Abstract
Analysis of the concentration of free amino acids in biological samples is useful in clinical diagnostics. However, currently available methods are time consuming, potentially delaying diagnosis. Therefore, the development of more rapid analytical tools is needed. In this study, a chemiluminescence detection method for amino acids was developed, and the conditions for the enzyme reaction and assay were examined. For the recognition of each amino acid (here, serine, proline, glycine, asparagine, leucine, and histidine), the corresponding aminoacyl-tRNA synthetase (aaRS) was employed, and multiple enzymatic reactions were combined with a luminol chemiluminescence reaction. This method provided selective quantification from 1 to 20 μM for serine, proline, glycine, and leucine; 1 to 60 μM for asparagine; and 1 to 150 μM for histidine. This assay, which utilized aaRSs for the detection of amino acids, could be useful for simple and rapid analysis of amino acids in clinical diagnostics.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Changes in the concentrations of free amino acids in urine and plasma are useful markers of disease status in clinical diagnostics. For example, reduced leucine concentrations in the blood indicate decreased liver function and delayed formation of glycogen. Additionally, lack of adequate leucine intake decreases insulin secretion, causing a rise in blood sugar levels and therefore promoting the development of diabetes [1]. Proline is an essential component of collagen peptide, which activates skin cells, and is expected to have beautifying effects on the skin [2]. Moreover, glycine functions to improve the negative symptoms of schizophrenia [3], and serine [4, 5], asparagine [6], and histidine [7] also have important effects in the human body [8–10]. Therefore, analysis of the concentrations of amino acids in biological samples may have diagnostic relevance.
High-performance liquid chromatography (HPLC) is generally used to analyze amino acids in biological fluids. However, HPLC analysis requires several hours and specialized equipment and is therefore not easy to use in many clinical settings. Therefore, the development of a simple, rapid analytical tool for measuring amino acid concentrations would be useful for clinical diagnostics. In previous studies, we have begun to develop an enzymatic detection system for amino acids such as serine, histidine, and lysine [8–10] based on aminoacyl-tRNA synthetase (aaRS) as the molecular recognition element; these biosensors showed selective responses to the corresponding amino acids. However, further optimization of this system is needed.
In this study, we further developed an amino acid-sensing method using seryl-tRNA synthetase (SerRS; serine-specific aaRS), prolyl-tRNA synthetase (ProRS; proline-specific aaRS), glycyl-tRNA synthetase (GlyRS; glycine-specific aaRS), asparaginyl-tRNA synthetase (AsnRS; asparagine-specific aaRS), leucyl-tRNA synthetase (LeuRS; leucine-specific aaRS), and histidyl-tRNA synthetase (HisRS; lysine-specific aaRS) as amino acid recognition elements in combination with luminol chemiluminescence detection. The quantitative performance and selectivity of the method were evaluated, and the optimal enzyme reaction and detection conditions were determined.
Experimental
Materials
Amino acids, potassium chloride (KCl), magnesium chloride hexahydrate (MgCl2), luminol, sodium hydroxide, and sodium hydrogen carbonate were purchased from Wako Pure Chemicals (Osaka, Japan). HEPES, ATP, sodium pyruvate, thiamine pyrophosphate (TPP), flavin adenine dinucleotide disodium salt hydrate (FAD), inorganic pyrophosphatase, and peroxidase from horseradish (HRP) were purchased from Sigma-Aldrich Japan (Tokyo, Japan). Pyruvate oxidase was purchased from Asahikasei Farmer (Tokyo, Japan). SerRS, ProRS, GlyRS, and AsnRS were commissioned from Ambis Corp. (Okinawa, Japan). LeuRS and HisRS were commissioned from Takara Bio Inc. (Shiga, Japan). The chemicals were commercial reagents of the highest grade and were used without further purification.
aaRS Multiple Enzymatic Reactions
The reaction mixture containing SerRS (500 μg mL−1) solutions of each amino acid ranging in concentration from 0 to 100 μM, 0.2 mM ATP, 10 mM KCl, and 5 mM MgCl2 was dissolved in 15 mM HEPES-NaOH (pH 8.0), heated at 80 °C for 30 min, and cooled on ice. The SerRS reaction mixture (40 μL) and the reaction mixture (10 μL) containing 0.5 unit mL−1 inorganic pyrophosphatase, 20 units mL−1 pyruvate oxidase, 2.0 mM sodium pyruvate, 5.0 mM MgCl2, 300 μM TPP, and 0.08 μM FAD were dissolved in 50 mM HEPES-NaOH (pH 6.8) and injected into a Nunc 96-well plate (No. 236108, Thermo Fisher Scientific K.K., Waltham, MA, USA). The assay mixture was reacted at 40 °C for 30 min in a microplate reader (Synergy 4, BioTek Instruments, Inc., Winooski, VT, USA).
Chemiluminescence Detection
The luminol solution (100 μL) containing 60 μM luminol and 5.0 unit mL−1 HRP in 800 mM carbonate (NaHCO3-NaOH) buffer (pH 9.0) was injected into the assay mixture using the injector attachment of the microplate reader. Chemiluminescence was detected for 3 s by the microplate reader. Data represent the average of three measurements and the standard deviation is indicated by error bars.
Assay for Proline, Glycine, Lysine, Asparagine, Leucine, and Histidine
For the analyses of proline, glycine, lysine, asparagine, leucine, and histidine, each corresponding aaRS—ProRS (806 μg mL−1), GlyRS (525 μg mL−1), AsnRS (804 μg mL−1), LeuRS (810 μg mL−1), and HisRS (458 μg mL−1)—was used as the amino acid recognition aaRS enzyme, respectively, and assayed in the same manner described above except for SerRS addition.
Results and Discussion
aaRS catalyzes the biosynthesis of proteins and peptides in the body and is thought to provide accurate recognition of corresponding amino acids [11–14]. In our system, aaRS first recognized its corresponding amino acid in the presence of ATP, and aminoacyl-AMP and pyrophosphate were released (Eq. 1). Following multiple enzymatic reactions with inorganic pyrophosphatase (Eq. 2) and pyruvate oxidase (Eq. 3), the released hydrogen peroxide (H2O2) reacted with the luminol reagents (Eq. 4), and the light emitted by the peroxidase-catalyzed luminol reaction was measured spectrophotometrically. The amino acid concentration in the sample was calculated from the intensity of the luminol chemiluminescence. Chemiluminescence detection methods are used for quantitative detection and biosensing technologies in various fields [15, 16]. Therefore, the superior recognition abilities of aaRSs for corresponding amino acids were utilized for the sensing of each amino acid via chemiluminescence detection in this study.
Assays for Serine, Proline, Glycine, Asparagine, and Leucine
Figure 1 shows the selectivity test for SerRS (a), ProRS (b), GlyRS (c), AsnRS (d), and LeuRS (e) in a 25-μM mixture of all 20 natural amino acids. In the presence of corresponding amino acids for each aaRS, high chemiluminescence intensities were obtained; in contrast, lower signals were observed when other amino acids were used. Thus, these data indicated that each aaRS selectively and precisely recognized the corresponding amino acid from analysis of 20 natural amino acids. Notably, different chemiluminescence intensities were observed for each aaRS, even when the amino acid concentration was constant at 25 μM because the binding constant of each aaRS differed.
Selectivity of seryl-tRNA synthetase (SerRS) (a), prolyl-tRNA synthetase (ProRS) (b), glycyl-tRNA synthetase (GlyRS) (c), asparaginyl-tRNA synthetase (AsnRS) (d), and leucyl-tRNA synthetase (LeuRS) (e) for 20 natural amino acids. Twenty natural amino acids, each at a concentration of 25 μM, were added to the reaction mixture, and the chemiluminescence intensity was measured with a microplate reader. Data represent the average of three measurements, and the error bars indicate standard deviations
Next, we examined the quantitative performance of each aaRS for each amino acid. The chemiluminescence intensity increased with increasing amino acid concentrations, and the calibration curve showed good correlations for SerRS (measureable concentration range 5–25 μM; correlation coefficient: r = 0.951), ProRS (1–15 μM, r = 0.972), GlyRS (1–20 μM, r = 0.930), AsnRS (1–60 μM, r = 0.951), and LeuRS (1–30 μM, r = 0.912) (Supplemental data 1).
Interestingly, the chemiluminescence intensity of the reaction of LeuRS for leucine was much lower than that of the other aaRSs. Thus, we next examined the effects of reaction time on the chemiluminescence intensity. We found that the chemiluminescence signal for leucine increased from 30 to 90 min, while those for the other amino acids were almost unchanged (Fig. 2). Analysis of the chemiluminescence intensity of reference amino acids revealed that there was no increase in background intensity. Therefore, the 90-min reaction time did not have adverse effects, such as phosphate release by decomposition of ATP, on the results.
Time dependence of the chemiluminescence intensity in the leucyl-tRNA synthetase (LeuRS) enzyme reaction for leucine and reference amino acids. Each at a concentration of 25 μM was added to the reaction mixture, and the chemiluminescence intensity was measured with a microplate reader. Data represent the average of three measurements, and the error bars indicate standard deviations. Gray bars 30 min, black bars 90 min
Optimization of aaRS Reaction Conditions in Chemiluminescence Detection
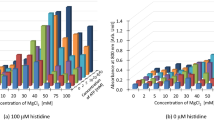
As shown in Figs. 1 and 2, this detection method could be used to measure the target amino acids selectively. However, some challenges were noted, including high background intensity, narrow range of measurable concentration, and low intensity. Therefore, we next sought to optimize the reaction conditions. Figure 3a shows the effects of different ATP concentrations on the chemiluminescence intensity. Although we had previously used 0.2 mM ATP for the reaction (including in the conditions in the previous section) [10], we observed a dramatic increase in the signal of the calibration curve when more than 1 mM ATP was used; therefore, we selected 2 mM ATP for further experiments. Additionally, analysis of the optimal concentration of sodium pyruvate revealed that the addition of 2.5 mM sodium pyruvate also caused a dramatic increase in the signal of the calibration curve (Fig. 3b). In contrast, no obvious changes to background intensity, range of measurable amino acid concentrations, or signal intensity were noted for magnesium chloride and potassium chloride. Therefore, we selected 5 mM magnesium chloride and 10 mM potassium chloride concentrations, as used previously (data not shown).
Chemiluminescence Detection of Histidine in HisRS
We then examined the quantitative performance of HisRS for sensing histidine using the optimized conditions. The reaction mixture, which contained HisRS (458 μg mL−1) solutions of each amino acid ranging from 0 to 300 μM, 2 mM ATP, 2.5 mM sodium pyruvate, 10 mM KCl, and 5 mM MgCl2 dissolved in 15 mM HEPES-NaOH (pH 8.0), was heated at 80 °C for 30 min. The chemiluminescence intensity increased with higher histidine concentrations, and the measurable range of histidine concentrations was obviously widened in comparison with the results of other amino acids, presented above, and the results of our previous paper, which showed a measurable concentration range from 1 to 30 μM [10]. Indeed, the calibration curve showed a good correlation (r = 0.938) between 1 and 150 μM histidine (Fig. 4).
The selectivity of HisRS for histidine was examined using 100 μM of 20 natural amino acids (Fig. 5). The background intensity was obviously reduced, and the chemiluminescence intensity increased by approximately 7.1-fold in the mixture containing histidine. Thus, HisRS selectively and precisely detected histidine in a mixture of 20 natural amino acids.
Selectivity test for 20 natural amino acids during histidine sensing. Twenty natural amino acids, each at a concentration of 100 μM, were added to the reaction mixture, and the chemiluminescence intensity was measured with a microplate reader. Data represent the average of three measurements, and the error bars indicate standard deviations
Conclusion
In this study, we presented a chemiluminescence detection method for serine, proline, glycine, asparagine, leucine, and histidine using a combination of enzymatic reactions and a luminol reaction. For the recognition of each amino acid, corresponding aaRSs were used and corresponding amino acids were detected selectively. Moreover, optimization of reaction conditions increased the detection range for histidine from 1 to 30 to 1–150 μM in this study. These measurable ranges of each amino acid, as shown in this study, were satisfied with the blood amino acid levels.
The glucose sensors with a spectrophotometric detection system are sold and utilized for the diagnosis of diabetes. The protocols and detection method demonstrated in this article would be applied to the existing biosensors. We are aiming at portable amino acid analysis applications that would examine levels of 20 natural amino acids simply and rapidly with the use of aaRSs, as an alternative to the conventional analysis tools in clinical diagnostics.
References
Valerio, A., D’Antona, G., & Nisoli, E. (2011). Branched-chain amino acids, mitochondrial biogenesis, and healthspan: an evolutionary perspective. Aging, 3, 464–478.
Ohara, H., Ichikawa, S., Matsumoto, H., Akiyama, M., Fujimoto, N., Kobayashi, T., & Tajima, S. (2010). Collagen-derived dipeptide, proline-hydroxyproline, stimulates cell proliferation and hyaluronic acid synthesis in cultured human dermal fibroblasts. Journal of Dermatology, 37, 330–338.
Bannai, M., Kawai, N., Nagao, K., Nakano, S., Matsuzawa, D., & Shimizu, E. (2011). Oral administration of glycine increases extracellular serotonin but not dopamine in the prefrontal cortex of rats. Psychiatry and Clinical Neurosciences, 65, 142–149.
Yang, J.-H., Wada, A., Yoshida, K., Miyoshi, Y., Sayano, Y., Esaki, K., Kinoshita, M., Tomonaga, S., Azuma, N., Watanabe, M., Hamase, K., Zaitsu, K., Machida, T., Messing, A., Itohara, S., Hirabayashi, Y., & Furuya, S. (2010). Brain-specific Phgdh deletion reveals a pivotal role for L-serine biosynthesis in controlling the level of D-serine, an NMDA receptor co-agonist, in adult brain. Journal of Biological Chemistry, 285, 41380–41390.
Koning, T. J., Snell, K., Duran, M., Berger, R., Poll-The, B.-T., & Surtees, R. (2003). L-serine in disease and development. Biochemical Journal, 371, 653–661.
Lancha, A. H., Jr., Poortmans, J. R., & Pereira, L. O. (2009). The effect of 5 days of aspartate and asparagine supplementation on glucose transport activity in rat muscle. Cell Biochemistry and Function, 27, 552–557.
Yan, S. L., Wu, S. T., Yin, M. C., Chen, H. T., & Chen, H. C. (2009). Protective effects from carnosine and histidine on acetaminophen-induced liver injury. Journal of Food Science, 74, H259–H265.
Kugimiya, A., & Matsuzaki, E. (2014). Microfluidic analysis of serine levels using seryl-tRNA synthetase coupled with spectrophotometric detection. Applied Biochemistry and Biotechnology, 174, 2527–2536.
Kugimiya, A., & Takamitsu, E. (2013). Spectrophotometric detection of histidine and lysine using combined enzymatic reactions. Materials Science and Engineering C, 33, 4867–4870.
Kugimiya, A., Fukada, R., & Funamoto, D. (2013). A luminol chemiluminescence method for sensing histidine and lysine using enzyme reactions. Analytical Biochemistry, 443, 22–26.
Sekine, S., Nureki, O., Dubois, D. Y., Bernier, S., Chenevert, R., Lapointe, J., Vassylyev, D. G., & Yokoyama, S. (2003). ATP binding by glutamyl-tRNA synthetase is switched to the productive mode by tRNA binding. EMBO Journal, 22, 676–688.
Sekine, S., Shichiri, M., Bernier, S., Chênevert, R., Lapointe, J., & Yokoyama, S. (2006). Structural bases of transfer RNA-dependent amino acid recognition and activation by glutamyl-tRNA synthetase. Structure, 14, 1791–1799.
Ohtsuki, T., Watanabe, Y., Takemoto, C., Kawai, G., Ueda, T., Kita, K., Kojima, S., Kaziro, Y., Nyborg, J., & Watanabe, K. (2001). An “elongated” translation elongation factor Tu for truncated tRNAs in nematode mitochondria. Journal of Biological Chemistry, 276, 21571–21577.
Han, J. M., Jeong, S. J., Park, M. C., Kim, G., Kwon, N. H., Kim, H. K., Ha, S. H., Ryu, S. H., & Kim, S. (2012). Leucyl-tRNA synthetase is an intracellular leucine sensor for the mTORC1-signaling pathway. Cell, 149, 410–424.
Brustein, V. P., Cavalcanti, C. L. B., Melo, M. R., Jr., Correia, M. T. S., Beltrão, E. I. C., & Carvalho, L. B., Jr. (2012). Chemiluminescent detection of carbohydrates in the tumoral breast diseases. Applied Biochemistry and Biotechnology, 166, 268–275.
Tan, H., & Song, Z. (2014). Human saliva-based quantitative monitoring of clarithromycin by flow injection chemiluminescence analysis: a pharmacokinetic study. Applied Biochemistry and Biotechnology, 172, 1320–1331.
Acknowledgments
This work was supported by the Program for Promotion of Basic and Applied Researches for Innovations in Bio-oriented Industry (BRAIN).
Author information
Authors and Affiliations
Corresponding author
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Supplemental data 1
(PPTX 86 kb)
Rights and permissions
About this article
Cite this article
Kugimiya, A., Fukada, R. Chemiluminescence Detection of Serine, Proline, Glycine, Asparagine, Leucine, and Histidine by Using Corresponding Aminoacyl-tRNA Synthetases as Recognition Elements. Appl Biochem Biotechnol 176, 1195–1202 (2015). https://doi.org/10.1007/s12010-015-1639-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-015-1639-6