Abstract
Zeaxanthin carotenoids are class of commercially important natural products and diverse biomolecules produced by plants and many microorganisms. Bacteria often produce a cocktail of polar and nonpolar carotenoids limiting their industrial applications. Marine members of the family Flavobacteriaceae are known to produce potential carotenoids such as astaxanthin and zeaxanthin. A few bacterial species have been reported for the predominant production zeaxanthin. Here, we report the molecular identification of the zeaxanthin as a major carotenoid produced by two novel bacteria (YUAB-SO-11 and YUAB-SO-45) isolated from sandy beaches of South West Coast of India and the effect of carbon sources on the production of zeaxanthin. The strains were identified based on the 16S rRNA gene sequencing as a member of genus Muricauda. The closest relatives of YUAB-SO-11 and YUAB-SO-45 were Muricauda aquimarina (JCM 11811T) (98.9 %) and Muricauda olearia (JCM 15563T) (99.2 %), respectively, indicating that both of these strains might represent a novel species. The highest level of zeaxanthin production was achieved (YUAB-SO-11, 1.20 ± 0.11 mg g−1) and (YUAB-SO-45, 1.02 ± 0.13 mg g−1) when cultivated in marine broth supplemented with 2 % NaCl (pH 7) and incubated at 30 °C. Addition of 0.1 M glutamic acid, an intermediate of citric acid cycle, enhanced the zeaxanthin production as 18 and 14 % by the strains YUAB-SO-11 and YUAB-SO-45 respectively. The zeaxanthin showed in vitro nitric oxide scavenging, inhibition of lipid peroxidation, and 2,2-diphenyl-1-picryl hydrazyl scavenging activities higher than the commercial zeaxanthin. The results of this study suggest that two novel strains YUAB-SO-11 and YUAB-SO-45 belonging to genus Muricauda produce zeaxanthin as a predominant carotenoid, and higher production of zeaxanthin was achieved on glutamic acid supplementation. The pigment showed good in vitro antioxidant activity, which can be exploited further for commercial applications.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Zeaxanthin (C40H56O2), an important xanthophyll carotenoid found in green leafy vegetables and fruits is regularly consumed in the diet. Zeaxanthin has been reported for its multiple applications in food and feed industry due to its potential antioxidant activity, immune modulating, and anticancer capabilities [1, 2]. Hence, there is an increasing demand for zeaxanthin in pharmaceutical, cosmetics, and functional food industries [3, 4].
Marigold (Tagetes erecta) is the chief source of zeaxanthin, but due to the presence of oleoresins and secondary metabolites, the quantity of extractable zeaxanthin is reduced and in turn increases the cost of downstream process [5]. Although chemical synthesis can supply zeaxanthin in commercial scale, it is not preferred over the natural sources due to the undesirable stereoisomers of zeaxanthin and also due to their lower activity [4]. Alternatively, marine microorganisms are promising source for xanthophyll production. Marine flavobacterial members contribute strongly to the carotenoid production particularly xanthophyll carotenoids, namely, astaxanthin, cantaxanthin, and zeaxanthin that are of high commercial and nutritional value. A comparative study has shown that the bioavailability of zeaxanthin produced from Flavobacterium multivorum is two to three times higher than that produced from marigold flowers [6].
Zeaxanthin has been identified as a major pigment in some flavobacterial members, such as F. multivorum [7], Mesoflavibacter zeaxanthinifaciens [8], Zeaxanthinibacter enoshimensis [9], and Muricauda lutaonensis [10]. However, efforts towards utilizing these bacteria as a source of commercial production of zeaxanthin are meager. Major limitation of employing bacterial strains for large-scale commercial scale-up is that they often produce mixed carotenoids as that of plant source and purification becomes expensive due to multiple steps. Hence, there is a significant demand for novel bacterial strain producing zeaxanthin as single pigment in predominant quantities, which could be a probable candidate to suite industrial scale production of zeaxanthin [3, 11]. It has been found that the genera Muricauda constitute a group of bacteria producing distinct yellow to golden-yellow pigmentation and our group had earlier reported the biosynthesis of zeaxanthin by M. lutaonensis CC-HSB-11T [10, 12]. It is interesting to note that it could produce high amounts of all-trans-zeaxanthin as major carotenoid, and hence, members of this genus can be exploited for zeaxanthin production.
The main challenge in using bacterial source for zeaxanthin production is to reduce the cost and simultaneously increase the zeaxanthin yield by increasing biomass production. In carotenoid producing microbes, intermediates of citric acid cycle are known to play an important role in metabolic reactions under aerobic conditions, forming a carbon skeleton for carotenoid and lipid synthesis [7]. Supplementation of citric acid intermediates such as malic acid and α-ketogluterate at the concentration of 10 mM were found to be simple and cost-effective methods for achieving hyperproduction of carotenoid in Flavobacterium sp. [1, 7].
Zeaxanthin has been identified as a potential free radical scavenger. Free radicals such as peroxides and nitric oxides, which are produced during normal/impaired metabolic processes, are associated with loss of cell function and tissue damage. Under normal circumstances, these radicals are eliminated by the natural antioxidant defense systems. However, this delicate balance between the generation and elimination would be lost under certain pathological conditions resulting in health disorders and diseases [13]. A continuous demand for natural and nontoxic antioxidant alternatives in pharmaceutical and food industry exists. In the present work, two novel isolates, YUAB-SO-11 and YUAB-SO-45, belonging to genus Muricauda of the family Flavobacteriaceae, competent of producing zeaxanthin as a predominant carotenoid and its overproduction using citric acid intermediates for industrial scale up were reported. In addition, a detailed evaluation of in vitro antioxidant activity of the carotenoid was also carried out.
Materials and Methods
Chemicals and Reagents
Standard zeaxanthin and 2,2-diphenyl-1-picryl hydrazyl (DPPH) were purchased from Sigma Aldrich (St. Louis, USA), marine agar 2216 (Difco), 2-deoxyribose-d-ribose, naphtyl ethylenediamine dihydrochloride (NED), ethylene diaminetetraacetic acid, sodium nitroprusside, sulphanilamide and thiobarbutric acid (TBA) were purchased from HiMedia (India). Acetonitrile, ascorbic acid, ammonium heptamolybdate, acetic acid, methanol, dimethylsulfoxide, ferric chloride, hydrogen peroxide, potassium ferricyanide, ortho-phosphoric acid, potassium dihydrogen phosphate and dipotassium hydrogen, phosphate, trichloro acetic acid (TCA), tetrahydrofurane, and sulfuric acid were purchased from Merck (India), and all the reagents were of analytical grade.
Isolation and Characterization of Strains
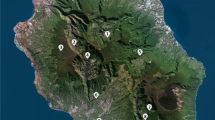
Strain YUAB-SO-11 and YUAB-SO-45 were isolated from coastal sands of Ullal and Cochin, South West Coast of India, respectively. One gram of the soil was taken in 10 mL of sterile water and homogenized for 10 min at 100 rpm using Ultra-Turrax workstation (IKA, Germany) and allowed to settle for 15 min. The supernatant was serially diluted, and 100 μL of sample was spread on marine agar plates and incubated at 32 °C for 48 h. Strains exhibiting yellow pigmentations were isolated, purified and stored at −80 °C for further analysis. Gram staining was performed as described previously [14]. Biochemical characterization and carbon source utilization were carried out using Enterobacteriaceae identification kit Hi 25 (Hi-Media, India) as per the manufacturer’s instructions.
Taxonomic Identification of the Isolate by 16S rRNA Gene-based Phylogeny
Taxonomic identification of YUAB-SO-11 and YUAB-SO-45 was carried out using 16S rRNA gene sequencing as described earlier [15]. For this, DNA was extracted from the 48 h bacterial cultures using genomic DNA extraction kit (MoBio Inc.). PCR amplification of the 16S rRNA gene was carried out using 16S rRNA gene universal primer pair 3F/9R [16, 17]. BigDye terminator cycle sequencing kit was used for sequencing, and determination of the nucleotide sequence of PCR product was performed by an automatic DNA sequencer (ABI PRISM 310, Applied Biosystems, USA). Sequence data was aligned and compared with available standard sequences of bacterial lineage in the National Center for Biotechnology Information GenBank using Basic Local Alignment Search Tool. Further analysis of the sequences was performed using the software Molecular Evolutionary Genetic Analysis version 4.0 [18] after multiple alignment of data by Clustal_X [19] to derive the exact phylogenetic position of the isolate. Reliability of the phylogenetic tree was evaluated based on the bootstrap values [20]. Sequences were submitted to GenBank (YUAB-SO-11, JQ257008 and YUAB-SO-45, JQ346699).
Growth Kinetics
Optimum bacterial growth was determined by inoculating YUAB-SO-11 and YUAB-SO-45 strains (108 CFU mL−1) to 100 mL flask containing 20 mL marine broth and incubated at 32 °C for 96 h under shaking (150 rpm). OD600 was determined at intervals of 24 h using a spectrophotometer. Cell dry weight measurement was carried out after sampling 1 mL broth and centrifuging it at 6,500 rpm for 15 min, followed by washing twice with sterile Milli-Q water. The cell pellets were dried to constant weight in low temperature oven (60 ± 5 °C) to avoid denaturation of the carotenoid.
Isolation and Identification of the Pigment
Cells of YUAB-SO-11 and YUAB-SO-45 were harvested by centrifugation at 6,500 rpm for 15 min and washed twice with Milli-Q water and lyophilized. The finely ground biomass was resuspended in a mixture of dimethyl sulfoxide and methanol [1:1 (v/v)] and agitated overnight in dark at 50 °C. The pigment was separated from the cell suspension by centrifugation (12,000 rpm for 10 min at 4 °C). This process was repeated until complete extraction of the pigment was ensured. The extract was evaporated under nitrogen gas in dark and redissolved in 95 % (v/v) ethanol. The pigment obtained was diluted and analyzed spectrophotometrically in a full-scan mode (250–700 nm UV-vis 1800, Shimadzu, Japan). The chromatographic separation of polar and nonpolar carotenoids was carried out by high-performance liquid chromatography (HPLC) as previously described [10]. For liquid chromatography, a diode array detector (L-2455, Hitachi) attached to reversed-phase column (CAPCELL PAK C18 MG S-5, 35 × 4.6 mm, 5 μm particle size; Shiseido, Tokyo, Japan) connected through a guard column (Phenomenex) maintained at 35 °C was employed, coupled to a HPLC pump (L-2130, Hitachi) equipped with an autosampler (AS-4000) was used for purification. For the identification of zeaxanthin, mass spectrometry was performed in a Thermo Finnigan LTQ linear ion trap mass spectrometer (Thermo LTQ XL, USA) connected to a Thermo Scientific Surveyor LC Plus system. An APcI source operated in the positive ion mode during analysis under the following conditions: sheath gas flow (N2), 50 arbitrary units; auxiliary gas flow (N2), 10 arbitrary units; source voltage, 6 kV; and capillary temperature, 300 °C. For the data-dependent mass spectrometry (MS)/MS experiments, precursor ion detected in the full MS scan selected with an isolation width of 2 m/z unit for collision-induced dissociation with the collision energy of 25 eV. Zeaxanthin was identified based on its absorption spectrum, retention time and m/z values with reference to authentic zeaxanthin standard (Sigma Aldrich, St. Louis, MO, USA).
Effect of pH, Temperature, and Salinity on Zeaxanthin Production
Strains YUAB-SO-11 and YUAB-SO-45 were subjected to grow under different pHs (3.0–12.0), temperature (25–40 °C), and salinity (NaCl, 2-20 % w/v) for the maximum zeaxanthin production in marine broth. The strains were incubated at 30 ± 2 °C, 125 rpm for 5 days. All the experiments were carried out in triplicates.
Effect of Citric Acid Cycle Intermediates and Carbon Source on Zeaxanthin Production
Effect of citric acid cycle intermediates and carbon source on the zeaxanthin production was analyzed according to [7]. Glutamic acid (0.1 M), sodium acetate (0.1 M), and mannitol (0.1 M) as carbon source were supplemented to marine broth. The growth and zeaxanthin yield were calculated in comparison with growth on marine broth (control).
In Vitro Antioxidant Activity of Zeaxanthin Isolated Form YUAB-SO-11 and YUAB-SO-45
Assessment of the antioxidant potential of zeaxanthin from YUAB-SO-11 and YUAB-SO-45 was carried out by extracting zeaxanthin from 50 mg lyophilized biomass in methanol. The concentration of the extracted zeaxanthin was determined prior to the assay from a standard graph plotted using known concentration of the zeaxanthin. Following standard assays were carried out.
Lipid Peroxidation, Nitric Oxide, and Free Radical Scavenging Assays
Inhibition of lipid peroxidation by zeaxanthin was determined by lipid peroxides generated by thiobarbutiric acid reactive substances using egg yolk as oxydizable substrate as previously described [21]. Briefly, 250 μL of egg yolk homogenate (1:40) diluted in phosphate buffer (0.1 M, pH 7.4), and 250 μL of the zeaxanthin (1–20 μg mL−1) were mixed. Lipid peroxidation was initiated by addition of ferrous sulfate (400 μL, 25 mM), incubated for 60 min at 37 °C. Reaction was terminated by the addition of TCA (1 mL, 20 %) and TBA (1 mL, 0.8 %). The resulting mixture was shaken and incubated in boiling water bath for 15 min, allowed to cool and centrifuged at 6,000 rpm for 10 min. The absorbance was measured at 532 nm. The concentration required for 50 % inhibition is considered as IC50. The percentage inhibition was calculated using the equation
where
- A c :
-
Absorbance of control
- A t :
-
Absorbance of test
Nitric Oxide Scavenging Assay
Nitric oxide scavenging assay was carried out as described earlier [22]. Briefly, sodium nitroprusside (500 μL, 25 mM) in phosphate-buffered saline (pH 7.4) was mixed with 500 μL of the zeaxanthin at concentrations (1–10 μg mL−1) in methanol. Mixture was incubated at 25 °C for 150 min. After incubation, 500 μL of the solution was reacted with equal volume of Griess reagent [1 % (w/v) sulphanalic acid, 2 % (v/v) orthophosphoric acid and 0.1 % (w/v) NED]. The absorbance of the chromophore formed during diazotization of nitrate with sulphanalic acid and subsequent coupling with NED was read at 548 nm. Percentage inhibition of nitric oxide by the pigment was calculated with reference to control (without pigment).
DPPH Radical Scavenging Assay
The radical-scavenging activity of the zeaxanthin against DPPH was assessed as outlined previously [23]. Different concentrations of pigment was dissolved in 1 mL methanol, mixed with equal volume of DPPH (0.1 mM in methanol), and incubated in dark for 20 min. The absorbance was read at 517 nm. IC50 concentration was determined using percentage inhibition equation.
Statistical Analysis
All quantitative experiments were carried out in triplicate, and mean was calculated. The effect of environmental factors and TCA intermediates was calculated by one-way analysis of variance and post hoc analysis by Tukey’s t test using software Statistica (Stat soft Inc., 1998).
Results
Isolation, Identification, and Characterization of the Zeaxanthin Producing Marine Bacteria
A total of 49 bacterial colonies were isolated from the coastal sand samples collected from the south west coast of India. Among the isolates collected, two distinctly yellow, translucent, rod-shaped bacteria designated YUAB-SO-11 and YUAB-SO-45 (Fig. 1) were selected and identified by 16S rRNA gene sequencing. Phylogenetic analysis based on the comparisons revealed that the strains YUAB-SO-11 and YUAB-SO-45 belong to the genus Muricauda (Fig. 2). Strain YUAB-SO-11 showed close resemblance (98.9 %) to M. aquimarina (JCM 11811T) and 98.5 % to M. lutimaris (KCTC 22173T). The strain YUAB-SO-45 showed 99.2 % similarity to Muricauda olearia (JCM 15563T) and 96.2 to M. lutimaris (KCTC 22173T). Muricauda sp. YUAB-SO-11 and M. olearia YUAB-SO-45 were strictly aerobic, nonmotile, nonspore forming, Gram-negative and short rod shaped. The selected strains produced yellow to orange pigmented, circular convex colonies on marine agar at 32 °C after 48 h. Both the strains were negative for H2S production, Voges–Proskauer, methyl red, indole, urease, phenyl alanine deamination and malonate utilization, positive for ONPG, lysine, orinithine, and esculin hydrolysis were positive. Detailed biochemical and physiological characterization of the strains in comparison with closely related Muricauda species is provided as Table 1.
Indicating phylogenetic analysis based on 16S rRNA gene sequences showing the relationship of the isolate YUAB-SO-11 (accession number, JQ257008) and YUAB-SO-45 (accession number, JQ346699) to other members of genus Muricauda. Distances and clustering with the neighbor joining method were determined using the software package MEGA version 4. Bar 0.01 substitutions per nucleotide
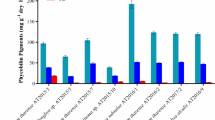
The strains YUAB-SO-11 and YUAB-SO-45 showed maximum growth and zeaxanthin production 1.20 ± 0.11 and 1.02 ± 0.13 mg g−1 at 48 and 72 h respectively (Fig. 3).
Identification and Characterization of the Carotenoid
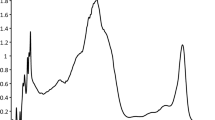
From the spectrophotometric analysis, absorption profile of the crude carotenoid from YUAB-SO-11 and YUAB-SO-45 in methanol showed a characteristic similarity to standard zeaxanthin (Fig. 4a). HPLC with diode-array detection analysis of the crude extracts showed a distinct predominant peak with RT of 5.9 min (68 % of total area), which was similar to that of the standard all-trans-zeaxanthin. In addition, the UV–visible absorption cross-section of peak 1 was also identical to that of the standard all-trans-zeaxanthin as visualized via diode array detector, having characteristic vibronic spectrum with λ max of 450 nm and associated typical shoulder peaks. Furthermore, the mass spectrum of the peak 1 of YUAB-SO-11 exhibited a parent ion [M + H] + at m/z 569 and collusion-induced disassociation fragments of m/z 551 and 419, identical to that of standard all-trans-zeaxanthin (Fig. 4b) and peak 1 of YUAB-SO-45 showed similar (parent ion m/z 569) and (collusion-induced disassociation fragments of m/z 551 and 419, respectively) (Fig. 4c). The m/z values of the compounds corresponding to peak 2 (RT, 11.8; 14.4 ± 1.3 % of total carotenoids) and −3 (RT, 10.9; 7.59 ± 0.7 % of total carotenoids) were similar to that of all-trans-zeaxanthin; however, these two compounds were predicted to be 9-cis-zeaxanthin and 9′-cis-lutein, respectively, based on the previously published results [9].
a UV–visible absorption spectrum of crude methanol extract of YUAB-SO-11 and YUAB-SO-45 and zeaxanthin standard (ZS) solubilized in methanol. b LC-MS/MS analysis and MS/MS spectrum of YUAB-SO-11 zeaxanthin with a parent peak at m/z 569, and subsequent fragements at m/z 551 and m/z 416. c LC-MS/MS analysis and MS/MS spectrum of YUAB-SO-45 zeaxanthin with a parent peak at m/z 569, and subsequent fragements at m/z 551 and m/z 416 and YUAB-SO-45 zeaxanthin
Effect of pH, Temperature, and Salinity on Zeaxanthin Production
Culture conditions such as media pH, salinity, and temperature greatly influenced the growth and zeaxanthin production by both the strains. pH 7, 2 % NaCl, and 30 °C temperature were found to be optimum for zeaxanthin production (Fig.5a–c). Both the isolates did not show any growth at 20 or 45 °C, 1 or 10 % of NaCl, or at pH 4 or 11.
Effect of TCA Cycle Intermediates and Carbon Source on Zeaxanthin Production
When the media was supplemented with 0.1 M glutamic acid, significant increase (p < 0.01) in cell density and zeaxanthin yield was obtained for both the strains. Glutamic acid (0.1 M) supplementation increased the yield by 18.4 % in YUAB-SO-11 (1.47 ± 0.02 mg g−1) and 14 % in YUAB-SO-45 (1.18 ± 0.10 mg g−1) compared to control conditions (Fig. 6). Supplementation of media with mannitol and sodium acetate could increase the cell biomass, but no significant increase in zeaxanthin yield was obtained.
In Vitro Antioxidant Activities of Zeaxanthin Pigment of YUAB-SO-11 and YUAB-SO-45
Zeaxanthin isolated from YUAB-SO-11 and YUAB-SO-45 exhibited significant antioxidant activity. Inhibition of lipid peroxidation, scavenging of nitric oxide and DPPH was effectively observed. Table 3 describes the concentration of YUAB-SO-11 and YUAB-SO-45 zeaxanthin required for 50 % inhibition of free radicals. The antioxidant activity of the zeaxanthin was dose dependent with a linear increase in the antioxidant activity with increase in the carotenoid concentration.
Discussion
Microbial source is being considered as an attractive alternative for the production of carotenoids due to the difficulty in extraction and purification of the carotenoids from plant and other natural sources [2]. Zeaxanthin biosynthesis in Flavobacteria is predicted to follow a sequential biochemical reactions primarily catalyzed by the enzymes geranyl–geranyl pyrophosphate synthase (CrtE), phytoene synthase (CrtB), phytoene desaturase (CrtI), lycopene-beta-cyclase (CrtY), and β-carotene hydroxylase (CrtZ) [26]. Recently, whole-genome sequence of marine Flavobacteria showed the presence of β-carotene hydroxylases (CrtZ) gene. The CrtZ is the key enzyme, which catalyzes the formation of zeaxanthin from β-carotene, via hydroxylation of the latter [2].
Few marine flavobacterial members have been identified to produce zeaxanthin as a predominant carotenoid [8–10]; however, these studies are limited to identification and quantitation of the zeaxanthin carotenoid. The marine environments consist of a large pool of bacteria, which has been left undiscovered due to the lack of sufficient knowledge pertaining to natural strains capable of producing zeaxanthin as predominant carotenoid and very few marine flavobaterial have been studied in terms of zeaxanthin purification, identification and overproduction by media optimization studies for industrial production. Marine Flavobacteria produce a stable form of zeaxanthin known as all-trans-zeaxanthin on the absence of carotenoid isomerase gene [26–28]. Due to the stable nature of all-trans-zeaxanthin, it could be industrially advantageous compared to others. The present study was carried out to describe novel, genetically nonengineered marine microbial resource available in the south west coast of India potential of producing zeaxanthin as the primary carotenoid.
The phylogenetic analysis showed that strain YUAB-SO-11 clustered with M. aquimarina (JCM 11811T) with more than 98 % 16S rRNA gene similarity and YUAB-SO-45 clustered with M. olearia (JCM 15563T) with 99 % 16S rRNA gene similarity, but both the strains showed different physiological and biochemical properties compared to that of M. aquimarina and M. olearia, respectively, indicating that they might represent novel species within the genus Muricauda. Presently, there are seven validly published type strains in the genus Muricauda, and these two might be new addition to the genus.
Identification and characterization of the orange yellow carotenoid using LC-MS/MS analysis revealed both the isolates (YUAB-SO-11 and YUAB-SO-45) produced zeaxanthin as a predominant pigment. Quantification of the carotenoid revealed the strains produced YUAB-SO-11 (1.20 ± 0.11) and YUAB-SO-45 (1.02 ± 0.13) mg g−1 zeaxanthin. Apart from F. multivorum no other flavobacterial members have been investigated in detail for overproduction of zeaxanthin yield. In the present study, we investigated that environmental factors such as pH, temperature, and salinity significantly affect the microbial growth and zeaxanthin production in Muricauda sp. YUAB-SO-11 and M. olearia YUAB-SO-45. These results are similar to the earlier reports on different microorganisms for carotenoid biosynthesis [7, 29]. Zeaxanthin production is generally optimum at pH 7, but the biomass production may vary with pH as in case of Sphingomonas natatoria KODA 19-6 T that produced optimum zeaxanthin at pH 7, but biomass production was optimum at pH 6 [30]. The optimum temperature for zeaxanthin production by the two strains was found to be 32 °C; similarly, earlier study showed S. natotoria KODA19-6T produced optimum zeaxanthin at 30 °C [30]. However, in M. lutaonensis the optimum growth and zeaxanthin production was reported at 40 °C [10]. The optimum yield of zeaxanthin was within 48 h for YUAB-SO-11, but it was 72 h for YUAB-SO-45. Hence, YUAB-SO-11 finds an advantage over others, namely, M. lutaonensis CC-HSB11T and S. natotoria KODA19-6T that require either longer incubation time and/or higher temperature for optimum yield (Table 2).
In carotenoid-producing microbes, citric acid cycle intermediates are known to play an important role in metabolic reactions under aerobic conditions, forming a carbon skeleton for carotenoid and lipid synthesis [7]. Supplementing the media with glutamic acid significantly increased the zeaxanthin yield by 18 % in YUAB-SO-11(1.47 ± 0.02 mg g−1) and 14 % YUAB-SO-45 (1.18 ± 0.11 mg g−1). This can be attributed to high respiratory and citric acid activity associated with the production of large quantities of reactive oxygen species stimulating increased carotenogenesis or due to the increase in the pool of oxaloacetate, which decorboxylates to pyruvate and increases the acetyl CoA pool, the starting substance for isoprenoid synthesis [7] (Table 3).
Zeaxanthin is well known for its antioxidant activity and free radical scavenging activities by different modes of action. Recently, an isomer meso-zeaxanthin was reported for significant antioxidant activities [31]. Biosynthesis of carotenoid by yeast (Paffia rhodozyma), algae (Hematococcus pluvialis), and Dunaliella bardawil are attributed to the production of active oxygen species [32]. These pigments are known to prevent chronic diseases such as cataracts, age-related macular degeneration, and atherosclerosis. The zeaxanthin from both these strains showed a significant antioxidant activity compared to ascorbic acid. The pigments were capable of scavenging nitric oxide, a free radical produced in biological system, which reacts with superoxide to form peroxynitrate and causes the oxidative damage and lipid peroxidation thereby damaging cellular components [33].
In conclusion, the present study characterized Muricauda sp. YUAB-SO-11 and M. olearia YUAB-SO-45 belonging to the genus Muricauda, which will be a new addition to the list of zeaxanthin producing marine flavobacterial isolates described so far. In addition, higher biomass and zeaxanthin production were achieved on glutamic acid supplementation and zeaxanthin isolated from both the strains showed higher antioxidant activities, hence could be explored for commercial importance. Therefore, Muricauda sp.YUAB-SO-11 and M. olearia YUAB-SO-45 offer an edge over the others for economically sustainable industrial exploitation of zeaxanthin.
References
Augusti, P. R., Conterato, G. M. M., Somacal, S., Sobieski, R., Quatrin, A., Maurer, L., et al. (2009). Astaxanthin reduces oxidative stress, but not aortic damage in atherosclerotic rabbits. Journal of Cardiovascular Pharmacology and Therapeutics, 14, 314–322.
Bhosale, P., & Bernstein, P. S. (2005). Microbial xanthophylls. Applied Microbiology and Biotechnology, 68, 445–455.
Asker, D., Awad, T. S., Beppu, T., & Ueda, K. (2012). Novel zeaxanthin-producing bacteria isolated from radioactive hot spring water. Methods in Molecular Biology, 892, 99–131.
Sajilata, M. G., Singhal, R. S., & Kamat, M. Y. (2008). The carotenoid pigment zeaxanthin a review. Comprehensive Reviews in Food Science and Food Safety, 7, 29–49.
Hadden, W. L., Watkins, R. H., Levy, L. W., Regalado, E., Rivadeneira, D. M., van Breemen, R. B., et al. (1999). Carotenoid composition of marigold (Tagetes erecta) flower extract used as nutritional supplement. Journal of Agricultural and Food Chemistry, 47, 4189–4194.
Gierhart, D. L. (1995). Zeaxanthin-containing compositions produced by Flavobacterium multivorum. US patent 5,427,783 (assignee, Applied Food Biotechnology Inc.; date of issue, June 27 1995.).
Bhosale, P., & Bernstein, P. S. (2004). Beta-carotene production by Flavobacterium multivorum in the presence of inorganic salts and urea. Journal of Industrial Microbiology and Biotechnology, 31, 565–571.
Asker, D., Beppu, T., & Ueda, K. (2007). Mesoflavibacter zeaxanthinifaciens gen. nov., sp. nov., a novel zeaxanthin-producing marine bacterium of the family Flavobacteriaceae. Systematic and Applied Microbiology, 30, 291–296.
Asker, D., Beppu, T., & Ueda, K. (2007). Zeaxanthinibacter enoshimensis gen. nov., sp. nov., a novel zeaxanthin-producing marine bacterium of the family Flavobacteriaceae, isolated from seawater off Enoshima Island, Japan. International Journal of Systematic and Evolutionary Microbiology, 57, 837–843.
Hameed, A., Arun, A. B., Ho, H. P., Chang, C. M., Rekha, P. D., Lee, M. R., et al. (2011). Supercritical carbon dioxide micronization of zeaxanthin from moderately thermophilic bacteria Muricauda lutaonensis CC-HSB-11T. Journal of Agriculture and Food Chemistry, 59, 4119–4124.
Hameed, A., Shahina, M., Lin, S. Y., Sridhar, K. R., Young, L. S., Lee, M. R., et al. (2012). Siansivirga zeaxanthinifaciens gen. nov., sp. nov., a novel zeaxanthin-producing member of the family Flavobacteriaceae isolated from coastal seawater of Taiwan. FEMS Microbiology Letters, 333, 37–45.
Arun, A. B., Chen, W. M., Lai, W. A., Chao, J. H., Rekha, P. D., Shen, F. T., et al. (2009). Muricauda lutaonensis sp. nov., a moderate thermophile isolated from a coastal hot spring. International Journal of Systematic and Evolutionary Microbiology, 59, 2738–2742.
Xu, R., Shang, N., & Li, P. (2011). In vitro and in vivo antioxidant activity of exopolysaccharide fractions from Bifidobacterium animalis RH. Anaerobe, 17, 226–231.
Gerhardt, P., Murray, R. G. E., Costilow, R. N., Nester, E. W., Wood, W. A., & Krieg, N. R. (1981). Manual of methods for general bacteriology. Washington: American Society for Microbiology.
Kämpfer, P., Dreyer, U., Neef, A., Dott, W., & Busse, H. J. (2003). Chryseobacterium defluvii sp. nov., isolated from wastewater. International Journal of Systematic and Evolutionary Microbiology, 53, 93–97.
Brosius, J., Palmer, M. L., Kennedy, P. J., & Noller, H. F. (1978). Complete nucleotide sequence of a 16S ribosomal RNA gene from Escherichia coli. Proceedings of the National Academy of Sciences, 75, 4801–4805.
Edwards, U., Rogall, T., Blöcker, H., Emde, M., & Böttger, E. C. (1989). Isolation and direct complete nucleotide determination of entire genes. Characterization of a gene coding for 16S ribosomal RNA. Nucleic Acids Research, 17, 7843–7853.
Tamura, K., Dudley, J., Nei, M., & Kumar, S. (2007). MEGA 4: Molecular Evolutionary Genetics Analysis (MEGA) Software Version 4.0. Molecular Biology and Evolution, 24, 1596–1599.
Thompson, J. D., Gibson, T. J., Plewniak, F., Jeanmougin, F., & Higgins, D. G. (1997). The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Research, 25, 4876–4882.
Felsenstein, J. (1985). Confidence limits on phylogenies: An approach using the bootstrap. Evolution, 39, 783–791.
Zhang, E. X., & Yu, L. J. (1997). Studies on polysaccharide from Sargassum thunbergii for its ability to scavenge active oxygen species. Chinese Journal of Marine Drugs, 3, 1–4.
Nakagawa, T., & Yokozawa, T. (2002). Direct scavenging of nitric oxide and superoxide by green tea. Food and Chemical Toxicology, 40, 1745–1750.
Aquino, R., Morelli, S., Lauro, M. R., Abdo, S., Saija, A., & Tomaino, A. (2001). Phenolic constituents and antioxidant activity of an extract of Anthurium versicolor leaves. Journal of Natural Products, 64, 1019–1023.
Hwang, C. Y., Kim, M. H., Bae, G. D., Zhang, G. I., Kim, Y. H., & Cho, B. C. (2009). Muricauda olearia sp. nov., isolated from crude-oil-contaminated seawater, and emended description of the genus Muricauda. International Journal of Systematic and Evolutionary Microbiology, 59, 1856–1861.
Yoon, J. H., Lee, M. H., Oh, T. K., & Park, Y. H. (2005). Muricauda flavescens sp. nov. and Muricauda aquimarina sp. nov., isolated from a salt lake near Hwajinpo Beach of the East Sea in Korea, and emended description of the genus Muricauda. International Journal of Systematic and Evolutionary Microbiology, 55, 1015–1019.
Armstrong, G. A. (1997). Genetics of eubacterial carotenoid biosynthesis: A colorful tale. Annual Reviews of Microbiology, 51, 629–659.
Kirchman, D. L. (2002). The ecology of Cytophaga–Flavobacteria in aquatic environments. FEMS Microbiology Ecology, 39, 91–100.
Martinez, A., Bradley, A. S., Waldbauer, J. R., Summons, R. E., & DeLong, E. F. (2007). Proteorhodopsin photosystem gene expression enables photophosphorylation in a heterologous host. Proceedings of the National Academy of Sciences, 104, 5590–5595.
Sarada, R., Tripathi, U., & Ravishankar, G. A. (2002). Influence of stress on astaxanthin production in Haematococcus pluvialis grown under different culture conditions. Process Biochemistry, 37, 623–627.
Thawornwiriyanun, P., Tanasupawat, S., Dechsakulwatana, C., Techkarnjanaruk, S., & Suntornsuk, W. Identification of newly zeaxanthin-producing bacteria isolated from sponges in the gulf of Thailand and their zeaxanthin production. Applied Biochemistry and Biotechnology, 167, 2357–2368.
Firdous, A. P., Preethi, K. C., & Kuttan, R. (2010). Antioxidant potential of meso-zeaxanthin a semi synthetic carotenoid. Food Chemistry, 119, 1096–1101.
Krinsky, N. I., Landrum, J. T., & Bone, R. A. (2003). Biologic mechanisms of the protective role of lutein and zeaxanthin in the eye. Annual Reviews of Nutrition, 23, 171–201.
Osborne, N. N., Casson, R. J., Wood, J. P., Chidlow, G., Graham, M., & Melena, J. (2004). Retinal ischemia: Mechanisms of damage and potential therapeutic strategies. Progress in Retinal and Eye Research, 23, 91–147.
Acknowledgments
This research work was kindly supported by a grant from Board Research of Nuclear Science BRNS 28/34/2009.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Prabhu, S., PD, R., Young, CC. et al. Zeaxanthin Production by Novel Marine Isolates from Coastal sand of India and its Antioxidant Properties. Appl Biochem Biotechnol 171, 817–831 (2013). https://doi.org/10.1007/s12010-013-0397-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-013-0397-6