Abstract
Mantle cell lymphoma (MCL) is an aggressive lymphoid neoplasm, incurable with current therapies. The t(11;14)(q13;q32) involving cyclin D1 is considered the first oncogenic hit found in virtually all MCLs. However, additional secondary genomic alterations are essential for complete transformation. MCLs are genetically very unstable with several genetic alterations associated with its high proliferative behavior involving several oncogenic pathways. Furthermore, SOX11 is overexpressed in the majority of conventional MCLs (cMCL), including cyclin D1-negative cases, but absent in non-nodal leukemic MCL with indolent clinical behavior (nnMCL). Recent data have revealed the potential oncogenic role of SOX11 in MCL biology, highlighting its implication in tumor aggressiveness and progression. This review addresses the implication of SOX11 overexpression and frequent genetic lesions, cooperating with cyclin D1 underlying the pathogenesis of this aggressive disease.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Mantle cell lymphoma (MCL) is a highly aggressive and incurable mature B-cell neoplasm, with a short median survival of only 4–6 years; it represents 5–10% of all non-Hodgkin’s lymphomas (NHL) with a male predominance and a median age of ~60 years. Most MCL cases present with disseminated disease, with bone marrow (BM), peripheral blood (PB), nodal and extranodal infiltration at diagnosis. MCLs have an aggressive disease course and frequent relapses despite the good respond to initial treatment, remaining incurable with current therapies [1, 2]. However, a subset of MCL patients (~20%) follows an indolent clinical course with long survival and does not need treatment at diagnosis (nnMCL) [3,4,5]. Several morphologic variants can be recognized; most MCL cases present with conventional monomorphic lymphoid proliferation, whereas other variants, generally associated with a more aggressive disease, include the blastoid and pleomorphic variants [1]. The primary cytogenetic alteration of MCL is the prototypical t(11;14)(q13;q32) translocation that leads to overexpression of the cyclin D1 protein [6], which is usually followed by the acquisition of additional alterations [7]. Besides the clinical parameters (age, performance status, lactate dehydrogenase level, and white blood cell count, among others), the strongest biologic prognostic factor in MCL is the enhanced tumor cell proliferation, mainly measured by immunohistochemical evaluation of the proliferation marker Ki-67 [8].
Interestingly, most MCLs express high levels of the neural transcriptional factor SOX11 (SRY (Sex determining region-Y)-box11). SOX11 was identified to be exclusively overexpressed in MCL as compared to other NHLs, but absent in a small subset of leukemic nnMCL cases with an indolent clinical course [3, 9, 10]. nnMCL has recently been recognized as a different clinical and biological subtype of MCL that, contrary to conventional aggressive MCL (cMCL), are clinically characterized by leukemic non-nodal disease, hypermutated immunoglobulin (IGHV) genes, stable karyotypes, low proliferation rates, lack of SOX11 expression, and better overall survival (OS) compared to cMCL [3, 5, 10,11,12,13]. The lack of SOX11 in this subset of MCL suggests that, besides its prognostic biomarker value, SOX11 may play an important pathogenic role in aggressive MCL. In line with this, recent in vitro studies in MCL cell lines and in vivo experiments using xenotransplanted mouse models have demonstrated specific functions of SOX11, highlighting its oncogenic role in MCL pathogenesis [14••, 15••, 16, 17, 18••].
Thus, cMCL have SOX11 overexpression influencing immune evasion and tumor microenvironment signaling pathways and genetic alterations mainly affecting genes involved in tumor proliferation, cell survival, and several B-cell pathways. All these mechanisms together may tightly collaborate and ultimately contribute to cellular transformation and lymphomagenesis. This review will examine the role of cooperating concomitant cyclin D1 deregulation, SOX11 overexpression, and frequent genetic lesions in MCL.
SOX11 Transcription Factor
SOX11 belongs to the SOX gene family that encode for transcriptional factors characterized by containing a high mobility group (HMG) DNA-binding domain. SOX genes bind to the consensus sequence 5′-(A/T)(A/T)CAA(A/T)G-3′ and induce several DNA conformational changes that allow other regulatory elements to bind together, facilitating the formation of transcriptional complexes essential for several development processes. In humans, about 20 SOX genes have been identified that are divided into eight subgroups, according to the degree of homology within and outside the DNA binding-HMG domain [19]. SOX11 belongs to subgroup C, with high homology to SOX4 and SOX12, which are essential in organogenesis and have overlapping roles in neural development and neurite growth [20].
SOX11 in Human Tumors
SOX11 is widely expressed during embryogenesis but absent in the majority of adult tissues. SOX11 is strongly expressed in most medulloblastomas [21] and malignant glioblastomas [22] and has prognostic value in epithelial ovarian tumors [23] and high grade breast cancer tumors [24]. SOX4 and SOX11 share transcriptional targets and functions and present 89% of identity within their HMG domains [25]. Nevertheless, contrary to SOX4, which is crucial for T- and B-lymphopoiesis, SOX11 is not known to have lymphopoietic functions and is not expressed in normal lymphoid tissues, progenitors, or normal B-cells. Nonetheless, SOX11 has been found to be overexpressed in virtually all aggressive MCLs and, in lower levels, in some Burkitt lymphomas and acute lymphoblastic leukemias, on the contrary, SOX11 is absent in other lymphoid neoplasm or nnMCL [3, 9, 26, 27]. Therefore, SOX11 has been recognized as a MCL diagnostic biomarker. The new specific SOX11-C1 monoclonal mouse antibody has improved the detection of SOX11-nuclear staining by immunohistochemistry [28, 29]. The use of SOX11-C1 has allowed the detection and quantification of SOX11 expression by flow cytometry using tumor samples of PB and BM, improving the diagnosis and prediction of outcome in MCL [30]. Moreover, a quantitative polymerase chain reaction (qPCR) technique to detect SOX11 mRNA levels has similarly demonstrated that high levels of SOX11 mRNA in MCL are related to poor OS [12, 13]. In addition, new techniques based on sensitive qPCR assays for SOX11 have facilitated the diagnosis and stratification of MCL patients [30, 31]. Although there are some controversies in its prognostic value, lack of SOX11 expression has been associated with indolent behavior and favorable prognosis in MCL. Therefore, accurate detection of SOX11 may be helpful to identify nnMCL patients, which could benefit from a “wait and watch” strategy avoiding overtreatment. Consistently, several groups have identified a subset of MCL with indolent clinical course, non-nodal but leukemic presentation and stable karyotype, that carry the typical t(11;14) but lack SOX11 expression [3, 10,11,12]. These results suggest that lack of SOX11 expression is a feature of nnMCL. In contrast, other groups have reported in rather small series that the absence of nuclear SOX11 in MCL with lymph node presentation is associated with shorter OS [32,33,34]. Nevertheless, the majority of SOX11-negative MCL cases of this last study showed strong p53 positivity (due to genetic mutations), which has been shown to be strongly associated with shorter OS in MCL. Of note, some leukemic MCL with lack of SOX11 developed progressive disease, and interestingly, all have increased genomic complexity and 17p loss, suggesting that the acquisition of 17p/TP53 alterations may represent a mechanism of tumor progression in MCL [12, 35], even in the absence of SOX11. In summary, SOX11 expression alone should not be used as a marker for disease aggressiveness, but, together with high number of genetic alterations or TP53 overexpression should be considered in the routine work up of MCL (Table 1).
Oncogenic Roles of SOX11 in MCL
All these findings suggest that besides its diagnostic value, SOX11 may play a relevant role in tumorigenesis and the aggressive progression of MCL. Recent in vitro studies in MCL cell lines and in vivo experiments using xenotransplanted mouse models have demonstrated the potential oncogenic role of SOX11 in MCL, showing that it is able to promote tumor growth of MCL cells by blocking terminal B-cell differentiation through paired box 5 (PAX5) and to induce angiogenesis via platelet-derived growth factor alpha (PDGFA) signaling activation, suggesting that SOX11 in MCL is a bona fide oncogene [14••, 15••]. On the contrary, by similar experimental approaches (SOX11-knockdown and SOX11-forced expression in MCL cell line models), other authors sustain that SOX11 acts as a tumor suppressor by preventing MCL cell proliferation through deregulation of genes involved in the cell cycle regulatory pathways such as components of the RB-E2F pathways (E2F1, CDKN2A), the TGF-β pathway (TGFβR1, SMAD2/3 and TAB1/2) [17], or by directly regulating the expression of several genes of the WNT/β-catenin pathways, repressing the WNT signaling in MCL [18••, 36]. However, SOX11-positive primary MCL tumors are characterized by a high proliferative rate and bad prognosis, as a consequence of the high number of genetic lesions that lead to the deregulation of several pathways (cell proliferation and tumor growth simultaneously) found in the majority of the SOX11-positive cMCLs. Thus, additional functional studies are needed to clarify these apparent contradictory results.
Recently, we have demonstrated that SOX11-knockdown reduces engrafted tumor growth in vivo, consistent with the indolent clinical course of human SOX11-negative nnMCL [14••]. We and others have demonstrated that SOX11 is regulating a large number of genes implicated in oncogenic pathways including proliferation, cell cycle control and apoptosis, B-cell differentiation, angiogenesis, and tumor microenvironment features in MCL [14••, 15••, 16, 18••, 37]. SOX11 directly binds to the regulatory regions of PAX5 and BCL6, two crucial transcription factors for early B-cell development and late differentiation (PAX5) and for entrance into the germinal center (BCL6). SOX11-KD reduces PAX5 expression in MCL cell lines, promoting the shift from a mature B-cell into the initial plasmatic differentiation phenotype by maintaining BLIMP1 repression, both in vitro and in vivo [14••, 38]. Simultaneously, SOX11 represses B-Cell CLL/Lymphoma 6 (BCL6) expression in vitro, essential for GC reaction. These findings support the observations that SOX11-negative primary MCL cases are usually associated with IGHV-mutated MCL and memory like B-cell molecular phenotype and suggest that SOX11-negative MCL may originate from cells that have experienced the GC microenvironment and post-GC differentiation [11, 16] (Fig. 1). Additionally, we observed that SOX11-positive xenografts and human MCL primary tumors presented increased expression of several pro-angiogenic factors and increased microvessel density, compared to its SOX11-negative counterparts [15••]. Our group found that SOX11 binds to the regulatory region of platelet-derived growth factor alpha (PDGFA), promoting its expression. PDGFA is the major pro-angiogenic factor secreted at the SOX11-positive conditioned media promoting in vitro angiogenesis in human vascular endothelial cells (HUVEC) tube formation, cell proliferation and migration. Those angiogenic effects were inhibited when HUVEC cells were treated with imatinib or a specific PDGFRA-neutralizing antibody, demonstrating that SOX11/PDGFA axis is directly involved in MCL angiogenesis. On the contrary, SOX11-negative xenografts and human MCL primary tumors presented with low PDGFA expression, lack of vessels and high necrotic areas which points towards reduced angiogenesis in the indolent non-nodal clinical feature of human nnMCL tumors (Fig. 1).
Hypothetical model of SOX11 overexpression and frequent genetic lesions, cooperating with cyclin D1 underlying the pathogenesis of the two different subsets of MCL. Stepwise progression of conventional SOX11-positive MCL (upper part) and leukemic non-nodal SOX11-negative MCL (lower part). The postulated cell of origin (pre-B-cell) acquires the initial oncogenic translocation of CCND1 (or its variants): these cells clonally expand and colonize the mantle zone (MZ) areas of the lymphoid follicles. The entrance into the germinal center (GC) and terminal B-cell differentiation may be blocked if these cells express SOX11, which determines the two different subsets of MCL. SOX11 overexpression causes B-cell development alterations by prolonging PAX5 expression and consequently Blimp1 inactivation that blocks further terminal differentiation of mature B-cells (upper part). Simultaneously, SOX11 represses BCL6 transcription, blocking the entrance of the cells into the GC, preventing IGHV mutation and further differentiation into terminal plasmacytic/memory B-cells (unmutated-IGHV MCL). Pre B-cells with the t(11;14) lacking SOX11 are express BCL6 and are able to colonize the MZ area of the lymphoid follicles and, enter the GC, undergo IGHV mutations and proceed to a post-GC differentiation (mutated-IGHV MCL) (lower part). Cells with high levels of cyclin D1 due to the translocation and high levels of SOX11 (conventional MCL) will further acquire a plethora of secondary genomic alterations and mutations targeting important pathways leading to high proliferation of cells and aggressive clinical behavior. Moreover, SOX11 promotes angiogenesis via PDGFA pathway activation in MCL facilitating tumor growth and frequent lymph node involvement. On the contrary, SOX11-negative MCL tumors (non-nodal MCL) have low angiogenesis due to the low levels of PDGFA that may be responsible for its prolonged localization only in PB, BM, and spleen and may also explain the clinical features of leukemic nnMCL and its stable indolent clinical evolution. IGHV-mutated/SOX11-negative cases with moderate/high cyclin D1 levels but no SOX11 expression may remain with stable karyotypes, low proliferation rate, no lymph node or extranodal involvement (except spleen) and indolent clinical course. However, a subset of SOX11-negative cases could progress and have a more aggressive clinical behavior by acquiring additional genetic alteration such as TP53 inactivation
Moreover, we observed that imatinib treatment reduced SOX11-positive tumor growth and angiogensis in xenoengrafted mice, to the same levels as SOX11-negative ones [15••]. These findings support the treatment of aggressive refractory human MCL with drugs with antiangiogenic effects such as lenalidomide or new inhibitors of the PDGFRA signaling to prevent angiogenesis and tumor growth in aggressive MCL tumors. Together, these findings demonstrate the oncogenic role of SOX11 in the pathogenesis of MCL and open new therapeutic perspectives for the treatment of aggressive MCL.
CCND1 Rearrangement: The Primary Genetic Alteration in MCL
The primary genetic alteration in MCL is the chromosomal translocation t(11;14)(q13;q32). This rearrangement occurs at the pre-B-cell stage of differentiation in the BM and juxtaposes the CCND1 gene to the IGH gene, resulting in constitutive overexpression of the cyclin D1 protein. In occasional cases, the rearrangement involves the immunoglobulin light chains, IGL or IGK. Additionally to this translocation, other mechanisms lead to increased cyclin D1 levels, which include secondary rearrangements or point mutations in the 3’UTR or 5'UTR regions [39, 40] or amplification of the translocated allele of CCND1 [41, 42]. These additional alterations are associated with high cyclin D1 protein levels, high proliferation, and poor clinical outcome. Together, all these convergent mechanisms highlight the pivotal relevance of cyclin D1 in initial MCL pathogenesis. The main role of cyclin D1 is the regulation of cell cycle and G1-S phase transition, through retinoblastoma (RB1) phosphorylation. Moreover, besides its important role in cell cycle, cyclin D1 may also have other independent oncogenic functions, such as transcription regulation [43] and chromatin remodeling [44], and may affect DNA damage repair mechanisms and chromosome stability [45,46,47]. However, these other functions are less studied in MCL cells and warrant further characterization.
Other Primary Genetic Alterations Affecting Cyclin D Genes
Some rare MCLs lacking the CCND1 translocation and cyclin D1 expression have been reported and recognized as a special subtype of MCL, sharing biological and clinical features, as well as gene expression profiles [48, 49]. Of note, more than half of these cases showed CCND2 rearrangements, mainly with IG light chains, and only a single MCL case has been reported with IGH/CCND3 rearrangement [50]. Hence, most of these cyclin D1-negative MCL show high levels of cyclin D2 or cyclin D3. Overall, these results suggest that cyclin D2 or cyclin D3 overexpression in MCL without t(11;14) may represent an alternative mechanism to cyclin D1 overexpression and may have similar oncogenic properties as cyclin D1 in MCL pathogenesis. The recognition of this MCL variant without cyclin D1 may be difficult, and immunohistochemistry for cyclin D2 or D3 is not recommended [27]. On the other hand, qPCR techniques to detect high levels of cyclin D2 or D3 have been used. In that sense, the detection of SOX11 in all cyclin D1-negative MCL has been of great diagnostic help [27, 49], representing a new useful tool to recognize these MCL cases. Overall, the existence of high levels of SOX11, both in cyclin D1-postive and cyclin D1-negative MCLs (with or without CCND2 rearrangement), suggests that SOX11, besides being a diagnostic biomarker, may also have an important role in MCL pathogenesis collaborating with cyclin D proteins.
Potential Cooperation of Cyclin D1 and SOX11
Despite the fact that deregulation of the cell cycle through high levels of cyclin D1 may be a crucial first step towards MCL pathogenesis, contrary to other human oncogenes, cyclin D1 has been shown to have only a weak oncogenic potential. Several lines of evidence support this notion: first, around 1–2% of healthy individuals carry the t(11;14) rearrangement in PB cells, without developing MCL [51]; second, forced cyclin D1 in MCL cell lines does not have a major effect on proliferation or cell survival [52]; and third, transgenic mice overexpressing cyclin D1 do not develop B-cell lymphomas [53]. Of note, cyclin D1 transgenic models require further oncogenic hits (such as Myc overexpression) for generating lymphomas and the generated tumors were more similar to those generated by overexpression of Myc alone than to human MCL. Moreover, in human primary MCL tumors, MYC alterations can be occasionally found, but are rare. Cyclin D1 transgenic mice have a normal development and their B- and T-lymphocytes do not have developmental alterations.
Another murine model able to generate highly aggressive B-cell lymphomas resembling MCL was when cyclin D1 was expressed constitutively in the nucleus [54]. The analysis of these tumors revealed that they also had high levels of Myc and concomitant alterations of other key players in cell survival and apoptosis pathways, such as TP53 or BCL2, also frequently found in human primary blastoid MCL. In addition, two other models of blastoid MCL were described, through the overexpression of MYC and interleukin-14α [55] and interleukin 3-dependent BaF3 lymphocytes coupled with inducible cyclin D1 expression [56]. The latter model, besides having increased cyclin D1 levels, showed copy number alterations quite similar to human MCL and link the function of cyclin D1 to cell survival through interaction with BAX, independent of its role in cell cycle. However, the first murine model that better recapitulates conventional (not blastoid) MCL combined cyclin D1 overexpression with deletion of BIM, a proapoptotic BCL2 member [57]. BIM has been found to be homozygously deleted in several MCL cell lines and downregulated in a small subset of MCL samples [42, 58]. These data suggest that, together with cyclin D1 overexpression, BIM disruption in B-lymphocytes predisposes to lymphomagenesis through coupled cell cycle and apoptosis deregulation. However, none of these murine models overexpressing cyclin D1 fully recapitulates the MCL phenotype, and they do not have the common genetic alterations found in primary MCL. Interestingly, Eμ-Cyclin D1 transgenic mice were injected with pristane (as a B-cell mitogenic stimulator) and they developed lymphomas with a pattern of dissemination, cell morphology, and phenotype reminiscent of human MCL [59]. Interestingly, these lymphomas were not generated in younger mice, suggesting an age-dependence for lymphoma development similar to human MCL, with a median age at diagnostic around 65 years and increasing incidence with aging. Some processes related with aging (e.g., shortening of telomeres, increase of genomic instability) may increase the susceptibility of Cyclin D1 mice to lymphomas induced by pristane, which may allow the acquisition of additional alterations, similar to human MCL which acquire genomic alterations in a multi-step process.
Recently, an interesting animal model was reported with ATM-deficiency in B-cells coupled with Cyclin D1 overexpression [60]. In this model, highly similar to human MCL, which have ATM mutations in roughly 50% of cases, the high Cyclin D1 and low ATM levels accelerated and increased the incidence of lymphomas (mostly pre-germinal center lymphomas) with morphological features similar to human MCL. Interestingly, the tumor cells of this model had focal deletion of Tp53, Cdkn2a, Mll2, and Rb (also frequently altered in human MCL, as discussed below). However, in none of these previous murine models SOX11 was studied, as they were developed before the recognition of SOX11 as a biomarker for MCL.
Given that SOX11 is expressed in the vast majority of MCLs and especially in the aggressive cases, Kuo PY and coworkers [61] have recently generated the first SOX11 transgenic mouse model with B-cell specific tissue expression. They demonstrated that SOX11 overexpression leads to aberrant expansion of early B-cells with an MCL phenotype, especially in PB and spleen. Interestingly, to evaluate the cooperation of SOX11 and high cyclin D1 levels in MCL pathogenesis they overexpressed both cyclin D1 and SOX11 in a double transgenic mice model, which considerably enhanced the aberrant MCL phenotype more than ten-fold compared with SOX11 and CCND1 single-transgenic mice. Together, these results demonstrate that SOX11 directly collaborates with cyclin D1 in early MCL pathogenesis. It will be interesting to study the secondary molecular alterations that these tumors acquire, and if they also recapitulate those of human MCL, including the frequent TP53 and ATM aberrations in aggressive tumors with high genomic instability. Interestingly, what all these models suggest is that cyclin D1 deregulation may be necessary, but not sufficient, and secondary hits are mandatory to generate MCL-like tumors. Also, the results highlight that both murine models as well as cell line models [62] may represent invaluable resources to study and understand MCL, as well as to test novel therapeutic agents and combination therapies. In addition, it will be of interest to develop mouse models of cyclin D2 or cyclin D3 and determine their collaboration with SOX11, and other genetic alterations as early events cooperating in MCL pathogenesis.
Secondary Genetic Alterations in MCL
All the observations suggest that cyclin D1 overexpression (or cyclin D2 and cyclin D3) is necessary but not sufficient for neoplastic transformation in MCL, and by its own, only cyclin D1 overexpression may not produce MCL tumors with aggressive behavior. In that sense, the highly complex karyotypes and the similar profile of secondary chromosomal alterations that the cyclin D1-positive and cyclin D1-negative MCL with SOX11 expression display, may collaborate with cyclin D and SOX11 overexpression in MCL aggressiveness. On the contrary, the nnMCL subset of cases, which lack SOX11 expression (and its related oncogenic features), have cyclin D1 overexpression, but no other genetic alterations, in line with its indolent clinical behavior. Interestingly, the levels of cyclin D1 in nnMCL are usually lower than in cMCL [3].
Overall, more than 90% of MCLs display highly altered genomes, with gains/amplifications and homozygous/heterozygous losses, as well as other non-recurrent chromosomal rearrangements. Furthermore, the blastic variants have the highest number of alterations [7, 63]. Interestingly, these secondary genetic alterations, which occur in B-cells already harboring high cyclin D1 and SOX11 levels, accumulate in genes affecting several pathways. Several chromosomal and DNA array based techniques in MCL have revealed frequent losses (1p, 2q, 6q, 8p, 9p, 9q, 10p, 11q, 13q, and 17p) and gains (3q, 7p, 8q, 10p, 15q and 18q) [7] (Table 1). Some target genes of these alterations related with cell cycle regulation are well known, for instance the loss of TP53 (at 17p) (Fig. 1). Other cell cycle related targets of deleted regions are CDKN2A and CDKN2B (at 9p21, and usually with biallelic loss), RB1 (at 13q14), CUL4A and ING1 (at 13q34), and four members of the Hippo signaling pathway also related to increased proliferation and worse OS (LATS1, LATS2, MOBKL2B, and MOBKL2A). In addition, several gains, amplifications and translocations affect components of the cell cycle, MYC (at 8q24), similar to murine models, BM1 (at 10p13), CDK4, and MDM2 (at 12q13), as well as IMP3 (at 7p21). Alterations affecting cell survival and apoptosis mainly involve amplification and/or overexpression of the anti-apoptotic BCL2 (at 18q21), and biallelic loss of the proapoptotic BIM (2q13). NF-kB pathway can also be affected by losses of its inhibitors TNFAIP1 (at 6q), and BIRC3 (at 11q). Deregulation of DNA damage response pathway is often accomplished through frequent ATM deletion (at 11q, co-deleted with BIRC3). Interestingly, cell cycle, DNA damage response, and cell survival pathways are altered not only in primary MCL but also in cyclin D1 transgenic models.
Mutational Landscape in MCL
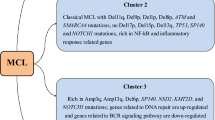
Recently, several studies using next-generation sequencing have been performed in MCL series, including whole-exome sequencing (WES), whole-genome sequencing (WGS), RNA-sequencing, and targeted sequencing. Interestingly, besides the tumor suppressor genes already known to be frequently deleted and mutated in MCL, ATM (41–61%) and TP53 (14–31%), other interesting genes were identified (Table 1). CCND1 mutations (14–34%) have also been recently related to ibrutinib resistance [64]. Important and novel mechanisms identified in MCL include the truncating and activating mutations of NOTCH1 and NOTCH2 associated with dismal prognosis and blastoid variants, epigenetic modifiers such as KTM2D (MLL2), KTM2C (MLL3) and NSD2 (WHSC1), transcription factors (MEF2B), and less frequently, CARD11, BIRC3, NFKBIE, TRAF3, involved in NF-kB signaling pathway, as well as UBR5, and S1PR1 genes [65••, 66,67,68,69,70] (Fig. 1). Interestingly, our study [65••] was the only one that studied cMCL together with a subset of nnMCL, which allowed us to observe that mutations were not equally distributed in both subgroups. cMCL with aggressive behavior and SOX11-positivity accumulated mutations in all genes, while the nnMCL only had mutations in CCND1, TP53, and TLR2. These results are consistent with the notion of the convergent cooperation between cyclin D1, SOX11, and alterations of genes related to important pathways that result in genetically unstable tumors with short OS and resistance to treatment. Conversely, the nnMCL subset with moderate cyclin D1 expression, lack of SOX11, and lack of alterations in oncogenic pathways are in line with their indolent clinical course, suggesting that this subset of patients may benefit from watch and wait strategies or delayed or less intensive treatments.
Perspectives for MCL Targeted Treatments
Although it is not the scope of this review, the amount of information generated in the past years could be of help for the design of targeted therapies. Due to the poor prognosis, especially at relapse, and the chemoresistance of most MCL patients, there is an urgent need for new treatment options and to study potential mechanisms involved in cell survival and drug resistance. Significantly, the mutational studies of the recent years led to the identification of druggable target genes or B-cell pathways that may allow the development of new personalized therapeutic approaches that may help to improve MCL prognosis. It is still in its early phases, but the incorporation of molecular information into clinical trials will be of paramount importance. Some of the deregulated pathways that could be targeted in MCL include the cell cycle (CDK inhibitors), apoptosis (BCL2 inhibitors), DNA damage, and the NOTCH pathway (anti-NOTCH antibodies). Moreover, B-cell receptor signaling pathway has been revealed as a promising target (Ibrutinib), as well as the NF-kB pathway, and other pathways that are targeted by PI3K and mTOR inhibitors or histone methyltransferases or demethylases. In addition to targeted treatments for mutated genes and pathways, also SOX11 functional effects could be targeted using antiangiogenic treatments (lenalidomide or specific PDGFRA-neutralizing antibodies). Although combinations and optimization of these treatments seem promising, still a lot of research in the context of clinical trials is required.
Conclusions
Overall, the findings presented in this review suggest that the IG/CCND1 or IG/CCND2 translocation in pre B-cells is the first oncogenic hit to drive B-cells to the initial steps of MCL. However, cyclin D1 overexpression is not enough to transform these cells and the collaboration of SOX11 overexpression may be necessary. The SOX11 transcription factor directly regulates several oncogenic pathways including proliferation, cell cycle control and apoptosis, B-cell differentiation, angiogenesis, and tumor microenvironment features in MCL, among others. SOX11 expression may synergize with the proliferation rate of mature naïve B-cell with t(11;14) that together with an acquired angiogenic support via direct PDGFA pathway activation may facilitate tumor growth and frequent lymph node involvement, becoming disseminated and potentially very aggressive. SOX11 overexpression may also block the terminal B-cell differentiation program in MCL, usually associated with relevant oncogenic mechanism in lymphoid neoplasm [71]. High expression of cylin D1 and SOX11 are generally accompanied with a high genomic instability with characteristic profiles of gains and losses and specific mutations that interestingly again involve cell cycle, DNA damage, cell survival mechanism, NOTCH, NF-kB and chromatin modifiers. The frequent genomic alterations together with cyclin D1 and SOX11 may contribute to the characteristic high proliferative rate and bad prognosis of aggressive cMCL. Moreover, high number of genomic alterations or p53 overexpression due to TP53 mutations have been associated with a more aggressive clinical behavior and adverse evolution, including the nnMCL subtype. Moreover, the acquisition of genetic alterations (mainly inactivation of CDNKA and TP53 and mutations of TP53, NOTCH1, and NOTCH2) may contribute to the progression to a more aggressive blastoid variants with an adverse prognosis of MCL. Whereas the nnMCL lacking SOX11 and its oncogenic associated pathways, lower levels of cyclin D1 and lack of genomic alterations may remain relatively stable. Together, these findings demonstrate that SOX11 and increased acquisition of genomic alterations collaborate with cyclin D1 in the pathogenesis of MCL. A deeper study of the genomic alterations and associated signaling pathways will further open new potential therapeutic perspectives for the treatment of aggressive MCL.
References
Papers of particular interest, published recently, have been highlighted as: •• Of major importance
Swerdlow S, Campo E, Harris N et al (Eds.). WHO Classification of Tumours of Haematopoietic and Lymphoid Tissues. IARC: Lyon 2008.
Swerdlow SH, Campo E, Pileri SA, Harris NL, Stein H, Siebert R, et al. The 2016 revision of the World Health Organization classification of lymphoid neoplasms. Blood. 2016;127(20):2375–90.
Fernandez V, Salamero O, Espinet B, Sole F, Royo C, Navarro A, et al. Genomic and gene expression profiling defines indolent forms of mantle cell lymphoma. Cancer Res. 2010;70(4):1408–18.
Martin P, Chadburn A, Christos P, Weil K, Furman RR, Ruan J, et al. Outcome of deferred initial therapy in mantle-cell lymphoma. J Clin Oncol. 2009;27(8):1209–13.
Ondrejka SL, Lai R, Smith SD, Hsi ED. Indolent mantle cell leukemia: a clinicopathological variant characterized by isolated lymphocytosis, interstitial bone marrow involvement, kappa light chain restriction, and good prognosis. Haematologica. 2011;96(8):1121–7.
Bosch F, Jares P, Campo E, Lopez-Guillermo A, Piris MA, Villamor N, et al. PRAD-1/cyclin D1 gene overexpression in chronic lymphoproliferative disorders: a highly specific marker of mantle cell lymphoma. Blood. 1994;84(8):2726–32.
Royo C, Salaverria I, Hartmann EM, Rosenwald A, Campo E, Bea S. The complex landscape of genetic alterations in mantle cell lymphoma. Semin Cancer Biol. 2011;21(5):322–34.
Hoster E, Rosenwald A, Berger F, Bernd HW, Hartmann S, Loddenkemper C, et al. Prognostic value of Ki-67 index, cytology, and growth pattern in mantle-cell lymphoma: results from randomized trials of the European mantle cell lymphoma network. J Clin Oncol. 2016;34(12):1386–94.
Ek S, Dictor M, Jerkeman M, Jirstrom K, Borrebaeck CA. Nuclear expression of the non B-cell lineage Sox11 transcription factor identifies mantle cell lymphoma. Blood. 2008;111(2):800–5.
Espinet B, Ferrer A, Bellosillo B, Nonell L, Salar A, Fernandez-Rodriguez C, et al. Distinction between asymptomatic monoclonal B-cell lymphocytosis with cyclin D1 overexpression and mantle cell lymphoma: from molecular profiling to flow cytometry. Clin Cancer Res. 2014;20(4):1007–19.
Navarro A, Clot G, Royo C, Jares P, Hadzidimitriou A, Agathangelidis A, et al. Molecular subsets of mantle cell lymphoma defined by the IGHV mutational status and SOX11 expression have distinct biologic and clinical features. Cancer Res. 2012;72(20):5307–16.
Royo C, Navarro A, Clot G, Salaverria I, Gine E, Jares P, et al. Non-nodal type of mantle cell lymphoma is a specific biological and clinical subgroup of the disease. Leukemia. 2012;26(8):1895–8.
Meggendorfer M, Kern W, Haferlach C, Haferlach T, Schnittger S. SOX11 overexpression is a specific marker for mantle cell lymphoma and correlates with t(11;14) translocation, CCND1 expression and an adverse prognosis. Leukemia. 2013;27(12):2388–91.
•• Vegliante MC, Palomero J, Perez-Galan P, Roue G, Castellano G, Navarro A, et al. SOX11 regulates PAX5 expression and blocks terminal B-cell differentiation in aggressive mantle cell lymphoma. Blood. 2013;121(12):2175–85. First work to demonstrate the potential oncogenic role of SOX11 in MCL. It shows that in in vivo xenotransplanted mice models SOX11-knockdown (KD) reduces tumor growth. SOX11 acts as an oncogene by blocking the terminal B-cell differentiation by directly binding PAX5
•• Palomero J, Vegliante MC, Rodriguez ML, Eguileor A, Castellano G, Planas-Rigol E, et al. SOX11 promotes tumor angiogenesis through transcriptional regulation of PDGFA in mantle cell lymphoma. Blood. 2014;124(14):2235–47. SOX11 directly regulates PDGFA expression, promoting angiogenesis in MCL. The low levels of PDGFA on SOX11-negative xenograft and human MCL primary tumors could explain the lack of vessels and high necrotic areas which support the indolent non-nodal clinical feature of human SOX11-negative nnMCL tumors
Palomero J, Vegliante MC, Eguileor A, Rodriguez ML, Balsas P, Martinez D, et al. SOX11 defines two different subtypes of mantle cell lymphoma through transcriptional regulation of BCL6. Leukemia. 2016;30(7):1596–9.
Gustavsson E, Sernbo S, Andersson E, Brennan DJ, Dictor M, Jerkeman M, et al. SOX11 expression correlates to promoter methylation and regulates tumor growth in hematopoietic malignancies. Mol Cancer. 2010;9:187.
•• Kuo PY, Leshchenko VV, Fazzari MJ, Perumal D, Gellen T, He T, et al. High-resolution chromatin immunoprecipitation (ChIP) sequencing reveals novel binding targets and prognostic role for SOX11 in mantle cell lymphoma. Oncogene. 2015;34(10):1231–40. First SOX11ChIP-sequencing.The authors identify several SOX11 target genes that regulate WNT pathway
Wegner M. All purpose sox: the many roles of sox proteins in gene expression. Int J Biochem Cell Biol. 2010;42(3):381–90.
Penzo-Mendez AI. Critical roles for SoxC transcription factors in development and cancer. Int J Biochem Cell Biol. 2010;42(3):425–8.
Lee CJ, Appleby VJ, Orme AT, Chan WI, Scotting PJ. Differential expression of SOX4 and SOX11 in medulloblastoma. J Neuro-Oncol. 2002;57(3):201–14.
Weigle B, Ebner R, Temme A, Schwind S, Schmitz M, Kiessling A, et al. Highly specific overexpression of the transcription factor SOX11 in human malignant gliomas. Oncol Rep. 2005;13(1):139–44.
Sernbo S, Gustavsson E, Brennan DJ, Gallagher WM, Rexhepaj E, Rydnert F, et al. The tumour suppressor SOX11 is associated with improved survival among high grade epithelial ovarian cancers and is regulated by reversible promoter methylation. BMC Cancer. 2011;11:405.
Shepherd JH, Uray IP, Mazumdar A, Tsimelzon A, Savage M, Hilsenbeck SG, et al. The SOX11 transcription factor is a critical regulator of basal-like breast cancer growth, invasion, and basal-like gene expression. Oncotarget. 2016;7(11):13106–21.
Dy P, Penzo-Mendez A, Wang H, Pedraza CE, Macklin WB, Lefebvre V. The three SoxC proteins—Sox4, Sox11 and Sox12—exhibit overlapping expression patterns and molecular properties. Nucleic Acids Res. 2008;36(9):3101–17.
Dictor M, Ek S, Sundberg M, Warenholt J, Gyorgy C, Sernbo S, et al. Strong lymphoid nuclear expression of SOX11 transcription factor defines lymphoblastic neoplasms, mantle cell lymphoma and Burkitt's lymphoma. Haematologica. 2009;94(11):1563–8.
Mozos A, Royo C, Hartmann E, De JD, Baro C, Valera A, et al. SOX11 expression is highly specific for mantle cell lymphoma and identifies the cyclin D1-negative subtype. Haematologica. 2009;94(11):1555–62.
Soldini D, Valera A, Sole C, Palomero J, Amador V, Martin-Subero JI, et al. Assessment of SOX11 expression in routine lymphoma tissue sections: characterization of new monoclonal antibodies for diagnosis of mantle cell lymphoma. Am J Surg Pathol. 2014;38(1):86–93.
Nordstrom L, Andreasson U, Jerkeman M, Dictor M, Borrebaeck C, Ek S. Expanded clinical and experimental use of SOX11—using a monoclonal antibody. BMC Cancer. 2012;12:269.
Hamborg KH, Bentzen HH, Grubach L, Hokland P, Nyvold CG. A highly sensitive and specific qPCR assay for quantification of the biomarker SOX11 in mantle cell lymphoma. Eur J Haematol. 2012;89(5):385–94.
Simonsen AT, Sorensen CD, Ebbesen LH, Bodker JS, Bentzen HH, Nyvold CG. SOX11 as a minimal residual disease marker for mantle cell lymphoma. Leuk Res. 2014;38(8):918–24.
Nordstrom L, Sernbo S, Eden P, Gronbaek K, Kolstad A, Raty R, et al. SOX11 and TP53 add prognostic information to MIPI in a homogenously treated cohort of mantle cell lymphoma—a Nordic lymphoma group study. Br J Haematol. 2014;166(1):98–108.
Wang X, Asplund AC, Porwit A, Flygare J, Smith CI, Christensson B, et al. The subcellular Sox11 distribution pattern identifies subsets of mantle cell lymphoma: correlation to overall survival. Br J Haematol. 2008;143(2):248–52.
Nygren L, Baumgartner WS, Klimkowska M, Christensson B, Kimby E, Sander B. Prognostic role of SOX11 in a population-based cohort of mantle cell lymphoma. Blood. 2012;119(18):4215–23.
Sander B, Quintanilla-Martinez L, Ott G, Xerri L, Kuzu I, Chan JK, et al. Mantle cell lymphoma—a spectrum from indolent to aggressive disease. Virchows Arch. 2016;468(3):245–57.
Nordstrom L, Andersson E, Kuci V, Gustavsson E, Holm K, Ringner M, et al. DNA methylation and histone modifications regulate SOX11 expression in lymphoid and solid cancer cells. BMC Cancer. 2015;15:273.
Wang X, Bjorklund S, Wasik AM, Grandien A, Andersson P, Kimby E, et al. Gene expression profiling and chromatin immunoprecipitation identify DBN1, SETMAR and HIG2 as direct targets of SOX11 in mantle cell lymphoma. PLoS One. 2010;5(11):e14085.
Ribera-Cortada I, Martinez D, Amador V, Royo C, Navarro A, Bea S, et al. Plasma cell and terminal B-cell differentiation in mantle cell lymphoma mainly occur in the SOX11-negative subtype. Mod Pathol. 2015;28(11):1435–47.
Wiestner A, Tehrani M, Chiorazzi M, Wright G, Gibellini F, Nakayama K, et al. Point mutations and genomic deletions in CCND1 create stable truncated cyclin D1 mRNAs that are associated with increased proliferation rate and shorter survival. Blood. 2007;109(11):4599–606.
Seto M, Yamamoto K, Iida S, Akao Y, Utsumi KR, Kubonishi I, et al. Gene rearrangement and overexpression of PRAD1 in lymphoid malignancy with t(11;14)(q13;q32) translocation. Oncogene. 1992;7(7):1401–6.
Gruszka-Westwood AM, Atkinson S, Summersgill BM, Shipley J, Elnenaei MO, Jain P, et al. Unusual case of leukemic mantle cell lymphoma with amplified CCND1/IGH fusion gene. Genes Chromosomes Cancer. 2002;33(2):206–12.
Bea S, Salaverria I, Armengol L, Pinyol M, Fernandez V, Hartmann EM, et al. Uniparental disomies, homozygous deletions, amplifications, and target genes in mantle cell lymphoma revealed by integrative high-resolution whole-genome profiling. Blood. 2009;113(13):3059–69.
Coqueret O. Linking cyclins to transcriptional control. Gene. 2002;299(1–2):35–55.
Aggarwal P, Vaites LP, Kim JK, Mellert H, Gurung B, Nakagawa H, et al. Nuclear cyclin D1/CDK4 kinase regulates CUL4 expression and triggers neoplastic growth via activation of the PRMT5 methyltransferase. Cancer Cell. 2010;18(4):329–40.
Li Z, Jiao X, Wang C, Shirley LA, Elsaleh H, Dahl O, et al. Alternative cyclin D1 splice forms differentially regulate the DNA damage response. Cancer Res. 2010;70(21):8802–11.
Jirawatnotai S, Hu Y, Livingston DM, Sicinski P. Proteomic identification of a direct role for cyclin d1 in DNA damage repair. Cancer Res. 2012;72(17):4289–93.
Casimiro MC, Di SG, Crosariol M, Loro E, Dampier W, Ertel A, et al. Kinase-independent role of cyclin D1 in chromosomal instability and mammary tumorigenesis. Oncotarget. 2015;6(11):8525–38.
Fu K, Weisenburger DD, Greiner TC, Dave S, Wright G, Rosenwald A, et al. Cyclin D1-negative mantle cell lymphoma: a clinicopathologic study based on gene expression profiling. Blood. 2005;106(13):4315–21.
Salaverria I, Royo C, Carvajal-Cuenca A, Clot G, Navarro A, Valera A, et al. CCND2 rearrangements are the most frequent genetic events in cyclin D1(-) mantle cell lymphoma. Blood. 2013;121(8):1394–402.
Wlodarska I, Dierickx D, Vanhentenrijk V, Van RK, Pospisilova H, Minnei F, et al. Translocations targeting CCND2, CCND3, and MYCN do occur in t(11;14)-negative mantle cell lymphomas. Blood. 2008;111(12):5683–90.
Lecluse Y, Lebailly P, Roulland S, Gac AC, Nadel B, Gauduchon P. t(11;14)-positive clones can persist over a long period of time in the peripheral blood of healthy individuals. Leukemia. 2009;23(6):1190–3.
Klier M, Anastasov N, Hermann A, Meindl T, Angermeier D, Raffeld M, et al. Specific lentiviral shRNA-mediated knockdown of cyclin D1 in mantle cell lymphoma has minimal effects on cell survival and reveals a regulatory circuit with cyclin D2. Leukemia. 2008;22(11):2097–105.
Lovec H, Grzeschiczek A, Kowalski MB, Moroy T. Cyclin D1/bcl-1 cooperates with myc genes in the generation of B-cell lymphoma in transgenic mice. EMBO J. 1994;13(15):3487–95.
Gladden AB, Woolery R, Aggarwal P, Wasik MA, Diehl JA. Expression of constitutively nuclear cyclin D1 in murine lymphocytes induces B-cell lymphoma. Oncogene. 2006;25(7):998–1007.
Ford RJ, Shen L, Lin-Lee YC, Pham LV, Multani A, Zhou HJ, et al. Development of a murine model for blastoid variant mantle-cell lymphoma. Blood. 2007;109(11):4899–906.
Beltran E, Fresquet V, Martinez-Useros J, Richter-Larrea JA, Sagardoy A, Sesma I, et al. A cyclin-D1 interaction with BAX underlies its oncogenic role and potential as a therapeutic target in mantle cell lymphoma. Proc Natl Acad Sci U S A. 2011;108(30):12461–6.
Katz SG, Labelle JL, Meng H, Valeriano RP, Fisher JK, Sun H, et al. Mantle cell lymphoma in cyclin D1 transgenic mice with Bim-deficient B cells. Blood. 2014;123(6):884–93.
Mestre-Escorihuela C, Rubio-Moscardo F, Richter JA, Siebert R, Climent J, Fresquet V, et al. Homozygous deletions localize novel tumor suppressor genes in B-cell lymphomas. Blood. 2007;109(1):271–80.
Smith MR, Joshi I, Jin F, Al-Saleem T. Murine model for mantle cell lymphoma. Leukemia. 2006;20(5):891–3.
Yamamoto K, Lee BJ, Li C, Dubois RL, Hobeika E, Bhagat G, et al. Early B-cell-specific inactivation of ATM synergizes with ectopic CyclinD1 expression to promote pre-germinal center B-cell lymphomas in mice. Leukemia. 2015;29(6):1414–24.
Kuo PY, Jiang Z, Perumal D, Leshchenko VV, Lagana A, Katz SG, et al. SOX11 Cooperates with CCND1 in Mantle Cell Lymphoma Pathogenesis. Blood [126], 1253. 1–1-2015. Ref Type: Abstract
Smith MR, Joshi I, Pei J, Slifker M, Jin F, Testa JR, et al. Murine mantle cell lymphoma model cell line. Leukemia. 2013;27(7):1592–4.
Hartmann EM, Campo E, Wright G, Lenz G, Salaverria I, Jares P, et al. Pathway discovery in mantle cell lymphoma by integrated analysis of high-resolution gene expression and copy number profiling. Blood. 2010;116(6):953–61.
Mohanty A, Sandoval N, Das M, Pillai R, Chen L, Chen RW, et al. CCND1 mutations increase protein stability and promote ibrutinib resistance in mantle cell lymphoma. Oncotarget 2016.
•• Bea S, Valdes-Mas R, Navarro A, Salaverria I, Martin-Garcia D, Jares P, et al. Landscape of somatic mutations and clonal evolution in mantle cell lymphoma. Proc Natl Acad Sci U S A. 2013;110(45):18250–5. First whole-exome and whole-genome sequencing study in MCL, including nnMCL cases. We identified mutations in several new pathways potentially involved in MCL pathogenesis
Wu C, de Miranda NF, Chen L, Wasik AM, Mansouri L, Jurczak W, et al. Genetic heterogeneity in primary and relapsed mantle cell lymphomas: Impact of recurrent CARD11 mutations. Oncotarget 2016.
Zhang J, Jima D, Moffitt AB, Liu Q, Czader M, Hsi ED, et al. The genomic landscape of mantle cell lymphoma is related to the epigenetically determined chromatin state of normal B cells. Blood. 2014;123(19):2988–96.
Kridel R, Meissner B, Rogic S, Boyle M, Telenius A, Woolcock B, et al. Whole transcriptome sequencing reveals recurrent NOTCH1 mutations in mantle cell lymphoma. Blood. 2012;119(9):1963–71.
Meissner B, Kridel R, Lim RS, Rogic S, Tse K, Scott DW, et al. The E3 ubiquitin ligase UBR5 is recurrently mutated in mantle cell lymphoma. Blood. 2013;121(16):3161–4.
Rahal R, Frick M, Romero R, Korn JM, Kridel R, Chun CF, et al. Pharmacological and genomic profiling identifies NF-kappaB-targeted treatment strategies for mantle cell lymphoma. Nat Med. 2014;20(1):87–92.
Mandelbaum J, Bhagat G, Tang H, Mo T, Brahmachary M, Shen Q, et al. BLIMP1 is a tumor suppressor gene frequently disrupted in activated B cell-like diffuse large B cell lymphoma. Cancer Cell. 2010;18(6):568–79.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
Sílvia Beà and Virginia Amador declare they have no conflict of interest.
Human and Animal Rights and Informed Consent
This article does not contain any studies with human or animal subjects performed by any of the authors.
Additional information
This article is part of the Topical Collection on Lymphomas
Rights and permissions
About this article
Cite this article
Beà, S., Amador, V. Role of SOX11 and Genetic Events Cooperating with Cyclin D1 in Mantle Cell Lymphoma. Curr Oncol Rep 19, 43 (2017). https://doi.org/10.1007/s11912-017-0598-1
Published:
DOI: https://doi.org/10.1007/s11912-017-0598-1