Abstract
AmpC β-lactamase is a cephalosporinase, which exhibits resistance against all existing β-lactam antibiotics except carbapenems. Their occurrence in many bacterial pathogens poses a threat to public health and is a growing concern in the medical world. The ampC gene is highly inducible in the presence of β-lactam antibiotics and can be expressed in high levels due to mutation. This inducible expression is regulated by several functional genes. Several studies on functional relationship of these genes and its resistance mechanisms are carried out but it still lacks comprehensible evidences. Thus, in our current study, we used computational gene networks to analyze ampC gene. Based on its interaction type, co-expression, Gene Ontology, and text mining, a functional interaction network is constructed. Around 247 functional genes in 15 different bacterial genus have a functional association with ampC gene. It is predicted that 19.8 % ampD, 13.3 % frdD, 8.5 % gcvA, 2.4 % ampR, and 55.7 % of other functional partners are associated with ampC gene. Our present study provides a glimpse about the functional gene network of ampC gene and also provides the integrated evidence for ampC gene in regulating the β-lactamase production and its role in antibiotic resistance.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Over decades, the β-lactam antibiotics are being widely used to treat the bacterial infections. However, the resistance shown by the bacterial pathogens toward the β-lactam antibiotics is one of the serious problems worldwide [1]. Bacteria persist several modes of resistance and one of the most important modes is the synthesis of β-lactamase [2]. β-lactamase hydrolyzes the β-lactam ring of β-lactam antibiotics and makes it ineffective [3, 4].
The amp genes are first discovered in Enterobacter cloacae [5]. Among various amp genes, the ampC gene is chromosomally encoded to produce cephalosporinase which exhibits resistance to a wide variety of β-lactam antibiotics including penicillin, narrow and broad spectrum cephalosporins, and also β-lactamase inhibitors [6]. AmpC β-lactamase is placed in “class C” of Ambler molecular classification and “Group 1” of Bush functional classification. This is the first reported bacterial enzyme that destroyed penicillin [7]. Due to their inducibility and expression in response to certain β-lactam antibiotics, AmpC β-lactamase is a clinically important enzyme [8]. AmpC β-lactamases are present in Gram-negative bacteria including in human pathogens such as Acinetobacter spp., Aeromonas sp., Citrobacter freundii, Enterobacter sp., Escherichia coli, Pseudomonas aeruginosa, and Yersinia enterocolitica [9, 10]. Chromosomal AmpC β-lactamases are inducible. Plasmid-mediated AmpC β-lactamases are formed through the transfer of chromosomal genes and having similar substrate profile [11, 12]. Thus, it is a major threat for the successful treatment of bacterial infections.
The presence of β-lactam antibiotics or gene mutation can induce the expression level from 100 to 1,000 fold [13, 14] resulting in the β-lactam antibiotic resistance [15]. This inducible expression of AmpC β-lactamase is regulated by several regulatory genes through cell wall recycling pathway along with ampC gene. Several studies reveal that ampC induction pathway requires three major amp gene products namely, the AmpG permease, the AmpD cytoplasmic amidases, and the transcriptional regulator AmpR [5, 14, 16–18].
In the present study, in silico functional interaction network analysis of ampC gene is done using various integrated evidence-based approaches such as network, pathway, functional enrichment analysis, and multiple sequence analysis.
Network visualization is done to identify the ampC gene, associated mutations, and its functional partners’ role in antibiotic resistance through regulation of β-lactamase production.
Gene networks generally depict a large number of interactions. They provide information on the physiological state of an organism. Interaction types can be studied by the construction of biochemical networks at various levels. Significant biological information can be extracted from the literature mining [19, 20]. This ampC gene network study provides the knowledge on associated genes/expressed proteins which are involved in regulation of ampC gene expression and in the synthesis of β-lactamases. The constructed ampC gene network also provides a valuable insight about the associated functional partners and their interactions in the regulation of β-lactamases production.
Materials and Methods
STRING Network Analysis
We used “search tool for the retrieval of interacting genes/proteins” (STRING 9.0) for the study, a pre-computed database resource, involved in the analysis of gene/protein interactions. Gene was represented as “node”, while the interactions between any two genes/proteins were represented as an “edge”. There were direct (physical) and indirect (functional) interactions/associations. The associations were derived from various sources such as high-throughput experimental data, mining of databases, literature, and analyses of co-expressed genes. Interactions between target gene and their closely related functional partners in the network were determined as combined confidence score. STRING provides a probabilistic confidence score for all associations. The scores were given by comparing the group of associations with manually created classification scheme of KEGG database. Each score represents a given association that provides information about the functional linkage between two proteins, i.e., least specific between a pair of proteins annotated in the same pathway. Majority of different scores of interaction or associated data from STRING were highlighted separately; in addition, a combined score is also calculated when the support for a given association is more than one. Combined score indicates the higher confidence. The confidence score values ranged from the lowest to highest. The highest confidence score was in the range of 0.9–1.0, high confidence score was of 0.7–0.9, medium confidence score was of 0.4–0.7, and low confidence score was up to 0.4 [21–27].
Pathway Databases and Sources
Penicillin and cephalosporin biosynthesis pathway for associated genes and other related information was retrieved from Kyoto Encyclopedia of Genes and Genomes (KEGG) database. Protein functions were described in the metabolic pathway database of KEGG [28].
Functional Enrichment Analysis by Gene Ontology
Gene Ontology (GO) and annotations were collected from the UniProt [29–31]. STRING-based GO was grouped by using the p value. The p values and functional annotations such as biological process, molecular function, and cellular component for functional partners were extracted [21–27].
Multiple Sequence Analysis
The molecular evolutionary genetics analysis software [MEGA] (version 4) was used for multiple sequence alignment [32, 33]. The MEGA was an integrated tool designed for comparative analysis of gene sequences and inferring phylogenetic trees by estimating the rates of molecular evolution. All AmpC protein sequences are subjected to ScanPROSITE web server [34] and Motif search tool [PROSITE pattern and ProDom] [35–38] to identify the pattern present in the sequences.
Construction of Gene Interaction Network
Graphical network model was generated by using Cytoscape software. Cytoscape was a free software package used for visualizing, modeling, and analyzing molecular & genetic interaction networks. It supports several algorithms for the layout of networks such as spring-embedded layout, hierarchical layout, and circular layout. The large network can be visualized using Cytoscape version 2.8.3 [39].
Results
Network Analysis of ampC Gene Using STRING
This network analysis on ampC gene provides a clear view on the mechanism of the functional genes in β-lactamase induction. The association of ampC gene with other functional genes/proteins is analyzed using STRING tool. Taking only the highest (0.9–1.0) and high (0.7–0.9) confidence score values into consideration, 15 different bacterial genus and 247 functional genes/proteins are filtered. Hence, we preferred these organisms for further analysis.
The results reveal that, among the functional genes, 21.9 % (54) share the highest confidence score and 78.1 % (193) share high confidence score. Among them, 65.1 % (161) genes are directly interacting, while 34.8 % (86) functional partners are indirectly interacting (sub network) with ampC gene by 34 interconnecting genes. Around 15 bacterial genus (human and nonhuman pathogens) included in network are Escherichia, Legionella, Pseudomonas, Rickettsia, Salmonella, Shewanella, Shigella, Yersinia, Acinetobacter, Aeromonas, Burkholderia, Mycobacterium, Vibrio, Enterobacter, and Laribacter. It is identified that out of 247 functional partners, 49 (19.8 %) are for ampD, 33 (13.3 %) are for frdD, 21(8.5 %) are for gcvA, 6 (2.4 %) for ampR, and 138 (55.7 %) for other genes, respectively. The results are listed in Table 1, 2 and Supplementary Table 1 which also contains descriptions of the functional partners collected from UniProt and National Center for Biotechnology Information (NCBI). Graphical model of ampC gene network and overall percentage of the functional partners are represented in Figs. 1 and 2.
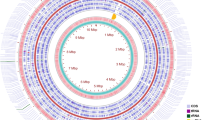
a A graphical illustration of gene network represents the interaction between the target gene and functional partners (gene/protein). b [Enlarge view] Hexagon shape in center indicates the AmpC gene (target gene) and interacting functional partners (color nodes). Interconnecting genes (sub network) are represented in triangle shape. Genes and gene products involved in GO terms (blue color diamond shape nodes) and KEGG pathways (yellow color round rectangle shape nodes) are highlighted (Color figure online)
Pathway Enrichment Analysis for Functional Partners
KEGG pathway enrichment analysis is carried out for all 247 functional partners. The results indicate that the most of genes and gene products shared a common two-component system pathway [signal transduction systems (reference pathway ko02020)]. A majority of associated genes such as ampD, frdD, gcvA, and ampR are involved in cell wall recyclic pathway.
The next significant pathway is penicillin and cephalosporin biosynthesis pathway (reference pathway ko00311). From penicillin and cephalosporin biosynthesis pathway, it is found that the 30 functional partners are involved in the two reactions i.e., K01467 and K01434 (ko00311), respectively. 24 out of 30 genes are involved in β-lactamase synthesis (K01467) which is responsible for β-lactam resistance. The remaining 6 genes are involved in penicillin amidases synthesis (K01434). The list of functional partners is provided in Table 3. The functional partners are highlighted (yellow color round rectangle shape) in Fig. 1. Cell wall recyclic pathway and penicillin & cephalosporin biosynthesis pathway are represented in Figs. 3 and 4.
AmpC beta-lactamase induction. a In cell wall recycling pathway, peptidoglycan releases the murein and get degraded into GlcNAc-anhMurNAc-tripeptide in periplasmic space. AmpG permease transports the muropeptide, GlcNAc-anhMurNAc-tripeptide into cytoplasm. Cytosolic enzyme N-β-acetylglucosaminidase cleaves GlcNAc-anhMurNAc-tripeptide into GlcNAc and anhMurNAc-tripeptide. AmpD produces amidases that hydrolyze the anhMurNAc-tripeptide into anhMurNAc and tripeptide. Then, tripeptide re-enter into the murein biosynthesis. b Mutation in ampD leads to accumulate large amounts of anhMurNAc-tripeptide in cytoplasm and it acts as signaling molecules for transcriptional regulator AmpR which in turn triggers the beta-lactamase production. The muropeptides can induce the beta-lactamase expression by binding with the regulator AmpR
GO Enrichment Analysis for Functional Partners
GO enrichment analysis is performed using STRING tool and UniProt database. 607 GO terms which are involved in cellular component, biological, and molecular function of 247 functional partners are collected. Out of 607 GO terms, 264 are in molecular function, 262 are in biological process, and 81 are in cellular component, respectively. However, for 21 functional partners, GO terms are unavailable. These functional partners are not included for further analysis.
Among all GO terms, the functional genes (264) which are involved in molecular function are 61 are for N-acetylmuramoyl-l-alanine amidase activity [GO:0008745], 29 are for sequence-specific DNA-binding transcription factor activity [GO:0003700], 28 are for beta-lactamase activity [GO:0008800], 25 are for DNA-binding [GO:0003677], 17 are for penicillin-binding [GO:0008658], 7 are for penicillin amidase activity [GO:0008953], 3 are for hydrolase activity, acting on carbon–nitrogen (but not peptide) bonds, in linear amides [GO:0016811], and 94 are for other molecular processes, respectively.
The functional genes (262) which are involved in biological process are as follows: 51 are for peptidoglycan catabolic process [GO:0009253], 35 are for transcription, DNA-dependent [GO:0006351], 33 are for fumarate metabolic process [GO:0006106], 17 are for antibiotic catabolic process [GO:0017001], 11 are for response to antibiotic [GO:0046677], 9 are for beta-lactam antibiotic catabolic process [GO:0030655], and 106 are for other biological processes, respectively.
Furthermore, the functional genes which are involved in cellular component are as follows: 30 are for integral to membrane [GO:0016021], 30 are for plasma membrane [GO:0005886], 13 are for cytoplasm, and 8 are for other cellular components, respectively.
The p value for the GO terms is obtained from STRING dataset. 94 GO terms have statistically significant values (p value ≤ 0.05). The significant GO terms (94) are then compared with complete GO term list (607). Interestingly, it is observed that 8 sets of GO terms have significant p value, remaining 7 sets of GO terms are found to be insignificant (p value ≥ 0.05). In spite of statistical criteria of insignificance, by taking GO terms also as one of the criteria, they are considered as functionally important in β-lactamase synthesis.
Based on p value (≤0.05), the percentage for the predominant functions in the network is calculated which is in the order of 29.8 % for peptidoglycan catabolic process [GO:0009253], 28.9 % for N-acetylmuramoyl-l-alanine amidase activity [GO:0008745], 18.1 % for beta-lactamase activity [GO:0008800], 11.7 % for antibiotic catabolic process [GO:0017001], 8.5 % for integral to membrane [GO:0016021], and 3.1 % for oxidoreductase activity [GO:0016491]. Apart from the above significant percentage values, other processes like DNA-binding [GO:0003677], transferase activity [GO:0016740], aromatic amino acid family metabolic process [GO:0009072], plasma membrane [GO:0005886], sequence-specific DNA-binding transcription factor activity [GO:0003700], carbohydrate metabolic process [GO:0005975], anaerobic respiration [GO:0009061], fermentation [GO: 0006113], electron carrier activity [GO:0009055], and hydrolase activity, acting on carbon–nitrogen (but not peptide) bonds, in linear amides [GO:0016811] are also involved in minimal percentages.
The results reveal that more than one function is shared by most of the associated genes. The associated genes in network are highly involved in regulating the expression of AmpC β-lactamase via, the three processes cellular component, biological process, and molecular function. List of annotated functional partners with significant (p value (≤0.05)) and insignificant GO terms is depicted in Table 4. The functional partners are highlighted (blue color diamond shape) in Fig. 1.
Multiple Sequence Analysis
Around 61 AmpC protein sequences of Acinetobacter sp., Burkholderia cenocepacia, Burkholderia xenovorans, Enterobacter sp. 638, E. coli, E. fergusonii, P. aeruginosa, P. entomophila, P. putida, R. felis, Salmonella enterica Choleraesuis, Shewanella sp., Shigella sp., Legionella pneumophila Corby, Legionella pneumophila Philadelphia, and Laribacter hongkongensis are retrieved from UniProt for the analysis. MEGA software is used for multiple alignments to explore the associated features among AmpC proteins at sequence level. Motif search tool [PROSITE pattern and ProDom] is used to scan the pattern in AmpC β-lactamase. The characteristic motif for class C hydrolase beta-lactamase is found in region having residues 1–58, β-lactamase class-C active site is found in region having residues 91–98, and beta-lactamase hydrolase cephalosporinase precursor signal plasmid porin is found in region having residues 238–390, respectively. These functionally conserved residues as well as probable substitutions are analyzed from multiple alignments.
Discussion
Network Analysis of ampC Gene Using STRING
From the STRING results, based on combined confidence score, 247 associated genes from 15 different genus sharing interaction with ampC gene are selected. STRING utilizes an exclusive scoring framework. The scoring system is based on benchmarks of different types of associations against a common reference set to produce a single confidence score per prediction [40]. In this network, both target gene and their functional partners are represented as “nodes”. They are connected by “colored edges” such as blue for co-occurrence, black for co-expression, deep pink for experiments, sky blue for databases, and green for text mining [21–27]. The probability of interaction between any two nodes indicates the strength of their functional relationship. It also reveals the possibility of genes to operate in the same or similar pathways. In addition to 30 genes which are identified from KEGG database (KEGG ko00311), GO for each functional partner (both human pathogen and nonhuman pathogenic bacteria) is extracted. Most of these associated genes or proteins in the network are directly or indirectly involved in the β-lactamase induction by regulating the expression of ampC gene. Overall results suggest that 19.8 % ampD, 13.3 % frdD, 8.5 % gcvA, 2.4 % of ampR, and 55.7 % functional partners are highly associated with ampC gene and it is represented in Fig. 2.
The results also reveal that the ampD (19.8 %) gene is in close interaction with ampC gene particularly in E. coli, P. aeruginosa, Acinetobacter ADP1, Y. enterocolitica, and Shigella sp. This ampD gene plays a vital role as a negative regulator of AmpC expression [41–44], ampD encodes for N-acetyl-anhydromuranmyl-l-alanine amidase [GO:0008745] and peptidoglycan catabolic process [GO:0009253] which are involved in peptidoglycan recyclic pathway. It also regulates the expression of AmpC β-lactamase [44, 45]. There are four important steps involved in recyclic pathway. They are Step I: ampG encodes an inner membrane permease for GlcNAc-1,6-anhydromuropeptides, which are peptidoglycan catabolites. Step II: GlcNAc-1,6-anhydromuropeptides are transformed to 1,6 anhydromuropeptides by NagZ. Step III: Interaction of 1,6-anhydromuropeptide with the LysR-type transcriptional regulator AmpR, which induces ampC gene to synthesis β-lactamase. Step IV: 1,6-anhydromuropeptides are processed by the N-acetyl-anhydromuramyl-l-alanine amidase ampD which blocks the AmpC induction [46–48]. Moreover, the mutations within the structural gene of ampD can lead to overproduction of AmpC β-lactamase [48, 49] and accumulation of anhydromuramyl pentapeptide, which acts as signal for β-lactamase induction [48, 50].
Subsequently, 2.4 % of ampR (Transcriptional regulator AmpR) and 8.5 % of gcvA regulate AmpC expression (transcription, DNA-dependent [GO:0006351]). This acts as a transcriptional regulator and a positive regulator for gene expression of β-lactamase (ampC). The ampR gene is located adjacent to ampC gene. It is divergently transcribed in C. freundii and E. cloacae, and P. aeruginosa [5]. This AmpR protein activates transcription by binding directly to the upstream promoter region of the ampC DNA. Certain studies also suggest that AmpR induces ampC by binding to anhydro N-acetylmuramyl peptides, which are cytosolic catabolites of peptidoglycan that gets accumulated when exposed to β-lactam antibiotics [51]. In addition, gcvA gene also binds with the ampR-binding region of ampC and activates transcription. Thus, gcvA mimics the activated state of ampR and provides a cross-talk between DNA-binding proteins of different inducible enzyme systems [52].
Another significant interaction is 13.3 % of frdD (fumarate reductase subunit D) which is involved in anchoring the catalytic components [GO:0005886] and fumarate metabolic process [GO:0006106]. Studies suggest that fumarate reductase (frd) operon, in C. freundii is located next to ampC gene which is separated by 1100 base pairs [53]. In another study, promoter for the ampC gene is located within the last gene of the fumarate reductase (frd) operon in E. coli, and ampC attenuator serves as the terminator for transcription of this preceding operon [54]. Although its locus is near to ampC, induction of β-lactamase by this gene still remains unclear.
These five genes ampC, ampR, ampD, frdD, and gcvA regulate the level of transcription both in the presence and absence of β-lactamase inducers. The inducers are the β-lactam antibiotics like cefoxitin and imipenem [15].
The remaining 55.7 % of functional partners are gene products, which exhibit various functions in regulating ampC gene such as N-acetylmuramyl-l-alanine amidase, negative regulator of AmpC, AmpD, beta-lactamase TEM precursor, beta-lactamase, transcriptional factor, LysR family transcriptional regulator, beta-lactamase expression regulator ampD, negative regulator of beta-lactamase expression, transcriptional regulator ampR, peptidase S45, and penicillin amidases that are listed in Table 1 and Supplementary Table 1.
Collectively, the results from STRING conclude that these associated gene and gene products are involved in triggering the ampC gene expressions and in overproduction of β-lactamase resulting in increased resistance to β-lactam antibiotics.
Pathway Enrichment Analysis for Functional Partners
Pathway enrichment analysis reveals that these functional partners are involved in major pathways such as two-component pathway, peptidoglycan recyclic pathway, and penicillin & cephalosporin biosynthesis pathway (KEGG ko00311) (Figs. 3, 4). In various bacterial species, beta-lactam resistance is regulated by two-component system. Two-component pathway comprises signal transduction that enables the adaptability in response to environmental changes [55]. It contains two signal transducers, sensor protein-histidine kinase and the response regulator. The response regulator regulates the gene expression and cellular physiology [56]. Thus, two-component pathway permits the cells to sense and respond by inducing changes in transcription. Studies suggest that overexpression of the response regulators confers resistance to number of chemical compounds and many antibiotics [57]. From the results, it is observed that the functional partners (bla, EFER_1643, PA14_72760, blaOXA-50f, LPC_1045, lpg1618, tem-1, Shew185_0813, Sbal195_0845, Sbal223_0838, Sputcn32_3157, Shewana3_3440, Shewmr4_0694, cphA, ampH, Bmul_6008, Bmul_3689, Bphyt_4009, lpl1405, lpp1588, blaC, PSPA7_6316, Sputw3181_0786, Shewmr7_3328, blaA, oxa-51, and blaTEM) are involved in two-component pathway.
Since, peptidoglycan recyclic pathway in bacteria involves in recycling a significant proportion of the peptidoglycan components of their cell walls, which maintains cell integrity by sustaining internal osmotic pressure and keeps the regular bacterial shape. The recent researches clearly depict that there remains a direct linkage between beta-lactamase induction and cell wall metabolism among Gram-negative bacteria. In specific, muropeptides that are released during peptidoglycan recyclic pathway induce the expression of β -lactamase and hence this recycling pathway may serve as a signaling vehicle, which can be used as an novel target site for developing new antibacterial drugs or in supplementing the existing therapies [58, 59].The majority of functional partners namely ampD, ampR genes, and their gene products are associated with the cell wall recyclic pathway (Fig. 3) [48].
KEGG database results suggest that 30 functional partners are involved in penicillin & cephalosporin biosynthesis pathway (Fig. 4). Among them, 24 functional partners are involved in β-lactamase (K01467) pathway. The analysis depicts that the associated genes stimulate beta-lactamase synthesis which in turn acts on penicillin. In the same pathway, six more genes that are involved in penicillin amidases synthesis (K01434) break the side chain of penicillin and 6-aminopenicillanate is liberated as an end product. This indicates that these reactions are involved in cleavage of β-lactam antibiotics. Since, this penicillin and cephalosporin biosynthesis pathway provides an overview of the diversity of ways that organisms can biosynthesize β-lactam antibiotics [28, 60].
GO Enrichment Analysis for Functional Partners
Enrichment analysis is carried out to gain insights to the functional roles for identified functional partners and to highlight their functional mechanisms at the network level. The enriched GO terms are evaluated specifically for the biological process. The GO terms are extracted from UniProt [29–31] with significant p value.
From the results, several functional genes or proteins having multiple functions are revealed. Among which, seven functional partners (Sputcn32_3157, Shewana3_3440, Shewmr4_0694, Shewmr7_3328, lpp1588, Sbal_3522, and ampH) are involved in all the three functions namely antibiotic catabolic process [GO:0017001], penicillin-binding [GO:0008658], and beta-lactamase activity [GO:0008800].
Apart from these, ten more associated genes are involved in antibiotic catabolic process [GO:0017001], twenty one more functional partners are involved in beta-lactamase activity [GO:0008800], ten functional partners are involved in penicillin-binding [GO:0008658], eight functional partners are involved in beta-lactam antibiotic catabolic process [GO:0030655], and eleven functional partners are involved in response to antibiotic [GO:0046677] (whereas seven functional partners are common in beta-lactam antibiotic catabolic process and response to antibiotic function).
Three genes (ampD, SBO_4393, and A1S_1851) are involved in hydrolase activity, acting on carbon–nitrogen (but not peptide) bonds, in linear amides. Seven genes (SbBS512_E4877, AB57_2186, ABBFA_001601, ABAYE1713, Pmen_1414, Pput_1147, and Bcep18194_B0916) are involved in penicillin amidase activity [GO: 0008953] (Table 4; Fig. 1).
Functional analysis reveals that GO terms of biological and molecular processes are highly represented in ampC gene interaction network. Particularly, the functional partner’s occurrences within selected organisms are more variable. These predictions suggest that these proteins may be indirectly involved in β-lactamase stimulation. A summary of the biological, molecular, and cellular functions of these genes or proteins indicate the functional importance in the organisms. Therefore, the constructed gene or protein network would be helpful in discovering the functional association among the associated proteins [61]. The GO and pathway-based association analysis allow us to expand the knowledge about association of the gene and the mechanisms of network.
Multiple Sequence Analysis
The PROSITE pattern represents “[FY] - E - [LIVM] - G - S - [LIVMG] - [SA] - K” which indicates β-lactamase class-C active site (residues 91–98). ProDom suggested two motifs, first motif belongs to class C hydrolase beta-lactamase (residues 1–58) and second motif represents hydrolase cephalosporinase precursor signal plasmid porin (residues 238–390). Subsequently, we compared PROSITE pattern with the alignment results. The comparison results suggest that Phe (F) and Lys (K) residues are conversed in class C, and AmpC β-lactamase, Lys (K), and Ser (S) seem to play vital role in orientating the active site by electrostatic interaction. Electrostatic interaction between S and K creates a net positive potential in the catalytic site and predicated as the binding site of β-lactam antibiotics [62].
MSA reveals that all strains of E. coli, Shigella sp., Salmonella sp., and Enterobacter sp. strains share the highly conserved sequences and play an important role in the activity of AmpC β-lactamases. The residues that vary from one another in sequences are also identified in strain of Acinetobacter sp., Burkholderia sp., Shewanella sp., Rickettsia sp., Pseudomonas sp., Legionella spp., and Vibrio sp. The results suggest that the amino acid substitution occurred in the active site regions. Residues in the active sites regions are either structurally or functionally important for the β-lactamase activity.
In nutshell, AmpC β-lactamase sequences exhibit variability and only a few residues are conserved. These motifs represent characteristic significance of β-lactamases and can be further developed for better diagnostics and therapy.
Conclusion
This study provides comprehensive evidence on ampC gene and their functionally associated genes in inducing β-lactamase synthesis. The generated ampC gene networks emphasize that β-lactamase induction is accompanied by various regulatory genes. Thus, our study overviewed on the identified genes/proteins from various organisms and their role in regulatory mechanism of β-lactamase induction via biological process, molecular function, cellular process, pathway, and text mining. This constructed gene network provides critical information about functional relationships among biological pathways. It also provides information on the diverse biological process which includes gene functions and complex cellular mechanisms of β-lactamase induction. This would help in better understanding the functions of association partners and their impact on β-lactamase induction. The multiple sequence alignment of AmpC proteins will be useful to study the amino acid variation. To conclude, this study might provide new dimension to understand the regulatory mechanisms of β-lactamases production thereby discovering the inhibitors targeting beta-lactamase induction pathway to prevent the emerging of beta-lactam resistance and improve the efficacy of clinical beta-lactam antibiotics. This study provides useful information for researchers exploring in the field of β-lactamase-mediated antibiotic resistance and also for researchers in drug discovery and development.
Abbreviations
- STRING tool:
-
Search tool for the retrieval of interacting genes/proteins
- KEGG:
-
Kyoto encyclopedia of genes and genomes
- GO:
-
Gene Ontology
- NCBI:
-
National Center for Biotechnology Information
References
Taneja, N., Rao, P., Arora, J., & Dogra, A. (2008). Occurrence of ESBL & Amp-C beta-lactamases & susceptibility to newer antimicrobial agents in complicated UTI. Indian Journal of Medical Research, 127(1), 85–88.
Wilke, M. S., Lovering, A. L., & Strynadka, N. C. (2005). Beta-lactam antibiotic resistance: A current structural perspective. Current Opinion in Microbiology, 8(5), 525–533.
Majiduddin, F. K., Materon, I. C., & Palzkill, T. G. (2002). Molecular analysis of beta-lactamase structure and function. International Journal of Medical Microbiology, 292(2), 127–137.
Abraham, E. P., & Chain, E. (1940). An enzyme from bacteria able to destroy penicillin. Nature, 146, 837.
Balasubramanian, D., Schneper, L., Merighi, M., Smith, R., Narasimhan, G., Lory, S., et al. (2012). The regulatory repertoire of Pseudomonas aeruginosa AmpC ß-lactamase regulator AmpR includes virulence genes. PLoS One, 7(3), 34067–34088.
Bush, K., Jacoby, G. A., & Medeiros, A. A. (1995). A functional classification scheme for β-lactamases and its correlation with molecular structure. Antimicrobial Agents and Chemotherapy, 39(6), 1211–1233.
Jacoby, G. A. (2009). AmpC beta-lactamases. Clinical Microbiology Reviews, 22(1), 161–182.
Moya, B., Juan, C., Alberti, S., Perez, J. L., & Oliver, A. (2008). Benefit of having multiple ampD genes for acquiring beta-lactam resistance without losing fitness and virulence in Pseudomonas aeruginosa. Antimicrobial Agents and Chemotherapy, 52(10), 3694–3700.
Philippon, A., Arlet, G., & Jacoby, G. A. (2002). Plasmid-determined AmpC-type beta-lactamases. Antimicrobial Agents and Chemotherapy, 46(1), 1–11.
Singhal, S., Mathur, T., Khan, S., Upadhyay, D. J., Chugh, S., Gaind, R., et al. (2005). Evaluation of methods for AmpC beta-lactamase in Gram negative clinical isolates from tertiary care hospitals. Indian Journal of Medical Microbiology, 23(2), 120–124.
Thomson, K. S. (2001). Controversies about extended spectrum and AmpC beta-lactamases. Emerging Infectious Diseases, 7(2), 333–336.
Barnaud, G., Arlet, G., Verdet, C., Gaillot, O., Lagrange, P. H., & Philippon, A. (1998). Salmonella enteritidis: AmpC plasmid-mediated inducible β-lactamases (DHA-1) with an ampR gene from Morganella morganii. Antimicrobial Agents and Chemotherapy, 42(9), 2352–2358.
Bagge, N., Ciofu, O., Hentzer, M., Campbell, J. I., Givskov, M., & Hoiby, N. (2002). Constitutive high expression of chromosomal β-lactamase in Pseudomonas aeruginosa caused by a new insertion sequence (IS1669) located in ampD. Antimicrobial Agents and Chemotherapy, 46(11), 3406–3411.
Lodge, J. M., & Piddock, L. J. (1991). The control of class I beta-lactamase expression in Enterobacteriaceae and Pseudomonas aeruginosa. Journal of Antimicrobial Chemotherapy, 28(2), 167–172.
Moya, B., Dotsch, A., Juan, C., Blázquez, J., Zamorano, L., Haussler, S., et al. (2009). Beta-lactam resistance response triggered by inactivation of a nonessential penicillin-binding protein. PLoS Pathogens, 5(3), 1000353–1000363.
Bartowsky, E., & Normark, S. (1993). Interactions of wild-type and mutant AmpR of Citrobacter freundii with target DNA. Molecular Microbiology, 10(3), 555–565.
Lodge, J. M., Busby, S. J. W., & Piddock, L. J. (1993). Investigation of the Pseudomonas aeruginosa ampR gene and its role at the chromosomal ampC beta-lactamase promoter. FEMS Microbiology Letters, 111, 315–320.
Lister, P. D., Wolter, D. J., & Hanson, N. D. (2009). Antibacterial-resistant Pseudomonas aeruginosa: Clinical impact and complex regulation of chromosomally encoded resistance mechanisms. Clinical Microbiology Reviews, 22(4), 582–610.
Natarajan, J., Berrar, D., Dubitzky, W., Hack, C., Zhang, Y., DeSesa, C., et al. (2006). Text mining of full-text journal articles combined with gene expression analysis reveals a relationship between sphingosine-1-phosphate and invasiveness of a glioblastoma cell line. BMC Bioinformatics, 7, 373–389.
Selga, E., Oleaga, C., Ramirez, S., De Almagro, M. C., Noe, V., & Ciudad, C. J. (2009). Networking of differentially expressed genes in human cancer cells resistant to methotrexate. Genome Medicine, 1(9), 83–99.
Snel, B., Lehmann, G., Bork, P., & Huynen, M. A. (2000). STRING: A web-server to retrieve and display the repeatedly occurring neighbourhood of a gene. Nucleic Acids Research, 28(18), 3442–3444.
Von Mering, C., Huynen, M., Jaeggi, D., Schmidt, S., Bork, P., & Snel, B. (2003). STRING: A database of predicted functional associations between proteins. Nucleic Acids Research, 31(1), 258–261.
Von Mering, C., Jensen, L. J., Snel, B., Hooper, S. D., Krupp, M., Foglierini, M., et al. (2005). STRING: Known and predicted protein–protein associations integrated and transferred across organisms. Nucleic Acids Research, 33, 433–437.
Von Mering, C., Jensen, L. J., Kuhn, M., Chaffron, S., Doerks, T., Kruger, B., et al. (2007). STRING 7: Recent developments in the integration and prediction of protein interactions. Nucleic Acids Research, 35, 358–362.
Jensen, L. J., Kuhn, M., Stark, M., Chaffron, S., Creevey, C., Muller, J., et al. (2009). STRING 8: A global view on proteins and their functional interactions in 630 organisms. Nucleic Acids Research, 37, 412–416.
Szklarczyk, D., Franceschini, A., Kuhn, M., Simonovic, M., Roth, A., Minguez, P., et al. (2011). The STRING database in 2011: Functional interaction networks of proteins globally integrated and scored. Nucleic Acids Research, 39(1), 561–568.
Franceschini, A., Szklarczyk, D., Frankild, S. M., Kuhn, M., Simonovic, A., Roth, J., et al. (2013). STRING v9.1: Protein–protein interaction networks, with increased coverage and integration. Nucleic Acids Research, 41, 808–815.
Kanehisa, M. (1997). A database for post-genome analysis. Trends in Genetics, 13(9), 375–376.
Apweiler, R., Bairoch, A., Wu, C. H., Barker, W. C., Boeckmann, B., Ferro, S., et al. (2004). UniProt: The universal protein knowledge base. Nucleic Acids Research, 32(1), 115–119.
UniProt-Consortium. (2010). The universal protein resource (UniProt) in 2010. Nucleic Acids Research, 38, 142–148.
Jain, E., Bairoch, A., Duvaud, S., Phan, I., Redaschi, N., Suzek, B. E., et al. (2009). Infrastructure for the life sciences: design and implementation of the UniProt website. BMC Bioinformatics, 10, 136–154.
Tamura, K., Dudley, J., Nei, M., & Kumar, S. (2007). MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Molecular Biology and Evolution, 24, 1596–1599.
Kumar, S., Stecher, G., Peterson, D., & Tamura, K. (2012). MEGA-CC: Computing core of molecular evolutionary genetics analysis program for automated and iterative data analysis. Bioinformatics, 28, 2685–2686.
De Castro, E., Sigrist, C. J., Gattiker, A., Bulliard, V., Langendijk-Genevaux, P. S., Gasteiger, E., et al. (2006). ScanPROSITE: Detection of PROSITE signature matches and ProRule-associated functional and structural residues in proteins. Nucleic Acids Research, 34, 362–365.
Falquet, L., Pagni, M., Bucher, P., Hulo, N., Sigrist, C. J., Hofmann, K., et al. (2002). The PROSITE database, its status in 2002. Nucleic Acids Research, 30(1), 235–238.
Sigrist, C. J. A., De Castro, E., Cerutti, L., Cuche, B. A., Hulo, N., Bridge, A., et al. (2013). New and continuing developments at PROSITE. Nucleic Acids Research, 41, 344–347.
Corpet, F., Gouzy, J., & Kahn, D. (1999). Recent improvements of the ProDom database of protein domain families. Nucleic Acids Research, 27, 263–267.
Sonnhammer, E. L., & Kahn, D. (1994). Modular arrangement of proteins as inferred from analysis of homology. Protein Science, 3, 482–492.
Shannon, P., Markiel, A., Ozier, O., Baliga, N. S., Wang, J. T., Ramage, D., et al. (2003). Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Research, 13(11), 2498–2504.
Raman, K. (2010). Construction and analysis of protein–protein interaction networks. Automated experimentation, 2, 2–11.
Lindberg, F., Lindquist, S., & Normark, S. (1987). Inactivation of the ampD gene causes semiconstitutive overproduction of the inducible Citrobacter freundii beta-lactamase. Journal of Bacteriology, 169(5), 1923–1928.
Lindquist, S., Lindberg, F., & Normark, S. (1989). Binding of the Citrobacter freundii AmpR regulator to a single DNA site provides both autoregulation and activation of the inducible ampC beta-lactamase gene. Journal of Bacteriology, 171(7), 3746–3753.
Dietz, H., Pfeifle, D., & Wiedemann, B. (1996). Location of N-acetylmuramyl-l-alanyl-d-glutamylmesodiaminopimelic acid, presumed signal molecule for beta-lactamase induction, in the bacterial cell. Antimicrobial Agents and Chemotherapy, 40(9), 2173–2177.
Schmidtke, A. J., & Hanson, N. D. (2008). Role of ampD homologs in overproduction of AmpC in clinical isolates of Pseudomonas aeruginosa. Antimicrobial Agents and Chemotherapy, 52(11), 3922–3927.
Tans-Kersten, J., Gay, J., & Allen, C. (2000). Ralstonia solanacearum AmpD is required for wild type bacterial wilt virulence. Molecular Plant Pathology, 1(3), 179–185.
Zamorano, L., Reeve, T. M., Juan, C., Moya, B., Cabot, G., Vocadlo, D. J., et al. (2011). AmpG inactivation restores susceptibility of pan-beta-lactam-resistant Pseudomonas aeruginosa clinical strains. Antimicrobial Agents and Chemotherapy, 55(5), 1990–1996.
Jacobs, C., Frere, J. M., & Normark, S. (1997). Cytosolic intermediates for cell wall biosynthesis and degradation control inducible beta-lactam resistance in Gram-negative bacteria. Cell, 88(6), 823–832.
Normark, S. (1995). Beta-Lactamase induction in gram-negative bacteria is intimately linked to peptidoglycan recycling. Microbial Drug Resistance, 1(2), 111–114.
Ni, M., & Zhang, D. (2005). Analysis of AmpC beta-lactamase gene in Pseudomonas aeruginosa. Journal of Huazhong University of Science and Technology, 25(1), 17–19.
Jones, R. N. (1998). Important and emerging β-lactamase-mediated resistances in hospital-based pathogens: The Amp C enzymes. Diagnostic Microbiology and Infectious Disease, 31(3), 461–466.
Balcewich, M. D., Reeve, T. M., Orlikow, E. A., Donald, L. J., Vocadlo, D. J., & Mark, B. L. (2010). Crystal structure of the AmpR effector binding domain provides insight into the molecular regulation of inducible ampC beta-lactamase. Journal of Molecular Biology, 400(5), 998–1010.
Everett, M., Walsh, T., Guay, G., & Bennett, P. (1995). GcvA, a LysR-type transcriptional regulator protein, activates expression of the cloned Citrobacter freundii ampC beta-lactamase gene in Escherichia coli: Cross-talk between DNA-binding proteins. Microbiology, 141(2), 419–430.
Lindberg, F., Westman, L., & Normark, S. (1985). Regulatory components in Citrobacter freundii ampC beta-lactamase induction. Proceedings of the National Academy of Sciences of the United States of America, 82(14), 4620–4624.
Grundstrom, T., & Jaurin, B. (1982). Overlap between ampC and frd operons on the Escherichia coli chromosome. Proceedings of the National Academy of Sciences of the United States of America, 79(4), 1111–1115.
Mizuno, T. (1998). His-Asp phosphotransfer signal transduction. Journal of Biochemistry, 123(4), 555–563.
Hoch, J. A., & Varughese, K. I. (2001). Keeping signals straight in phosphorelay signal transduction. Journal of Bacteriology, 183(17), 4941–4949.
Hirakawa, H., Nishino, K., Yamada, J., Hirata, T., & Yamaguchi, A. (2003). Beta-lactam resistance modulated by the overexpression of response regulators of two-component signal transduction systems in Escherichia coli. Journal of Antimicrobial Chemotherapy, 52(4), 576–582.
Zeng, X., & Lin, J. (2013). Beta-lactamase induction and cell wall metabolism in Gram-negative bacteria. Frontiers in Microbiology, 4, 128–137.
Johnson, J. W., Fisher, J. F., & Mobashery, S. (2013). Bacterial cell-wall recycling. Annals of the New York Academy of Sciences, 1277, 54–75.
Martín, J. F., Casqueiro, J., Kosalková, K., Marcos, A. T., & Gutiérrez, S. (1999). Penicillin and cephalosporin biosynthesis: mechanism of carbon catabolite regulation of penicillin production. Antonie van Leeuwenhoek, 75(1–2), 21–31.
Anitha, P., Anbarasu, A., & Ramaiah, S. (2014). Computational gene network study on antibiotic resistance genes of Acinetobacter baumannii. Computers in Biology and Medicine, 48, 17–27.
Singh, R., Saxena, A., & Singh, H. (2009). Identification of group specific motifs in beta-lactamase family of proteins. Journal of Biomedical Science, 16(1), 109–116.
Acknowledgments
A.A and S.R. gratefully acknowledge the Indian council of Medical Research (ICMR), Government of India Agency for the research grant [IRIS ID: 2014-0099]. The authors would also like to thank the management of VIT University for providing the necessary facilities to carry out this research project.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Anitha, P., Bag, S., Anbarasu, A. et al. Gene and Protein Network Analysis of AmpC β Lactamase. Cell Biochem Biophys 71, 1553–1567 (2015). https://doi.org/10.1007/s12013-014-0379-5
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12013-014-0379-5