Abstract
High-grade serous ovarian carcinoma (HGSOC) accounts for the majority of the ovarian cancer deaths, but over the last years little improvement in overall survival has been achieved. HGSOC is a molecularly and clinically heterogeneous disease. At genomic level, it represents a C-class malignancy having frequent gene losses (NF1, RB1, PTEN) and gains (CCNE1, MYC). HGSOC shows a simple mutational profile with TP53 nearly always mutated and with other genes mutated at low frequency. Importantly, 50 % of all HGSOCs have genetic features indicating a homologous recombination (HR) deficiency. HR deficient tumors are highly sensitive to PARP inhibitor anticancer agents, which exhibit synthetic lethality with a defective HR pathway. Transcriptionally, HGSOCs can be grouped into different molecular subtypes with distinct biology and prognosis. Molecular stratification of HGSOC based on these genomic features may result in improved therapeutic strategies.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Epithelial ovarian cancer (EOC) is the most lethal gynecological malignancy in western countries and accounts for almost two-thirds of ovarian cancer deaths [1, 2••]. The majority of patients are diagnosed at an advanced stage due to a lack of symptoms and the paucity of effective methods to detect the disease early [3]. High-grade serous ovarian carcinomas (HGSOCs) account for 60–80 % of the women diagnosed with epithelial ovarian cancer, and most deaths are associated with this subtype [4]. The disease is referred to as ovarian cancer, but clinical and experimental data indicate that secretory epithelial cells of the distal fallopian tube (FTSECs) are the likely progenitors of the majority of HGSOCs [2••]. Currently, the standard therapy combines maximal cytoreductive surgery followed by platinum-taxane chemotherapy [3, 5••]. HGSOC is associated with initial chemotherapy responsiveness; however, most tumors recur and become increasingly chemotherapy resistant, resulting in an overall 5-year survival probability of 31 % [5••, 6]. Importantly, the debulking surgery remains the key since the extent of debulking profoundly influences prognosis in EOC; suboptimal cytoreductive surgery (residual tumor size of greater than 2 cm) is indeed associated with poor survival in advanced EOC and with a specific molecular subtype characterized by desmoplastic stromal reaction, as recently reported by Liu and colleagues [7]. The evolution of surgical techniques and chemotherapy regimens over the past three decades has resulted in improvements in ovarian cancer treatment [8]. However, in spite of the fact that changes in both chemotherapy schedules and administration had increased response rates of over 80 %, most of the patients eventually relapse. One of the major disappointments in the field of ovarian cancer research is the failure of currently established therapies to induce a cure at diagnosis, even in chemosensitive tumors [8]. HGSOC has shown little improvement in overall survival (OS) for decades, whereas improvements in OS were achieved for a number of other common solid malignancies [2••, 9].
This lack of successful treatment strategies led researchers in the recent years to measure molecular aberrations of HGSOC in order to identify new therapeutic targets or biomarkers that could improve treatment options for ovarian cancer patients. The emerging view from this work is that high-grade serous ovarian carcinoma shows a heterogeneous molecular and cellular biology with no predominant pathway deregulated in most of the patients [8]. Therefore, individualized patient selection for treatment decision-making becomes increasingly essential.
Here, I review recent advances in our understanding of the molecular features of HGSOC with possible therapeutic implications. Table 1 summarized the key molecular features of high-grade serous ovarian carcinoma (HGSOC).
Genetic Alterations of High-Grade Serous Ovarian Carcinoma (HGSOC)
Somatic Mutations and HR Pathway Alterations
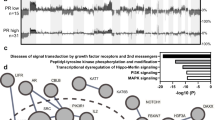
In 2011, two important studies shed light on the genetic alterations driving ovarian cancer [5••, 13•]. The Cancer Genome Atlas (TCGA) Research Network employed a variety of high-throughput technologies to systematically catalogue molecular aberrations in about 500 HGSOC cases. Cheung and colleagues instead took a more functional approach where they performed systematic loss-of-function studies in cell lines of ovarian (and other) cancers to identify genes essential for tumor proliferation and survival. In particular, they aimed to find genes that were both essential for the ovarian cancer cells but also amplified or differentially overexpressed in primary tumors and ovarian cancer cell lines.
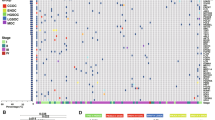
Consistent with previous reports, the TP53 gene is nearly always mutated in HGSOC (97 % of the tumors), and the TP53 mutation occurs early in the genesis of ovarian cancer [14, 15]. The next most frequently affected genes are BRCA1 and BRCA2, which had germline mutations in about 9 and 8 % of the TCGA tumor samples, respectively, and showed somatic mutations in additional 3 % of cases [5••]. Researchers from the TCGA also identified mutations in other genes at lower (<6 %) but statistically significant rates, such as NF1, RB1, CDK12, FAT3, CSMD3, and GABRA6. Additional studies are required to confirm these mutation frequencies in independent patient cohorts. The cyclin-dependent kinase CDK12 represents an interesting novel hit, not previously associated with ovarian cancer. Considering its mutation pattern can be classified as a tumor suppressor gene. Subsequent functional studies reported that CDK12 kinase domain mutations impede catalytic activity and cells with catalytically inactive CDK12 have defect in homologous recombination (HR). Similarly, disabling CDK12 in ovarian cancer cells disrupts HR and also sensitizes cells to DNA cross-linking agents and poly ADP-ribose polymerase (PARP) inhibitors [16, 17•]. This was explained by the indirect role of CDK12 affecting the transcription of key factors of the HR pathway [18, 19]. Taken together, these studies underline a possible role of CDK12 as biomarker for treatment. Still, further studies are needed to refine the mutation frequency of CDK12 in a larger patient population.
Defects in HR DNA repair pathway occur in up to 50 % of HGSOC cases. HR defects are mainly caused by germline, somatic, and epigenetic mutations (silencing through methylation) in BRCA1 and BRCA2 and less frequently by mutations in other components of the HR pathway [20]. Specifically, germline BRCA1 and BRCA2 mutations are the most common alterations and are present in up to 22 % of the HGSOC, whereas somatic BRCA1 and BRCA2 mutations have been identified in 6 to 7 % of HGSOC [10••, 21, 22]. The majority of the BRCA1 and BRCA2 mutations (81 and 72 %, respectively) are also followed by loss of heterozygosity, which indicates a biallelic inactivation [10••]. Mutations in BRCA1/2 genes are mainly frameshift insertions or deletions and occur in all functional domains of BRCA1/2 genes [10••]. Other HR pathway alterations include mutations in many Fanconi anemia (FA) genes (mainly PALB2, FANCA, FANCI, FANCL, and FANCC), in core HR restriction site associated DNA (RAD) genes (RAD50, RAD51, RAD51C, and RAD54L) and in DNA damage response genes involved in HR (ATM, ATR, CHEK1, and CHEK2). Alterations in genes that modulate the HR pathway can also indirectly cause HR deficiency. Mendes-Pereira and colleagues proposed that phosphatase and tensin homolog (PTEN) deficiency could be linked to transcriptional downregulation of RAD51 and therefore cause a HR defective phenotype [23]. EMSY overexpression and amplification is another mechanism of HR deficiency, seen in 17 % of HGSOCs. This could be due to the interaction of EMSY with the transactivation domain of BRCA2, which leads to inhibition of its transcriptional activity [24]. However, the exact role of EMSY alteration in HR deficiency is still controversial.
In addition to HR pathway alterations, other DNA repair pathways, such as nucleotide excision repair (NER) and mismatch repair (MMR), can be affected through missense mutations or homozygous deletions in several genes (ERCC2-6, DDB1, XPC, RFC1, RAD23B, MNAT1); this is the case of approximately 8 % of HGSOC as reported by the TCGA [5••, 25].
Identification of reliable biomarkers that could capture the diverse mechanisms of HR deficiency represents a very important step for clinical treatment decisions [10••]. Among others, tumor gene expression profiles of BRCAness or targeted mutational profiling of HR genes are promising in identifying tumors with altered HR pathway. An example is BROCA, a targeted deep sequencing assay that identifies all types of mutations of key HR genes and that can predict an improved primary response to platinum chemotherapy and longer overall survival in epithelial ovarian cancer [26, 27]. Patients with HR defective tumors are associated with enhanced sensitivity to platinum-based chemotherapy; therefore, platinum sensitivity has been used as “clinical biomarker” of HR deficiency [10••, 28]. Clinically, patients with BRCA1/2 mutations show significantly higher response rates and prolonged progression-free survival (PFS) after platinum-based chemotherapy. The importance of HR deficiency and platinum sensitivity is also highlighted by the fact that restoration of proficient HR by different mechanisms (such as secondary mutations restoring BRCA1/2 protein functionality or epigenetic reversion of BRCA1 promoter hypermethylation) induces platinum resistance [10••]. What makes HR deficient tumors clinically very interesting is their high sensitivity to poly ADP-ribose polymerase inhibitors (PARPis) a novel class of anticancer agents, which exhibit synthetic lethality with a defective HR pathway [25, 29, 30]. The strong activity of PARPis in HR-deficient epithelial ovarian cancer made this treatment the first molecular-targeted therapy approved in this disease. In December 2014, the FDA approved the PARP inhibitor olaparib for use in epithelial ovarian cancer patients with germline BRCA1/2 mutations after three or more chemotherapy regimens. Still, patients with HR-deficient/no-BRCA mutated tumors cannot benefit of these agents outside clinical trials [10••]. That highlights the importance of identifying biomarkers of HR deficiency in patients that could benefit from the PARPis without having BRCA1/2 mutations.
The molecular mechanism behind the synthetic lethal interaction of PARPis with HR deficiency is still under investigation, and several models were proposed so far. One of the most accredited is the essential role of PARP1 in the base excision repair (BER), which would be inhibited by PARPis leading to DNA double-strand breaks (DSBs) that are normally repaired by HR. In HR-deficient cells, these lesions will remained unrepaired and therefore will cause cytotoxicity [10••, 31]. Other models that could explain the PARPis-HR synthetic lethality are the role of PARP1 in inhibiting NHEJ repair activity, the trapping of PARP1 on DNA damage sites, the ability of PARP1 of recruiting DNA repair proteins, and last but not least, the PARP1 function in the alternative end joining (alt-EJ) DSB repair pathway, on which HR-deficient cells rely for survival [32–34]. Recently, D’Andrea and colleagues showed that there is an inverse correlation between HR activity and DNA polymerase θ (Polθ also known as POLQ) expression in EOCs. Specifically, knockdown of Polθ in HR-proficient cells upregulates HR activity and RAD51 assembly, while knockdown of Polθ in HR-deficient EOCs enhances cell death [32]. This work identified Polθ as a novel therapeutic target for PARPis.
However, it is important to remember that epithelial ovarian tumors that are sensitive to platinum-based treatment due to alterations in another DNA repair pathway than HR, namely the nucleotide excision repair (NER) pathway, may exhibit resistance to PARP inhibition as recently showed by Ceccaldi and colleagues [25].
Copy Number Alterations
With the exception of TP53, BRCA1, and BRCA2, mutations in oncogenes or tumor suppressor genes are relatively uncommon in HGSOC [2••]. Instead, HGSOC is mostly characterized by chromosomal instability with a large burden of copy number gains and losses making this type of cancer a good example of a chromosomally unstable malignancy, so-called C-class [2••, 35–37]. The TCGA researchers identified amplifications in CCNE1, MYC, and MECOM genes in at least 20 % of cases. Moreover, they found that areas of focal amplification encode the receptor for activated C-kinase (ZMYND8), a p53 target gene (IRF2BP2), a telomerase catalytic subunit (TERT), and, interestingly, an embryonic development gene (PAX8) that was also reported by Cheung and colleagues [13•]. Interestingly, PAX8 was the only gene in the Cheung study that hit the three selection criteria used by the authors: it was amplified primary ovarian tumors, overexpressed in ovarian cancer cell lines, and scored as “essential” by all three scoring methods. Of importance, PAX8 suppression induced apoptosis and viability reduction (>50 %) only in ovarian cell lines with PAX8 amplification and/or overexpression. Immunohistochemistry suggests that it is expressed in 90–100 % of serous, clear-cell, and endometrioid subtypes. PAX8 encodes a transcription factor involved, among others, in the development of the female reproductive tract. Its expression is restricted to secretory cells of the fallopian tube epithelium, which is the point of origin for serous ovarian carcinoma [13•].
The TCGA study reported also 50 focal deletions, and among others in known tumor suppressor genes PTEN, RB1, and NF1. Recently, Patch and colleagues showed that gene breakage inactivates not only tumor suppressors RB1, NF1, and PTEN but also RAD51B and contributes to acquired chemotherapy resistance in HGSOC [38••]. Additionally, the group of Brenton confirmed that PTEN loss is a common event in HGSOC and defines a subgroup with significantly worse prognosis, suggesting the rational use of drugs to target PI3K and androgen receptor pathways for HGSOC [39].
As previously discussed, approximately 50 % of HGSOC shows a defective HR DNA repair pathway. This phenotype is most likely a key determinant of platinum and PARPi sensitivity in HGSOC. However, the other half of HGSOC, which have no evidence of HR deficiency, is poorly characterized. Interestingly 20 % of this group has focal amplification of Cyclin E1 (CCNE1), which is an early event in HGSOC tumorigenesis. Clinically, tumors with CCNE1 gene amplification are associated with poor overall survival and primary treatment failure [40]. It has been showed that HGSOC cell lines with CCNE1 amplified undergo cell cycle arrest or apoptosis when cyclin E1 or its binding partner cyclin-dependent kinase 2 (CDK2) is lost [2••, 41]. This leads to the possibility of using CDK2 as novel therapeutic target in CCNE1-amplified tumors. Preclinical studies showed indeed that ovarian cell lines with CCNE1 amplification are selectively sensitive to CDK2 suppression using small-molecule inhibitors [40]. Nevertheless, resistance to CDK2 inhibitors may occur due to upregulation of CDK2 target protein and through preexisting cellular polyploidy [40]. Intriguingly, a recent study provides an explanation for the observed mutual exclusivity of CCNE1 amplification and BRCA1/2 loss in HGSOC, namely the intolerance of CCNE1 amplified cell lines of BRCA1 suppression [42]. CCNE1 overexpression promotes uncontrolled S-phase entry and therefore disrupts DNA replication and increased genomic instability. This can make cells dependent on an intact HR repair pathway with functional BRCA1. Additionally, cells that harbor CCNE1 amplification show dependency on members of the ubiquitin pathway as well. Based on these findings, the same study proposed a novel therapeutic approach where platinum-resistant tumors could be treated with bortezomib, which inhibits both the proteasome and the HR pathway [42].
As described in the previous section (“Somatic Mutations and HR Pathway Alterations”), CDK12 was found statistically significantly mutated gene in the TCGA patient cohort, and cells with inactivated CDK12 showed high sensitivity to DNA damaging as PARPis. Only very recently, Popova and colleagues reported that ovarian tumors with CDK12 inactivation show a specific genomic instability pattern consisting of a >200 tandem duplications (TDs) up to 10 Mb [18]. This so-called CDK12 TD-plus phenotype is observed in 4 % of the HGSOC samples included in their study (∼650 cases).
mRNA and miRNA Expression Analysis
Large-scale genomics studies have identified novel molecular subtypes of HGSOC that are characterized by distinct biology and are prognostically relevant with potential therapeutic implications. First, a gene expression analysis of HGSOC and endometrioid OCs from the Australian Ovarian Cancer Study identified six different subtypes, of which four subtypes represented higher grade and advanced stage cancers of serous and endometrioid morphology [43]. Tothill and colleagues called these four subtypes C1 (high stromal response), C2 (high immune signature), C4 (low stromal response), and C5 (mesenchymal, low immune signature). These four subtypes were then validated by the TCGA group using an independent set of 489 HGSOCs and renamed as proliferative, immunoreactive, differentiated, and mesenchymal. The proliferative subtype is characterized by high expression of proliferative markers (MCM2, PCNA) and transcription factors (HMGA2, SOX11) and low expression of ovarian tumor markers (MUC1 and MUC16) [5••]. Patients with these tumors displayed a significant trend toward early relapse and short OS [43]. In addition, this group shows a decrease in the rate of MYC amplification and RB1 deletion compared to the other subtypes. The immunoreactive subtype expresses T-cell chemokine ligands (CXCL11 and CXCL10) and the receptor CXCR3. The immune subtype may benefit from use of immune checkpoint inhibitors also in light of what Zhang and colleagues reported that a subset of HGSOC shows intratumoral infiltration with activated lymphocytes, such as CD8+ T-cells [44]. The differentiated subtype is associated with high expression of MUC1 and MUC16, and it represents an advanced stage of development reflected by the expression of SLP1, the secretory fallopian tube marker. The mesenchymal subtype is characterized by a de-differentiated phenotype with overexpression of the WNT signaling genes, developmental transcription factors, and reduced expression of E-cadherin, which is suggestive of epithelial-mesenchymal transition [43]. It displays high expression of HOX genes and markers of increased stromal components [5••]. Interestingly, Tothill and colleagues observed low numbers of intratumoral CD3+ T-cell numbers for subtypes C1 and C5, both of which are associated with poorer prognosis compared with other groups. When the TCGA investigated whether there were mutations significantly associated with the subtypes, only CDK12 mutations appeared to be nearly significantly enriched in the differentiated and in the immunoreactive subtypes (corrected p value = 0.055) [5••]. This result might be related to the rather homogeneous mutation spectrum of HGSOC with relatively uncommon mutations in oncogenes or tumor suppressor genes and the majority of the tumors having mutations in TP53 gene.
However, in spite of clear biological differences between the molecular subtypes and the variation in outcome observed in HGSOC patients (also based on stage and residual disease), the TCGA subtypes did not differ significantly in survival time. Therefore, the TCGA network created a classification tool termed CLOVAR (Classification of Ovarian Cancer) that combines the original TCGA subtype gene signatures with survival gene expression signatures and allows stratifying HGSOCs based on survival and treatment outcome [4]. Using CLOVAR, the authors could identify different risk groups, and the group with worse outcome showed a median survival of 23 months and a platinum resistance rate of 63 %. Still, CLOVAR needs to be prospectively validated also including other prognostic factors such as BRCA1/2 mutation status and residual disease. More recently, Konecny and colleagues proposed a de novo classification for molecular subtyping the HGOCs. They confirmed the presence of four stable molecular TCGA subtypes in two independent patient cohorts, and more importantly they added a prognostic value to them [45••]. The immunoreactive-like subtype had the best survival, whereas the proliferative-like and the mesenchymal-like subtypes had a significantly worse survival. However, both the TGCA and the Konecny studies showed that individual HGSOC samples could be assigned to more than one subtype at the time when using alternative ways of sample clustering and statistics. This points out that the same tumor can express multiple subtype signatures at different levels of activation, or, in other words, that HGOC does not consist of mutually exclusive expression subtypes, as it is more the case of other cancer types (i.e., breast, colon, or glioblastoma) [45••]. As Verhaak and colleagues interestingly underlined, infiltrating cells have a major impact on the expression patterns of HGSOC, and purified HGSOC cells could express novel signatures which are now masked by tumor microenvironment signals [4]. Taken together, it emerges that HGSOC is a highly heterogeneous disease both at molecular and clinical level. Therefore, a classification approach that could take into account a spectrum with overlapping phenotypes may be more informative for stratifying HGSOCs with clinical relevance. This can be the reason why these molecular subtypes have not yet been integrated into the clinical setting. Thus, before these subtypes can be effectively used in the clinic, more extensive validation of their prognostic value and additionally, an accurate assessment of their predictive value (i.e., association with treatment response) are required.
Recent studies tried to map the micro RNA (miRNA) regulatory network in HGOC and have identified specific miRNA that regulate the epithelial-mesenchymal transition in the mesenchymal subtype [5••, 46–48]. The TCGA group identified three miRNA subtypes (1, 2, 3) which partially overlapped with the messenger RNA (mRNA) expression subtypes. The miRNA 1 had a significantly longer survival and overlapped with the proliferative subtype, whereas the miRNA 2 showed similarities with the mesenchymal subtype. Yang and colleagues investigated further the mesenchymal HGSOC subtype integrating mRNA, miRNA, DNA copy number and methylation data from the TCGA cohort (N = 459 cases), and selected a set of 219 genes targeted by 19 miRNAs, which included EMT (epithelial-mesenchymal transition) inducers such as SNAI2 and ZEB2. The 19 miRNAs were dowregulated and inhibited EMT. Using these 219 genes, the authors were able to group three independent patient groups in so-called iM (mesenchymal) and iE (epithelial) subtypes that significantly differ in overall survival. Eight key miRNAs were the master regulators for the iM subtype, which robustly associated with poor OS in HGSOC patients. Additionally, among these eight miRNAs, the miR-506 was extensively characterized and proved to be a potent EMT inhibitor. They showed that miR-506 directly targets SNAI2 that induces EMT by repressing E-cadherin [49]. Interestingly, miR-506 induced E-cadherin expression and reduced tumor growth when it was delivered into lipid-based nanoparticles in orthotopic mouse models [47]. This highlights a potential therapeutic value of miR-506, which will need further investigation also with regard to treatment sensitivity. A subsequent study identified another miRNA, miR-181a that promoted TGFβ-mediated EMT by repressing its functional target, SMAD7 [46]. The authors used the P-SMAD2 protein levels to assess the activation status of the TGFβ-pathway. In patients, higher levels of P-SMAD2 significantly correlated with shorter median progression-free interval (PFI) and survival, whereas concomitant decrease in both miR-181a and P-SMAD2 expression was associated to significantly prolonged PFI and OS. Moreover, the authors observed that miR-181a expression is increased in recurrent HGSOC clinical tumors compared with primary chemotherapy-naïve tumors and that miR-181a expression is inversely correlated with SMAD7 expression. Taken together, the study emphasized the importance of TGFβ-pathway in HGSOC and its role in contributing to poor patient outcome.
Another group profiled microRNA expression in serous ovarian carcinoma with the attempt of finding predictive markers of treatment response [48]. They focused their attention on the miR-484 that appeared to be significantly dowregulated in non-responders tumors. Interestingly, miR-484 targets the VEGF signaling by both modulating the VEGFB protein of tumor cells and VEGFR2 receptor of tumor-associated endothelial cells. The modulation of both VEGFB and VEGFR2 leads to better control of the tumor neovascularization, which plays an important role in ovarian cancer progression. These findings support once again the importance of neoangiogenesis in HGSOC patients. HGSOC express high levels of proangiogenic proteins, and this contributes to the formation of ascites in patients [2••]. Many groups explored the activity of antiangiogenic agents in HGSOC, such as the VEGF-specific monoclonal antibody bevacizumab. Clinical trials showed that bevacizumab increases progression-free survival (PFS) in ovarian patients in combination with chemotherapy in a first-line setting and also in platinum-resistant recurrent HGSOC [50–52]. However, low response rate, high toxicity, de novo resistance, and no evidence of increase in overall survival (OS), downsized the use of bevacizumab in a standard clinical setting. As Bowtell and colleagues rightly pointed out, greater effort should be placed on identifying predictive biomarkers of response using tissue and blood samples of patients enrolled in clinical trials [2••]. A recent attempt came from the transcriptional profiling of samples collected during the ICON7 clinical trial. In this study, researchers identified a molecular subgroup of HGSOC that highly expressed immune response genes and that showed a shorter PFS and OS after receiving bevacizumab maintenance therapy compared to those who did not received bevacizumab maintenance therapy [53]. Therefore, the use of miRNA (e.g., miR-484) and mRNA molecular signatures could potentially identify those patients that will effectively benefit from the addition of antiangiogenic compounds.
Conclusions
High-grade serous ovarian cancer remains one of the most lethal gynecologic malignancies in women. A detailed characterization at a molecular level is essential for developing new treatment strategies to add to the preexisting standard therapy. Many studies in recent years, and particularly the TCGA network group, addressed this issue and have begun to unravel the high degree of clinical and molecular heterogeneity of this disease. What is emerging is that an accurate patient stratification based on molecular features such as HR deficiency, PTEN loss, CCNE1 amplification, and molecular subtypes will contribute to patient stratification for selected drugs.
References
Papers of particular interest, published recently, have been highlighted as • Of importance •• Of major importance
Ross JS, Ali SM, Wang K, Palmer G, Yelensky R, Lipson D, et al. Comprehensive genomic profiling of epithelial ovarian cancer by next generation sequencing-based diagnostic assay reveals new routes to targeted therapies. Gynecol Oncol. 2013;130(3):554–9.
Bowtell DD, Bohm S, Ahmed AA, Aspuria PJ, Bast Jr RC, Beral V, et al. Rethinking ovarian cancer II: reducing mortality from high-grade serous ovarian cancer. Nat Rev Cancer. 2015;15(11):668–79. In this Opinion article the authors outline seven research priorities which should improve outcome and possibly reduce incidence of such a deadly disease as high grade serous ovarian carcinoma.
Itamochi H, Kigawa J. Clinical trials and future potential of targeted therapy for ovarian cancer. Int J Clin Oncol. 2012;17(5):430–40.
Verhaak RG, Tamayo P, Yang JY, Hubbard D, Zhang H, Creighton CJ, et al. Prognostically relevant gene signatures of high-grade serous ovarian carcinoma. J Clin Invest. 2013;123(1):517–25.
Integrated genomic analyses of ovarian carcinoma. Nature. 2011;474(7353):609–15. This is the first comprehensive molecular characterization of high grade serous ovarian carcinoma that integrates mutation, copy number, expression and methylation data.
Jemal A, Siegel R, Ward E, Hao Y, Xu J, Thun MJ. Cancer statistics, 2009. CA Cancer J Clin. 2009;59(4):225–49.
Liu Z, Beach JA, Agadjanian H, Jia D, Aspuria PJ, Karlan BY, et al. Suboptimal cytoreduction in ovarian carcinoma is associated with molecular pathways characteristic of increased stromal activation. Gynecol Oncol. 2015.
Yap TA, Carden CP, Kaye SB. Beyond chemotherapy: targeted therapies in ovarian cancer. Nat Rev Cancer. 2009;9(3):167–81.
Coward JI, Middleton K, Murphy F. New perspectives on targeted therapy in ovarian cancer. Int J Women’s Health. 2015;7:189–203.
Konstantinopoulos PA, Ceccaldi R, Shapiro GI, D’Andrea AD. Homologous recombination deficiency: exploiting the fundamental vulnerability of ovarian cancer. Cancer Discov. 2015;5(11):1137–54. In this interesting review several aspects of the homologous recombination (HR) deficiency of epithelial ovarian carcinoma are described with the intent of highlighting potential biomarkers of HR in addition to the well characterized BRCA1/2 mutation status. In addition they emphasized the importance of the de novo and acquired resistance observed in tumors followed by PARP inhibition.
Cerami E, Gao J, Dogrusoz U, Gross BE, Sumer SO, Aksoy BA, et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2012;2(5):401–4.
Gao J, Aksoy BA, Dogrusoz U, Dresdner G, Gross B, Sumer SO, et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal. 2013;6(269):l1.
Cheung HW, Cowley GS, Weir BA, Boehm JS, Rusin S, Scott JA, et al. Systematic investigation of genetic vulnerabilities across cancer cell lines reveals lineage-specific dependencies in ovarian cancer. Proc Natl Acad Sci U S A. 2011;108(30):12372–7. The authors performed systematic loss-of-function studies to identify essential genes in different cancer cell types. Suppression of PAX8, a gene found amplified in 16% of high grade serous ovarian carcinoma, selectively induced apoptotic cell death of ovarian cancer cells. High-throughput in vitro functional studies should be considered for identification of tumor vulnerabilities, which are potentially targetable.
Bowtell DD. The genesis and evolution of high-grade serous ovarian cancer. Nat Rev Cancer. 2010;10(11):803–8.
Ahmed AA, Etemadmoghadam D, Temple J, Lynch AG, Riad M, Sharma R, et al. Driver mutations in TP53 are ubiquitous in high grade serous carcinoma of the ovary. J Pathol. 2010;221(1):49–56.
Joshi PM, Sutor SL, Huntoon CJ, Karnitz LM. Ovarian cancer-associated mutations disable catalytic activity of CDK12, a kinase that promotes homologous recombination repair and resistance to cisplatin and poly(ADP-ribose) polymerase inhibitors. J Biol Chem. 2014;289(13):9247–53.
Bajrami I, Frankum JR, Konde A, Miller RE, Rehman FL, Brough R, et al. Genome-wide profiling of genetic synthetic lethality identifies CDK12 as a novel determinant of PARP1/2 inhibitor sensitivity. Cancer Res. 2014;74(1):287–97. The authors provide evidences that CDK12 deficiency is a clinically relevant biomarker of PARP1/2 inhibitor sensitivity. This study investigated further the role of CDK12 as novel biomarker for high grade serous ovarian carcinoma targeted therapy.
Popova T, Manie E, Boeva V, Battistella A, Goundiam O, Smith NK, et al. Ovarian cancers harboring inactivating mutations in CDK12 display a distinct genomic instability pattern characterized by large tandem duplications. Cancer Res. 2016.
Ekumi KM, Paculova H, Lenasi T, Pospichalova V, Bosken CA, Rybarikova J, et al. Ovarian carcinoma CDK12 mutations misregulate expression of DNA repair genes via deficient formation and function of the Cdk12/CycK complex. Nucleic Acids Res. 2015;43(5):2575–89.
Walsh T, Lee MK, Casadei S, Thornton AM, Stray SM, Pennil C, et al. Detection of inherited mutations for breast and ovarian cancer using genomic capture and massively parallel sequencing. Proc Natl Acad Sci U S A. 2010;107(28):12629–33.
Alsop K, Fereday S, Meldrum C, deFazio A, Emmanuel C, George J, et al. BRCA mutation frequency and patterns of treatment response in BRCA mutation-positive women with ovarian cancer: a report from the Australian Ovarian Cancer Study Group. J Clin Oncol Off J Am Soc Clin Oncol. 2012;30(21):2654–63.
Pal T, Permuth-Wey J, Betts JA, Krischer JP, Fiorica J, Arango H, et al. BRCA1 and BRCA2 mutations account for a large proportion of ovarian carcinoma cases. Cancer. 2005;104(12):2807–16.
Mendes-Pereira AM, Martin SA, Brough R, McCarthy A, Taylor JR, Kim JS, et al. Synthetic lethal targeting of PTEN mutant cells with PARP inhibitors. EMBO Mol Med. 2009;1(6–7):315–22.
Hughes-Davies L, Huntsman D, Ruas M, Fuks F, Bye J, Chin SF, et al. EMSY links the BRCA2 pathway to sporadic breast and ovarian cancer. Cell. 2003;115(5):523–35.
Ceccaldi R, O’Connor KW, Mouw KW, Li AY, Matulonis UA, D’Andrea AD, et al. A unique subset of epithelial ovarian cancers with platinum sensitivity and PARP inhibitor resistance. Cancer Res. 2015;75(4):628–34.
Pennington KP, Walsh T, Harrell MI, Lee MK, Pennil CC, Rendi MH, et al. Germline and somatic mutations in homologous recombination genes predict platinum response and survival in ovarian, fallopian tube, and peritoneal carcinomas. Clin Cancer Res Off J Am Assoc Cancer Res. 2014;20(3):764–75.
Konstantinopoulos PA, Spentzos D, Karlan BY, Taniguchi T, Fountzilas E, Francoeur N, et al. Gene expression profile of BRCAness that correlates with responsiveness to chemotherapy and with outcome in patients with epithelial ovarian cancer. J Clin Oncol Off J Am Soc Clin Oncol. 2010;28(22):3555–61.
Yang D, Khan S, Sun Y, Hess K, Shmulevich I, Sood AK, et al. Association of BRCA1 and BRCA2 mutations with survival, chemotherapy sensitivity, and gene mutator phenotype in patients with ovarian cancer. JAMA. 2011;306(14):1557–65.
Farmer H, McCabe N, Lord CJ, Tutt AN, Johnson DA, Richardson TB, et al. Targeting the DNA repair defect in BRCA mutant cells as a therapeutic strategy. Nature. 2005;434(7035):917–21.
Bryant HE, Schultz N, Thomas HD, Parker KM, Flower D, Lopez E, et al. Specific killing of BRCA2-deficient tumours with inhibitors of poly(ADP-ribose) polymerase. Nature. 2005;434(7035):913–7.
McCabe N, Turner NC, Lord CJ, Kluzek K, Bialkowska A, Swift S, et al. Deficiency in the repair of DNA damage by homologous recombination and sensitivity to poly(ADP-ribose) polymerase inhibition. Cancer Res. 2006;66(16):8109–15.
Ceccaldi R, Liu JC, Amunugama R, Hajdu I, Primack B, Petalcorin MI, et al. Homologous-recombination-deficient tumours are dependent on poltheta-mediated repair. Nature. 2015;518(7538):258–62.
Li M, Yu X. Function of BRCA1 in the DNA damage response is mediated by ADP-ribosylation. Cancer Cell. 2013;23(5):693–704.
Langelier MF, Pascal JM. PARP-1 mechanism for coupling DNA damage detection to poly(ADP-ribose) synthesis. Curr Opin Struct Biol. 2013;23(1):134–43.
Wang ZC, Birkbak NJ, Culhane AC, Drapkin R, Fatima A, Tian R, et al. Profiles of genomic instability in high-grade serous ovarian cancer predict treatment outcome. Clin Cancer Res Off J Am Assoc Cancer Res. 2012;18(20):5806–15.
Sohn I, Jung WY, Sung CO. Somatic hypermutation and outcomes of platinum based chemotherapy in patients with high grade serous ovarian cancer. Gynecol Oncol. 2012;126(1):103–8.
Ciriello G, Miller ML, Aksoy BA, Senbabaoglu Y, Schultz N, Sander C. Emerging landscape of oncogenic signatures across human cancers. Nat Genet. 2013;45(10):1127–33.
Patch AM, Christie EL, Etemadmoghadam D, Garsed DW, George J, Fereday S, et al. Whole-genome characterization of chemoresistant ovarian cancer. Nature. 2015;521(7553):489–94. This is one of the first studies that have analyzed in depth using transcriptome, methylome and microRNA expression analysis paired primary and relapse high-grade serous ovarian carcinoma samples to highlight new mechanisms of platinum-based chemotherapy resistance.
Martins FC, Santiago I, Trinh A, Xian J, Guo A, Sayal K, et al. Combined image and genomic analysis of high-grade serous ovarian cancer reveals PTEN loss as a common driver event and prognostic classifier. Genome Biol. 2014;15(12):526.
Etemadmoghadam D, Au-Yeung G, Wall M, Mitchell C, Kansara M, Loehrer E, et al. Resistance to CDK2 inhibitors is associated with selection of polyploid cells in CCNE1-amplified ovarian cancer. Clin Cancer Res Off J Am Assoc Cancer Res. 2013;19(21):5960–71.
Etemadmoghadam D, George J, Cowin PA, Cullinane C, Kansara M, Australian Ovarian Cancer Study G, et al. Amplicon-dependent CCNE1 expression is critical for clonogenic survival after cisplatin treatment and is correlated with 20q11 gain in ovarian cancer. PLoS One. 2010;5(11):e15498.
Etemadmoghadam D, Weir BA, Au-Yeung G, Alsop K, Mitchell G, George J, et al. Synthetic lethality between CCNE1 amplification and loss of BRCA1. Proc Natl Acad Sci U S A. 2013;110(48):19489–94.
Tothill RW, Tinker AV, George J, Brown R, Fox SB, Lade S, et al. Novel molecular subtypes of serous and endometrioid ovarian cancer linked to clinical outcome. Clin Cancer Res Off J Am Assoc Cancer Res. 2008;14(16):5198–208.
Zhang L, Conejo-Garcia JR, Katsaros D, Gimotty PA, Massobrio M, Regnani G, et al. Intratumoral T cells, recurrence, and survival in epithelial ovarian cancer. N Engl J Med. 2003;348(3):203–13.
Konecny GE, Wang C, Hamidi H, Winterhoff B, Kalli KR, Dering J, et al. Prognostic and therapeutic relevance of molecular subtypes in high-grade serous ovarian cancer. J Natl Cancer Inst. 2014;106(10). The authors robustly validated the four molecular subtypes of high grade serous ovarian carcinoma previously described using three independent patient cohorts in order to add a significant prognostic value to the subtypes.
Parikh A, Lee C, Joseph P, Marchini S, Baccarini A, Kolev V, et al. microRNA-181a has a critical role in ovarian cancer progression through the regulation of the epithelial-mesenchymal transition. Nat Commun. 2014;5:2977.
Yang D, Sun Y, Hu L, Zheng H, Ji P, Pecot CV, et al. Integrated analyses identify a master microRNA regulatory network for the mesenchymal subtype in serous ovarian cancer. Cancer Cell. 2013;23(2):186–99.
Vecchione A, Belletti B, Lovat F, Volinia S, Chiappetta G, Giglio S, et al. A microRNA signature defines chemoresistance in ovarian cancer through modulation of angiogenesis. Proc Natl Acad Sci U S A. 2013;110(24):9845–50.
Kurrey NK, Amit K, Bapat SA. Snail and slug are major determinants of ovarian cancer invasiveness at the transcription level. Gynecol Oncol. 2005;97(1):155–65.
Perren TJ, Swart AM, Pfisterer J, Ledermann JA, Pujade-Lauraine E, Kristensen G, et al. A phase 3 trial of bevacizumab in ovarian cancer. N Engl J Med. 2011;365(26):2484–96.
Burger RA, Brady MF, Bookman MA, Fleming GF, Monk BJ, Huang H, et al. Incorporation of bevacizumab in the primary treatment of ovarian cancer. N Engl J Med. 2011;365(26):2473–83.
Pujade-Lauraine E, Hilpert F, Weber B, Reuss A, Poveda A, Kristensen G, et al. Bevacizumab combined with chemotherapy for platinum-resistant recurrent ovarian cancer: the AURELIA open-label randomized phase III trial. J Clin Oncol Off J Am Soc Clin Oncol. 2014;32(13):1302–8.
Gourley G. Molecular subgroup of high-grade serous ovarian cancer (HGSOC) as a predictor of outcome following bevacizumab. J Clin Oncol. 2014;32.
Acknowledgments
The author would like to thank Professor René Bernards (NKI/AVL Amsterdam) and Astrid J Bosma (NKI/AVL Amsterdam) for critical reading of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
Lorenza Mittempergher declares that she has no conflict of interest.
Human and Animal Rights and Informed Consent
This article does not contain any studies with human or animal subjects performed by any of the authors.
Additional information
This article is part of the Topical Collection on Translational Oncology
Rights and permissions
About this article
Cite this article
Mittempergher, L. Genomic Characterization of High-Grade Serous Ovarian Cancer: Dissecting Its Molecular Heterogeneity as a Road Towards Effective Therapeutic Strategies. Curr Oncol Rep 18, 44 (2016). https://doi.org/10.1007/s11912-016-0526-9
Published:
DOI: https://doi.org/10.1007/s11912-016-0526-9