Abstract
Schizophrenia (SZ) is a debilitating psychotic disorder of unknown etiology, and the diagnosis is essentially based on clinical symptoms. So it is urgent to find an objective and feasible clinical diagnostic index for SZ. MicroRNA array was performed in peripheral blood mononuclear cells (PBMCs) obtained from young SZ patients and gender-, age-, and ethnicity-matched healthy controls. Then, real-time quantitative reverse transcription polymerase chain reaction (qRT-PCR) was used to verify the top 10 microRNAs (miRNAs) with the highest fold change values in 55 SZ patients and 28 healthy controls, and 9 miRNAs demonstrate significant differences in expression levels (P < 0.01). Receiver operating characteristic (ROC) curve analysis showed that the combining area under the ROC curve (AUC) of the nine miRNAs was 0.973 (95 % confidence interval (CI): 0.945–1.000). miRNA target gene prediction and functional annotation analysis showed that there were significant enrichments in several gene ontology (GO) biological process and Kyoto encyclopedia of genes and genomes (KEGG) pathways associated with nervous system and brain functions, suggesting that the differentially expressed miRNAs may be involved in mechanism of SZ. We conclude that altered expression of miRNAs in PMBCs might be involved in young SZ pathogenesis and may serve as noninvasive biomarker for SZ diagnosis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Schizophrenia (SZ) is a severe, chronic, and disabling mental disorder that affects about 1 % of the world’s population throughout life (Saha et al. 2005). Until today, the diagnosis of SZ is mainly based on clinical symptoms varying greatly between individuals. Moreover, there are considerable symptomatic overlap between some mental disorders (Haller et al. 2014), which may lead to misdiagnosis or different diagnosis between doctors. So it is urgent to find an objective and feasible clinical diagnostic index for SZ.
microRNAs (miRNAs) are endogenous small noncoding RNAs that negatively regulate gene expression by translational repression or mRNA cleavage at the post-transcriptional level. Accumulating evidence indicates that miRNAs are involved in a wide variety of biological processes, including development, differentiation, proliferation, apoptosis, invasion, and metastasis (Bartel 2009; Zhuo et al. 2013; Moreno-Moya et al. 2014), and play an important role in brain development and function (Bian and Sun 2011; Iyengar et al. 2014; Smalheiser 2014). There is also growing evidence showing that miRNA expression profiles are altered in psychiatric disorders such as SZ, bipolar disorder, autism spectrum disorders (Miller and Wahlestedt 2010; Xu et al. 2010; Song et al. 2014), suggesting that miRNAs may contribute to the etiology and pathogenesis of psychiatric disorders. Since 2008, many studies have reported that specific miRNA expression can be detected in peripheral blood in many diseases. From then on, circulating miRNAs have become a hot issue of current research. Because of its characteristics of high stability, noninvasiveness, easy accessibility, high measurement accuracy, and cost-effectiveness (Ajit 2012), circulating miRNAs may serve as promising biomarkers of many diseases. Although the research on circulating miRNAs as biomarkers is still at a groping stage, hundreds of papers have already been published on the subject in various diseases such as cancer and cardiovascular diseases (Wittmann and Jäck 2010; Ajit 2012; Xu et al. 2012). However, there is a very limited number of study on the miRNA expression profile in peripheral blood of psychiatry diseases. In terms of schizophrenia, the only study we could found was conducted by Lai et al., who identified a seven-miRNA signature (hsa-miR-34a, miR-449a, miR-564, miR-548d, miR-572, and miR-652 upregulated; miR-432 downregulated) in blood mononuclear leukocytes which could distinguish SZ patients from normal controls (Lai et al. 2011).
Given that the onset of schizophrenia generally occurs between the late teens and the mid-30s (American Psychiatric Association 2013) and that the expression of miRNA is influenced by age (Noren Hooten et al. 2010; Pena-Chilet et al. 2014), we speculate that young SZ patients would be a good sample to identify altered miRNAs as biomarker for diagnosis. The present study screened the differently expressed miRNAs in peripheral blood mononuclear cells (PBMCs) from young SZ patients and healthy controls using miRNA microarray, and then the candidate miRNAs were validated in a larger sample of cases and controls using real-time quantitative reverse transcription polymerase chain reaction (qRT-PCR). Finally, target genes were predicted by bioinformatics tools and were subjected to gene ontology (GO) analysis and Kyoto encyclopedia of genes and genomes (KEGG) pathway analysis.
Materials and Methods
Patients
A total of 55 SZ patients between 18 and 30 years old who met the criteria of the Diagnostic and Statistical Manual of Mental Disorders, 4th Edition for schizophrenia, were enrolled from No.102 Hospital of the People’s Liberation Army from December 2012 to May 2013. Patients were either first onset or drug naive from any antidepressant for at least 3 months before enrollment. Patients with severe medical diseases, other psychiatric disorders, structural brain disorders, mental retardation, mood incongruent psychotic symptoms, and primary substance abuse were excluded. Patients who had brain injury causing traumatic amnesia longer than 24 h and who received blood transfusion within 1 month or electroconvulsive therapy within 6 months were also excluded from the study.
Twenty-eight healthy controls without any family history of major psychiatric disorders (SZ, bipolar disorder, and major depressive disorder) were recruited. All healthy controls were without any history of blood transfusion or severe traumatic event within 1 month. Patients and healthy controls were matched in gender, age, and ethnicity on a ratio of 2:1. The study was approved by the local ethics committee. Written informed consent was obtained from all subjects.
Blood Collection and RNA Extraction
Whole blood (5 ml) was collected in EDTA anticoagulant tube from each subject and processed within 1 h. PBMCs were isolated through density gradient centrifugation and stored at −80 °C until use. Total RNAs were extracted from the PBMCs with the mirVana™miRNA Isolation Kit (Ambion, LOT:1406120 AM1561) according to the manufacturer’s protocol. To ensure a robust analysis for the following procedures, samples with an RNA integrity number (RIN) inferior to 8 were excluded.
miRNA Microarray Expression Profiling
RNA samples from three SZ patients (male, 20 years; male, 21 years; female, 19 years) and three controls (male, 20 years; male, 21 years; female, 19 years) were used for miRNA microarray profiling. miRNA expression was measured by Affymetrix miRNA 3.0 array (Affymetrix, Santa Clara, CA, USA). The sample labeling, microarray hybridization, and washing were performed based on the manufacturer’s standard protocols. Briefly, total RNA were tailed with poly A and then labeled with Biotin. Afterward, the labeled RNAs were hybridized onto the microarray. Having washed and stained the slides, the arrays were scanned by the Affymetrix Scanner 3000 (Affymetrix). The scanned images were analyzed using Expression Console software (version 1.3.1, Affymetrix).
Real-Time Quantitative Reverse Transcription PCR
According to microarray results, the top 10 miRNAs with the highest expression changes were chosen for further validation with qRT-PCR. Blood samples from 55 SZ patients and 28 controls were used to validate the candidate miRNAs. Total RNAs were isolated from the PBMCs using Trizol reagent (Invitrogen®, USA) for quantitative detection of miRNA. Complementary DNA was synthesized using the Reverse Transcription TaqMan MicroRNA Reverse Transcription Kit and miRNA-specific stem-loop primers (Applied Biosystems, Inc., USA, P/N: 4366596) according to the manufacturer’s instructions. Real-time PCR was performed using Applied Biosystems 7900HT Real-Time PCR System (Applied Biosystems, Inc., USA). The 5× RT primers (miRNA-specific stem-loop primers) and 20× miRNA-specific PCR primer/probe mix were supplied by the TaqMan MicroRNA Assays (Applied Biosystems, Inc.). Data were collected using the SDS 2.3 software (Applied Biosystems, Inc.). After normalized to RNU48, the expression levels of miRNAs were calculated using the 2−ΔΔCt method (Schmittgen and Livak 2008).
miRNA Target Prediction and Pathway Analysis
The predicted target genes of the differentially expressed miRNAs were obtained using two open access databases (miRDB, http://mirdb.org/miRDB/index.html; and DIANA-microT v5.0, http://diana.imis.athena-innovation.gr/DianaTools/index.php?r=microT_CDS/index). The genes co-identified by both programs were considered as potential target genes of a given miRNA. We next performed GO and KEGG pathway analyses using the public database FunNet (http://www.funnet.ws/) to explore the functional annotation of candidate target genes.
Statistical Analysis
Statistical analyses were carried out using Statistical Package for Social Sciences (SPSS) for Windows 22.0, DataAssist 3.0, and Graphpad Prism 5.01. Demographic variables were compared between patients and controls with chi-square test for qualitative variables and t test for quantitative variables. Expression levels of miRNAs were compared using the Mann–Whitney U test. miRNA data are presented as fold change relative to the control group (control = 1). Receiver operating characteristic (ROC) curves and the area under the ROC curve (AUC) were analyzed to assess specificity and sensitivity of single miRNA and their combination using multiple logistic regression analysis. All statistical tests were two-tailed and P values of <0.05 were considered to indicate significant differences.
Results
Clinical Characteristics of the Patients
The mean age (mean ± SD) of patients and health controls was 21.7 ± 4.09 and 22.1 ± 4.00 years, respectively. All the subjects were of Han nationality, and there were no significant differences in age, sex, or residential locations between SZ patients and healthy controls (Table 1).
Microarray Expression Analysis
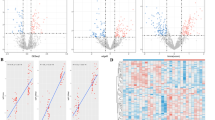
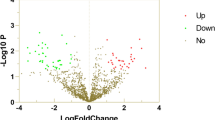
There were 33 miRNAs significantly differentially expressed in three SZ patients compared with three controls (fold change ≧ 2; P < 0.05); all but one (hsa-miR-4701-3p) of which were upregulated (Table 2). Heat map was generated to visualize the results of hierarchical clustering, where a general distinction between samples of SZ and control were clearly observed (Fig. 1).
Heat map showing 33 differentially expressed miRNAs in PBMCs from SZ patients (n = 3) and controls (n = 3). Rows represent miRNA species, and columns represent individual blood sample. The relative miRNA expression is depicted according to the color scale. Red indicates upregulation; green indicates downregulation. The numbers with SZ denote schizophrenia; numbers with NC denote normal controls (Color figure online)
Real-Time qRT-PCR Validation
To validate the results of the microarray assay, 10 miRNAs (miR-1273d, miR-1303, miR-21, miR-3064-5p, miR-3131, miR-3687, miR-3916, miR-4428, miR-4725-3p, and miR-5096, all of which were upregulated) with the highest fold change values were examined in a larger sample (55 cases and 28 controls) using qRT-PCR method. Using RNU48 as normalization control, all the 10 miRNAs were upregulated in SZ patients compared with normal controls, showing the same tendency with the microarray results, and 9 of them demonstrate significant difference (P < 0.01) except miR-3916 (Fig. 2).
To evaluate the potential of the 10 miRNAs as biomarkers of SZ, we performed ROC analysis of data from the qRT-PCR results. ROC curve analysis showed that the AUC of nine miRNAs (miR-1273d, miR-1303, miR-21, miR-3064-5p, miR-3131, miR-3687, miR-4428, miR-4725-3p, miR-5096) could distinguish SZ cases from normal controls with an AUC ranging from 0.699 to 0.886 (all P < 0.01), while the AUC of miR-3916 did not reach statistical significance (P = 0.128). By using logistic regression approach, we examined the predictive power of the combined ROC of the nine miRNAs. The AUC of the combined ROC curve was 0.973 (95 % confidence interval (CI) 0.945–1.000) with 89.29 % sensitivity and 94.55 % specificity (Fig. 3).
miRNA Target Analysis
Using miRDB and DIANA-microT v5.0, a total number of 1457 genes of the nine differentially expressed miRNAs were identified. Then, the microRNA–gene network was established to outline the interactions of miRNAs and the targets (Fig. 4). GO biological process analysis indicate that the target genes have participated in a wide variety of physiological and pathophysiological processes, such as regulation of transcription, signal transduction, cell adhesion, ion transport, blood coagulation, synaptic transmission, axon guidance, small GTPase mediated signal transduction, interspecies interaction between organisms, and nerve growth factor receptor signaling pathway. Among which, biological processes related to brain development and function are listed in Table 3, which may be involved in the pathophysiology of SZ. Likewise, KEGG pathway analysis showed a significant enrichment in several important pathways related to neuronal brain function, such as neurotrophin signaling pathway, Jak-STAT signaling pathway, Wnt signaling pathway, Axon guidance, ErbB signaling pathway, chemokine signaling pathway, TGF-beta signaling pathway, and Notch signaling pathway (Table 4).
Discussion
Peripheral blood is an ideal tissue for basic and clinical research for its easy obtainment. Generally, peripheral blood samples can be divided into two categories, PBMCs and plasma/serum, which were widely applied in blood-based miRNA research. Our previous study had demonstrated that aberrant microRNA expression could be detected in both plasma and PBMCs and that the expression level of miR-30e is more significant in plasma than in PBMCs (Sun et al. 2014). But, in the present study, we chose to use PBMCs for the microarray and PCR experiment for several reasons. First, there is more and more evidence indicating close relationship between inflammation and SZ (Nikkila et al. 2001; Miller et al. 2011; Fineberg and Ellman 2013; Miller et al. 2013), and most of the blood-based miRNA research studies having relation to inflammation were conducted in PBMCs (Lai et al. 2011; Yao et al. 2011; Ma et al. 2014; Munshi et al. 2014). Second, several studies have indicated that transcriptional alterations in PBMCs may reflect the molecular and cellular changes in the brain (Fisar and Raboch 2008; Cattaneo et al. 2010). Furthermore, evidence suggests that PBMCs may share a common miRNA expression pattern with the brain (Liang et al. 2007).
In this study, microarray was first used to screen dysregulated miRNAs in the PBMCs of young SZ patients. As shown in Table 2, a total of 33 miRNAs were identified to be differentially expressed, in which 32 miRNAs were upregulated and only 1 miRNA (hsa-miR-4701-3p) was downregulated. Next, we chose the top 10 miRNAs with the highest fold change values (miR-1273d, miR-1303, miR-21, miR-3064-5p, miR-3131, miR-3687, miR-3916, miR-4428, miR-4725-3p, miR-5096; all of which were up-regulated) for qRT-PCR validation and 9 of them (except miR-3916) reached statistical significance in SZ patients compared with normal controls. To further explore the diagnostic potential of these nine miRNAs as biomarkers for SZ diagnosis, receiver operating characteristic (ROC) curves were constructed on the basis of the nine miRNA expression levels between two groups. When the expression levels of these nine miRNAs were subjected to combined analysis by multiple logistic regression, the ROC curve reflected a higher ability to differentiate patients with SZ from healthy controls (AUC value 0.973, 95 % CI 0.945–1.000), demonstrating that the diagnostic accuracy of the nine-miRNA signature is an effective biomarker for SZ diagnosis.
Among the nine differentially expressed miRNAs, miR-21 was the most frequently reported one. The function of miR-21 is complex and has elicited considerable interest in diverse fields including embryonic development (Ramachandra et al. 2008), tumorigenesis (Wang et al. 2014), fibrosis (Zhang et al. 2013), and immune reaction (Smigielska-Czepiel et al. 2013). Until now, the majority of the studies of miRNA-21 were related to various human cancers such as breast cancer (Mar-Aguilar et al. 2013), lung cancer(Tang et al. 2013), colorectal cancer (Kanaan et al. 2012), and gastric cancer(Li et al. 2012). However, several recent studies have found that miRNA-21 played an important role in many physiological and pathological processes of the central nervous system. In a rat stroke model, Buller et al. (2010) found that miR-21 levels had been markly improved in neurons of the ischemic boundary zone, and overexpression of miR-21 in neurons significantly reduced Faslg protein levels which is known to contribute to ischemic injury of neurons (Barone et al. 1997). In another study, miR-21 was found exerting the function of reducing neuronal apoptosis through activating the PTEN-Akt signaling pathway (Han et al. 2014). Montalban et al. (2014) reported that miR-21 could enhance neurotrophin signaling and control neuronal differentiation induced by Ngf. In the same study, miR-21 was able to preserve the neurite network and to support viability of the neurons in a situation mimicking neurodegeneration.
In several studies, miR-1303 was reported to be concerned with cancers such as breast cancer, gastric cancer, colorectal cancer, and hepatocellular carcinoma. So far, the other seven miRNAs differentially expressed in our study have scarcely been reported before. All these nine miRNAs were found to be related with SZ for the first time. This may provide new clues for SZ research.
The microRNA–gene network showed that there were 138 genes targeted by at least two miRNAs in which 1 gene (EIF2C1) was targeted by four miRNAs and 11 genes (CLIC6, DCAF7, DGKB, DSEL, ESRRG, GLDN, KCNA1, LPP, PCGF5, RAB22A, ZNF445) were targeted by three miRNAs, suggesting that these miRNAs may have potential roles in the pathogenesis of SZ. Indeed, CLIC6, a member of the intracellular chloride channel family, was proven to be involved in dopamine D(2)-like receptor complex (Griffon et al. 2003). Kcna1, a gene involved in voltage-gated ion channels, was supposed to play a role in mediating antipsychotic drug effects (Duncan et al. 2008).
In order to gain insight into the function of miRNAs, GO term and KEGG pathway annotation were applied to their target gene pool. GO enrichment analysis showed that biological processes regulated by differentially expressed miRNAs including diverse terms, and some important terms, such as synaptic transmission, nerve growth factor receptor signaling pathway, axon guidance, Notch signaling pathway, and regulation of long-term neuronal synaptic plasticity, had direct relationship with central nervous system (CNS) and brain functions. These biological processes may contribute to the etiopathogenesis of SZ based on recent data from many studies (Kalkman 2009; Chen et al. 2011; Earls et al. 2012; Masana et al. 2012; Ikeda et al. 2013). KEGG pathway analysis revealed a close linkage between signaling pathways identified and SZ, as shown in Table 4. For example, neurotrophin 3 levels in serum were found significantly decreased in SZ patients compared to controls, suggesting that the neurotrophin 3 signaling system may play a role in the pathophysiology of SZ (Vargas et al. 2008). JAK-STAT signaling has been proven to play a role in treatment of schizophrenia with olanzapine (Singh et al. 2007). Wnt signaling is a highly conserved pathway that plays a prominent role in the CNS, and an increasing number of evidence suggest a close connection between Wnt signaling and the pathogenesis and therapeutics of schizophrenia (Singh 2013; Peng et al. 2014). NTNG1 is a well-known axon guidance factor, whose allelic variation was proven to be a very important contributor to the risk for schizophrenia (Ohtsuki et al. 2008; Wilcox and Quadri 2014). Besides, ErbB signaling pathway, Chemokine signaling pathway, TGF-beta signaling pathway, and Notch signaling pathway are also found to be implicated in the pathologic mechanisms of SZ (Chertkow et al. 2007; Ohtsuki et al. 2008; Kerns et al. 2010; Frydecka et al. 2013).
There are several limitations of this study that should be mentioned. First, our sample size was relatively small; therefore, further validation in larger cohorts is needed to better evaluate the sensitivity and specificity of the nine-miRNA signature as biomarkers. Second, the predicted genes and their regulation function should be validated by future functional experimental research.
In conclusion, we have identified nine miRNAs (miR-1273d, miR-1303, miR-21, miR-3064-5p, miR-3131, miR-3687, miR-4428, miR-4725-3p, miR-5096) that are upregulated in PBMC of young SZ patients and may serve as useful noninvasive biomarkers for SZ diagnosis. The bioinformatics analysis suggested that these miRNAs may be involved in the pathogenesis of SZ.
References
Ajit SK (2012) Circulating microRNAs as biomarkers, therapeutic targets, and signaling molecules. Sensors 12:3359–3369
American Psychiatric Association (2013) Diagnostic and statistical manual of mental disorders. 5th ed. doi:10.1176/appi.books.9780890425596
Barone FC, Arvin B, White RF, Miller A, Webb CL, Willette RN, Lysko PG, Feuerstein GZ (1997) Tumor necrosis factor-alpha. A mediator of focal ischemic brain injury. Stroke 28:1233–1244
Bartel DP (2009) MicroRNAs: target recognition and regulatory functions. Cell 136:215–233
Bian S, Sun T (2011) Functions of noncoding RNAs in neural development and neurological diseases. Mol Neurobiol 44:359–373
Buller B, Liu X, Wang X, Zhang RL, Zhang L, Hozeska-Solgot A, Chopp M, Zhang ZG (2010) MicroRNA-21 protects neurons from ischemic death. FEBS J 277:4299–4307
Cattaneo A, Sesta A, Calabrese F, Nielsen G, Riva MA, Gennarelli M (2010) The expression of VGF is reduced in leukocytes of depressed patients and it is restored by effective antidepressant treatment. Neuropsychopharmacology 35:1423–1428
Chen SY, Huang PH, Cheng HJ (2011) Disrupted-in-Schizophrenia 1-mediated axon guidance involves TRIO-RAC-PAK small GTPase pathway signaling. Proc Natl Acad Sci U S A 108:5861–5866
Chertkow Y, Weinreb O, Youdim MB, Silver H (2007) Gene expression changes in peripheral mononuclear cells from schizophrenic patients treated with a combination of antipsychotic with fluvoxamine. Prog Neuropsychopharmacol Biol Psychiatry 31:1356–1362
Duncan CE, Chetcuti AF, Schofield PR (2008) Coregulation of genes in the mouse brain following treatment with clozapine, haloperidol, or olanzapine implicates altered potassium channel subunit expression in the mechanism of antipsychotic drug action. Psychiatr Genet 18:226–239
Earls LR, Fricke RG, Yu J, Berry RB, Baldwin LT, Zakharenko SS (2012) Age-dependent microRNA control of synaptic plasticity in 22q11 deletion syndrome and schizophrenia. J Neurosci 32:14132–14144
Fineberg AM, Ellman LM (2013) Inflammatory cytokines and neurological and neurocognitive alterations in the course of schizophrenia. Biol Psychiatry 73:951–966
Fisar Z, Raboch J (2008) Depression, antidepressants, and peripheral blood components. Neuro Endocrinol Lett 29:17–28
Frydecka D, Misiak B, Beszlej JA, Karabon L, Pawlak-Adamska E, Tomkiewicz A, Partyka A, Jonkisz A, Kiejna A (2013) Genetic variants in transforming growth factor-beta gene (TGFB1) affect susceptibility to schizophrenia. Mol Biol Rep 40:5607–5614
Griffon N, Jeanneteau F, Prieur F, Diaz J, Sokoloff P (2003) CLIC6, a member of the intracellular chloride channel family, interacts with dopamine D(2)-like receptors. Brain Res Mol Brain Res 117:47–57
Haller CS, Padmanabhan JL, Lizano P, Torous J, Keshavan M (2014) Recent advances in understanding schizophrenia. F1000prime reports 6:57. doi:10.12703/P6-57
Han Z, Chen F, Ge X, Tan J, Lei P, Zhang J (2014) miR-21 alleviated apoptosis of cortical neurons through promoting PTEN-Akt signaling pathway in vitro after experimental traumatic brain injury. Brain Res 1582:12–20
Ikeda M et al (2013) Genetic evidence for association between NOTCH4 and schizophrenia supported by a GWAS follow-up study in a Japanese population. Mol Psychiatry 18:636–638
Iyengar BR, Choudhary A, Sarangdhar MA, Venkatesh KV, Gadgil CJ, Pillai B (2014) Non-coding RNA interact to regulate neuronal development and function. Front Cell Neurosci 8:47
Kalkman HO (2009) Altered growth factor signaling pathways as the basis of aberrant stem cell maturation in schizophrenia. Pharmacol Ther 121:115–122
Kanaan Z, Rai SN, Eichenberger MR, Roberts H, Keskey B, Pan J, Galandiuk S (2012) Plasma miR-21: a potential diagnostic marker of colorectal cancer. Ann Surg 256:544–551
Kerns D, Vong GS, Barley K, Dracheva S, Katsel P, Casaccia P, Haroutunian V, Byne W (2010) Gene expression abnormalities and oligodendrocyte deficits in the internal capsule in schizophrenia. Schizophr Res 120:150–158
Lai C-Y, Yu S-L, Hsieh MH, Chen C-H, Chen H-Y, Wen C-C, Huang Y-H, Hsiao P-C, Hsiao CK, Liu C-M (2011) MicroRNA expression aberration as potential peripheral blood biomarkers for schizophrenia. PLoS One 6:e21635
Li BS, Zhao YL, Guo G, Li W, Zhu ED, Luo X, Mao XH, Zou QM, Yu PW, Zuo QF, Li N, Tang B, Liu KY, Xiao B (2012) Plasma microRNAs, miR-223, miR-21 and miR-218, as novel potential biomarkers for gastric cancer detection. PLoS One 7:e41629
Liang Y, Ridzon D, Wong L, Chen C (2007) Characterization of microRNA expression profiles in normal human tissues. BMC Genomics 8:166
Ma X, Zhou J, Zhong Y, Jiang L, Mu P, Li Y, Singh N, Nagarkatti M, Nagarkatti P (2014) Expression, regulation and function of microRNAs in multiple sclerosis. Int J Med Sci 11:810–818
Mar-Aguilar F, Mendoza-Ramirez JA, Malagon-Santiago I, Espino-Silva PK, Santuario-Facio SK, Ruiz-Flores P, Rodriguez-Padilla C, Resendez-Perez D (2013) Serum circulating microRNA profiling for identification of potential breast cancer biomarkers. Dis Markers 34:163–169
Masana M, Santana N, Artigas F, Bortolozzi A (2012) Dopamine neurotransmission and atypical antipsychotics in prefrontal cortex: a critical review. Curr Top Med Chem 12:2357–2374
Miller BH, Wahlestedt C (2010) MicroRNA dysregulation in psychiatric disease. Brain Res 1338:89–99
Miller BJ, Buckley P, Seabolt W, Mellor A, Kirkpatrick B (2011) Meta-analysis of cytokine alterations in schizophrenia: clinical status and antipsychotic effects. Biol Psychiatry 70:663–671
Miller BJ, Culpepper N, Rapaport MH, Buckley P (2013) Prenatal inflammation and neurodevelopment in schizophrenia: a review of human studies. Prog Neuropsychopharmacol Biol Psychiatry 42:92–100
Montalban E, Mattugini N, Ciarapica R, Provenzano C, Savino M, Scagnoli F, Prosperini G, Carissimi C, Fulci V, Matrone C, Calissano P, Nasi S (2014) MiR-21 is an Ngf-modulated microRNA that supports Ngf signaling and regulates neuronal degeneration in PC12 cells. Neuromol Med 16:415–430
Moreno-Moya JM, Vilella F, Simon C (2014) MicroRNA: key gene expression regulators. Fertil Steril 101:1516–1523
Munshi SU, Panda H, Holla P, Rewari BB, Jameel S (2014) MicroRNA-150 is a potential biomarker of HIV/AIDS disease progression and therapy. PLoS One 9:e95920
Nikkila HV, Muller K, Ahokas A, Rimon R, Andersson LC (2001) Increased frequency of activated lymphocytes in the cerebrospinal fluid of patients with acute schizophrenia. Schizophr Res 49:99–105
Noren Hooten N, Abdelmohsen K, Gorospe M, Ejiogu N, Zonderman AB, Evans MK (2010) microRNA expression patterns reveal differential expression of target genes with age. PLoS One 5:e10724
Ohtsuki T, Horiuchi Y, Koga M, Ishiguro H, Inada T, Iwata N, Ozaki N, Ujike H, Watanabe Y, Someya T, Arinami T (2008) Association of polymorphisms in the haplotype block spanning the alternatively spliced exons of the NTNG1 gene at 1p13.3 with schizophrenia in Japanese populations. Neurosci Lett 435:194–197
Pena-Chilet M, Martinez MT, Perez-Fidalgo JA, Peiro-Chova L, Oltra SS, Tormo E, Alonso-Yuste E, Martinez-Delgado B, Eroles P, Climent J, Burgues O, Ferrer-Lozano J, Bosch A, Lluch A, Ribas G (2014) MicroRNA profile in very young women with breast cancer. BMC Cancer 14:529
Peng Y, Xu Y, Cui D (2014) Wnt signaling pathway in schizophrenia. CNS Neurol Disord Drug Targets 13:755–764
Ramachandra RK, Salem M, Gahr S, Rexroad CE 3rd, Yao J (2008) Cloning and characterization of microRNAs from rainbow trout (Oncorhynchus mykiss): their expression during early embryonic development. BMC Dev Biol 8:41
Saha S, Chant D, Welham J, McGrath J (2005) A systematic review of the prevalence of schizophrenia. PLoS Med 2:e141
Schmittgen TD, Livak KJ (2008) Analyzing real-time PCR data by the comparative C(T) method. Nat Protoc 3:1101–1108
Singh KK (2013) An emerging role for Wnt and GSK3 signaling pathways in schizophrenia. Clin Genet 83:511–517
Singh RK, Shi J, Zemaitaitis BW, Muma NA (2007) Olanzapine increases RGS7 protein expression via stimulation of the Janus tyrosine kinase-signal transducer and activator of transcription signaling cascade. J Pharmacol Exp Ther 322:133–140
Smalheiser NR (2014) The RNA-centred view of the synapse: non-coding RNAs and synaptic plasticity. Philos Trans R Soc Lond Ser B Biol Sci 369(1652). doi:10.1098/rstb.2013.0504
Smigielska-Czepiel K, van den Berg A, Jellema P, Slezak-Prochazka I, Maat H, van den Bos H, van der Lei RJ, Kluiver J, Brouwer E, Boots AM, Kroesen BJ (2013) Dual role of miR-21 in CD4+ T-cells: activation-induced miR-21 supports survival of memory T-cells and regulates CCR7 expression in naive T-cells. PLoS One 8:e76217
Song HT, Sun XY, Zhang L, Zhao L, Guo ZM, Fan HM, Zhong AF, Niu W, Dai YH, Zhang LY, Shi Z, Liu XP, Lu J (2014) A preliminary analysis of association between the down-regulation of microRNA-181b expression and symptomatology improvement in schizophrenia patients before and after antipsychotic treatment. J Psychiatr Res 54:134–140
Sun XY, Lu J, Zhang L, Song HT, Zhao L, Fan HM, Zhong AF, Niu W, Guo ZM, Dai YH, Chen C, Ding YF, Zhang LY (2014) Aberrant microRNA expression in peripheral plasma and mononuclear cells as specific blood-based biomarkers in schizophrenia patients. J Clin Neurosci. doi:10.1016/j.jocn.2014.08.018
Tang D, Shen Y, Wang M, Yang R, Wang Z, Sui A, Jiao W, Wang Y (2013) Identification of plasma microRNAs as novel noninvasive biomarkers for early detection of lung cancer. Eur J Cancer Prev 22:540–548
Vargas HE, Gama CS, Andreazza AC, Medeiros D, Stertz L, Fries G, Palha J, Cereser KM, Berk M, Kapczinski F, Belmonte-de-Abreu PS (2008) Decreased serum neurotrophin 3 in chronically medicated schizophrenic males. Neurosci Lett 440:197–201
Wang Y, Gao X, Wei F, Zhang X, Yu J, Zhao H, Sun Q, Yan F, Yan C, Li H, Ren X (2014) Diagnostic and prognostic value of circulating miR-21 for cancer: a systematic review and meta-analysis. Gene 533:389–397
Wilcox JA, Quadri S (2014) Replication of NTNG1 association in schizophrenia. Psychiatr Genet 24:266–268
Wittmann J, Jäck H-M (2010) Serum microRNAs as powerful cancer biomarkers. Biochim Biophys Acta Rev Cancer 1806:200–207
Xu B, Karayiorgou M, Gogos JA (2010) MicroRNAs in psychiatric and neurodevelopmental disorders. Brain Res 1338:78–88
Xu J, Zhao J, Evan G, Xiao C, Cheng Y, Xiao J (2012) Circulating microRNAs: novel biomarkers for cardiovascular diseases. J Mol Med 90:865–875
Yao R, Ma Y, Du Y, Liao M, Li H, Liang W, Yuan J, Ma Z, Yu X, Xiao H, Liao Y (2011) The altered expression of inflammation-related microRNAs with microRNA-155 expression correlates with Th17 differentiation in patients with acute coronary syndrome. Cell Mol Immunol 8:486–495
Zhang Z, Zha Y, Hu W, Huang Z, Gao Z, Zang Y, Chen J, Dong L, Zhang J (2013) The autoregulatory feedback loop of microRNA-21/programmed cell death protein 4/activation protein-1 (MiR-21/PDCD4/AP-1) as a driving force for hepatic fibrosis development. J Biol Chem 288:37082–37093
Zhuo Y, Gao G, Shi JA, Zhou X, Wang X (2013) miRNAs: biogenesis, origin and evolution, functions on virus-host interaction. Cell Physiol Biochem 32:499–510
Author information
Authors and Affiliations
Corresponding authors
Additional information
Hui-min Fan, Xin-yang Sun, and Wei Niu agreed to share the first authorship position together.
Hui-min Fan, Xin-yang Sun and Wei Niu contributed equally to this work.
Rights and permissions
About this article
Cite this article
Fan, Hm., Sun, Xy., Niu, W. et al. Altered microRNA Expression in Peripheral Blood Mononuclear Cells from Young Patients with Schizophrenia. J Mol Neurosci 56, 562–571 (2015). https://doi.org/10.1007/s12031-015-0503-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12031-015-0503-z