Abstract
To investigate the impact of florfenicol on soil microbial community structure and diversity, an indoor florfenicol exposure model was established. Soil samples were collected at different concentrations of florfenicol (0, 0.05, 0.5, 5, and 50 mg/kg) on days 0, 7, 30, and 60. High-throughput sequencing was employed to examine the changes in soil microbial community structure, diversity, and abundance. The analysis revealed a total of 31874 operational taxonomic units (OTUs) in the soil samples, averaging 9524 OTUs per sample. The number of soil microbial community OTUs declined with increasing florfenicol concentration. The microbial species richness and diversity showed a decreasing trend at day 7, while the treatment group with 50 mg/kg florfenicol at 60 days exhibited the lowest richness index. Examination of the soil microbial community structure identified 50 phyla and 1303 genera. At the phylum level, the abundance of Actinobacteria and Bacteroides decreased with increasing florfenicol concentration. Similarly, at the genus level, some of the dominant genera displayed a decline in abundance with the rise of florfenicol concentration. Cluster analysis demonstrated significant temporal and concentration variability. The results indicate differences in the composition, diversity, dominance, abundance, and evenness of soil bacterial communities among different groups. Florfenicol has a significant negative impact on the structure and diversity of soil microbial communities. The findings of this study provide a scientific basis for the rational use of florfenicol in livestock farming to maintain a healthy and stable soil microecological environment.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Antibiotics serve not only as essential medications for preventing and treating bacterial infections but also as growth promoters extensively employed in livestock farming and aquaculture practices. After antibiotics are administered to animals, most of them are discharged into the environment in their original form, resulting in antibiotic residues in farming areas and discharged wastewater. Research has revealed that approximately 30–90% of antibiotics consumed by livestock are excreted through manure and urine (Qiu et al. 2021), which can subsequently enter the soil through manure application. The utilization of over 150 different antibiotics has been documented (Tasho and Cho 2016). Florfenicol, a broad-spectrum antimicrobial agent classified as an aminol, has gained widespread usage in Chinese livestock industry. Florfenicol exerts its effects by targeting the 50S subunit of the bacterial ribosome, thereby inhibiting trans-peptidase activity and impeding protein synthesis through the blockage of protein peptide chain elongation (Zeng et al. 2019). If left untreated, large quantities of livestock manure containing antibiotics can be directly discharged into agricultural lands, rivers, and lakes, thereby polluting the ecological environment. Studies have reported the presence of florfenicol in soil samples from Shandong and Jiangsu provinces in China, with concentrations of 0.302 μg/L and 11 μg/L, respectively (Wang et al. 2021). Furthermore, in eastern China, animal manure-amended soil exhibited the presence of thirteen antibiotics, including florfenicol (Wei et al. 2016). Florfenicol is primarily excreted through the kidneys after ingestion by animals, with higher concentrations found in urine. Due to its predominant excretion in its original form, it retains biological activity. Once introduced into the soil, it can be adsorbed by soil particles, continuously migrate, settle, and accumulate, posing toxicity to non-target organisms, generating threats such as the emergence of drug-resistant strains, and potentially causing a decrease in microbial biomass, reduced activity, and alterations in community structure. The frequency of florfenicol use in farms can impact the drug's accumulation in adjacent soil, and higher concentrations may exert greater inhibitory effects on soil microorganisms. However, this inhibition may diminish over time.
Microorganisms play a crucial role in soil ecosystems, contributing significantly to improving soil fertility and participating in the material cycle and energy flow in nature (Coban et al. 2022). However, the introduction of antibiotics into soil can have detrimental effects on the viability, community structure, metabolic function, and population size of soil microorganisms. These impacts can disrupt the soil nutrient cycle and self-purification capacity by altering biodiversity, biomass, and biota (Fang et al. 2014). Hu et al. reported that high concentrations of florfenicol inhibited soil microorganisms (Hu et al. 2011). Zhang et al. Similarly, demonstrated that prolonged exposure to elevated concentrations of florfenicol in Sediments suppressed the biological activity of soil microbial communities (Zhang et al. 2023). In this study, high-throughput sequencing techniques (Lo Giudice et al. 2021) were employed to examine the impact of florfenicol on the structure, diversity, and abundance of bacterial communities. To achieve this, different concentrations of florfenicol were introduced into the soil, and the samples were subsequently analyzed.
Materials and methods
Reagents
Florfenicol, purchased from North China Pharmaceutical Company Limited (Lot No. 13021322); Soil DNA Kit was purchased from Omega Bio-tek Incorporation (Lot No. D5625010000L22T024).
Study design
Soil samples were collected from the plant plot at the Guangdong Ocean University Horticulture Practice Base. The surface layer of soil was manually removed, and subsequent soil samples were acquired from a depth of 10–20 cm below ground level. The collected samples were then subjected to a series of procedures, including drying, screening using a 4 mm diameter sieve, and removal of debris prior to being packed into plastic containers. The soil specimens were organized into 5 distinct groups, with each group consisting of 3 replicates. Florfenicol was introduced to the soil at concentrations of 0, 0.05, 0.5, 5, and 50 mg/kg, denoted as S1 to S5, respectively. Subsequent sampling events were conducted on days 0, 7, 30, and 60 (referred to as D0, D7, D30, and D60), following a continuous incubation within an artificial climatic chamber. The chamber was programmed to maintain a temperature of (28 ± 1)°C, a humidity level of 75% ± 7%, intermittent light exposure with a 12-h on/off cycle, and a light intensity of 1333 lx.
DNA extraction
Total DNA was extracted from the soil samples using the Soil DNA Kit according to the manufacturer's instructions. Following extraction, the concentration and purity of the obtained DNA were evaluated using the Nanodrop UV–Vis spectrophotometer.
High throughput sequencing and data analysis
The extracted DNA samples were submitted to Illumina MiSeq sequencing. To optimize the data quality, QIIME was employed, which involved splicing the overlapping regions at the ends, eliminating sequences containing "N" in the splicing results, removing spliced sequences shorter than 200 bp, and discarding chimeric sequences. The remaining valid sequences were subjected to Operational Taxonomic Units (OTUs) clustering analysis at a 97% similarity threshold. Species taxonomic annotation was performed using the Silva_138 16S rRNA database. Based on OTUs results, alpha diversity indices, such as Shannon and Chao1, reflecting species abundance and diversity, were calculated by random draw leveling of the sample sequences. (Un)weighted UniFrac analysis was employed to assess whether there were significant microbial community differences among the samples. β diversity analysis was conducted using R language. Non-metric multidimensional scaling method (NMDS) is based on the Bray–Curtis inter-sample distance matrix and serves as a visualization tool for the β diversity graph. The Unweighted Pair Group Method with Arithmetic Mean (UPGMA) clustering tree was constructed using the unweighted pair group average method in hierarchical clustering.
Results
Analysis of OTUs
Upon exclusion of low-quality data, the valid sequence counts obtained from the soil samples ranged from 79,020 to 654,877. Clustering analysis of the sequences yielded a total of 31,874 OTUs from the soil samples, with an average of 9524 OTUs per sample. A notable trend was observed with the addition of florfenicol, wherein the number of soil bacterial OTUs decreased compared to the control group. Moreover, a dose-dependent relationship was evident, with higher concentrations of florfenicol leading to a decrease in bacterial OTUs. Among the treated groups, the S5D7 group exhibited the lowest OTUs count at 5949. Additionally, Venn diagrams were constructed based on the OTUs results, highlighting that the soil samples treated with a florfenicol concentration of 50 mg/kg displayed the lowest number of shared OTUs (Fig. 1A). Notably, the soil samples collected at 7 days of incubation demonstrated the lowest number of OTUs in common (Fig. 1B).
Effects of different concentrations of florfenicol on operational taxonomic units (OTUs) of bacteria (venn diagram) in soil. (A) Effects of different concentrations of florfenicol on OTUs of bacteria in soil samples at 7 days. (B) Effect of sampling time on OUTs of bacteria in group S5 samples. Circles of different colors represent distinct samples/groups, and the numbers in the diagram denote the unique or shared OTU counts for each sample/group. Each petal in the petal diagram represents a sample/group, with the numbers on the petals indicating the sample-specific OTU counts. The white circle in the middle represents the shared OTU counts across all samples/groups
Rarefaction curve
Rarefaction curve serve as a valuable tool for evaluating the adequacy of sequencing efforts in capturing the full spectrum of observed taxa within a given soil sample, providing an indirect reflection of sample species richness. A plateau or leveling-off of the rarity curve indicates that the sequencing depth has effectively covered the entirety of species present in the sample (Gao et al. 2021). Figure 2 illustrates the relationship between the number of detected OTUs and the increasing number of valid sequences. Notably, the curve gradually reaches a saturation point, suggesting that the sequencing data sufficiently represents the bacterial taxa present within the soil samples, thereby enabling reliable estimations.
Effects of different concentrations of florfenicol on dilution curve of operational taxonomic units (OTUs) in the soil model. The x-axis represents the effective sequence count extracted per sample (Sequences Per Sample), and the y-axis represents the number of observed OTUs. Each curve in the graph represents an individual sample, differentiated by different colors. As the sequencing depth increases, the number of OTUs also increases. When the curve levels off, it indicates that with an increase in the amount of extracted data, the detected OTU count no longer rises. At this point, the sequencing data volume is considered reasonable
Alpha diversity index analysis
Microbial richness index, such as the Chao1 index and ACE index, along with microbial diversity index, including the Shannon index and Simpson index, are commonly used to assess microbial community characteristics. A higher Chao1 index value signifies a larger total species count, while a greater ACE index indicates higher species richness within the community. Moreover, elevated Shannon and Simpson index values correspond to increased community diversity (Lozupone and Knight 2008). As demonstrated in Table 1, the sample coverage values exceeded 92% for all samples, confirming sufficient sequencing depth for evaluating bacterial biodiversity across the soil samples. On the 7th day, the increase in florfenicol concentration was accompanied by a more pronounced decreasing trend in species richness and diversity. With the progression of time, the S1 and S2 groups exhibited a consistent decline in species richness and diversity. Thus, overall, the inhibitory effect of florfenicol on bacterial species richness and diversity was observed exclusively in samples from groups S1, S2, and D7, whereby the sample S3D7 demonstrated the lowest value for the diversity index, and the sample S5D7 displayed the lowest value for the richness index.
Bacterial community structure analysis
The RDP classifier method implemented in Qiime software was employed to classify the representative sequences of the obtained OTUs at the species level. The resulting taxonomic information was utilized to analyze the bacterial community structure. Figure 3 provides a visual representation of the taxonomic composition at the phylum level. A total of 50 phyla were identified within the soil samples, with the dominant bacteria observed as Proteobacteria, Acidobacteriota, Actinobacteriota, and Bacteroidota. Notably, Proteobacteria emerged as the predominant group across all soil samples, exhibiting relative abundances ranging from 36.62% to 59.05%. The relative abundance of Proteobacteria was significantly higher in the S3D0 and S4D0 samples compared to other groups. Acidobacteria, on the other hand, displayed relative abundances ranging from 6.48% to 20.02%. In contrast to the Proteobacteria, the relative abundance of Acidobacteria in the S3D0 and S4D0 samples exhibited a decreasing trend. The relative abundance of Actinomycetes and Anaplasma ranged from 7.04% to 13.04% and 4.79% to 18.53%, respectively. Notably, the abundance of these samples tended to decrease with increasing florfenicol concentrations. However, at 30 and 60 days, the abundance values of Actinobacteriota, Bacteroidota and Acidobacteriota increased when the florfenicol concentration reached 50 mg/kg.
Effects of different concentrations of florfenicol on microbial abundance at the phylum level of soil microbiota. The x-axis represents sample names or group names, while the y-axis represents the relative abundance of different species, and the legend includes the taxonomic names of species at different classification levels. "Other" represents the cumulative relative abundances of all phylum-level classifications beyond the top 30
The taxonomic analysis at the genus level revealed a total of 1303 genera within the soil samples, with the number of groups ranging from 662 to 903. Heatmap visualizations were generated, focusing on the Top30 genera at the species level. Figure 4 illustrates the composition of the soil bacterial community and its dominant species. Notably, Sphingomonas, RB41, Nitrospira, SC-1–84, Chitinophagaceae, Xanthobacteraceae, Vicinamibacteraceae, Bryobacter, Sphingomonadaceae and Gemmatimonadaceae emerged as the dominant bacteria at the genus level. While the overall composition remained similar across the different treatments, variations were observed in terms of mean relative abundance.
Effects of different concentrations of florfenicol on microbial abundance at the genus level of soil microbiota. The column names represent sample/group information, while the row names denote species names. The tree on the left side of the figure displays the species clustering, and the values corresponding to the different colors in each square of the middle heatmap are the relative abundance values of the species in each row
The initial relative abundance value of Acinetobacter spp.in the florfenicol 0.5 mg/kg group was 17.15% on day 0, and this value significantly decreased over time. Similarly, the relative abundance value of Pseudomonas spp. in the florfenicol 5 mg/kg group was 15.28% on day 0, but in the 50 mg/kg group, it decreased to 2.11%. The abundance value of Pseudomonas spp. was also influenced by time, with the S3 group exhibiting a decreasing trend in abundance value over time, although there was a slight rebound at 60 days. On the other hand, the abundance value of the Acinetobacter spp. was more prominently affected by time. Chitinophagaceae, Nitrospira, and Bryobacter exhibited decreased abundance as the concentration of florfenicol increased, while the higher concentration of florfenicol (50 mg/kg) led to increased abundance values over time. In the S3D7 sample group, Enterobacter and Sporosarcina exhibited higher abundances of 12.16% and 12.13%, respectively.
Multisample comparative analysis
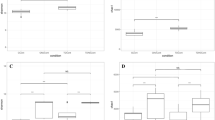
To assess the similarities and dissimilarities among different samples, the NMDS analysis and the UPGMA clustering method were employed. Figure 5 provides a graphical representation wherein each point represents a sample, and the distance between points indicates the degree of variation. A Stress value below 0.2 indicates that the NMDS accurately captures the variation between samples. Figure 5 demonstrates that soil samples with similar microbial community structures cluster together. For instance, S3D7 and S5D30, as well as S5D60, form distinct clusters, clearly separated from other treatment groups. The cluster analysis reveals notable temporal and concentration variability, with treatment groups from different sampling times or with greater concentration differences being further apart in the clustering pattern.
Non-metric multidimensional scaling (NMDS) and unweighted pair group method with arithmetic mean (UPGMA) were employed to reveal the similarity and dissimilarity of microbial communities among different soil samples. (a) Results of NMDS analysis. Each point represents a sample, and the distance between points reflects the degree of dissimilarity. (b) Results of UPGMA analysis. Each branch represents a sample, and the farther the distance, the greater the dissimilarity between the two samples
Discussion
Soil microorganisms play a vital role in soil formation, development, and material cycling, and their diversity and community structure composition are commonly employed as indicators for assessing soil quality. This study investigated the effects of different concentrations of florfenicol on soil microorganisms. The results revealed a negative correlation between the duration of florfenicol action and OTUs values, indicating that longer exposure periods led to decreased OTUs values. Soil α-diversity analysis demonstrated the detrimental impact of florfenicol on bacterial community diversity and abundance, with a notable effect observed at day 7, where increasing concentrations of florfenicol significantly reduced bacterial community abundance. Furthermore, the diversity of the bacterial community exhibited a declining trend with increasing concentrations of florfenicol at day 60.
This study further revealed that the soil bacterial community was primarily dominated by the Proteobacteria phylum, exhibiting a high relative abundance ranging from 36.62% to 59.05%. Following Proteobacteria, the Acidobacteriota phylum was also prevalent. These findings are similar to previous studies investigating soil bacterial diversity. For instance, a study conducted in the prairie of Oklahoma employed a near full-length 16S rRNA gene clone library, sequencing a total of 13,001 clones, and similarly identified the Proteobacteria phylum as the most abundant (Spain et al. 2009). Additionally, analyses of bacterial communities in the soil of the Cerrado Prairie demonstrated a high abundance of the Proteobacteria phylum, accompanied by a significant presence of Acidobacteriota spp. Furthermore, dominance of Proteobacteria spp. was also observed in the soil community of soybean cultivation (de Souza and Procópio 2021).
At the genus level, the most abundant taxa observed in both florfenicol-added and non-florfenicol-added soil samples were Sphingomonas, followed by Gemmatimonadaceae. The presence of Sphingomonas in high abundance is consistent with findings from a separate study involving isolation of bacterial strains from desert sandy soil samples collected in the Gurbantunggut Desert, located in northwestern China's Xinjiang region. In that study, two isolated strains were identified as belonging to the same species within the Sphingomonas genus (Dong et al. 2022). These results align with the outcomes obtained in the current experiment.
Florfenicol can alter the structure of the microbiota and can reduce the biodiversity of the microbiota by acting as a strong stressor (Zeng et al. 2019). The evaluation of microbial diversity using the ACE index, Shannon index, and Simpson index in this study revealed that soil samples supplemented with florfenicol exhibited reduced microbial diversity compared to those without florfenicol. This observation strongly suggests that florfenicol exerts inhibitory effects on the soil microbial community.
Soil microbial communities play a vital role in driving various ecosystem functions and ecological processes and are instrumental in maintaining biogeochemical cycles (Delgado-Baquerizo et al. 2016). The structure of bacterial communities is intricately linked to the soil environment, and any alterations in the soil environment can lead to corresponding changes in the bacterial community structure (Zhang et al. 2022). Antibiotics have a profound impact on microbial communities, primarily influencing community diversity, abundance, composition, and function (Qiu et al. 2023). Extensive research has indicated that antibiotics can modulate microbial biomass and community structure by affecting microbial enzyme activity and metabolic capacity (Wang et al. 2020). Florfenicol, an amidol class broad-spectrum antibiotic, possesses a prolonged half-life and can persist in soil for extended durations, remaining biologically active (Subbiah et al. 2011). Moreover, it exhibits high solubility and resistance to degradation (Dechene et al. 2014). Consequently, the accumulation of this antibiotic poses a significant concern with potential ecological implications.
Utilizing the high-throughput sequencing research method, this study investigated the impact of varying times and concentrations of florfenicol on soil microbial structure and diversity. The findings revealed that florfenicol not only altered the structure of the soil microbial community but also influenced species diversity and abundance. The inhibitory effect became more pronounced with increasing drug concentrations, with the most significant impact observed at 50 mg/kg of florfenicol. Additionally, the negative impact of florfenicol on soil microbial structure and diversity exhibited temporal variations, with the duration of florfenicol stress correlating with a greater impact. Consequently, it can be anticipated that residues of veterinary drugs in the aquaculture industry may result in persistent pollution in soil and water environments, posing long-term potential risks to human health and the entire ecosystem.
Conclusions
The results demonstrate that florfenicol significantly alter the soil microbial community structure and decrease microbial richness and diversity. Greater florfenicol concentrations exert more pronounced inhibitory effects.
Data availability
16S rDNA sequencing data can be found in the repository (National Center for Biotechnology Information, NCBI) at http://www.ncbi.nlm.nih.gov/bioproject/995195 (accession no., PRJNA995195).
References
Coban O, De Deyn GB, van der Ploeg M (2022) Soil microbiota as game-changers in restoration of degraded lands. Science 375:abe0725. https://doi.org/10.1126/science.abe0725
de Souza LC, Procópio L (2021) The profile of the soil microbiota in the Cerrado is influenced by land use. Appl Microbiol Biotechnol 105:4791–4803. https://doi.org/10.1007/s00253-021-11377-w
Dechene A, Rosendahl I, Laabs V, Amelung W (2014) Sorption of polar herbicides and herbicide metabolites by biochar-amended soil. Chemosphere 109:180–186. https://doi.org/10.1016/j.chemosphere.2014.02.010
Delgado-Baquerizo M, Maestre FT, Reich PB, Jeffries TC, Gaitan JJ, Encinar D, Berdugo M, Campbell CD, Singh BK (2016) Microbial diversity drives multifunctionality in terrestrial ecosystems. Nat Commun 7:10541. https://doi.org/10.1038/ncomms10541
Dong L, Li S, Lian WH, Wei QC, Mohamad OAA, Hozzein WN, Ahmed I, Li WJ (2022) Sphingomonas arenae sp. nov., isolated from desert soil. Int J Syst Evol Microbiol 72. https://doi.org/10.1099/ijsem.0.005195
Fang H, Han Y, Yin Y, Pan X, Yu Y (2014) Variations in dissipation rate, microbial function and antibiotic resistance due to repeated introductions of manure containing sulfadiazine and chlortetracycline to soil. Chemosphere 96:51–56. https://doi.org/10.1016/j.chemosphere.2013.07.016
Gao J, Pei H, Xie H (2021) Influence of Allyl Isothiocyanate on the Soil Microbial Community Structure and Composition during Pepper Cultivation. J Microbiol Biotechnol 31:978–989. https://doi.org/10.4014/jmb.2012.12016
Hu X, Xue Z, Wang S, Su F, Zhao K (2011) Studies on the effect of florfenicol on soil microbial activity. J Shandong Anim Sci Vet Med 32:3–5. https://doi.org/10.3969/j.issn.1007-1733.2011.02.002
Lo Giudice C, Pesole G, Picardi E (2021) High-Throughput Sequencing to Detect DNA-RNA Changes. Methods Mol Biol 2181:193–212. https://doi.org/10.1007/978-1-0716-0787-9_12
Lozupone CA, Knight R (2008) Species divergence and the measurement of microbial diversity. FEMS Microbiol Rev 32:557–578. https://doi.org/10.1111/j.1574-6976.2008.00111.x
Qiu M, Hu A, Huang YM, Zhao Y, He Y, Xu J, Lu Z (2021) Elucidating degradation mechanisms of florfenicol in soil by stable-isotope assisted nontarget screening. J Hazard Mater 403:123974. https://doi.org/10.1016/j.jhazmat.2020.123974
Qiu J, Chen Y, Feng Y, Li X, Xu J, Jiang J (2023) Adaptation of Rhizosphere Microbial Communities to Continuous Exposure to Multiple Residual Antibiotics in Vegetable Farms. Int J Environ Res Public Health 20:3137. https://doi.org/10.3390/ijerph20043137
Spain AM, Krumholz LR, Elshahed MS (2009) Abundance, composition, diversity and novelty of soil Proteobacteria. ISME J 3:992–1000. https://doi.org/10.1038/ismej.2009.43
Subbiah M, Mitchell SM, Ullman JL, Call DR (2011) β-lactams and florfenicol antibiotics remain bioactive in soils while ciprofloxacin, neomycin, and tetracycline are neutralized. Appl Environ Microbiol 77:7255–7260. https://doi.org/10.1128/AEM.05352-11
Tasho RP, Cho JY (2016) Veterinary antibiotics in animal waste, its distribution in soil and uptake by plants: A review. Sci Total Environ 563–564:366–376. https://doi.org/10.1016/j.scitotenv.2016.04.140
Wang J, Liu X, Dai Y, Ren J, Li Y, Wang X, Zhang P, Peng C (2020) Effects of co-loading of polyethylene microplastics and ciprofloxacin on the antibiotic degradation efficiency and microbial community structure in soil. Sci Total Environ 741:140463. https://doi.org/10.1016/j.scitotenv.2020.140463
Wang M, Wu J, Zhou T, Liang Y, Zheng L, Sun Y (2021) Effects of copper and florfenicol on nirS- and nirK-type denitrifier communities and related antibiotic resistance in vegetable soils. Ecotoxicol Environ Saf 213:112011. https://doi.org/10.1016/j.ecoenv.2021.112011
Wei R, Ge F, Zhang L, Hou X, Cao Y, Gong L, Chen M, Wang R, Bao E (2016) Occurrence of 13 veterinary drugs in animal manure-amended soils in Eastern China. Chemosphere 144:2377–2383. https://doi.org/10.1016/j.chemosphere.2015.10.126
Zeng Q, Liao C, Terhune J, Wang L (2019) Impacts of florfenicol on the microbiota landscape and resistome as revealed by metagenomic analysis. Microbiome 7:155. https://doi.org/10.1186/s40168-019-0773-8
Zhang P, Guan P, Hao C, Yang J, Xie Z, Wu D (2022) Changes in assembly processes of soil microbial communities in forest-to-cropland conversion in Changbai Mountains, northeastern China. Sci Total Environ 818:151738. https://doi.org/10.1016/j.scitotenv.2021.151738
Zhang T, Sun J, Peng J, Ding Y, Li Y, Ma H, Yu M, Ma Y (2023) Effects of Florfenicol on nirS-Type Denitrification Community Structure of Sediments in an Aquatic Microcosm Model. Antibiotics (basel) 12:1254. https://doi.org/10.3390/antibiotics12081254
Acknowledgements
This research was supported by the Natural Science Foundation of Guangdong Province of China (2023A1515012181); the Start-Up Research Project of Maoming Laboratory, Guangdong Laboratory for Lingnan Modern Agriculture (2021TDQD002); and the Program for Scientific Research Start-Up Funds of Guangdong Ocean University (060302052301).
Author information
Authors and Affiliations
Contributions
Conceptualization, Y.M.; funding acquisition, Y.M. and Y.L.(Yuanjia Liu); methodology, Y.L.(Yang Li) and S.L.; project administration, J.P. and Y.D.; investigation, Y.L.(Yang Li), X.X., H.M. and M.Y.; writing-original draft preparation, Y.L.(Yang Li); writing-review and editing, Y.M. and Y.L.(Yuanjia Liu); supervision, Y.M.; All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Conflicts of interest
The authors declare no conflict of interest with this manuscript.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, Y., Luo, S., Peng, J. et al. Impacts of florfenicol on soil bacterial community structure and diversity by high throughput sequencing analysis. Biologia 79, 1931–1938 (2024). https://doi.org/10.1007/s11756-024-01659-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11756-024-01659-9