. Previously, it was observed that spontaneous tumorigenesis in transgenic mice with wild-type p53 was induced by MDM4 overexpression [41]. MDM4 and MDM2 overexpression were correlated with an increase in circulating tumor cells in triple-negative breast cancers in a mouse model [42], while p53 mutations were correlated with MDM4 amplification and MDM2 overexpression in primary breast tumors [43]. Interestingly, MDM4 expression can be controlled by micro RNAs (miRNAs). MiRNAs are non-coding RNAs with approximately 30 base pairs that regulate protein levels post-transcriptionally [44]. Some miRNAs can bind in MDM4 mRNA 3'UTR, resulting in low levels of MDM4 and p53 function release [45,46,47]. Some miRNAs have an affinity with MDM4 mRNA affected by SNPs (Single Nucleotide Polymorphisms) located on 3'UTR [48,49,50,51]. Furthermore, MDM4 has an assortment of SNPs that can be used as biomarkers for different types of tumors since some miRSNPs interfere with gene expression and may contribute to clinical outcomes affecting cancer susceptibility [52, 53]. Herein, we reviewed the association studies between MDM4 SNPs and cancer, focusing on the controversy of those SNPs previously associated with cancer risk, solid tumors prognosis, and cancer survival. We propose to investigate some linkage SNPs in specific haplotypes that might help future association studies with MDM4 and cancer.
MDM4 SNPs associated with cancer risk
The MDM4 roles in tumorigenesis and cancer aggressiveness have been studied for the last few years, and some findings from MDM4 SNPs association studies with cancer in different populations can create insights in terms of understanding the MDM4 allelic establishment of cancer susceptibility. Interestingly, all MDM4 SNPs observed in these studies are in non-coding regions, except the rs116197192, located in exon 7 (Supplementary Table 1). The most studied MDM4 SNP, the rs4245739 (C > A), is in 3'UTR. It was predicted to change the affinity of three miRNAs: miR191-5p, miR-887, and miR-3669 [54]. This miRSNP disrupts miRNA binding sites and reduces the MDM4 oncogenic effects. Indeed, the allele rs4245739-C significantly affects the binding of miR-191-5p and miR-887 and decreases MDM4 expression in heterozygous genotype (A/C), while no effect was observed in homozygous genotype (A/A) in different cancer cell lines [49]. Regarding that, the localization of SNPs in distinct gene regions as promoter, exons, introns, and UTRs modulates gene expression in different ways, for example, by altering the splicing machinery and/ or long non-coding RNA (lncRNAs) binding sites and affects mRNA expression, which may play a functional role in cancer susceptibility [55]. However, most studies about MDM4 SNPs and their association with cancer do not provide information concerning their impact on gene expression and its alternative splicing. In addition, the genetic background of the analyzed population and the cancer characteristics might explain the conflicting results between functional and association studies. The rs4245739 A/A genotype was associated in the Norwegian population with the breast cancer increased risk, while the same genotype was associated with the ovarian cancer reduction risk in this population (Supplementary Table 1) [56, 57].
However, most studies described the rs4245739 A/A genotype or rs4245739-A allele as positively associated with many cancer types (ovarian, esophageal, breast, lung, oropharynx, and colorectal) in a diversity of European and Asian populations [46,47,48, 58,59,60,61,62,63,64]. Recently, Chen et al. [65] and Wang et al. [66] confirmed by meta-analysis the association between rs4245739-C and the reduction of cancer risk in the Asian ancestry population. On the other hand, few studies demonstrated that the rs4245739 C/C genotype is associated with increased cancer risk in different populations [67], but most of them correlated this genotype with negative estrogen receptor (ER-) breast cancer subtype in other populations [68, 69].
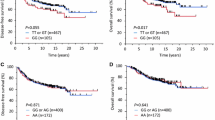
Some studies evaluated whether the MDM4 SNPs influence clinicopathological cancer aspects, such as overall survival (OS), disease-free survival (DFS), disease recurrence, and tumor aggressiveness. Specifically, the rs4245739 C/A and C/C genotypes in NSCLC (Non-Small Cell Lung Cancer) and colorectal cancer were significantly associated with better OS [46, 63]. On the other hand, the ovarian carcinoma patients with rs4245739 A/A genotype presented an increased death risk and a significant reduction in their OS compared with C allele carrier patients [48]. After chemoradiation in the HPV16 + patients with SCCOP (Squamous Cell Carcinoma of the Oropharynx), it was observed that patients with rs4245739 C/C and A/C genotypes had the risk of overall death, death due to disease, and disease recurrence reduced in comparison to the patients with rs4245739 A/A [62]. This MDM4 polymorphism is extensively studied since its 3'UTR is functionally related to altering target recognition of miRNAs by disrupting sequence complementarity [48]. As a regulatory region, the 3'UTR is indispensable for regular gene expression, and the MDM4 rs4245739 polymorphism could affect its protein translation. As MDM4 forms an MDM2 complex that promotes p53 degradation, resulting in cell cycle deregulation and creating a carcinogenesis environment, this MDM4 SNP has a suggestive relevance in this process [49].
In addition to this SNP, other associated SNPs are located in MDM4 UTRs but are no longer functionally analyzed. The genotype A/A of 3'UTR rs11801299 (G > A) was associated with lower OS, higher tumor invasion, and poor tumor differentiation in retinoblastoma Chinese patients in comparison to G/G genotype [70]. The G/A genotype was associated with gastric cancer susceptibility in the Chinese population [38]. Different findings in the American non-Hispanic population association studies reinforce the importance of population background genetic importance. In this population, the SCCOP patients with rs11801299 G/G genotype had worse DFS in comparison to patients with rs11801299 A/A or A/G genotypes as well as with MDM4 3'UTR rs10900598 (G > T/ C > A) T/T + T/G genotypes in comparison with G/G genotype [71]. The rs10900598 C/C genotype was associated with reduced NSCLC risk in the Chinese population, while a genotype synergic effect was demonstrated between rs10900598 C/C and rs4245739 C/C, both polymorphisms located in MDM4 3'UTR. The NSCLC patients with these genotypes had better OS than those without these genotypes’ combinations [46].
The functionally less known and analyzed SNPs in association studies were MDM4 5'UTR rs10900594 (G > C) and 3'UTR rs12039454 (T > C). These SNPs were analyzed together with the intronic MDM4 rs2369244 (G > C) in one single association study with breast cancer risk in the Ashkenazi Jewish population (Supplementary Table 1) [52]. Additional investigations will be required to determine the biological relevance of these SNPs. However, it has been demonstrated that the promoter region SNPs affect gene expression by altering promoter activity, transcription-factor binding, DNA methylation, and histone modification [72,73,74]. Indeed, MDM4 polymorphisms located in the 5'UTR region, such as rs10900594 and rs4252668 (T > C), might play an important role in its transcription that interferes with MDM4 oncogenic activity. In fact, Reincke et al. [75] suggested that the rs4252668 variant may have a regulatory function for being located at 5'UTR in MDM4. According to the SNP Nexus portal, rs4252668 is in a CpG island (CGI) with 69.3% C/G content and 358 base pairs in length [76]. DNA methylation occurs primarily in the CGIs of the promoter region. Therefore, SNPs in the promoter region, such as MDM4 rs4252668-T, might alter DNA methylation status, histone acetylation, chromatin modification, and gene silencing. Thus, epigenetic modifications are associated with disease and cancer development [77, 78]. In a study developed by Oliveira Reis et al. [79], the MDM4 polymorphisms rs4252668-T and rs116197192-G were associated with retinoblastoma increased risk in the Brazilian population. Interestingly, all patients had the rs4252668 T/T genotype, while the C/C genotype was at a higher frequency in the control group.
The MDM4 rs116197192 (A > G) in exon 7 gives rise to D153G substitution located within a predicted casein kinase II (CK2) site. Both protein kinases, CK1 and CK2, participate in various cellular processes, including DNA repair and cell cycle control, and phosphorylate Ser or Thr residues [75]. Phosphorylation on MDM4 Ser289 by CK1α increases its association with p53 with a profound impact on its p53 inhibition [80], but there is no report of MDM4 phosphorylation by CK2 at Thr150 or other sites [81]. According to TarBase v.8 [82], rs116197192 and rs4252668 are in miRNAs binding sites, and these interactions promote the downregulation of MDM4 in different cell lines and tissues. The allele change of these polymorphisms might increase cancer risk by modifying the regulation site, upregulating MDM4 expression, and inhibiting p53 activity. However, the SIFT portal [83] shows a prediction that the effect of change A to G in rs116197192 (D153G) is tolerated in the MDM4 function.
The polymorphism MDM4 rs4252707 (G > A) in intron 8 might modulate mRNA splicing activity and impact the production of the splice variants without exon nine as MDM4-A isoform. The rs4252707-A was associated with increased risk for non-glioblastoma glioma in the European and Chinese populations [84, 85]. The rs1563828 (A > G/ T > C) is an intronic variant in which T/T and A/A genotypes were described as associated with earlier ovarian tumor onset [52], higher breast tumor aggressiveness [53], and 1.3-fold increased risk of lymphatic vessel infiltration in breast cancer patients compared with C/C genotype (Supplementary Table 1) [61]. In contrast, Morvan et al. [86] described the rs1563828 A/A breast cancer patients with higher PFS (Progression-Free Survival) compared with G/G and A/G patients after chemotherapy.
The MDM4 rs1380576 (G > C) located in intron 1 is one of the MDM4 SNPs that usually is analyzed in association studies with cancer (Supplementary Table 1). The genotype C/C was associated with prostate cancer increased aggressiveness in the German population [35] and positive estrogen receptor (ER +) breast cancer increased risk in the Lithuanian population [64]. A protective effect was observed with the rs1380576 G/G genotype, which was associated with a significant breast cancer decrease in the Iranian population [87], lower retinoblastoma tumor aggressiveness [70], and gastric cancer reduction in the Chinese population [38]. The HapMap datasets from Asian and Chinese populations associated this genotype with lower MDM4 mRNA expression [38]. Interestingly, the rs1380576 G/G genotype was associated with many types of cancer risk reduction in the Asian population [65], and the C/C genotype was associated with reduced cancer risk compared with the G/G + G/C genotypes in the Asian ancestry population (Supplementary Table 1) [66]. Considering that high MDM4 expression may result in low levels of p53 and the activation of different cell cycle-promoting proteins [14, 42], this associated polymorphism could interfere with the reduction of MDM4 level and, consequently, it might contribute to reducing cancer development.
The MDM4 expression might be affected by quantitative trait loci alleles (eQTL). A particular MDM4 downstream intergenic variant in 1q32.1, the rs12133735-G allele (G > T), was related to a lower MDM4 expression; however, it was associated with increased risk for lung, and oropharyngeal cancer in the European population according to GWAS meta-analysis study [88]. The additional information extracted from the GTEx portal (ENSG00000198625.12) listed the aforementioned MDM4 SNPs associated with cancer risk and a significant MDM4 eQTL effect. The rs4245739, rs1380576, rs1563828, rs10900598, rs10900594, rs2369244, and rs12039454 are related to a negative MDM4 eQTL effect in several tissues, except rs11801299, which effect was seen only in blood. No eQTL impact was found for rs4252668 and rs116197192. According to NESDA NTR Condicional eQTL Catalog, the allelic effects in the blood of the top associated SNPs (rs4245739-C, rs2369244-G, rs10900594-G, and rs1380576-G) have significant negative MDM4 eQTL. In contrast, rs12039454-T, rs10900598-G, rs1563828-A, and rs11801299-A have a positive MDM4 eQTL effect in European ancestry individuals [89]. This data could explain why some alleles are associated with increased cancer risk as they can increase MDM4 expression and downregulate the p53 pathway.
Lack of association between MDM4 SNPs and cancer risk
The association studies represent an essential step in defining disease-mediating genetic variants. However, some studies failed to show an association between MDM4 SNPs and cancer risk or survival as a consequence of population-specific genotype frequencies, differences due to genetic background, and solid tumor heterogeneity (Supplementary Table 2) [57, 90,91,92]. Despite the potential for improving the ability to detect susceptibility/protection SNPs, the inconsistency of association data is still a feature of association studies [93,94,95,96,97]. One of the reasons for this problem might be the small sample size, as observed in some association studies herein analyzed [46, 85, 91, 92, 98]. The strategy to explore a large group of MDM4 SNPs in a genome regional association study that has never been studied before in the Chinese population was useful in informing allele and haplotype descriptions, despite the absence of significant results observed for seven MDM4 SNPs (Supplementary Table 2) [46, 85].
The cancer risk associations detected for some MDM4 SNPs herein studied were not reproducible in other association studies with different cancer types and populations (Supplementary Tables 1 and 2). The ethnic diversity leading to population stratification can bear significantly on the power of an association study. In the same vein, the association studies between MDM4 rs4245739 (C > A) and lung, colon, and prostate cancer risks showed different conclusions in the Norwegian [56] population in comparison to association studies with Asian ancestry populations.
A further explanation source of nonreplicable results might be the small effect of many genetic factors contributing to cancer formation. For instance, the impact of HPV infection, one of the environmental risk factors for oropharyngeal, head, and neck cancer development, could be stronger than the exclusive presence of some MDM4 genotypes. In fact, Wang et al. [92] showed a significant association between TP53, MDM2, and MDM4 SNPs and oral cancer risk only for HPV16 + patients when the risk genotypes of these three genes are together. No association was observed for each MDM4 SNPs (rs11801299, rs10900598, rs1380576) in HPV16 + or HPV16− patients. Thus, the relative risk of only MDM4 genetic variants might be small [99].
MDM4 SNPs in linkage disequilibrium and haplotypes
The characterization of multiple SNPs (marker alleles) on the same chromosome, which tends to be inherited together, is termed haplotype. The association analysis based on haplotypes may provide more power and accuracy in disease gene mapping than those based on single markers [100]. The linkage disequilibrium (LD) analysis based on underlying haplotypes varies significantly in different populations and reflects the pattern of inheritance over evolution [101]. Based on the MDM4 SNPs single data found in the association studies with cancer risk listed in Supplementary Tables 1 and 2, we inferred the haplotype configurations of 18 total SNPs in three different populations: Euro-American (CEU), Han Chinese (CHN), and Afro-American (ASW) using the linkage disequilibrium information from LD link tools [102].
Accordingly, we observed 12 different MDM4 haplotypes over three populations (Fig. 3A). The phylogenetic relationships within 12 MDM4 haplotypes were performed using a set of haplotypes dissimilarities to construct a hierarchical cluster (Fig. 3B). The most similar clusters were built by Ward’s minimum variance method and the resulted phylogenetic haplotype tree demonstrates 3 major clusters (Fig. 3B). Specifically, haplotype 2 differs from 11 to 12 ancestral haplotypes by 11 MDM4 SNPs alleles (Fig. 4). Haplotype 12 (5.7%) is unique to the ASW population, and it is represented in the smallest cluster (Fig. 3B) in which the haplotypes 1 and 4 diverge by rs12133735 allele (Fig. 4). The haplotype 1 is the most frequent in the CEU and CHN population and, the third most common in the ASW population, while the haplotype 4 is observed only in the CEU population (Fig. 3A). Interestingly, the CHN population showed more haplotype combinations than CEU and ASW populations, since haplotypes 9, 10, and 11 are exclusive to the CHN population and these haplotypes are represented together with the haplotype 3 in the second haplotype cluster (Fig. 3B). These two clusters separated the CHN and the ASW populations according to the MDM4 SNPs alleles. One of the differences between these two clusters (ASW x CHN) is the eQTL MDM4 SNP rs11801299-A allele, which was previously related to positive MDM4 expression [89] and herein it was found as a cancer risk in the CHN population.
Haplotypes frequencies in each population: CEU population (Utah residents with Northern and Western European ancestry), the CHN (Han Chinese in Beijing and Southern with East Asian ancestry), and the ASW population (African Ancestry in the Southwest United States) A and the dendrogram grouped by Euclidian distance and Ward.D linkage method as Hierarchical Clustering B using the packages dist() and hclust() in R programming language. The Euclidian distance (Height) organizes the haplotypes according to similarity relations to obtain a classification, while Ward’s method creates groups minimizing the variation within clusters. Haplotypes and LD correlation analysis were generated by the LD Link tools [102]
MDM4 SNPs and haplotypes in different populations. The MDM4 haplotypes frequencies were calculated for the CEU population (Utah residents with Northern and Western European ancestry), the CHN (Han Chinese in Beijing and Southern with East Asian ancestry), and the ASW population (African Ancestry in the Southwest United States). Haplotypes and LD correlation analysis were generated by the LD Link tools [102]
The third cluster is composed of more haplotypes (haplotypes 2, 8, 6, 5, and 7) with distinct frequencies along the three populations analyzed (Fig. 3A, B). The most frequent haplotype in this cluster is haplotype 7, with 32.8% in the ASW population and 16.4% in the CHN population, while absent in the CEU population. Interestingly, haplotype 7 is the only one that has the MDM4 SNP rs3014610-T allele (A > T) (Fig. 4). According to ENCODE portal [103], this SNP is related to a region with post-translationally modified histones (H4K20me1, H3K36me3) in tumor cell lines. Considering its high frequency in the ASW and CHN populations (Fig. 3A), haplotype 7 could be protective against cancer development. However, the rs3014610 intronic variant needs further investigation as it is not associated with cancer risk in the Chinese population (Supplementary Table 2). Haplotype 6 is unique to the CEU population, and it diverges from haplotypes 5, 7, and 8 by MDM4 SNPs rs4252668, rs16853949, rs10900594, and rs1380576 alleles. Haplotype 6 has rs4252668-C, and haplotype 8 has rs16853949-A alleles (Fig. 4), which are uncommon in the three populations (Supplementary Table 6) and might be related to conflicted association studies results herein observed (Supplementary Table 2). Additionally, the rs4252668 is in the transcriptional factors binding site and the rs16853949 is in regions with post-translationally modified histones in tumor cell lines, and both might impact MDM4 expression (ENCODE portal [103]). The last two SNPs are related to histone epigenetic modifications and eQTL effects that might affect cancer susceptibility directly or indirectly. Haplotype 2 is very common in the CEU and ASW populations and less frequent in the CHN population (Fig. 3A). The difference between haplotype 2 and the other haplotypes is the presence of MDM4 SNP rs4245739-C allele (Fig. 4), which functionally disrupts miRNA binding site, and has a significant negative MDM4 eQTL effect. Haplotype 2 is less frequent in the CHN population, and its distribution might affect the association studies in populations with different genetic backgrounds.
The haplotype‐based association methods are generally more powerful than methods based on single markers since it exploits LD information from multiple markers. D′ and r2 are LD measurements based on a pair of markers. LD patterns and haplotype frequencies diverge significantly in different populations. Indeed, we observed in the CEU and the CHN populations (Supplementary Tables 3 and 4, respectively) a strong linkage disequilibrium between MDM4 rs4252707 and rs11801299 (D′ = 1) and low LD in the ASW population (Supplementary Table 5). Haplotype 12 might result from this weak LD observed in the ASW population. Herein, we listed studies with no association between rs11801299 and cancer risks in the American non-Hispanic population (Supplementary Table 2).
The MDM4 rs4245739 is in strong LD with rs2169137 in the CEU and CHN populations. In the ASW population, these SNPs are highly correlated (r2 = 0.924). Interestingly, Haplotype 2 is characterized by the rs4245739-C and rs2169147-G alleles that were found in low frequency in the CHN population compared to the two other populations (Fig. 3A). The MDM4 SNP rs10900594 is strongly linked with rs1380576 in the CEU population and has a high correlation in the CHN and the ASW populations (r2 = 0.989 and r2 = 0.92, respectively). Curiously, both protective alleles (rs10900594-G and rs1380576-G) are in the same haplotype combination with the susceptibility alleles (rs1563828-A and rs12039454-T) located in the no coding region (intron 10 and 3'UTR, respectively), forming two contrasting blocks with a significant MDM4 eQTL SNPs (Fig. 4). The two first alleles are related to a negative eQTL effect and, the two last alleles have a positive MDM4 eQTL effect. It seems that along the length of MDM4, there are different ways to control gene expression since rs1380576 and rs12039454 SNPs are located at binding sites for various transcription factors. Additionally, the posttranslational modifications of histones alter chromatin structure, impacting transcriptional factors binding and increasing the oncogenic process. The histone H3 tail lysine and arginine residues suffer covalent modifications, like acetylation and methylation. Both modifications might occur on these four SNPs described above: H3K27me3, H3K4me1, H3K36me3, and H3K27ac (ENCODE portal [103]). These epigenetic modifications could activate an oncogene, such as MDM4, thus providing oncogenic reprogramming in tumor cells.
Conclusions
The results indicated that the MDM4 association studies with solid tumors suffer from the influence of the differences in population genetic structure. Here the single SNPs listed were useful in constructing the haplotype structures. The level of linkage between some eQTL alleles and SNPs was also useful in exploiting the MDM4 polymorphism in cancer. Our results suggest some additional haplotypes to refine the association studies results.
Data availability
All data analyzed during this study are included in this published article and its supplementary information file.
References
Shvarts A, Bazuine M, Dekker P, Ramos YFM, Steegenga WT, Merckx G, van Ham RCA, vander HouvenOordt W, vander Eb A, Jochemsen AG. Isolation and identification of the human homolog of a new p53-binding protein. Mdmx Genom. 1997;43(1):34–42. https://doi.org/10.1006/geno.1997.4775.
Shvarts A, Steegenga W, Riteco N, van Laar T, Dekker P, Bazuine M, van Ham R, vander HouvenOordt W, vander Eb A, Jochemsen A. MDMX a novel p53-binding protein with some functional properties of MDM2. EMBO J. 1996. https://doi.org/10.1002/j.1460-2075.1996.tb00919.x.
Sharp DA, Kratowicz SA, Sank MJ, George DL. Stabilization of the MDM2 oncoprotein by Interaction with the structurally related MDMX protein. J Biol Chem. 1999;274:38189–96. https://doi.org/10.1074/jbc.274.53.38189.
Tan BX, Liew HP, Chua JS, Ghadessy FJ, Tan YS, Lane DP, Coffill CR. Anatomy of Mdm2 and Mdm4 in evolution. J Mol Cell Biol. 2017;9:3–15. https://doi.org/10.1093/jmcb/mjx002.
Picksley SM, Lane DP. What the papers say: the p53-mdm2 autoregulatory feedback loop: a paradigm for the regulation of growth control by p53? BioEssays. 1993;15:689–90. https://doi.org/10.1002/bies.950151008.
Toledo F, Krummel KA, Lee CJ, Liu CW, Rodewald LW, Tang M, Wahl GM. A mouse p53 mutant lacking the proline-rich domain rescues Mdm4 deficiency and provides insight into the Mdm2-Mdm4-p53 regulatory network. Cancer Cell. 2006;9:273–85. https://doi.org/10.1016/j.ccr.2006.03.014.
Yu DH, Xu ZY, Mo S, Yuan L, Cheng XD, Qin JJ. Targeting MDMX for cancer therapy: rationale, strategies, and challenges. Front Oncol. 2020;10:1389. https://doi.org/10.3389/fonc.2020.01389.
Shadfan M, Lopez-Pajares V, Yuan Z-M. MDM2 and MDMX: alone and together in regulation of p53. Transl Cancer Res. 2012;1:88–9.
Francoz S, Froment P, Bogaerts S, de Clercq S, Maetens M, Doumont G, Bellefroid E, Marine J-C. Mdm4 and Mdm2 cooperate to inhibit p53 activity in proliferating and quiescent cells in vivo. Proc Natl Acad Sci. 2006;103:3232–7. https://doi.org/10.1073/pnas.0508476103.
Okamoto K, Taya Y, Nakagama H. Mdmx enhances p53 ubiquitination by altering the substrate preference of the Mdm2 ubiquitin ligase. FEBS Lett. 2009;583:2710–4. https://doi.org/10.1016/j.febslet.2009.07.021.
Wang X, Jiang X. Mdm2 and MdmX partner to regulate p53. FEBS Lett. 2012;586:1390–6. https://doi.org/10.1016/j.febslet.2012.02.049.
Malbert-Colas L, Ponnuswamy A, Olivares-Illana V, Tournillon AS, Naski N, Fåhraeus R. HDMX folds the nascent p53 mRNA following activation by the ATM kinase. Mol Cell. 2014;54:500–11. https://doi.org/10.1016/j.molcel.2014.02.035.
Mancini F, Moretti F. Mitochondrial MDM4 (MDMX): an unpredicted role in the p53-mediated intrinsic apoptotic pathway. Cell Cycle. 2009;8:3854–9. https://doi.org/10.4161/cc.8.23.10089.
Klein AM, Biderman L, Tong D, Alaghebandan B, Plumber SA, Mueller HS, van Vlimmeren A, Katz C, Prives C, Designed CP, Performed CK. MDM2, MDMX, and p73 regulate cell-cycle progression in the absence of wild-type p53. Proc Natl Acad Sci. 2021;18:2102420118. https://doi.org/10.1073/pnas.2102420118.
Jin Y, Zeng SX, Sun X-X, Lee H, Blattner C, Xiao Z, Lu H. MDMX promotes proteasomal turnover of p21 at G 1 and early S phases independently of, but in cooperation with, MDM2. Mol Cell Biol. 2008;28:1218–29. https://doi.org/10.1128/MCB.01198-07.
Wu J, Lu G, Wang X. MDM4 alternative splicing and implication in MDM4 targeted cancer therapies. Am J Cancer Res. 2021;11:5864–80.
David CJ, Manley JL. Alternative pre-mRNA splicing regulation in cancer pathways and programs unhinged. Genes Dev. 2010;24:2343–64. https://doi.org/10.1101/gad.1973010.
Bezzi M, Teo SX, Muller J, Mok WC, Sahu SK, Vardy LA, Bonday ZQ, Guccione E. Regulation of constitutive and alternative splicing by PRMT5 reveals a role for Mdm4 pre-mRNA in sensing defects in the spliceosomal machinery. Genes Dev. 2013;27:1903–16. https://doi.org/10.1101/gad.219899.113.
Rallapalli R, Strachan G, Cho B, Mercer WE, Hall DJ. A Novel MDMX transcript expressed in a variety of transformed cell lines encodes a truncated protein with potent p53 repressive activity. J Biol Chem. 1999;274(12):8299–308. https://doi.org/10.1074/jbc.274.12.8299.
Rallapalli R, Strachan G, Tuan RS, Hall DJ. Identification of a domain within MDMX-S that is responsible for its high affinity interaction with p53 and high-level expression in mammalian cells. J Cell Biochem. 2003;89:563–75. https://doi.org/10.1002/jcb.10535.
Bartel F, Schulz J, Böhnke A, Blümke K, Kappler M, Bache M, Schmidt H, Würl P, Taubert H, Hauptmann S. Significance of HDMX-S (or MDM4) mRNA splice variant overexpression and HDMX gene amplification on primary soft tissue sarcoma prognosis. Int J Cancer. 2005;117:469–75. https://doi.org/10.1002/ijc.21206.
Liu L, Fan L, Fang C, Zou ZJ, Yang S, Zhang LN, Li JY, Xu W. S-MDM4 mRNA overexpression indicates a poor prognosis and marks a potential therapeutic target in chronic lymphocytic leukemia. Cancer Sci. 2012;103:2056–63. https://doi.org/10.1111/cas.12008.
Dewaele M, Tabaglio T, Willekens K, Bezzi M, Teo SX, Low DHP, Koh CM, Rambow F, Fiers M, Rogiers A, Radaelli E, Al-Haddawi M, Tan SY, et al. Antisense oligonucleotide-mediated MDM4 exon 6 skipping impairs tumor growth. J Clin Investig. 2016;126:68–84. https://doi.org/10.1172/JCI82534.
Bardot B, Bouarich-Bourimi R, Leemput J, Lejour V, Hamon A, Plancke L, Jochemsen AG, Simeonova I, Fang M, Toledo F. Mice engineered for an obligatory Mdm4 exon skipping express higher levels of the Mdm4-S isoform but exhibit increased p53 activity. Oncogene. 2015;34:2943–8. https://doi.org/10.1038/onc.2014.230.
Lenos K, Grawenda AM, Lodder K, Kuijjer ML, Teunisse AFAS, Repapi E, Grochola LF, Bartel F, Hogendoorn PCW, Wuerl P, Taubert H, Cleton-Jansen A-M, Bond GL, et al. Alternate splicing of the p53 inhibitor HDMX offers a superior prognostic biomarker than p53 mutation in human cancer. Can Res. 2012;72:4074–84. https://doi.org/10.1158/0008-5472.CAN-12-0215.
Giglio S, Mancini F, Gentiletti F, Sparaco G, Felicioni L, Barassi F, Martella C, Prodosmo A, Iacovelli S, Buttitta F, Farsetti A, Soddu S, Marchetti A, et al. Identification of an aberrantly spliced form of HDMX in human tumors: a new mechanism for HDM2 stabilization. Can Res. 2005;65:9687–94. https://doi.org/10.1158/0008-5472.CAN-05-0450.
Prodosmo A, Giglio S, Moretti S, Mancini F, Barbi F, Avenia N, di Conza G, Schünemann HJ, Pistola L, Ludovini V, Sacchi A, Pontecorvi A, Puxeddu E, et al. Analysis of human MDM4 variants in papillary thyroid carcinomas reveals new potential markers of cancer properties. J Mol Med. 2008;86:585–96. https://doi.org/10.1007/s00109-008-0322-6.
Alatawi A, Kho S, Markey MP. MDM4 isoform expression in melanoma supports an oncogenic role for MDM4-A. J Skin Cancer. 2021. https://doi.org/10.1155/2021/3087579.
de Graaf P, Little NA, Ramos YFM, Meulmeester E, Letteboer SJF, Jochemsen AG. Hdmx protein stability is regulated by the ubiquitin ligase activity of Mdm2. J Biol Chem. 2003;278:38315–24. https://doi.org/10.1074/jbc.M213034200.
Chandler DS, Singh RK, Caldwell LC, Bitler JL, Lozano G. Genotoxic stress induces coordinately regulated alternative splicing of the p53 modulators MDM2 and MDM4. Can Res. 2006;66:9502–8. https://doi.org/10.1158/0008-5472.CAN-05-4271.
Jacob AG, O’Brien D, Singh RK, Comiskey DF, Littleton RM, Mohammad F, Gladman JT, Widmann MC, Jeyaraj SC, Bolinger C, Anderson JR, Barkauskas DA, Boris-Lawrie K, et al. Stress-induced isoforms of MDM2 and MDM4 correlate with high-grade disease and an altered splicing network in pediatric rhabdomyosarcoma. Neoplasia. 2013. https://doi.org/10.1593/neo.13286.
Riemenschneider MJ, Büschges R, Wolter M, Reifenberger J, Boström J, Kraus JA, Schlegel U, Reifenberger G. Amplification and overexpression of the MDM4 (MDMX) gene from 1q32 in a subset of malignant gliomas without TP53 mutation or MDM2 amplification. Can Res. 1999;59:6091–6.
Danovi D, Meulmeester E, Pasini D, Migliorini D, Capra M, Frenk R, de Graaf P, Francoz S, Gasparini P, Gobbi A, Helin K, Pelicci PG, Jochemsen AG, et al. Amplification of MDMX (or MDM4) directly contributes to tumor formation by inhibiting p53 tumor suppressor activity. Mol Cell Biol. 2004;24:5835–43. https://doi.org/10.1128/MCB.24.13.5835-5843.2004.
Laurie NA, Donovan SL, Shih CS, Zhang J, Mills N, Fuller C, Teunisse A, Lam S, Ramos Y, Mohan A, Johnson D, Wilson M, Rodriguez-Galindo C, et al. Inactivation of the p53 pathway in retinoblastoma. Nature. 2006;444:61–6. https://doi.org/10.1038/nature05194.
Sun T, Lee GSM, Oh WK, Pomerantz M, Yang M, Xie W, Freedman ML, Kantoff PW. Single-nucleotide polymorphisms in p53 pathway and aggressiveness of prostate cancer in a caucasian population. Clin Cancer Res. 2010;16:5244–51. https://doi.org/10.1158/1078-0432.CCR-10-1261.
Yu H, Wang LE, Liu Z, Wei S, Li G, Sturgis EM, Wei Q. Polymorphisms of MDM4 and risk of squamous cell carcinoma of the head and neck. Pharmacogenet Genom. 2011;21:388–96. https://doi.org/10.1097/FPC.0b013e32834632e4.
Swetzig WM, Wang J, Das GM. Estrogen receptor alpha ERα/ESR1 mediates the p53-independent overexpression of MDM4/MDMX and MDM2 in human breast cancer. Oncotarget. 2016. https://doi.org/10.18632/oncotarget.7533.
Wang MY, Jia M, He J, Zhou F, Qiu LX, Sun MH, Yang YJ, Wang JC, Jin L, Wang YN, Wei QY. MDM4 genetic variants and risk of gastric cancer in an eastern chinese population. Oncotarget. 2017. https://doi.org/10.18632/oncotarget.14666.
Zhang X, Yamamoto Y, Wang X, Sato M, Imanishi M, Sugaya A, Hirose M, Endo S, Moriwaki T, Yamato K, Hyodo I. MDM4 as a Prognostic factor for patients with gastric cancer with low expression of p53. Anticancer Res. 2021;41:1475–83. https://doi.org/10.21873/anticanres.14906.
Li L, Tan Y, Chen X, Xu Z, Yang S, Ren F, Guo H, Wang X, Chen Y, Li G, Wang H. MDM4 overexpressed in acute myeloid leukemia patients with complex karyotype and wild-type TP53. PLoS ONE. 2014;9:e113088. https://doi.org/10.1371/journal.pone.0113088.
Xiong S, Pant V, Suh YA, van Pelt CS, Wang Y, Valentin-Vega YA, Post SM, Lozano G. Spontaneous tumorigenesis in mice overexpressing the p53-negative regulator Mdm4. Can Res. 2010;70:7148–54. https://doi.org/10.1158/0008-5472.CAN-10-1457.
Gao C, Xiao G, Piersigilli A, Gou J, Ogunwobi O, Bargonetti J. Context-dependent roles of MDMX (MDM4) and MDM2 in breast cancer proliferation and circulating tumor cells. Breast Cancer Res. 2019;21:5. https://doi.org/10.1186/s13058-018-1094-8.
Yu Q, Li Y, Mu K, Li Z, Meng Q, Wu X, Wang Y, Li L. Amplification of Mdmx and overexpression of MDM2 contribute to mammary carcinogenesis by substituting for p53 mutations. Diagn Pathol. 2014;9:71. https://doi.org/10.1186/1746-1596-9-71.
Hoffman Y, Pilpel Y, Oren M. MicroRNAs and Alu elements in the p53-Mdm2-Mdm4 regulatory network. J Mol Cell Biol. 2014;3:192–7. https://doi.org/10.1093/jmcb/mju020.
Kim VN, Han J, Siomi MC. Biogenesis of small RNAs in animals. Nat Rev Mol Cell Biol. 2009;10:126–39. https://doi.org/10.1038/nrm2632.
Yang Y, Gao W, Ding X, Xu W, Liu D, Su B, Sun Y. Variations within 3’-UTR of MDM4 gene contribute to clinical outcomes of advanced non-small cell lung cancer patients following platinum-based chemotherapy. Oncotarget. 2017;8:16313–24. https://doi.org/10.18632/oncotarget.10771.
Gao F, Xiong X, Pan W, Yang X, Zhou C, Yuan Q, Zhou L, Yang M. A regulatory MDM4 genetic variant locating in the binding sequence of multiple MicroRNAs contributes to susceptibility of small cell lung cancer. PLoS ONE. 2015;10:e0135647. https://doi.org/10.1371/journal.pone.0135647.
Wynendaele J, Böhnke A, Leucci E, Nielsen SJ, Lambertz I, Hammer S, Sbrzesny N, Kubitza D, Wolf A, Gradhand E, Balschun K, Braicu I, Sehouli J, et al. An illegitimate microRNA target site within the 3′ UTR of MDM4 affects ovarian cancer progression and chemosensitivity. Can Res. 2010;70:9641–9. https://doi.org/10.1158/0008-5472.CAN-10-0527.
Stegeman S, Moya L, Selth LA, Spurdle AB, Clements JA, Batra J. A genetic variant of MDM4 influences regulation by multiple microRNAs in prostate cancer. Endocr Relat Cancer. 2015;22:265–76. https://doi.org/10.1530/ERC-15-0013.
Yu D, Xu Z, Cheng X, Qin J. The role of miRNAs in MDMX-p53 interplay. J Evid Based Med. 2021;14:152–60. https://doi.org/10.1111/jebm.12428.
Zhao J, Dong X, Tao S-J, Liu X-L, Li Z, Liu J-M, Chen Y. MDM4 is targeted by miR-449b-5p to promote the proliferation of endometrial carcinoma. Eur Rev Med Pharmacol Sci. 2020;24:11528–35. https://doi.org/10.26355/eurrev_202011_23794.
Singh Atwal G, Kirchhoff T, Bond EE, Montagna M, Menin C, Bertorelle R, Chiara Scaini M, Bartel F, Bö Hnke FA, Pempe C, Gradhand E, Hauptmann S, Offit K, et al. Altered tumor formation and evolutionary selection of genetic variants in the human MDM4 oncogene. Proc Natl Acad Sci. 2009;106:10236–41. https://doi.org/10.1073/pnas.0901298106.
Kulkarni DA, Vazquez A, Haffty BG, Bandera Ev, Hu W, Sun YY, Toppmeyer DL, Levine AJ, Hirshfield KM. A polymorphic variant in human MDM4 associates with accelerated age of onset of estrogen receptor negative breast cancer. Carcinogenesis. 2009;30:1910–5. https://doi.org/10.1093/carcin/bgp224.
Eeles RA, Olamaal AA, Benlloch S, Saunders EJ, Leongamornlert DA, Tymrakiewicz M, Ghoussaini M, Luccarini C, Dennis J, Jugurnauth-Little S, Dadaev T, Neal DE, Hamdy FC, et al. Identification of 23 new prostate cancer susceptibility loci using the iCOGS custom genotyping array. Nat Genet. 2013;45:385–91. https://doi.org/10.1038/ng.2560.
Deng N, Zhou H, Fan H, Yuan Y. Single nucleotide polymorphisms and cancer susceptibility. Oncotarget. 2017;8:110635–49. https://doi.org/10.18632/oncotarget.22372.
Gansmo LB, Romundstad P, Birkeland E, Hveem K, Vatten L, Knappskog S, Lønning PE. MDM4 SNP34091 (rs4245739) and its effect on breast-, colon-, lung-, and prostate cancer risk. Cancer Med. 2015;4:1901–7. https://doi.org/10.1002/cam4.555.
Gansmo LB, Bjørnslett M, Halle MK, Salvesen HB, Dørum A, Birkeland E, Hveem K, Romundstad P, Vatten L, Lønning PE, Knappskog S. The MDM4 SNP34091 (rs4245739) C-allele is associated with increased risk of ovarian—but not endometrial cancer. Tumor Biology. 2016;37:10697–702. https://doi.org/10.1007/s13277-016-4940-2.
Zhou L, Zhang X, Li Z, Zhou C, Li M, Tang X, Lu C, Li H, Yuan Q, Yang M. Association of a genetic variation in a miR-191 binding site in MDM4 with risk of esophageal squamous cell carcinoma. PLoS ONE. 2013;8:e64331. https://doi.org/10.1371/journal.pone.0064331.
Liu J, Tang X, Li M, Lu C, Shi J, Zhou L, Yuan Q, Yang M. Functional MDM4 rs4245739 genetic variant, alone and in combination with P53 Arg72Pro polymorphism, contributes to breast cancer susceptibility. Breast Cancer Res Treat. 2013;140:151–7. https://doi.org/10.1007/s10549-013-2615-x.
Jin X, Zhao W, Zheng M, Zhou P, Niu T. The role of MDM4 SNP34091 A>C polymorphism in cancer: a meta-analysis on 19,328 patients and 51,058 controls. Int J Biol Markers. 2017;32:e62–7. https://doi.org/10.5301/jbm.5000228.
Bauer M, Kantelhardt EJ, Stiewe T, Nist A, Mernberger M, Politt K, Hanf V, Lantzsch T, Uleer C, Peschel S, John J, Buchmann J, Weigert E, et al. Specific allelic variants of SNPs in the MDM2 and MDMX genes are associated with earlier tumor onset and progression in Caucasian breast cancer patients. Oncotarget. 2019;10:1975–92. https://doi.org/10.18632/oncotarget.26768.
Zhang Y, Sturgis EM, Wei P, Liu H, Wang Z, Ma Y, Liu C, Gu KJ, Wei Q, Li G. A genetic variant within MDM4 3′UTR miRNA binding site is associated with HPV16-positive tumors and survival of oropharyngeal cancer. Mol Carcinog. 2019;58:2276–85. https://doi.org/10.1002/mc.23116.
Zhao DM, Diao YE, Xu Q. Association of mdm4 gene rs4245739 polymorphism with the risk and clinical characteristics of colorectal cancer in a chinese han population. Pharmacogen Personal Med. 2020;13:673–8. https://doi.org/10.2147/PGPM.S260209.
Bartnykaitė A, Savukaitytė A, Ugenskienė R, Daukšaitė M, Korobeinikova E, Gudaitienė J, Juozaitytė E. Associations of mdm2 and mdm4 polymorphisms with early-stage breast cancer. J Clin Med. 2021;10:1–11. https://doi.org/10.3390/jcm10040866.
Chen J, Li X, Liu R, Xie Y, Liu Z, Xiong H, Li Y. The correlation of mouse double minute 4 (MDM4) polymorphisms (rs4245739, rs1563828, rs11801299, rs10900598, and rs1380576) with cancer susceptibility: a meta-analysis. Med Sci Monit. 2022;28:e935671. https://doi.org/10.12659/MSM.935671.
Wang Y, Yang Z, Chang X, Li J, Han Z. Five MDM4 gene polymorphisms on cancer risk: an updated systematic review and meta-analysis. Int J Biol Markers. 2021;36:17246008211033874. https://doi.org/10.1177/17246008211033874.
Kotarac N, Dobrijevic Z, Matijasevic S, Savic-Pavicevic D, Brajuskovic G. Association of KLK3, VAMP8 and MDM4 genetic variants within microRNA binding sites with prostate cancer evidence from serbian population. Pathol Oncol Res. 2020;26:2409–23. https://doi.org/10.1007/s12253-020-00839-7.
Garcia-Closas M, Couch FJ, Lindstrom S, Michailidou K, Schmidt MK, Brook MN, Orr N, Rhie SK, Riboli E, Feigelson HS, le Marchand L, Buring JE, Eccles D, et al. Genome-wide association studies identify four ER negative-specific breast cancer risk loci. Nat Genet. 2013;45:392–8. https://doi.org/10.1038/ng.2561.
Purrington KS, Slager S, Eccles D, Yannoukakos D, Fasching PA, Miron P, Carpenter J, Chang-claude J, Martin NG, Montgomery GW, Kristensen V, Anton-Culver H, Goodfellow P, et al. Genome-wide association study identifies 25 known breast cancer susceptibility loci as risk factors for triple-negative breast cancer. Carcinogenesis. 2014;35:1012–9. https://doi.org/10.1093/carcin/bgt404.
Yu F, Jiang Z, Song A. Association of rs11801299 and rs1380576 polymorphisms at MDM4 with risk, clinicopathological features and prognosis in patients with retinoblastoma. Cancer Epidemiol. 2019;58:153–9. https://doi.org/10.1016/j.canep.2018.12.010.
Lu Z, Sturgis EM, Zhu L, Zhang H, Tao Y, Wei P, Wei Q, Li G. Mouse double minute 4 variants modify susceptibility to risk of recurrence in patients with squamous cell carcinoma of the oropharynx. Mol Carcinog. 2018;57:361–9. https://doi.org/10.1002/mc.22760.
Fan H, Liu D, Qiu X, Qiao F, Wu Q, Su X, Zhang F, Song Y, Zhao Z, Xie W. A functional polymorphism in the DNA methyltransferase-3A promoter modifies the susceptibility in gastric cancer but not in esophageal carcinoma. BMC Med. 2010;8:12. https://doi.org/10.1186/1741-7015-8-12.
Shao N, Li J, Xu B, Wang Y, Lu X, Feng N. Role of the functional variant (−652T>G) in the XRCC4 promoter in prostate cancer. Mol Biol Rep. 2014;41:7463–70. https://doi.org/10.1007/s11033-014-3636-1.
Rintisch C, Heinig M, Bauerfeind A, Schafer S, Mieth C, Patone G, Hummel O, Chen W, Cook S, Cuppen E, Colomé-Tatché M, Johannes F, Jansen RC, et al. Natural variation of histone modification and its impact on gene expression in the rat genome. Genome Res. 2014;24:942–53. https://doi.org/10.1101/gr.169029.113.
Reincke S, Govbakh L, Wilhelm B, Jin H, Bogdanova N, Bremer M, Karstens JH, Dörk T. Mutation analysis of the MDM4 gene in German breast cancer patients. BMC Cancer. 2008;8:942–53. https://doi.org/10.1186/1471-2407-8-52.
Oscanoa J, Sivapalan L, Gadaleta E, Dayem Ullah AZ, Lemoine NR, Chelala C. SNPnexus: a web server for functional annotation of human genome sequence variation. Nucleic Acids Res. 2020;48:W185–92. https://doi.org/10.1093/nar/gkaa420.
Vohra M, Sharma AR, Prabhu BN, Rai PS. SNPs in sites for DNA methylation, transcription factor binding, and miRNA targets leading to allele-specific gene expression and contributing to complex disease risk: a systematic review. Public Health Genom. 2020;23:155–70. https://doi.org/10.1159/000510253.
Shyamala N, Kongettira CL, Puranam K, Kupsal K, Kummari R, Padala C, Hanumanth SR. In silico identification of single nucleotide variations at CpG sites regulating CpG island existence and size. Sci Rep. 2022;12:3574. https://doi.org/10.1038/s41598-022-05198-8.
de Oliveira Reis AH, de Carvalho INSR, de Sousa Damasceno PB, Ferman SE, Lucena E, Lopez-Camelo JS, Seuánez HN, Vargas FR. Influence of MDM2 and MDM4 on development and survival in hereditary retinoblastoma. Pediatric Blood Cancer. 2012;59:39–43. https://doi.org/10.1002/pbc.24014.
Spinello Z, Fregnani A, Quotti Tubi L, Trentin L, Piazza F, Manni S. Targeting protein kinases in blood cancer: focusing on CK1α and CK2. Int J Mol Sci. 2021;22:3716. https://doi.org/10.3390/ijms22073716.
Chen L, Li C, Pan Y, Chen J. Regulation of p53-MDMX interaction by casein kinase 1 alpha. Mol Cell Biol. 2005;25:6509–20. https://doi.org/10.1128/MCB.25.15.6509-6520.2005.
Karagkouni D, Paraskevopoulou MD, Chatzopoulos S, Vlachos IS, Tastsoglou S, Kanellos I, Papadimitriou D, Kavakiotis I, Maniou S, Skoufos G, Vergoulis T, Dalamagas T, Hatzigeorgiou AG. DIANA-TarBase v8: a decade-long collection of experimentally supported miRNA–gene interactions. Nucleic Acids Res. 2018;46:D239–45. https://doi.org/10.1093/nar/gkx1141.
Sim N-L, Kumar P, Hu J, Henikoff S, Schneider G, Ng PC. SIFT web server: predicting effects of amino acid substitutions on proteins. Nucleic Acids Res. 2012;40:W452–7. https://doi.org/10.1093/nar/gks539.
Melin BS, Barnholtz-Sloan JS, Wrensch MR, Johansen C, Il’yasova D, Kinnersley B, Ostrom QT, Labreche K, Chen Y, Armstrong G, Liu Y, Eckel-Passow JE, Decker PA, et al. Genome-wide association study of glioma subtypes identifies specific differences in genetic susceptibility to glioblastoma and non-glioblastoma tumors. Nat Genet. 2017. https://doi.org/10.1038/ng.3823.
Sun P, Yan F, Fang W, Zhao J, Chen H, Ma X, Song J. MDM4 contributes to the increased risk of glioma susceptibility in Han Chinese population. Sci Rep. 2018;8:11093. https://doi.org/10.1038/s41598-018-29468-6.
le Morvan V, Litière S, Laroche-Clary A, Ait-Ouferoukh S, Bellott R, Messina C, Cameron D, Bonnefoi H, Robert J. Identification of SNPs associated with response of breast cancer patients to neoadjuvant chemotherapy in the EORTC-10994 randomized phase III trial. Pharmacogen J. 2015;15:63–8. https://doi.org/10.1038/tpj.2014.24.
Hashemi M, Sanaei S, Hashemi SM, Eskandari E, Bahari G. Association of single nucleotide polymorphisms of the MDM4 gene with the susceptibility to breast cancer in a Southeast Iranian population sample. Clin Breast Cancer. 2018;18:e883–91. https://doi.org/10.1016/j.clbc.2018.01.003.
Lesseur C, Ferreiro-Iglesias A, McKay JD, Bossé Y, Johansson M, Gaborieau V, Landi MT, Christiani DC, Caporaso NC, Bojesen SE, Amos CI, Shete S, Liu G, et al. Genome-wide association meta-analysis identifies pleiotropic risk loci for aerodigestive squamous cell cancers. PLoS Genet. 2021;17:e1009254. https://doi.org/10.1371/journal.pgen.1009254.
Jansen R, Hottenga J-J, Nivard MG, Abdellaoui A, Laport B, de Geus EJ, Wright FA, Penninx BWJH, Boomsma DI. Conditional eQTL analysis reveals allelic heterogeneity of gene expression. Hum Mol Genet. 2017;26:1444–51. https://doi.org/10.1093/hmg/ddx043.
Khanlou ZM, Pouladi N, Feizi MH, Pedram N. Lack of associations of the MDM4 rs4245739 polymorphism with risk of thyroid cancer among Iranian-Azeri patients: a case-control study. Asian Pac J Cancer Prev. 2017;18:1133–8. https://doi.org/10.22034/APJCP.2017.18.4.1133.
Stumbrytė-Kaminskienė A, Gudlevičienė Ž, Dabkevičienė D, Mackevičienė I. Combined effect of HPV and several gene SNPs in laryngeal cancer. Medicina. 2020;56:81. https://doi.org/10.3390/medicina56020081.
Wang Z, Sturgis EM, Zhang Y, Huang Z, Zhou Q, Wei Q, Li G. Combined p53-related genetic variants together with HPV infection increase oral cancer risk. Int J Cancer. 2012;131:E251–8. https://doi.org/10.1002/ijc.27335.
Stumbryte A, Gudleviciene Z, Kundrotas G, Dabkeviciene D, Kunickaite A, Cicenas S. Individual and combined effect of TP53, MDM2, MDM4, MTHFR, CCR5, and CASP8 gene polymorphisms in lung cancer. Oncotarget. 2018;9:3214–29. https://doi.org/10.18632/oncotarget.22756.
Pedram N, Pouladi N, Feizi M, Montazeri V, Sakhinia E, Estiar M. Analysis of the association between MDM4 rs4245739 single nucleotide polymorphism and breast cancer susceptibility. Clin Lab. 2016;62:1303–8. https://doi.org/10.7754/Clin.Lab.2016.151128.
Anwar SL, Wulaningsih W, Watkins J. Profile of the breast cancer susceptibility marker rs4245739 identifies a role for miRNAs. Cancer Biol Med. 2017;14:387–95. https://doi.org/10.20892/j.issn.2095-3941.2017.0050.
Zhou R, Li Y, Wang N, Niu C, Huang X, Cao S, Huo X. MDM4 polymorphisms associated with the risk but not the prognosis of esophageal cancer in Cixian high-incidence region from northern China. Asia Pacific J Clin Oncol. 2022;18:e435–41. https://doi.org/10.1111/ajco.13746.
Wu GC, Zhang ZT. Genetic association of single nucleotide polymorphisms in P53 pathway with gastric cancer risk in a Chinese Han population. Med Oncol. 2015;32:1–5. https://doi.org/10.1007/s12032-014-0401-1.
Fang S, Krahe R, Lozano G, Han Y, Chen W, Post SM, Zhang B, Wilson CD, Bachinski LL, Strong LC, Amos CI. Effects of MDM2, MDM4and TP53 codon 72 polymorphisms on cancer risk in a cohort study of carriers of TP53 germline mutations. PLoS ONE. 2010;5:e10813. https://doi.org/10.1371/journal.pone.0010813.
Yu H, Sturgis EM, Liu Z, Wang LE, Wei Q, Li G. Modifying effect of MDM4 variants on risk of HPV16-associated squamous cell carcinoma of oropharynx. Cancer. 2012;118:1684–92. https://doi.org/10.1002/cncr.26423.
Cardon LR, Bell JI. Association study designs for complex diseases. Nat Rev Genet. 2001;2:91–9. https://doi.org/10.1038/35052543.
Slatkin M. Linkage disequilibrium—understanding the evolutionary past and mapping the medical future. Nat Rev Genet. 2008;9:477–85. https://doi.org/10.1038/nrg2361.
Machiela MJ, Chanock SJ. LDlink: a web-based application for exploring population-specific haplotype structure and linking correlated alleles of possible functional variants. Bioinformatics. 2015;31:3555–7. https://doi.org/10.1093/bioinformatics/btv402.
Luo Y, Hitz BC, Gabdank I, Hilton JA, Kagda MS, Lam B, Myers Z, Sud P, Jou J, Lin K, Baymuradov UK, Graham K, Litton C, et al. New developments on the encyclopedia of DNA elements (ENCODE) data portal. Nucleic Acids Res. 2020;48:D882–9. https://doi.org/10.1093/nar/gkz1062.
Acknowledgements
We acknowledge the Post-Graduation Program of Genetics from the Federal University of Parana for their support.
Funding
The scholarship and financial support were provided by the Brazilian National Council for Scientific and Technological Development (CNPq), Coordination for the Improvement of Higher Education Personnel (CAPES), Support Program for Centers of Excellence (PRONEX), and Araucaria Foundation to Support Scientific and Technological Development of the State of Parana. Pronex-FA/CNPq, 116/2018, CNPq, 406187/2016-9, Karin Braun-Prado.
Author information
Authors and Affiliations
Contributions
The authors contributed equally to this article.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that no conflicts of interest exist.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Almeida, G.M., Castilho, A.C., Adamoski, D. et al. MDM4: What do we know about the association between its polymorphisms and cancer?. Med Oncol 40, 61 (2023). https://doi.org/10.1007/s12032-022-01929-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12032-022-01929-z