Abstract
Filamentous fungi are prolific producers of carbohydrate-active enzymes (CAZymes) and important agents that carry out plant cell wall degradation in natural environments. The number of fungal species is frequently reported in the millions range, with a huge diversity and genetic variability, reflecting on a vast repertoire of CAZymes that these organisms can produce. In this study, we evaluated the ability of previously selected ascomycete and basidiomycete fungi to produce plant cell wall-degrading enzyme (PCWDE) activities and the potential of the culture supernatants to increase the efficiency of the Cellic® CTec2/HTec2 for steam-exploded sugarcane straw saccharification. The culture supernatant of Penicillium ochrochloron RLS11 showed a promising supplementation effect on Cellic® CTec2/HTec2, and we conducted the whole-genome sequencing and proteomic analysis for this fungus. The size of the assembled genome was 38.06 Mbp, and a total of 12,015 protein-coding genes were identified. The repertoire of PCWDE-coding genes was comparatively high among Penicillium spp. and showed an expansion in important cellulases and xylanases families, such as GH3, GH6, GH7, and GH11. The proteomic analysis indicated cellulases that probably enhanced the biomass saccharification performance of the Cellic® CTec2/HTec2, which included enzymes from GH3, GH6, and GH7 families.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Fungi are a diverse and large group of eukaryotes, comprising species adapted to almost all terrestrial and aquatic ecosystems [1]. The widespread and multitude of lifestyles of fungi entail a huge genetic variability, even inside the same genus [2]. This diversity is accompanied by biological importance, such as its prominent role in soil carbon cycling, pathogenesis, and plant symbiosis. These features may be highly dependent on the fungal ability to produce and secrete carbohydrate-active enzymes (CAZymes) that must act on highly heterogeneous substrates, and thus, a massive diversity regarding sequence and function is frequently observed for these enzymes [3].
Filamentous fungi significantly diverge in how they decompose lignocellulose, which is a consequence of their genetic background driving the production and secretion of plant cell wall-degrading enzymes (PCWDEs). Soft-rot fungi include ascomycetes and basidiomycetes, mainly employing hydrolases to depolymerize polysaccharides from the plant cell wall. Brown-rot species are basidiomycetes that can produce non-enzymatic oxidative systems, while white-rot fungi are prolific producers of lignin-degrading oxidoreductases [4]. Necrotrophic phytopathogenic fungi generally have an enriched repertoire of PCWDE-coding genes to rapidly and effectively decompose plant polysaccharides, while symbionts and biotrophic fungi generally possess fewer CAZyme-coding genes [5]. Although this classification of fungi provides a glimpse of plant cell wall-degrading capabilities, a closer look into the repertoire of such genes in fungal genomes may reveal considerable variations, even considering closely related species. This knowledge was enriched by many whole-genome sequences available and the comprehensive characterization of exoproteomes by mass spectrometry [6,7,8].
Despite being an industrially attractive process, the use of enzymes for plant biomass saccharification represents a significant proportion of the capital cost to generate value-added products from lignocellulose [9]. In this sense, there is a continuous effort to improve (hemi)cellulolytic cocktails, driven by protein engineering and enzyme screening from wild organisms, aiming to produce or discover more efficient biocatalysts.
The production and commercialization of (hemi)cellulase mixtures for industrial processes are dominated by companies such as Novozymes (Cellic® enzyme cocktails) and DuPont (Accellerase® and Spezyme® enzyme cocktails). Although the composition of commercial enzyme mixtures is not known in detail, they are primarily composed of Trichoderma sp. CAZymes that were possibly improved by protein engineering. The enzyme mixtures probably contain β-glucosidase from Aspergillus spp. and other proteins that enhance lignocellulose conversion obtained from various organisms, such as swollenins and LPMOs from the AA9 family [10].
Besides improving enzymatic cocktails using molecular tools, the inspection of high-performance fungal enzyme systems could provide important information regarding new or more efficient biocatalysts to supplement existing enzyme cocktails.
In this study, we evaluated the ability of filamentous fungi with previous ability to secrete of CAZymes to produce PCWDEs grown on pretreated sugarcane straw and the application of these enzymes in the saccharification of the same biomass. The saccharification assays were conducted using the Cellic® CTec2/HTec2 (Novozymes A/S) supplemented with crude fungal enzyme preparations to assess the ability of fungi to secrete proteins to improve the performance of the commercial enzyme mixture. Our analysis indicated that the culture supernatant of Penicillium ochrochloron strain RLS11 showed a promising supplementation effect, and we conducted whole-genome sequencing of this fungus to uncover its PCWDE-coding gene repertoire. Also, LC-MS/MS analysis was carried out to comprehensively characterize its exoproteome and identify the proteins that provided the good ability to supplement the Cellic® CTec2/HTec2 for sugarcane straw saccharification. The cellulases and xylanases of P. ochrochloron RLS11 identified here are promising targets for future experiments to improve modern commercial CAZyme preparations.
Materials and Methods
Fungal Strains and Culture Conditions for PCWDE Production
In this study, we evaluated the ability of nine pre-selected fungal isolates to produce PCWDEs with the potential to improve saccharification yields of Cellic® CTec2/HTec2. The fungal cultivation periods (10 days for ascomycetes and 15 days for basidiomycetes) were based on a previous screening study, which provided higher PCWDE activities (data not published).
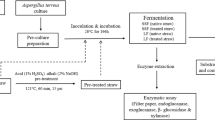
The ascomycete fungi (Trichoderma sp. J7, Penicillium ochrochloron RLS11, and Fusarium verticilliodes AZB) were obtained from the mycological culture collection of the Biochemical Analysis Laboratory, Bioagro, Universidade Federal de Viçosa (Brazil). The basidiomycete fungi (Pleurotus cornucopiae PLOCOR, Pleurotus floridanus PLO09, Pleurotus ostreatoroseus PLO13, Pholiota adiposa KGM, Pholiota nameko PH01, Hericium erinaceus HE) were kindly provided by the Laboratory of Mycorrhizal Association, Bioagro, Universidade Federal de Viçosa (Brazil). These fungal isolates were chosen based on previous screening studies evaluating CAZyme production and activity. Each fungus was cultivated on potato-dextrose-agar (PDA) plates for 7 days at 28 °C in the dark. Ten agar plugs (Ø = 5.0 mm) of PDA-containing mycelia were added to sterile Erlenmeyer flasks (250 mL) containing 100 mL of propagation medium with the following composition: yeast extract 10 g.L−1, peptone 20 g.L−1, and glucose 20 g.L−1. The flasks were kept under agitation (150 rpm) for 7 days at 28 °C in the dark.
For the production of PCWDEs, 5 mL of propagation medium from each fungal culture was added to sterile Erlenmeyer flasks (125 mL) containing 55 mL of mineral solution and 1.2 g of steam-exploded sugarcane straw (dry weight) as the sole carbon source. The mineral solution consisted of NH4NO3 2.0 g.L−1; K2HPO4 2.0 g.L−1; MgSO4 2.0 g.L−1; NaCl 1.0 g.L−1; CaCl2 2.0 g.L−1; citric acid 2.8 g.L−1; yeast extract 3.0 g.L−1; peptone 2.0 g.L−1; and 1.0 mL.L−1 of trace elements solution (EDTA 50.0 g.L−1; MnCl2 0.4 g.L−1; CoCl2 0.16 g.L−1; CuSO4 0.16 g.L−1; H3BO3 1.1 g.L−1; (NH4)6Mo7O24 0.13 g.L−1; FeSO4 0.5 g.L−1; and ZnSO4 2.2 g.L−1). The flasks were kept at 28 °C under the agitation of 250 rpm in the dark. The ascomycetes were grown for 10 days and the basidiomycetes for 15 days. The residual solids were filtered for extracellular enzyme recovery using a nylon filter followed by centrifugation at 15,000g for 10 min at 4 °C. The clarified supernatants were kept at 4 °C, and phenylmethanesulfonyl fluoride (PMSF) was immediately added to a final concentration of 1.0 mM. All experiments were performed in triplicate.
Plant Cell Wall-Degrading Enzyme Activities
Endo-β-1,4-glucanase (CMCase) and β-1,4-xylanase activities were measured with carboxymethyl cellulose (CMC) and beechwood xylan as substrates, respectively. The enzymatic reactions were conducted in test tubes with substrate suspension (1.0% w/v) in sodium acetate buffer (50 mM pH 5.0) and appropriately diluted enzyme preparation. The tubes were incubated at 50 °C for 15 min. The released reducing sugars were quantified using dinitrosalicylic acid reagent [11]. Enzymatic activity units (U) were determined by the correlation between absorbance at 540 nm and known concentrations of the analyte.
β-glucosidase, β-xylosidase, and cellobiohydrolase activities were quantified using 4-nitrophenyl sugar derivatives as substrates (β-D-glucopyranoside, β-D-xylopyranoside, and β-D-cellobioside, respectively). The enzymatic reactions were carried out on substrate suspension (0.5 mmol.L−1) in sodium acetate buffer (pH 5.0) and appropriately diluted enzyme preparation. After 15 min at 50 °C, the reactions were terminated by adding 1 M NaOH. The 4-nitrophenolate was quantified by the correlation between absorbance at 410 nm and known concentrations of the analyte.
Laccase activity was measured by monitoring the oxidation of the ABTS (2,2′-azino-bis(3-ethylbenzothiazoline-6-sulfonic acid)). The enzymatic reactions contained 300 μL of the appropriately diluted enzyme suspension, 600 μL of sodium acetate buffer (50 mM pH 5.0), and 100 μL of 10 mM ABTS. This mixture was incubated for 5 min at 50 °C. After that, the absorbance was taken at 420 nm. Laccase activity was calculated using the Lambert-Beer law, using a molar extinction coefficient of 3.6 × 104 M−1cm−1.
The manganese peroxidase activity (MnP) was carried out using 100 μL of 0.25 mol.L−1 sodium lactate, 50 μL of 2 mM MnSO4, 20 μL of 0.5% (m/v) bovine serum albumin, 100 μL of 0.1% (m/v) phenol red, 500 μL appropriately diluted enzyme suspension and 50 μL of 2 mM H2O2 in sodium succinate buffer (25 mM and pH 4.5). The mixture was incubated for 5 min at 30 °C, and the reaction was ended by the addition of 40 μL of 2 M NaOH. Control assays were conducted in the absence of Mn2+ by omitting the addition of MnSO4 in the reaction mixture. The absorbance was taken at 610 nm, subtracting the value of the control reaction. MnP activity was calculated using the Lambert-Beer law, using a molar extinction coefficient of the oxidized phenol red (0.022 M−1cm−1).
One unit of enzymatic activity was defined as the amount of enzyme that released 1 μmol of product per minute under assay conditions.
Steam-Exploded Sugarcane Straw Saccharification by Cellic® CTec2/HTec2 Supplementend with Crude Fungal Enzyme Preparations
The saccharification assays were conducted based on protein loading since complex fungal exoproteomes might contain poorly active enzymes on model substrates or even proteins that are not enzymes that positively impact biomass saccharification, such as swollenins. To access the effect of the fungal culture supernatants on Cellic® CTec2/HTec2 saccharification performance, we supplemented it with each supernatant individually and quantified the glucose release. The steam-exploded sugarcane straw had a minor amount of hemicelluloses (Table S4), leading to low xylose concentration after saccharification. Thus, the xylose yields were not reported.
For saccharification assays, Cellic® CTec2/HTec2 enzyme mixture (75% of CTec2 and 25% of HTec2) was applied alone and combined with fungal culture supernatants for steam-exploded sugarcane straw saccharification. Reactions were conducted in 25-mL Erlenmeyer flasks with 5 mL of reaction medium consisting of pretreated plant biomass 5.0% (w/v), 0.1 M sodium citrate buffer pH 5.0, and 1 mM sodium azide. The reactions contained 5.00 mg of protein per gram of dry matter (referred to as “Mix”), 5.50 mg of protein per gram of dry matter (“Mix + 10%”) or 6.25 mg of protein per gram of dry matter (“Mix + 25%”). The protein supplementations (Mix +10% and Mix + 25%) were conducted using the commercial mixture itself Cellic® CTec2/HTec2 (control) or the fungal culture supernatants (treatments). A detailed description of the protein loadings of each enzyme mixture used in sugarcane straw saccharification is given in supplementary file 1.
The flasks were incubated at 250 rpm and 50 °C for 72 h. Aliquots were taken and immediately boiled to protein denaturation, centrifuged at 16,000g for 15 min, and filtered using PVDF filters with 0.45 μm pore size. Glucose was detected in a high-performance liquid chromatography system Shimadzu 20A series equipped with Aminex HPX-87H column (300 × 7.8 mm) and a refractive index detector. The elution was performed with H2SO4 5 mM, with a flow rate of 0.6 mL.min−1 and column temperature of 55 °C. The glucose was quantified using a calibration curve that correlates peak areas and known concentrations of the analyte.
The experiment was performed in triplicate for each condition, and the statistical analyses were conducted in the R environment (version 3.5.3) [12]. The T-test was used for the comparison of means. The null hypothesis was defined as the control mean higher than the treatment mean, and the alternative hypothesis consisted of equal means or control mean less than treatment mean. The significance level was 5% for all conclusions.
DNA Extraction, Genome Sequencing, and Assembly
For genomic DNA extraction, P. ochrochloron RLS11 was cultured on PDA plates, and the mycelium was collected with a sterile scalpel and immediately frozen with liquid nitrogen. DNA isolation was carried out according to a method described elsewhere [13].
The genome was sequenced using Illumina Novaseq platform (Illumina Inc., San Diego, CA, USA). The library was prepared using TruSeq Nano DNA prep kit (Illumina Inc., San Diego, CA, USA) with 150 bp pair-end reads and 550 bp fragment length. Raw data filtering was performed with Trimmomatic v0.36 [14] using a 4-mer sliding window and mean Q equal to 15. Filtered reads were used for de novo genome assembly using SPAdes v.3.9.0 [15] using kmer sizes ranging from 27 to 123. The genome completeness was accessed with BUSCO v3 [16], using the Ascomycota and Eurotiomycetes single-copy orthologous genes.
This Whole Genome Shotgun project has been deposited at DDBJ/ENA/GenBank under the accession JAAVMA000000000 and BioProject number PRJNA614650.
Phylogenetic Analysis
Initially, its regions (internal transcribed spacer 1 – 5.8S – internal transcribed spacer 2) were searched in the assembled genome of P. ochrochloron RLS11 using BLASTn searches with sequences from Fungi RefSeq ITS project (NCBI BioProject: PRJNA177353). After identifying the fungal species that provided the best-hit (based on sequence identity), the RNA polymerase II subunit 2 (rpb2) and β-tubulin (tub) sequences were downloaded from Genbank and used in new BLASTn searches to identify these genes in the P. ochrochloron RLS11 assembly.
For phylogenetic analysis, reference sequences of rpb2, tub, and its regions were downloaded from GenBank (Table S1). These sequences were separately aligned with MAFFT v.7.305 [17] using the L-INS-i strategy. The three multiple sequence alignment files generated were combined with a custom Python script.
The jmodeltest v.2.1.10 [18] was used to infer the best substitution model under Akaike information criterion. Phylogenetic reconstruction using the maximum likelihood method was carried out in RaxML v.8.2.12 [19] using the GTR+I+G model with 2000 searches for the best-scoring tree. Maximum likelihood bootstrap proportions (MLBP) were calculated with 5000 replicates. Bayesian inference was carried out with MrBayes v.3.2.7 [20] using Markov Chain Monte Carlo (MCMC) algorithm. Six independent runs with four chains each were conducted for 2,000,000 generations, sampling every 100th generation. The initial 25% trees were discarded as burn-in phase, and the remaining trees were used for estimating Bayesian inference posterior probability (BIPP) values. Trees were visualized in FigTree v.1.4.4 [21].
Gene Prediction and Annotation
A combination of ab initio and similarity-based methods were applied to predict protein-coding genes in the P. ochrochloron RLS11 genome assembly. Augustus v3.2.2 [22] trained with Penicillium chrysogenum NRRL 1951 gene parameters and GeneMark-ES v4.38 [23] using self-training mode were used for ab initio gene predictions. The similarity-based gene predictions were based on RNA-seq data from Penicillium species downloaded from NCBI Sequence Read Archive (Table S2). Transcriptome assembly was conducted with Tophat/Cufflinks pipeline [24], and the coding sequences were used as input for Exonerate v.2.2.0 [25], which predicted its most likely position in the P. ochrochloron RLS11 assembly using spliced-sequence alignments. The EvidenceModeler v.1.1.1 [26] was used to merge the genes predicted by ab initio and similarity-based methods applying a weighted consensus metric.
Carbohydrate-active enzymes annotation was carried out with dbCAN2 [27] using HMMER, DIAMOND, and Hotpep predictions. Only CAZy families predicted by at least 2 tools were kept in the data set. The dbCAN2 predictions were refined with BLASTp (cutoff values: identity > = 60% and coverage > = 80%) and InterProScan best-hits (e-value < = 1e–30) aiming to annotate enzymes at activity level. This prediction was also applied to other Penicillium genomes used in this work (Table S3).
Mass Spectrometry-Based Proteomics of Penicillium Ochrochloron RLS11 Culture Supernatants
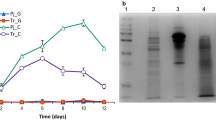
Proteins of the 5-day and 10-day culture supernatants were precipitated with ethanol (90% v/v) for 24 h at −20 °C. After centrifugation (9000 × g, 20 min, and 4 °C), the protein pellet was resuspended in 100 μL of a solution containing 7.0 M urea, 2.0 M thiourea, and 4.0% (w/v) CHAPS. A SDS-PAGE was conducted with 30 μg of proteins, and the run was stopped when proteins migrated from stacking gel to resolving gel. The unique protein bands were excised, discolored with methanol 50% and acetic acid 5.0%, treated with 0.05 mol.L−1 dithiothreitol and 0.10 mol.L−1 iodoacetamide, and subjected to in-gel trypsinization with 2 μg of trypsin (Trypsin Gold, Promega™). The peptides were desalted using stage-tips [28], dried in a vacuum, and reconstituted in 0.1% formic acid. One microliter of the peptide-containing solution was injected on PicoFrit Column (20 cm × ID75 μm, 5 μm particle size, New Objective) and analyzed on an Orbitrap Velos mass spectrometer (Thermo Fisher Scientific, Waltham, MA, USA) connected to the EASY-nLC system (Proxeon Biosystem, West Palm Beach, FL, USA). Protein identification was performed with MaxQuant v.1.6.3.3 [29] using a database of predicted protein sequences from P. ochrochloron RLS11, P. oxalicum 114-2, P. brasilianum, and P. rubens Wisconsin 54-1255. Quantification was performed using the label-free quantification (LFQ) method. A false discovery rate (FDR) of 1% and a minimum of 2 unique peptides were used as identification confidence parameters.
The experiment was conducted in triplicate for each condition (5-day and 10-day supernatants), and statistical and exploratory data analyses were conducted in the R environment (version 3.5.3) [12]. The T-test was used to compare the means (protein relative abundance). The null hypothesis was defined as equal means and the alternative hypothesis as different means. The significance level was 5% for all conclusions.
Results
Plant Cell Wall-Degrading Enzyme Activities in Fungal Culture Supernatants
Overall, the culture supernatants of basidiomycetes showed less cellulase and xylanase activities than the ascomycetes (Table 1). All basidiomycetes produced laccase activity, but only Pleurotus floridanus PLO09 produced manganese peroxidase in detectable amounts. This fungus and the Pleurotus ostreatoroseus PLO13 showed higher PCWDEs activities than other basidiomycetes evaluated in this work.
Regarding the ascomycetes, the Trichoderma sp. J7 produced a high xylanolytic activity, several orders of magnitude higher than the cellulase activities, while F. verticillioides AZB provided a more balanced set of cellulase and xylanase activities. In turn, the Penicillium ochrochloron RLS11 was by far the best producer of cellulases specific activities (CMCase, β-glucosidase and cellobiohydrolase), as well as a high xylanolytic specific activity. Hence, this fungus produced the highest proportion of proteins that are active enzymes, or enzymes with higher efficiency to catalyze reactions on model substrates. Therefore, the P. ochrochloron RLS11 showed the greatest potential for plant biomass saccharification among the fungi evaluated.
Steam-Exploded Sugarcane Straw Saccharification by Cellic® CTec2/HTec2 Supplemented with Fungal Culture Supernatants
Overall, the culture supernatants of basidiomycetes promoted a lower glucose release from steam-exploded sugarcane straw compared to the Cellic® CTec2/HTec2 alone, especially of Pholiota nameko PH01 and Pholiota adiposa PKGM (Fig. 1A). Increasing the protein loads from these fungi in the saccharification medium had a greater detrimental effect on Cellic® CTec2/HTec2 since it was detected with lower glucose concentration relative to the control (less protein loading — Mix). The culture supernatant of Pleurotus ostreatoroseus PLO13 was the only from basidiomycetes that showed a promising result at 10% supplementation, providing a glucose release equal to or greater than the control supplemented with the commercial enzyme mixture itself at the same protein loading (Mix + 10%). Nevertheless, the same effect was not observed for 25% supplementation (Mix + PLO13 25%). Thus, these results indicated that all evaluated basidiomycetes have poor ability to improve the commercial enzymatic mixture for saccharification.
Glucose concentration (g.L−1) in the reaction medium of steam-exploded sugarcane straw after saccharification. The reaction was conducted with 5% (w/v) dry matter for 72 h at 50 °C under 250 rpm agitation. The “Mix” refers to saccharification containing by the Cellic® CTec2/HTec2 alone (3:1 ratio) with 5.0 mg of protein per gram of dry matter (5 mg/g DM). The “Mix” was supplemented based on protein loading with the Cellic® CTec2/HTec2 itself or with the culture supernatants of basidiomycetes (15 days of cultivation) and ascomycetes (10 days of cultivation). A detailed description of the protein loadings of each enzyme mixture used in sugarcane straw saccharification is given in supplementary file 1. The symbol * represents p-value < 0.05
Similarly, the culture supernatants of Trichoderma sp. J7 and Fusarium verticilliodes AZB provided significant results at 10% protein supplementation but a lower glucose release when protein load was increased (25% protein supplementation) (Fig. 1B). At first glance, these crude enzymatic preparations have no components that enhance the efficiency of Cellic® CTec2/HTec2 for steam-exploded sugarcane straw saccharification.
On the other hand, the secretome of Penicillium ochrochloron RLS11 showed a promising result. Adding this fungal culture supernatant to the Cellic® CTec2/HTec2 mixture led to positive supplementation effects on both protein loading levels (10% and 25%), i.e., glucose release compatible with the commercial mixture alone at same protein loading (Fig. 1B). During growth, fungi generally produce and secrete proteins that do not participate in the biomass saccharification or even show a negative effect, such as proteases. Thus, the above results provided evidence that the Penicillium ochrochloron RLS11 secretome contains components that increase the saccharification performance of the Cellic® CTec2/HTec2.
Steam-Exploded Sugarcane Straw Saccharification by Cellic® CTec2/HTec2 Supplemented with Penicillium Ochrochloron RLS11 Culture Supernatants
To confirm the potential of P. ochrochloron RLS11 to improve the Cellic® CTec2/HTec2, we reconducted enzymatic saccharification assays supplementing the commercial mixture with culture supernatants of P. ochrochloron RLS11 obtained after 10 days of cultivation, as in the previous saccharification assay, and culture supernatants obtained in other cultivation periods (3, 5, and 7 days). The 10-day culture supernatant yielded the same result as the previous assay, but the other culture supernatants did not show the same supplementation effect (Fig. 2).
Glucose concentration (g.L−1) in the reaction medium of steam-exploded sugarcane straw after saccharification. The reaction was conducted with 5% (w/v) dry matter for 72 h at 50 °C under 250 rpm agitation. The “Mix” refers to saccharification containing by the Cellic® CTec2/HTec2 alone (3:1 ratio) with 5.0 mg of protein per gram of dry matter (5 mg/g DM). The “Mix” was supplemented based on protein loading with the Cellic® CTec2/HTec2 itself or with the culture supernatants of Penicillium sp. RLS11 (3-day, 5-day, 7-day, and 10-day culture supernatants). A detailed description of the protein loadings of each enzyme mixture used in sugarcane straw saccharification is given in supplementary file 1. The symbol * represents p-value < 0.05
This result motivated the conduction of proteomic analysis of the Penicillium RLS11 culture supernatants to gain insights into which proteins promote the good supplementation effect. Also, the genome sequencing of this fungus was interesting to gain knowledge on the genetic basis of its promising ability to supplement commercial enzyme mixtures and predict the sequence of genes to aid protein identification by proteomic analysis.
Penicillium Ochrochloron RLS11 Genome and Protein-Coding Gene Prediction
The P. ochrochloron RLS11 genome sequencing generated 71,026,620 reads with 151 base pairs (bp) each, totalizing 10,725,019,620 bp. After quality control and raw data filtering, 69,872,948 reads and 10,550,815,148 bp were kept in the data set, representing 98.4% of the total data generated. The assembled genome contained 991 contigs in 354 scaffolds (minimum length equal to 500 bp), with the largest scaffold of 2094 Kbp. The N50 and L50 parameters were 832,608 bp and 16, indicating a good genome assembly (Table 2). The total size of the assembled genome was approximately 38 Mbp, featuring in the upper range of known genome sizes of Penicillium species (https://mycocosm.jgi.doe.gov) [30].
We employed ab initio and similarity-based approaches to predict protein-coding genes in the P. ochrochloron RLS11 assembly. The final consensus gene set comprised 12,015 protein-coding genes, and a search for Ascomycota and Eurotiomycetes universal single-copy orthologs with BUSCO v3 yielded 99.2% and 97.7% of completeness, respectively, indicating good genome assembly and gene content.
Phylogenetic Analysis
Preliminary searches with sequences from Fungi RefSeq ITS bioproject using BLASTn provided best hits between P. ochrochloron RLS11 and sequences from other Penicillium ochrochloron strains, with 100% sequence identity. The combined multiple sequence alignment of rpb2, tub, and its gene sequences was used for phylogenetic reconstruction and confirmation of P. ochrochloron RLS11 taxonomic classification.
The maximum likelihood (ML) and Bayesian inference analysis generated the same tree topology, and MLBP/BIPP values were depicted on the ML tree (Fig. 3). All sections obtained were in accordance with previous studies and the currently accepted Penicillium sections [31]. Regarding the P. ochrochloron RLS11, it grouped in a clade containing Penicillium ochrochloron strains with reliable bootstrap proportions and posterior probabilities, and thus, we confidently classified this fungus as a Penicillium ochrochloron strain RLS11.
Best-scoring Maximum Likelihood tree of concatenated sequence alignments of RPB2 (RNA polymerase II subunit 2), β-tubulin, and ITS (internal transcribed spacer 1 – 5.8S – internal transcribed spacer 2) from species belonging to Penicillium sections Chrysogena, Citrina, and Lanata-divaricata. The tree is rooted in Aspergillus niger. MLBP above 70% (left) and BIPP above 95% (right) are indicated in black at the nodes. The Penicillium strain in boldface was used in this study. The Genbank accessions in given in Table S1
Comparative Analysis of CAZyme Repertoire Between P. Ochrochloron RLS11 and Other Penicillium Species
The number of genes encoding CAZymes in P. ochrochloron RLS11 genome was compared to the other Penicillium species. The Penicillium genus contains species with an excellent ability to depolymerize plant biomasses and thus are important sources of CAZymes [32] already used in commercial preparations [33].
Among the 12,015 protein-coding genes of P. ochrochloron RLS11, 737 were predicted as carbohydrate-active enzymes. Glycosyl-hydrolases were by far the most abundant family with 391 predicted genes, followed by glycosyl-transferases (121 genes), carbohydrate esterases (105 genes), auxiliary activity enzymes (90 genes), and polysaccharide-lyases (12 genes) (Table S5).
Overall, P. ochrochloron RLS11 showed more genes related to cellulose, xylan, and pectin depolymerization than other Penicillium species (Fig. 4). This PCWDE-coding gene repertoire was more similar to that of Penicillium subrubescens, a closely related species to P. ochrochloron (Fig. 3), and a good source of enzymes for plant biomass saccharification [34].
The putative inventory of carbohydrate-active enzymes (CAZymes) in Penicillium spp. genomes. CAZyme families were grouped according to their primary polysaccharide substrate, given on the right side (cellulose, pectin, xylan). The number of genes belonging to CAZy families in each genome is indicated inside the frames. The cladogram on the top is based on the number of genes per CAZy family. CAZy families marked with ** contain members acting on other polysaccharides in addition to that depicted (see http://www.cazy.org/). In some cases, the number of CAZyme-coding genes depicted may be slightly different from the true values since the genes were predicted using fully-automatic annotation tools (dbCAN2, BLASTp, InterProScan) and were not subjected to a careful curation by experts
P. ochrochloron RLS11 showed the largest number of protein-coding genes related to cellulose depolymerization (65 genes), outperforming all other Penicillium species analyzed (min = 27 genes, max = 56 genes, s.d. = 7). This included oxidative (AA9 and AA3_1 families) and hydrolytic enzymes (GH3, GH5_5, GH6, and GH7 families). Furthermore, P. ochrochloron RLS11 genome harbors the largest number of endo-β-1,4-xylanases (GH10, GH11, GH30_7 families), α-arabinofuranosidases (GH51), β-xylosidases/α-arabinofuranosidases (GH43_14), and acetylxylan/feruloyl esterases (CE1, CE3, CE4, CE5).
Proteomic Analysis of P. Ochrochloron RLS11 Culture Supernatants
We conducted LC-MS/MS analysis of P. ochrochloron RLS11 culture supernatants to identify which PCWDEs of the 10-day culture supernatant provided the supplementation of Cellic® CTec2/HTec2. We compared this sample to the 5-day supernatant (Tables S6 and S7), as it did not significantly supplementate the commercial enzyme cocktail (Fig. 2).
To detect quantitative changes in the abundance of PCWDEs between the samples, we first accessed the similarity of replicates using principal component analysis (Fig. 5A). The replicates were close in space using the PC1, mostly encompassing the data variance (39.4%). Only one replicate of the day 10 samples was apart from the others in the PC2, but this component represented less than 19% of the total variance. Thus, the overall similarity of samples agrees with the experimental design.
Exoproteome analysis by LC/MS-MS of 5-day and 10-day culture supernatants of P. ochrochloron RLS11. A Principal component analysis of 5-day culture supernatant (red dots) and 10-day culture supernatant (blue dots). PC1 explains 39.4% of the variance, while PC2 explains 18.6%. B Venn diagrams showing the number of proteins detected exclusively in each culture supernatant and the proteins found in both samples. C Volcano plot showing the significance (-log p-value > 1.30) and log2 fold-change of proteins with the 10-day culture supernatant proteome as reference. The red dots represent proteins with significantly higher relative abundance than the 5-day culture supernatant relative abundances, while the blue dots represent proteins with significantly lower relative abundance. Protein IDs: 1 – evm_model_NODE_48_1; 2 – evm_model_NODE_44_29; 3 – evm_model_NODE_1_426; 4 – evm_model_NODE_23_206; 5 – evm_model_NODE_1_379; 6 – evm_model_NODE_38_43; 7 – evm_model_NODE_26_63; 8 – evm_model_NODE_5_339; 9 – evm_model_NODE_12_211; 10 – evm_model_NODE_6_91
An overview of the number of proteins detected in each culture supernatant was given in the Venn diagram (Fig. 5B). A total of 151 and 120 unique proteins were detected in 10-day and 5-day culture supernatants, respectively. Although many exclusive proteins were detected, few corresponded to PCWDEs (Table 3). The 10-day culture supernatant had a higher number of unique cellulases, comprising β-glucosidases, exo- and endo-β-1,4-glucanases, which might contribute to the improvements of Cellic® CTec2/HTec2. A volcano plot was built to visualize the proteins with a significant fold change between samples, which indicated 18 proteins with higher abundance in the 10-day culture supernatant (Fig. 5C). These proteins comprised GH6 and GH7 exo-β-1,4-glucanases (cellobiohydrolases) and GH3 β-1,4-glucosidases (T-test, p-value < 0.05 and log2(fold change) > 1), promising targets for biochemical characterization and supplementation of commercial enzyme cocktails to increase plant biomass saccharification efficiency.
Although a qualitative analysis indicated high similarity between the two culture supernatants (Fig. 6A and 6B), especially regarding endo-β-1,4-xylanases (GH10 and GH11) and β-1,4-glucanases (GH5, GH6, and GH7), the 10-day sample was more enriched with total cellulases than the 5-day sample (10.9% vs. 4.1% of total relative abundance). On the other hand, the relative abundances of the xylan-degrading enzymes were similar (23.5% vs 22.4%, respectively). Such enzyme profiles may result from a higher proportion of cellulose in steam-exploded sugarcane straw (Table S4) and accumulation of cellulose over xylan during the cultivation period, leading to more expression of cellulases than hemicellulase genes.
Exoproteome analysis of culture supernatants of P. ochrochloron RLS11 by LC/MS-MS. Relative abundances of plant cell wall-degrading enzymes detected in the A 5-day culture supernatant and B 10-day culture supernatant of Penicillium ochrochloron RLS11 grown on steam-exploded sugarcane straw. The “Others” sector comprised proteins not annotated as CAZymes by dbCAN2. Protein IDs: 1 – evm_model_NODE_54_10; 2 – evm_model_NODE_1_379; 3 – evm_model_NODE_38_114; 4 – evm_model_NODE_5_339; 5 – evm_model_NODE_26_63; 6 – evm_model_NODE_38_43; 7 – evm_model_NODE_2_422; 8 – evm_model_NODE_8_18; 9 – evm_model_NODE_6_4; 10 – evm_model_NODE_11_34; 11 – evm_model_NODE_16_264; 12 – evm_model_NODE_1_425; 13 – evm_model_NODE_18_192; 14 – evm_model_NODE_18_43; 15 – evm_model_NODE_23_206; 16 – evm_model_NODE_23_208
Discussion
Recent estimates place the number of fungal species in the million range [35], and such magnitude can be linked to huge species diversity, genetic variability, and the widespread across the globe [36]. This led to various fungal lifestyles, which directly influence the repertoire of extracellular enzymes related to their nutritional strategies, such as CAZymes [5]. Therefore, the search for enzymes with superior biochemical-kinetic properties or even unknown enzyme activities is justified since many unexplored fungal species and CAZyme sequences variability may exist [37].
In this work, we first accessed the ability of nine fungal species from Ascomycota and Basidiomycota phyla to produce PCWDE activities when grown on steam-exploded sugarcane straw. We choose sugarcane straw as model substrate for fungal cultivation and saccharification assays since it is abundant in Brazil and easily recovered from commercial plantations.
Most basidiomycetes produced fewer enzyme activities than ascomycetes (Table 1). This could result from unappropriated growth conditions (stirred liquid cultures at high rotation speeds), leading to poor substrate colonization. On the other hand, the ascomycetes showed higher potential to produce cellulases and xylanases but no ability to produce lignin-degrading activities (e.g., laccase activity). Considering only the PCWDE activities detected in the culture supernatants, the enzyme mixtures produced by the ascomycetes were more suitable for saccharification of grass biomass since this material generally shows a high proportion of cellulose especially after the pretreatment. The biomass pretreatment generally removes hemicelluloses and removes/redistributes the lignin on the surface of the lignocellulosic material. Therefore, cellulases are the primary set of enzymes that drives the production of fermentable sugars from such biomasses.
Although the composition of commercial enzyme mixtures is not known in detail, they contain protein stabilizers, a significant proportion of protein-engineered (hemi)cellulases, and the absence of proteases. Thus, they are specifically tailored for biomass saccharification, making it unfair to directly compare such enzyme cocktails with crude fungal enzymatic preparations. For this reason, we opted to supplement the commercial enzyme mixture with the fungal culture supernatants to assess if they significantly enhance sugarcane straw saccharification. The Cellic® CTec2/HTec2 supplemented with the 10-day culture supernatant of P. ochrochloron RLS11 provided a comparable glucose release to the commercial mixture alone at the same protein loading. This suggested that P. ochrochloron RLS11 secretome contains components that increase the efficiency of the commercial enzyme mixture for biomass saccharification. Penicillium species are proving to be promising sources of PCWDEs [33, 38], producing a complete and balanced set of enzymes to catalyze polysaccharide conversion into fermentable sugars [34, 39, 40]. Several reports indicated that cellulases produced by Penicillium species exhibit better catalytic performance than widely used cellulases from Trichoderma reesei [41,42,43], showing a potential to improve the conversion of plant biomasses into fermentable sugars.
The genome of P. ochrochloron RLS11 contained a higher number of gene coding for cellulases and hemicellulases than many Penicillium species (Fig. 4). Such genetic features suggested that P. ochrochloron RLS11 might be very effective for saccharification of graminaceous feedstock, such as sugarcane straw/bagasse and corn stover, which are biomasses with high potential to support biorefineries [44].
The (hemi)cellulolytic enzyme system of P. ochrochloron RLS11 induced by steam-exploded sugarcane straw was primarily composed of hydrolases (families GH3, GH5, GH6, GH7, GH10, GH11) and a lower proportion of oxidative enzymes from AA3 and AA9 families (Fig. 6A and 6B). Such an enzyme profile seems common among Penicillium species since a similar trend was observed in other studies [40, 45]. Surprisingly, an endo-β-1,4-xylanase was the most abundant enzyme in the P. ochrochloron RLS11 exoproteome, although steam-exploded sugarcane straw was composed primarily of cellulose and lignin with small amounts of xylan. This could be due to transcription factors that induce xylanase-coding genes expression in cellulose medium, such as POX02484 of P. oxalicum HP7-1 [46], which have a homolog in P. ochrochloron RLS11 (evm_model_NODE_31_72).
To infer which PCWDEs in the 10-day supernatant of P. ochrochloron RLS11 are providing good supplementation on Cellic® CTec2/HTec2, we compared the proteomic profile of this sample to the 5-day sample, which did not show a significant supplementation effect. Since the saccharification assays were conducted based on protein loading, the proteomic profile of each sample is well correlated to glucose yields. Although homogeneous qualitatively, the relative abundance of PCWDEs between the culture supernatants of P. ochrochloron RLS11 varied substantially, which may be a consequence of modifications in the carbon source composition over the cultivation time. It is well known that the secretion of PCWDEs by fungi is tightly regulated and largely dependent on the carbon source available. As the fungus grows and modifies the surrounding environment, a change in the requirement of enzymes may occur, and the fungus adapts its metabolism for better growth performance. This can be evidenced by the greater abundance of cellulases in the 10-day culture supernatant than the 5-day (more than twice), while a similar trend was not observed for total xylan-degrading enzymes.
The 10-day culture supernatant possessed many unique cellulases (Table 3) and other enzymes with higher relative abundance (Fig. 5C, Fig. 6A and 6B). The exo-β-1,4-glucanases (cellobiohydrolases - CBH) from GH6 and GH7 families are fundamental hydrolases for plant biomass saccharification, and previous studies indicated that such enzymes from a Penicillium species possessed a better hydrolysis potential than CBH from Trichoderma reesei [42]. Thus, CBHs of P. ochrochloron RLS11 are exciting targets for subsequent experiments to confirm if these proteins increase saccharification yields by Cellic® CTec2/HTec2.
Also, β-1,4-glucosidases are essential to alleviate the inhibitory effects of cellobiose on CBHs and endo-β-1,4-glucanases and are necessary enzymes to improve enzymatic saccharification yields [47]. A β-1,4-glucosidase from the GH3 family (evm_model_NODE_5_339) was five times more abundant in the 10-day culture supernatant P. ochrochloron RLS11 and, thus, might be involved in the positive supplementation effect observed for this crude enzymatic extract.
These results are the first steps towards improving modern commercial enzyme preparations for plant biomass depolymerization and have shown the potential of poorly explored fungi to produce efficient PCWDEs. Also, biochemical and kinetic characterization of the cellulases identified here may provide more profound knowledge on the structure-function of enzymes with superior catalytic properties. Taking together, the results presented here may contribute to the development of biotechnology regarding the production of plant biomass-derived biofuels and biomaterials.
Conclusions
Poorly explored fungi are potent sources of carbohydrate-enzymes and can improve modern commercial enzyme mixtures, such as Cellic® CTec2/HTec2 for biomass saccharification.
Genomic, proteomic, and functional assays evidenced the ability of Penicillium ochrochloron RLS11 to produce CAZymes for plant biomass saccharification and provided a wealth of “omics” data that may be used for other biotechnological applications.
The P. ochrochloron RLS11 cellulases from GH3, GH6, and GH7 families are promising targets for following experiments to improve modern commercial enzyme mixtures for biomass saccharification.
Data Availability
This Whole Genome Shotgun project has been deposited at DDBJ/ENA/GenBank under the accession JAAVMA000000000 and BioProject number PRJNA614650. The predicted protein sequences from the genome mentioned above and raw LC-MS/MS data are available from the corresponding author on reasonable request
References
Blackwell, M. (2011). The fungi: 1, 2, 3 ... 5.1 million species? Am. J. Bot., 98, 426–438. https://doi.org/10.3732/ajb.1000298
Galagan, J. E., Henn, M. R., Ma, L. J., Cuomo, C. A., & Birren, B. (2005). Genomics of the fungal kingdom: Insights into eukaryotic biology. Genome Res., 15, 1620–1631. https://doi.org/10.1101/gr.3767105
Cantarel, B. L., Coutinho, P. M., Rancurel, C., Bernard, T., Lombard, V., & Henrissat, B. (2009). The carbohydrate-active enzymes database (CAZy): An expert resource for glycogenomics. Nucleic Acids Res., 37, D233–D238. https://doi.org/10.1093/nar/gkn663
Kameshwar, S. A. K., & Qin, W. (2017). Comparative study of genome-wide plant biomass-degrading CAZymes in white rot, brown rot and soft rot fungi. Mycology., 9, 93–105. https://doi.org/10.1080/21501203.2017.1419296
Zhao, Z., Liu, H., Wang, C., & Xu, J. R. (2013). Comparative analysis of fungal genomes reveals different plant cell wall degrading capacity in fungi. BMC Genomics., 14(274), 1–12. https://doi.org/10.1186/1471-2164-15-6
Couturier, M., Navarro, D., Favel, A., Haon, M., Lechat, C., Lesage-Meessen, L., Chevret, D., Lombard, V., Henrissat, B., & Berrin, J. G. (2016). Fungal secretomics of ascomycete fungi for biotechnological applications. Mycosphere., 7, 1546–1553. https://doi.org/10.5943/mycosphere/si/3b/6
Presley, G. N., Panisko, E., Purvine, S. O., & Schilling, J. S. (2018). Coupling secretomics with enzyme activities to compare the temporal processes of wood metabolism among white and brown rot fungi. Appl. Environ. Microbiol., 84, 1–12. https://doi.org/10.1128/AEM.000159-18
Rytioja, J., Hildén, K., Yuzon, J., Hatakka, A., de Vries, R. P., & Mäkelä, M. R. (2014). Plant-polysaccharide-degrading enzymes from basidiomycetes. Microbiol. Mol. Biol. Rev., 78, 614–649. https://doi.org/10.1128/mmbr.00035-14
Humbird, D., Davis, R., Tao, L., Kinchin, C., Hsu, D., Aden, A., Schoen, P., Lukas, J., Olthof, B., Worley, M., Sexton, D., & Dudgeon, D. (2011). Process design and economics for biochemical conversion of lignocellulosic biomass to ethanol: Dilute-acid pretreatment and enzymatic Hydrolysis of Corn Stover. Natl. Renew. Energy Lab., NREL/TP-5100-47764:1-147. https://doi.org/10.2172/1107470
Gusakov, A. V. (2011). Alternatives to Trichoderma reesei in biofuel production. Trends Biotechnol., 29, 419–425. https://doi.org/10.1016/j.tibtech.2011.04.004
Miller, G. L. (1959). Use of dinitrosalicylic acid reagent for determination of reducing sugar. Anal. Chem., 31, 426–428.
R Core Team. (2013). R: A language and environment for statistical computing. R Foundation for Statistical Computing http://www.R-project.org/
Doyle, J. J., & Doyle, J. L. (1990). Isolation of plant DNA from fresh tissue. Focus (Madison)., 12, 13–15.
Bolger, A. M., Lohse, M., & Usadel, B. (2014). Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics., 30, 2114–2120. https://doi.org/10.1093/bioinformatics/btu170
Bankevich, A., Nurk, S., Antipov, D., Gurevich, A. A., Dvorkin, M., Kulikov, A. S., Lesin, V. M., Nikolenko, S. I., Pham, S., Prjibelski, A. D., Pyshkin, A. V., Sirotkin, A. V., Vyahhi, N., Tesler, G., Alekseyev, M. A., & Pevzner, P. A. (2012). SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol., 19, 455–477. https://doi.org/10.1089/cmb.2012.0021
Simão, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V., & Zdobnov, E. M. (2015). BUSCO: Assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics., 31, 3210–3212. https://doi.org/10.1093/bioinformatics/btv351
Katoh, K., & Standley, D. M. (2013). MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Mol. Biol. Evol., 30, 772–780. https://doi.org/10.1093/molbev/mst010
Darriba, D., Taboada, G. L., Doallo, R., & Posada, D. (2015). jModelTest 2: More models, new heuristics and high-performance computing. Nat. Methods., 9(8)-772. https://doi.org/10.1038/nmeth.2109.jModelTest
Stamatakis, A. (2014). RAxML version 8: A tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics., 30, 1312–1313. https://doi.org/10.1093/bioinformatics/btu033
Ronquist, F., Teslenko, M., Van Der Mark, P., Ayres, D. L., Darling, A., Höhna, S., Larget, B., Liu, L., Suchard, M. A., & Huelsenbeck, J. P. (2012). Mrbayes 3.2: Efficient bayesian phylogenetic inference and model choice across a large model space. Syst. Biol., 61, 539–542. https://doi.org/10.1093/sysbio/sys029
Rambaut, A. (2018) FigTree v1.4.4, a graphical viewer of phylogenetic trees. Available at https://github.com/rambaut/figtree/releases.
Stanke, M., & Waack, S. (2003). Gene prediction with a hidden Markov model and a new intron submodel. Bioinformatics., 19, 215–225. https://doi.org/10.1093/bioinformatics/btg1080
Ter-Hovhannisyan, V., Lomsadze, A., Chernoff, Y. O., & Borodovsky, M. (2008). Gene prediction in novel fungal genomes using an ab initio algorithm with unsupervised training. Genome Res., 18, 1979–1990. https://doi.org/10.1101/gr.081612.108
Trapnell, C., Roberts, A., Goff, L., Pertea, G., Kim, D., Kelley, D. R., Pimentel, H., Salzberg, S. L., Rinn, J. L., & Patcher, L. (2013). Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc., 7, 562–578. https://doi.org/10.1038/nprot.2012.016
Slater, G. S. C., & Birney, E. (2005). Automated generation of heuristics for biological sequence comparison. BMC Bioinformatics., 6, 1–11. https://doi.org/10.1186/1471-2105-6-31
Haas, B. J., Salzberg, S. L., Zhu, W., Pertea, M., Allen, J. E., Orvis, J., White, O., Robin, C. R., & Wortman, J. R. (2008). Automated eukaryotic gene structure annotation using EVidenceModeler and the Program to Assemble Spliced Alignments. Genome Biol., 9, 1–22. https://doi.org/10.1186/gb-2008-9-1-r7
Zhang, H., Yohe, T., Huang, L., Entwistle, S., Wu, P., Yang, Z., Busk, P. K., Xu, Y., & Yin, Y. (2018). DbCAN2: A meta server for automated carbohydrate-active enzyme annotation. Nucleic Acids Res., 46, W95–W101. https://doi.org/10.1093/nar/gky418
Rappsilber, J., Mann, M., & Ishihama, Y. (2007). Protocol for micro-purification, enrichment, pre-fractionation and storage of peptides for proteomics using StageTips. Nat. Protoc., 2, 1896–1906. https://doi.org/10.1038/nprot.2007.261
Cox, J., & Mann, M. (2008). MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol., 26, 1367–1372. https://doi.org/10.1038/nbt.1511
Grigoriev, I. V., Nikitin, R., Haridas, S., Kuo, A., Ohm, R., Otillar, R., Riley, R., Salamov, A., Zhao, X., Korzeniewski, F., Smirnova, T., Nordberg, H., Dubchak, I., & Shabalov, I. (2014). MycoCosm portal: Gearing up for 1000 fungal genomes. Nucleic Acids Res., 42, 699–704. https://doi.org/10.1093/nar/gkt1183
Houbraken, J., & Samson, R. A. (2011). Phylogeny of Penicillium and the segregation of Trichocomaceae into three families. Stud. Mycol., 70, 1–51. https://doi.org/10.3114/sim.2011.70.01
Saini, R., Saini, J. K., Adsul, M., Patel, A. K., Mathur, A., Tuli, D., & Singhania, R. R. (2015). Enhanced cellulase production by Penicillium oxalicum for bio-ethanol application. Bioresour. Technol., 188, 240–246. https://doi.org/10.1016/j.biortech.2015.01.048
Gusakov, A. V., & Sinitsyn, A. P. (2012). Cellulases from Penicillium species for producing fuels from biomass. Biofuels., 3, 463–477. https://doi.org/10.4155/bfs.12.41
Mäkelä, M. R., Mansouri, S., Wiebenga, A., Rytioja, J., de Vries, R. P., & Hildén, K. S. (2016). Penicillium subrubescens is a promising alternative for Aspergillus niger in enzymatic plant biomass saccharification. N. Biotechnol., 33, 834–841. https://doi.org/10.1016/j.nbt.2016.07.014
Wu, B., Hussain, M., Zhang, W., Stadler, M., Liu, X., & Xiang, M. (2019). Current insights into fungal species diversity and perspective on naming the environmental DNA sequences of fungi. Mycology., 10, 127–140. https://doi.org/10.1080/21501203.2019.1614106
Tedersoo, L., Bahram, M., Põlme, S., Kõljalg, U., Yorou, N. S., Wijesundera, R., Ruiz, L. V., Vasco-Palacios, A. M., Thu, P. Q., Suija, A., Smith, M. E., Sharp, C., Saluveer, E., Saitta, A., Rosas, M., Riit, T., Rat, D., & Abarenkov, K. (2014). Global diversity and geography of soil fungi. Science., 346, 1052–1053. https://doi.org/10.1126/science.aaa1185
Helbert, W., Poulet, L., Drouillard, S., Mathieu, S., Loiodice, M., Couturier, M., Lombard, V., Terrapon, N., Turchetto, J., Vincentelli, R., & Henrissat, B. (2019). Discovery of novel carbohydrate-active enzymes through the rational exploration of the protein sequences space. Proc. Natl. Acad. Sci. U. S. A., 116, 6063–6068. https://doi.org/10.1073/pnas.1815791116
Vaishnav, N., Singh, A., Adsul, M., Dixit, P., Sandhu, S. K., Mathur, A., Puri, S. K., & Singhania, R. R. (2018). Penicillium : The next emerging champion for cellulase production. Bioresour. Technol. Reports., 2, 131–140. https://doi.org/10.1016/j.biteb.2018.04.003
Schneider, W.D.H., Gonçalves, T.A., Uchima, C.A., Reis, L. dos, Fontana, R.C., Squina, F.M., Dillon, A.J.P., Camassola, M. (2018) Comparison of the production of enzymes to cell wall hydrolysis using different carbon sources by Penicillium echinulatum strains and its hydrolysis potential for lignocelullosic biomass. Process Biochem. 66:162–170. https://doi.org/10.1016/j.procbio.2017.11.004
Song, W., Han, X., Qian, Y., Liu, G., Yao, G., Zhong, Y., & Qu, Y. (2016). Proteomic analysis of the biomass hydrolytic potentials of Penicillium oxalicum lignocellulolytic enzyme system. Biotechnol. Biofuels, 9, 1–15. https://doi.org/10.1186/s13068-016-0477-2
Chekushina, A. V., Dotsenko, G. S., & Sinitsyn, A. P. (2013). Comparing the efficiency of plant material bioconversion processes using biocatalysts based on Trichoderma and Penicillium verruculosum enzyme preparations. Catal. Ind., 5, 98–104. https://doi.org/10.1134/S2070050413010042
Taylor, L. E., Knott, B. C., Baker, J. O., Alahuhta, P. M., Hobdey, S. E., Linger, J. G., Lunin, V. V., Amore, A., Subramanian, V., Podkaminer, K., Xu, Q., Vanderwall, T. A., Schuster, L. A., Chaudhari, Y. B., Adney, W. S., Crowley, M. F., Himmel, M. E., Decker, S. R., & Beckham, G. T. (2018). Engineering enhanced cellobiohydrolase activity. Nat. Commun., 9, 1–10. https://doi.org/10.1038/s41467-018-03501-8
Thygesen, A., Thomsen, A. B., Schmidt, A. S., Jørgensen, H., Ahring, B. K., & Olsson, L. (2003). Production of cellulose and hemicellulose-degrading enzymes by filamentous fungi cultivated on wet-oxidised wheat straw. Enzyme Microb. Technol., 32, 606–615. https://doi.org/10.1016/S0141-0229(03)00018-8
van der Weijde, T., Alvim Kamei, C. L., Torres, A. F., Vermerris, W., Dolstra, O., Visser, R. G. F., & Trindade, L. M. (2013). The potential of C4 grasses for cellulosic biofuel production. Front. Plant Sci., 4, 1–18. https://doi.org/10.3389/fpls.2013.00107
Schneider, W. D. H., Gonçalves, T. A., Uchima, C. A., Couger, M. B., Prade, R., Squina, F. M., Dillon, A. J. P., & Camassola, M. (2016). Penicillium echinulatum secretome analysis reveals the fungi potential for degradation of lignocellulosic biomass. Biotechnol. Biofuels., 9, 1–26. https://doi.org/10.1186/s13068-016-0476-3
Zhao, S., Yan, Y. S., He, Q. P., Yang, L., Yin, X., Li, C. X., et al. (2016). Comparative genomic, transcriptomic and secretomic profiling of Penicillium oxalicum HP7-1 and its cellulase and xylanase hyper-producing mutant EU2106, and identification of two novel regulatory genes of cellulase and xylanase gene expression. Biotechnol Biofuels., 9, 1–17. https://doi.org/10.1186/s13068-016-0616-9
de Andrade, L. G. A., Maitan-Alfenas, G. P., Morgan, T., Gomes, K. S., Falkoski, D. L., Alfenas, R. F., & Guimarães, V. M. (2017). Sugarcane bagasse saccharification by purified β-glucosidases from Chrysoporthe cubensis. Biocatal. Agric. Biotechnol., 12, 199–205. https://doi.org/10.1016/j.bcab.2017.10.007
Acknowledgements
The authors thank the “Diretoria de Tecnologia de Informação” (DTI) at “Universidade Federal de Viçosa” for availability of the computational cluster and software used in this work, and the Mass Spectrometry Facility at Brazilian Biosciences National Laboratory (LNBio), CNPEM, Campinas, Brazil for their support on mass spectrometry analysis.
Funding
This work was supported by Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES) for research scholarship granted to Tulio Morgan, Conselho Nacional de desenvolvimento Científico e Tecnológico (CNPq), Fundação de Amparo à Pesquisa do Estado de Minas Gerais (FAPEMIG), Consórcio Pesquisa Café, and Bill and Melinda Gates Foundation.
Author information
Authors and Affiliations
Contributions
Tulio Morgan conceived the study, conducted fungal cultivation, enzymatic and saccharification assays, genome sequencing and analysis, proteomic analysis, data interpretation, and discussion, and drafted the manuscript. Daniel Luciano Falkoski conceived the study and participated in designing the methodology. Murillo Peterlini Tavares participated in proteomic analysis and data interpretation, and discussion. Mariana Bicalho Oliveira participated in fungal cultivation and enzymatic assays. Valéria Monteze Guimarães and Tiago Antônio de Oliveira Mendes participated in data interpretation and discussion and prepared the final manuscript version. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics Approval
Not applicable.
Consent to Participate
Not applicable.
Consent for Publication
Not applicable.
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Morgan, T., Falkoski, D.L., Tavares, M.P. et al. Penicillium Ochrochloron RLS11 Secretome Containing Carbohydrate-Active Enzymes Improves Commercial Enzyme Mixtures During Sugarcane Straw Saccharification. Appl Biochem Biotechnol 194, 2946–2967 (2022). https://doi.org/10.1007/s12010-022-03898-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-022-03898-5