Abstract
Ectomycorrhizal symbiosis promotes the growth of masson pine (Pinus massoniana) in low-phosphorus (low-P) conditions; however, the mechanism underlying this phenomenon has not yet been fully described. Here, we cloned four members of the Pht1 phosphate transporter protein family (GenBank accession: AMR43649.1 to AMR43652.1) encoding phosphate transporters in masson pine (PmPTs) by rapid amplification of cDNA ends (RACE) and characterized them in Boletus edulis and Pisolithus tinctorius colonized plants under low-P stress. PmPT1 to PmPT4 encoded 548, 548, 535, 535 amino acid polypeptides, respectively, containing the typical domain of the Pi:H+ symporter (PHS-transporter). Homology multiple sequence alignment indicated that these PmPTs were highly similar to phosphate transporters from other species. The polypeptides were characterized by high hydropathicity and contained 12 putative intra-membrane regions and 1 cytoplasmic loop. The temporal and spatial expression profiles showed higher expression of these PmPTs in ectomycorrhiza (ECM)-inoculated plants compared to non-inoculated ones. In addition, expressions of these PmPT members shared a similar pattern and might be intensively activated by low-P stress or inhibited under P excess. Interestingly, the ECM-colonized plants accumulated more phosphate compared to non-ECM-colonized specimens when exposed to low-P. Therefore, enhanced low-P tolerance in ECM-inoculated masson pine to low-P stress was at least partially dependent on the up-regulation of phosphate transporter genes, reflecting that the intimate interaction between plants and ECM fungi resulted in the improvement of P nutrition.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Masson pine (Pinus massoniana) is one of the most economically important trees in China, because it has been widely used for timber, pulp and resin production (Fan et al. 2014). It had the ability to grow fast thus produced high yield because it might accommodate to the hostile environments, including poor and droughty clay or sandy soils, mine areas and so on (Zhu et al. 2010). Masson pine was a typical ECM-colonized plant (Huang et al. 2014), resulting in the improvement of nutrient uptake including in inorganic phosphorus (Gao et al. 2009; Plassard and Dell 2010).
The phosphorus (P) is one of the key components that influence plant growth and development, and it is widely involved in biochemical synthesis, energy transfer, signal transduction and other metabolism processes. P uptake highly depends on the availability in soil and may also be considerably mediated by plant roots (Bucher et al. 2001). At the soil–root interface, the plant absorbs the soluble P directly through root epidermis and root hairs or via symbiosis mycorrhiza (Smith et al. 2003a, b). Finally, the concentrated P is absorbed and transported by the phosphate transporter, and P starvation is known to enhance synthesis of these carrier systems (Karandashov and Bucher 2005). One of the plant evolutionary strategies to improve availability and uptake of P is the establishment of symbiosis with arbuscular mycorrhiza (AM).
ECM symbiosis not only directly influences P availability in the soil, but also has a positive effect on the uptake of P in the host plant. Association with ECM helps plants enhancing the levels of available P in the rhizosphere, and, in turn, to improve P uptake. Previously, high alkalinity and acid phosphatase activity were investigated near the AM fungi, which were attributed to P hydrolyzation (Ezawa et al. 2002). Moreover, most roots in symbiosis with AM fungi absorbed P through the mycelium (Smith et al. 2003a, b, 2004), further improving P intake.
P transporters play an important role in P absorption in plants. P-induced transporter family genes had been found in many plants (Bucher 2007; Javot et al. 2007; Loth-Pereda et al. 2011). The induction in the expression of these P transporter genes highly indicated that this was an evolutionary conserved mechanism independent of plant species, which was consistent with evidences that the P-induced transporter family Pht1 has been cloned in a wide variety of plants, from Arabidopsis to perennial trees (Karthikeyan et al. 2002; Shin et al. 2004; Leggewie et al. 1997; Rausch et al. 2001; Paszkowski et al. 2002; Glassop et al. 2007; Tuskan et al. 2006). In particular, mycorrhiza formation in plants might induce the expression of genes encoding P transporters in Medicago truncatula (Harrison et al. 2002), Lotus japonicus (Maeda et al. 2006), Glycine max (Yosuke et al. 2012), Oryza sativa (Yang et al. 2012) and Linum usitatissimum (Walder et al. 2015), etc. Therefore, the growth and development improvement of plant with ECM symbiosis was presumably ascribed to the enhanced expression of P transporter genes.
To date, unfortunately, the mechanism underlying the improvement of P absorption had not yet well unraveled. Furthermore, no cue concerning the interaction between the ECM symbiosis and expression of phosphate transporter gene of masson pine had yet been available so far. Previously, we discovered an elite germplasm showing high tolerance to low-P status. To fully understand the involvement of ECM in P absorption of masson pine, currently, we cloned P transporter genes from the low-P tolerance masson pine germplasm and characterized their expression. Subsequently, to investigate the regulation of P transport via ECM, we quantified the expression of these transporter genes in the presence of ECM fungi and under different P soil concentrations, which might provide insights into the role of ECM fungi in promoting plant P utility.
Materials and methods
Plant growth conditions and stress treatments
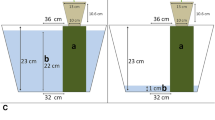
The 6-month-old ECM-free P. massoniana seedlings from Meishan, Sichuan, China were used to carry out P treatments. Yellow soil at pH 5.08 containing 5.71 g kg−1 organic matter, 0.43 g kg−1 total N, 38.38 mg kg−1 available N, 0.12 g kg−1 total P, 1.32 mg kg−1 available P, 4.23 g kg−1 total K, 28 mg kg−1 available K, 0.16 mg kg−1 available copper, 0.53 mg kg−1 available zincum, 4.26 mg kg−1 available ferrum, 4.52 mg kg−1 available manganese, 0.062 mg kg−1 available boron, 445 mg kg−1 exchangeable calcium, 33 mg kg−1 exchangeable magnesium was collected from 50 cm underground in Mengguan Forestry Farm, Guiyang, China. The soil was sprayed with 37% formaldehyde and covered with plastic film for a week for sterilization then impurities were eliminated. Subsequently, the soil was air-sterilized twice at 124–126 °C for 1 h. After being cooled, the soil was put on in culture pots (25 cm × 30 cm) sterilized with potassium permanganate for 24 h. Sterilized pearl stones were used as upper substrate. The fungal isolates used for our experiment came from Boletus edulis and Pisolithus tinctorius, which were donated by the Beijing University of Agriculture (China) and the Plant Physiology Laboratory, Guizhou University (China), respectively. 100 ml of 2-week-old mycelia suspensions and 10 ml of spore suspension (107–106 spores per seedling of B. edulis and P. tinctorius, respectively), and non-ECM-inoculated (NECM) seedlings was used as control. Inoculation was carried out in June 2013. The seedlings were transplanted in sterilized containers, and watered with the 1/2 Hoagland solution (Douds and Schenck 1990) added with different phosphorus concentrations (0, 50 or 5000 μM) at regular intervals of 1 month for 3 times. To minimize the error, the seedlings with uniform size and growth vigor were selected as trials, and 150 seedlings were subjected to phosphorus treatment. To illustrate the temporal expression of the genes, both ECM and non-ECM seedlings without P addition were harvested separately on the 1st, 6th, 12th, 18th, 24th, and 30th day. The seedlings treated with different P concentrations were harvested at the 180th day. The samples were flash frozen using liquid nitrogen, and kept at −80 °C for further experiments.
P content quantification and root structure observation
The P content in the solution was measured using the vanadium molybdate blue colorimetric method with absorbance read at 700 nm on a UV–Visible spectrophotometer (Analytik Jena, Germany). The seedlings were dried to constant weight at 70 °C for 12 h after de-enzymed at 90 °C for 30 min. Around 0.2 g of well-grounded powder sample was used to quantify P content according to the method published by Agricultural Standard of Industry (NY/T 2421-2013, China). P contents of roots, stems and leaves were titrated in molybdate–antimony–scandium color agent after being digested by concentrated sulfuric acid, with deviation <5%.
Fresh root structures were imaged under stereo microscope (Olympus Corp., Japan). Root surfaces were scanned using an S-3400 N scanning electron microscope (Hitachi Ltd., Japan). Root total length, diameter, lateral root length, number of lateral root, and root surface area were measured using Win-RHIZO Pro 2008a (Regent Instruments Inc., Canada). Root fresh weight and dry weight (DW) were recorded after imaging. Mycorrhizal dependence was quantified according to a previously reported formula (Plenchette et al. 1983):
Total RNA isolation
We extracted total RNA from masson pine using Concert Plant RNA Reagent (Invitrogen, USA) according to the supplier’s instructions. After treatment with DNase I (MBI Fermentus, Canada), we used reverse transcriptase M-MLV (RNase H-) (Tiangen Biotech, China) and oligo (dT)18 to synthesize the 1st cDNA strand for each RNA sample. Root total RNA was used for rapid amplification of cDNA ends (RACE) and expression analysis.
Full-length cloning of Pht1 genes
Degenerate PCR primers were designed for RACE applications against conserved regions of Pht1 transporters from Cryptomeria japonica (GenBank accession: BAE92291). Five micrograms of total RNA was used to synthesize RACE-Ready cDNAs with specific primers (Table 1) and Superscript II RT for SMART RACE cDNAs Amplification Kit (Clontech, Palo Alto, CA, USA) following supplier’s instructions. Based on a previously cloned EST sequence, our primers (Table 1) targeted the 3′ end and 5′ end of Pht1 cDNA in a gene-specific fashion. PCR program was performed with the procedure as denaturation at 94 °C for 3 min, followed by 30 amplification cycles at 94 °C for 30 s, 50 °C for 1 min and 72 °C for 1 min, and finally with an extension at 72 °C for 7 min. PCR products were purified from electrophoretic gels using a DNA gel extraction Kit (Takara, Japan). The fragment of interest was cloned into the pMD-19T vector (Takara, Japan) and transformed into Escherichia coli DH5α (Tiangen Biotech, China). After extraction, the recombinant plasmids were sequenced (Sangon Biotech, China). The 3′-RACE and 5′-RACE cDNA together with overlapping sequences from sequencing data to create a cDNA contig were used that we used to find the initiating translation codon (ATG) and ORF on PmPTs.
Bioinformatics analysis
The nucleotide sequence for genes using the NCBI BLAST program (http://blast.ncbi.nlm.nih.gov/) was predicted. We used tools on the Expasy (http://www.expasy.org/tools/pi_tool.html) and Phytozome (http://phytozome.jgi.doe.gov/pz/portal.html) web sites for the analysis of amino acid sequence and protein structure. The BLAST searches against the GenBank protein database for sequence homology analysis were used. Putative protein sequences were aligned using DNAMAN 8.0. A phylogenetic tree representing the relationship between the amino acid sequences with Clustal X 2 and MEGA 5 using the neighbor-joining method (Bootstrap values were from 1000 replications) was built. Protein topology was predicted using TM-pred (http://www.cbs.dtu.dk/services/TMHMM-2.0/). ProtScale provided hydrophobicity plot (http://web.expasy. org/protscale/). SWISS-MIDEL was used for the protein structure homology modeling (http://swissmodel.expasy.org).
Gene expression analysis
The qRT-PCR was employed to measure PmPTs expression in leaf, stem and root after treatments with different P concentrations in the presence or absence of ECM colonization and gene-specific primers (Table 1), The reactions were performed using SYBR Green Master Mix (Applied Biosystems, USA) on an Applied Biosystems 7500 System (Applied Biosystems, USA) following the supplier’s instructions. The qRT-PCR was carried out using the following procedure: initial denaturation at 95 °C for 30 s, followed by 40 PCR cycles of 95 °C for 20 s, 60 °C for 20 s and 72 °C for 20 s, and the amplications were performed in 3 technical replicates. Comparative threshold cycle (Ct) was used to determine gene expression, using the geometric mean of Ct values standard control genes (UBQ, GenBank accession: AF461687.1 and 18S, GenBank accession: X75080.1), as a normalization factor. Relative quantity (RQ) was calculated using the Cycler’s system with software. The relative expression using Pfaffi method (Pfaffl 2001) was calculated. Values reported here are means of three replicates. Furthermore, the treatment means were analyzed by one-way analysis of variance (ANOVA) via the SPSS statistical software version 18.0 (SPSS Inc., USA). Statistical significance was calculated using least-significant difference (LSD) test.
Results
Effects of ECM-fungal colonization on plant P content
To analyze the effect of ECM-fungal colonization on plant P content, we grew plants in the presence or absence of ECM fungi and treated with 0, 50 or 5000 μM P supplies, representing starved, low- or high-P treatment, respectively. The P content in roots, stems and leaves was quantified (Table 2). Using 0 and 50 μM P, the P content in leaf of ECM-colonized plants was significantly higher than of NECM plants. ECM-colonized plants showed low P content in stems, while presented no differences in P content of roots, irrespective of P treatments and ECM-colonized or not. Total P contents in ECM-colonized plants were higher than that of NECM plants under low-P condition. Moreover, P allocation ratio in shoots of ECM-colonized plants was higher than that of the NECM.
Effects of ECM-fungal colonization at different P concentrations on root structure
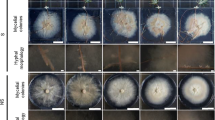
After 6 months of incubation with the mixed fungi, extraradical mycelia might be clearly observed using stereo microscopy (Fig. 1a). Also, epitaxial hyphae were found on the epidermis using scanning electron microscopes (Fig. 1b), which in contrast was not found in roots of NECM plants (Fig. 1c). Interestingly, more hyphae were investigated on the root surface from low-P environment (Fig. 1b) compared with that from high-P treatment (Fig. 1d). Additionally, the mycelia length of 0 μM P treatment (ca. 3.5 mm) was longer than that of 5000 μM P (ca. 3.0 mm). Therefore, the ectomycorrhiza might substantially enlarge the soil contact area of masson pine root, probably leading to the improvement of P uptake.
Compared to NECM plants, ECM-colonized plants showed superior growth vigor and increased biomass production (Table 3). The root/shoot ratio, plant height and stem diameter of ECM plants were somewhat higher in comparison with the NECM ones and the mycorrhizal dependence was 46.33–48.75% in root of ECM-colonized plants. The total root length and lateral root length of ECM-colonized with 0 and 50 μM P gave higher values in comparison with other treatments. However, the main roots became shorter and lateral roots grew more densely as subjected to 5000 μM P. After 50 µM P treatment, the number of lateral root and root surface area of ECM-colonized plants were enhanced by 1.78 and 1.3 times, respectively, compared with the NECM plants.
Sequence and phylogenetic analysis
Four Pht1 genes were isolated from ECM-inoculated masson pine roots, designated as PmPT1, 2, 3 and 4 (GenBank accessions: AMR43649.1, AMR43650.1, AMR43651.1 and AMR43652.1, respectively). These genes contained no introns, with 2388, 2053, 2315 and 2108-bp long full-length cDNA sequences whose corresponding ORF lengths were 1647, 1647, 1608 and 1608 bp, respectively. The predicted respective protein masses were 60.05, 60.05, 58.67 and 58.66 kDa with an isoelectric point (pI) of 8.91, 8.62, 8.98 and 8.48, respectively. Sequence analysis revealed that the putative PmPTs protein sequences were highly similar to phosphate transporters from C. japonica (BAE92291.1, 76–77%), Selaginella moellendorffii (XP_002962968.1, 75–76%), Nicotiana tabacum (AAF74025.1, 74–76%), Camellia oleifera (AFU07481.1, 75–78%) and Theobroma cacao (XP_007050310.1, 77–78%) (Fig. 2). The Amino acid sequence analysis showed 1.46% diversity between PmPT1 and PmPT2, only 0.56% between PmPT3 and PmPT4. NCBI alignment indicated that PmPT1, 2, 3 and 4 presented features of the MFS-transporter family.
Multiple alignments of PmPT1, 2, 3 and 4 to Pht1 from other species. CjPT (C. japonica, BAE92291.1), SelmPT (S. moellendorffii, XM_002962968.1), NtPT (N. tabacum, AAF74025.1), CoPT (C. oleifera, AFU07481.1), TcPT (T. cacao, XP_007050310.1). The 13-amino-acid consensus sequence residue regions (GGDYPLSATIMSE) were marked with a red asterisk
Multiple sequence alignment showed that the four PmPT proteins carried a 29-amino-acid consensus sequence (TLCFFRFWLGFGIGGDYPLSATIMSEYAN) containing the Pht1 signature region (Karandashov and Bucher 2005) (Fig. 2). In addition, hydropathy plots of the four PmPTs predicted them to be integral membrane proteins, as they displayed high hydropathicity and contained 12 putative transmembrane regions with one cytoplasmic loop.
Based on the amino acid sequences, a phylogenetic tree was constructed for PmPTs using the neighbor-joining method (Fig. 3), which indicated that all PmPTs belonged to members of the Pht1 family. The Pht1 family is composed of five subfamilies with clade I (Dicots), clade II (Gymnosperms), clade III (Monocots), clade IV (Mycorrhizal-specific Pht1 family) and clade V (Fungal). PmPTs were similar to their homologues in C. japonica (BAE92291.1) and S. moellendorffii (XM_002962922.1). This was not surprising as all Pht1 members in gymnosperms were placed in clade II. Interestingly, all Pht1 transporters of clade IV were closely related to those in plant species colonized by AM, including M. truncatula, Lycopersicon esculentum, O. sativa, Solanum melongena and Triticum aestivum. Meanwhile, these proteins were far away from those in clade II.
Molecular phylogenetic tree of Pht1 family members in plants and fungi (bootstrap analysis with 1000 replicates). C. japonica (CjPT); S. moellendorffii (SelmPT); L. esculentum (LePT1, LePT2, LePT3, LePT4, LePT6, LePT7, LePT8); Solanum tuberosum (StPT1, StPT3, StPT4, StPT5); Lotus japonicus (LjPT3); M. truncatula (MtPT4); S. melongena (SmPT1, SmPT3, SmPT4, SmPT5); Citrus trifoliata (CtPT1,CtPT2, CtPT3, CtPT4, CtPT7); N. tabacum (NtPT1, NtPT2, NtPT3); T. aestivum (TRIae); C. oleifera (CoPT1); Glycine max (GmPT1, GmPT2, GmPT7, GmPT10, GmPT11, GmPT13); T. cacao (T cPT4); Manihot esculenta (MePT1, MePT6); Vernicia fordii (VfPT); Capsicum frutescens (CfPT1, CfPT3, CfPT4, CfPT5); Populus trichocarpa (PtPT1, PtPT2, PtPT3, PtPT4); Populus euphratica (PePT4, PePT5, PePT9); Hevea brasiliensis (HbPT1); Zea mays (ZmPT1, ZmPT2, ZmPT3, ZmPT4, ZmPT6); O. sativa (OsPT1, OsPT2, OsPT3, OsPT4, OsPT5, OsPT6, OsPT7, OsPT8, OsPT9, OsPT10, OsPT11, OsPT12, OsPT13; Hordeum vulgare (HvPT1, HvPT4, HvPT5, HvPT6, HvPT7, HvPT8, HvPT9, HvPT10, HvPT11); Sesamum indicum (SiPT4); Ophiostoma piceae (OpPT); Glomus versiforme (GvPT); Hebeloma cylindrosporum (HcPT1, HcPT2); Saccharomyces cerevisiae (PHO84); Gigaspora margarita (GiPT); Pholiota nameko (PnPT); Laccaria bicolor (LbPT1, LbPT2)
Expression analysis of the PmPTs
The temporal expressions of PmPTs were measured in plants colonized by ECM fungi (Fig. 4). Expressions of PmPTs were relatively stable in ECM-free plants, and expression levels in roots of ECM-colonized plant were higher than those of NECM plants. PmPT1, 2, 3 and 4 were expressed on the 1st day and they were widely present in the root system of masson pine. Using the expressions of the 1st day as reference, we found that the expressions of PmPT1, 2, 3 and 4 changes over time, from the 1st day to the 30th day. Compared with 1st day, the expression levels of PmPTs in ECM-colonized plants increased slightly on 6th to 24th days and these levels were still higher than those of the control. After the 24th day, the expression levels of PmPT1, 2, 3 and 4 were obvious up-regulated in ECM colonized roots.
The temporal expression profiles of PmPT1, 2, 3 and 4 in ECM-colonized and NECM roots without P supply by qRT-PCR. The samples were collected from roots colonized by ECM fungi during the 1st, 6th, 12th, 18th, 24th, 30th. ECM: ECM-inoculated plants; NECM: Non-ECM-inoculated plants; Data are shown as the mean ± SD (n = 3)
PmPTs were widely expressed in roots, stems and leaves of masson pine (Fig. 5). In ECM plants, PmPTs were spacially expressed in roots, followed by leaves, while the lowest expression was investigated in the stems. However, in the P-starved plants (0 μM), abundant transcripts were detected in all organs, conversely, 5000 μM P completely inhibited the expression of these genes. Furthermore, ECM-colonized plants demonstrated obviously higher expression in PmPTs compared with the NECM ones. Under P-free treatment (0 μM), roots of ECM-colonized plants grown showed significant enchantment in PmPT1, 2, 3 and 4 expression (p < 0.05), which was about sevenfold as high as that of the NECM plants; however, the expression in ECM leaves was approximately 1–2 times lower than that in roots. The similar expression trends were also investigated in plants treated with 50 μM P.
Discussion
Ectomycorrhiza involved in low-P tolerance of masson pine
ECM fungi can promote the growth of host plants by facilitating nutrient and water uptake, and the normal growth of many tree species depends on ECM fungi in nature (Smith and Read 2008). As a gymnosperm, masson pine is a typical ECM-colonized plant, in which the lack of mycorrhizal fungi causes hypogenesis and even impaired growth (Gao et al. 2009). Currently, ECM-colonized plants demonstrated the superiority in growth vigor, e.g., total DW, plant height, total root length, etc. (Table 3), and the tissues including root, stem and leaf of ECM-inoculated plants contained more P compared to the NECM. Moreover, the differences of growth parameters between ECM-inoculated and NECM plants under low-P status were more obvious than that under high-P treatments (Table 3). The similar result was also obtained in other trees (Jones et al. 1990; Zavišić et al. 2016). Therefore, the growth improvement of masson pine inoculated by ECM symbiosis might be highly due to promoting P uptake, especially under low-P status.
The effect of ECM on root P absorption might, however, be also mediated by an ECM-induced reshaping of root architecture. P is mostly absorbed from the soil by diffusion, typically across distances of 1–2 mm from the roots (Barber and Mackay 1986). Thus, the main morphological factor regulating P uptake is the root system status, distribution and topology (Hodge 2008; Hodge et al. 2009). Although ECM fungi did not penetrate their host cell walls, it might cover radicle surface where they exchanged soluble nutrients (Harrison 2005), and their mycelia could extend as far as 10 cm away from the root (Tibbett and Sanders 2002; Rakshit and Bhadoria 2009); thus, a root system with a mycorrhizal network has a substantially bigger surface area for nutrient absorption including P, which has been documented in other species (Schnepf et al. 2008; Ananthi et al. 2011; Zavišić et al. 2016). Due to the exposure to P-deficient condition, currently, ECM-colonized plants demonstrated longer roots, more lateral roots, and larger root surface area compared to NECM ones (Table 3). In addition, dense and long epitaxial hyphae were observed on roots of the ECM plants (Fig. 1a), particularly in low-P status. Extraradical hyphae extended outward from the mantle into the soil and increased the surface area and distance of colonized roots. The beneficial effect of ECM on masson pine plant under low-P condition might be considerably ascribed to the improvement of the absorption system, leading to promoting P uptake, which were also described in other forest trees (Plassard and Dell 2010; Cairney 2011). The just-mentioned involvement of ECM symbiosis in P uptake superficially explained the higher P content in masson pine as subjected to P deficiency; however, the precise molecular role of ECM fungi in P uptake of masson pine has not yet been unraveled so far.
Transcriptional responses of phosphate transporter to low-P stress
The Pht1 family phosphate transporter widely existed in fungi, gymnosperm and angiosperm. In the present case, the four P transporter genes from masson pine plants co-colonized with both B. edulis and P. tinctorius were cloned. These transporters shared the conserved sequence GGDYPLSATIMSE, typical of the Pht1 family (Fig. 3), a member of the major facilitator super-family, composed of monomeric proteins with 12 transmembrane domains (Guan and Kaback 2006). The phylogenetic analysis demonstrated that PmPTs, CjPT (C. japonica) and SelmPT (S. moellendorffii) were grouped into same sub-clade, implying that they probably derived from the same ancestor. Further, the amino acid sequences were quite similar between PmPT1 and PmPT2, as well as between PmPT3 and PmPT4 (Fig. 2), suggesting functional redundancy within the member of each pair.
P transporter genes of plant may be classified into two categories in plants based on their expression features in response to mycorrhizal symbiosis. The first is mycorrhiza-inducible P transporters expressing either in colonized or non-colonized plants, but usually shows higher expression in ECM-colonized plants; the other is the mycorrhiza-specific P transporters, which only expresses in mycorrhizal-colonized plants (Harrison et al. 2002; Daisuke et al. 2006). The present evidences indicated that ECM colonization enhanced the expression of PmPT1, 2, 3 and 4, their expression levels significantly increased in ECM-inoculation roots under low-P status. Moreover, total P contents in ECM-colonized plants were higher than that of NECM plants under low-P condition. Therefore, we inferred that PmPTs were differentially expressed in roots to efficiently transport soil P, and that the total plant P uptake might depend on the expression regulation of P transporter genes, whose expression might be mediated by the integrated signals from ECM colonization and P availability in soil. Phosphate transporters are often divided into high- and low-affinity groups. Low-affinity transporters are constitutively expressed and high-affinity ones are typically expressed in low-P conditions (Rae and Smith 2002). Most Pht1 family members are high-affinity P transporters, whose expression can be induced by low-P supplies (Smith et al. 2003a, b). Our data indicate that low-P (0 or 50 μM) up-regulated expression of PmPTs, conversely, high-P (5000 μM) level inhibited it, further suggesting that ECM facilitated the P availability in soil for plant absorption under low available P status. The expression patterns of PmPTs were compatible with those of the typical high-affinity transporters of other Pht1 family members (Smith et al. 2003a, b, 2004), reflecting that the Pht1 family members in masson pine also belonged to high-affinity transporters. The available P concentration of ≤50 μM treatment herein was presumably within the affinity range of PmPTs, and high-P (5000 μM) trial exceeded highly their affinity range.
As subjected to low-P stress, PmPTs’ expression in both NECM and ECM-colonized seedlings increased, mirroring that low-P supply was the most important factor for their expressions. In addition, PmPTs expression and P content in ECM-colonized seedlings were significantly higher than those in NECM seedlings, further implying that ECM symbiosis was capable of promoting the expression of PmPTs under low-P conditions. Additionally, the expression of PmPTs positively correlated to root surface area of masson pine (Table 3; Fig. 5), similar to previously reported in rice (Li et al. 2001). Therefore, the ECM-triggered growth enhancement in masson pine might be at least partially ascribed to the up-regulations of PmPTs expression, especially under low-P status, which had been first reported in this species so far. However, the subtle mechanism behind ECM regulations of PmPTs expression will be addressed in our further work.
Author contribution statement
Design of experiments: T.Z, XP.W, GJ.D. Analysis and interpretation: T.Z. Data collection: T.Z, XP.W, GJ.D. Provide the materials: XP.W, GJ.D. Writing the article: T.Z. Critical revision of the article: XP.W, GJ.D. Final approval of the article: XP.W.
Abbreviations
- Low-P:
-
Low phosphorus
- ECM:
-
Ectomycorrhiza
- NECM:
-
Non-ECM-inoculated
- AM:
-
Arbuscular mycorrhiza
- ORF:
-
Open reading frame
- RACE:
-
Rapid amplification of cDNA ends
- PmPT:
-
Phosphate transporter of masson pine (Pinus massoniana)
- PHS:
-
Phosphate–H+ symporter
References
Ananthi T, Amanullah MM, Subramanian KS (2011) Influence of fertilizer levels and mycorrhiza on root colonization, root attributes and yield of hybrid maize. Madras Agric J 98(1–3):56–61. doi:10.1007/BF01049351
Barber SA, Mackay AD (1986) Root growth and phosphorus and potassium uptake by two corn genotypes in the field. Fertil Res 10(3):217–230. doi:10.1007/BF01049351
Bucher M (2007) Functional biology of plant phosphate uptake at root and mycorrhiza interfaces. New Phytol 173(1):11–26. doi:10.1111/j.1469-8137.2006.01935.x
Bucher M, Rausch C, Daram P (2001) Molecular and biochemical mechanisms of phosphorus uptake into plants. J Plant Nutr Soil Sci 164(2):209–217. doi:10.1002/1522-2624(200104)164:2<209::AID-JPLN209>3.0.CO;2-F
Cairney JWG (2011) Ectomycorrhizal fungi: the symbiotic route to the root for phosphorus in forest soils. Plant Soil 344(1):51–71. doi:10.1007/s11104-011-0731-0
Daisuke M, Kanae A, Keita I, Chechetka SA, Ayaka H, Yasuhiro O, Yuichi D, Katsura I, Shingo H (2006) Knockdown of an arbuscular mycorrhiza-inducible phosphate transporter gene of Lotus japonicus suppresses mutualistic symbiosis. Plant cell Physiol 47(7):807–817. doi:10.1093/pcp/pcj069
Douds DD, Schenck NC (1990) Cryopreservation of spores of vesicular–arbuscular mycorrhizal fungi. New Phytol 115(4):647–667. doi:10.1111/j.1469-8137.1990.tb00498.x
Ezawa T, Smith SE, Smith FA (2002) P metabolism and transport in AM fungi. Plant Soil 244(1–2):221–230. doi:https://doi.org/10.1093/pcp/pcj069
Fan F, Cui B, Zhang T, Ding G, Wen X (2014) LTR-retrotransposon activation, IRAP marker development and its potential in genetic diversity assessment of masson pine (Pinus massoniana). Tree Genet Genomes 10(1):213–222. doi:10.1007/s11295-013-0677-x
Gao Y, Wu X, Su M (2009) Effects of ectomycorrhizal seedlings of masson pine on absorption and utilization of N, P and K. J Nanjing For Univ (china) 33(4):77–80
Glassop D, Godwin RM, Smith SE, Smith FW (2007) Rice phosphate transporters associated with phosphate uptake in rice roots colonised with arbuscular mycorrhizal fungi. Botany 85(7):644–651. doi:10.1139/B07-070
Guan L, Kaback HR (2006) Lessons from lactose permease. Annu Rev Biophys Biomol Struct 35(1):67–91. doi:10.1146/annurev.biophys.35.040405.102005
Harrison MJ (2005) Signaling in the arbuscular mycorrhizal symbiosis. Annu Rev Microbiol 59:19–42. doi:10.1146/annurev.micro.58.030603.123749
Harrison MJ, Dewbre GR, Jinyuan L (2002) A phosphate transporter from medicago truncatula involved in the acquisition of phosphate released by arbuscular mycorrhizal fungi. Plant Cell 14(10):2413–2429. doi:10.1105/tpc.004861
Hodge A (2008) Root decisions. Plant Cell Environ 32(6):628–640. doi:10.1111/j.1365-3040.2008.01891.x
Hodge A, Berta G, Doussan C, Merchan F, Crespi M (2009) Plant root growth, architecture and function. Plant Soil 321(1–2):153–187. doi:10.1007/s11104-009-9929-9
Huang J, Nara K, Zong K, Wang J, Xue S, Peng K, Shen Z, Lian C (2014) Ectomycorrhizal fungal communities associated with Masson pine (Pinus massoniana) and white oak (Quercus fabri) in a manganese mining region in Hunan Province, China. Fungal Ecol 22(8):589–602. doi:10.1016/j.funeco.2014.01.001
Javot H, Pumplin N, Harrison MJ (2007) Phosphate in the arbuscular mycorrhizal symbiosis: transport properties and regulatory roles. Plant Cell Environ 30(3):310–322. doi:10.1111/j.1365-3040.2006.01617.x
Jones MD, Durall DM, Tinker PB (1990) Phosphorus relationships and production of extrametrical hyphae by two types of willow ectomycorrhizas at different soil phosphorus levels. New Phytol 115(2):259–267. doi:10.1111/j.1469-8137.1990.tb00451.x
Karandashov V, Bucher M (2005) Symbiotic phosphate transport in arbuscular mycorrhizas. Trends Plant Sci 10(1):22–29. doi:10.1016/j.tplants.2004.12.003
Karthikeyan AS, Varadarajan DK, Mukatira UT, D'Urzo MP, Damsz B, Raghothama KG (2002) Regulated expression of Arabidopsis phosphate transporters. Plant Physiol 130(1):221–233. doi:10.1104/pp.020007
Leggewie G, Willmitzer L, Riesmeier JW (1997) Two cDNAs from potato are able to complement a phosphate uptake-deficient yeast mutant: identification of phosphate transporters from higher plants. Plant Cell 9(3):381–392. doi:10.1105/tpc.9.3.381
Li HB, Xia M, Wu P (2001) Effect of phosphorus deficiency stress on rice lateral root growth and nutrient absorption. Acta Bot Sin 43(11):1154–1160
Loth-Pereda V, Orsini E, Courty PE, Lota F, Kohler A, Diss L, Blaudez D, Chalot M, Nehls U, Bucher M, Martin F (2011) Structure and expression profile of the phosphate Pht1 transporter gene family in mycorrhizal Populus trichocarpa. Plant Physiol 156(4):2141–2154. doi:10.1104/pp.111.180646
Maeda D, Ashida K, Iguchi K, Chechetka SA, Hijikata A, Okusako Y, Deguchi Y, Lzui K, Hata S (2006) Knockdown of an arbuscular mycorrhiza-inducible phosphate transporter gene of Lotus japonicus suppresses mutualistic symbiosis. Plant cell physiol 47(7):807–817. doi:10.1093/pcp/pcj069
Paszkowski U, Kroken S, Roux C, Briggs SP (2002) Rice phosphate transporters include an evolutionarily divergent gene specifically activated in arbuscular mycorrhizal symbiosis. Proc Natl Acad Sci 99(20):13324–13329. doi:10.1073/pnas.202474599
Pfaffl MW (2001) A new mathematical model for relative quantification in real-time RT–PCR. Nucl Acids Res 29(9):e45
Plassard C, Dell B (2010) Phosphorus nutrition of mycorrhizal trees. Tree Physiol 30(9):1129–1139. doi:10.1093/treephys/tpq063
Plenchette C, Fortin JA, Furlan V (1983) Growth responses of several plant species to mycorrhizae in a soil of moderate p-fertility. Plant Soil 70(2):211–217. doi:10.1007/BF02374781
Rae AL, Smith FW (2002) Localisation of expression of a high-affinity sulfate transporter in barley roots. Planta 215(4):565–568. doi:10.1007/s00425-002-0777-6
Rakshit A, Bhadoria PS (2009) Influence of arbuscular mycorrhizal hyphal length on simulation of P influx with the mechanistic model. Afr J Microbiol Res 3(1):001–004
Rausch C, Daram P, Brunner S, Jansa J, Laloi M, Leggewie G, Amrhein N, Bucher M (2001) A phosphate transporter expressed in arbuscule-containing cells in potato. Nature 414(6862):462–470. doi:10.1038/35106601
Schnepf A, Roose T, Schweiger P (2008) Impact of growth and uptake patterns of arbuscular mycorrhizal fungi on plant phosphorus uptake—a modelling study. Plant Soil 312(1–2):85–99. doi:10.1007/s11104-008-9749-3
Shin H, Shin HS, Dewbre GR, Harrison MJ (2004) Phosphate transport in Arabidopsis: Pht1; 1 and Pht1; 4 play a major role in phosphate acquisition from both low-and high-phosphate environments. Plant J 39(4):629–642. doi:10.1111/j.1365-313X.2004.02161.x
Smith SE, Read D (2008) Mycorrhizal symbiosis. 13–41. doi:10.1016/B978-012370526-6.50003-9
Smith FW, Mudge SR, Rae AL, Glassop D (2003a) Phosphate transport in plants. Plant Soil 248(1–2):71–83. doi:10.1023/A:1022376332180
Smith SE, Smith FA, Jakobsen I (2003b) Mycorrhizal fungi can dominate phosphate supply to plants irrespective of growth responses. Plant Physiol 133(1):16–20. doi:10.1104/pp.103.024380
Smith SE, Smith FA, Jakobsen I (2004) Functional diversity in arbuscular mycorrhizal (AM) symbioses: the contribution of the mycorrhizal P uptake pathway is not correlated with mycorrhizal responses in growth or total P uptake. New Phytol 162(2):511–524. doi:10.1111/j.1469-8137.2004.01039.x
Tibbett M, Sanders FE (2002) Ectomycorrhizal symbiosis can enhance plant nutrition through improved access to discrete organic nutrient patches of high resource quality. Ann Bot 89(6):783–789. doi:10.1093/aob/mcf129
Tuskan GA, Difazio S, Jansson S, Bohlmann J, Grigoriev I, Hellsten U et al. (2006) The genome of black cottonwood, populus trichocarpa (torr.&gray). Science 313(5793):1596–1604. doi:10.1126/science.1128691
Walder F, Brulé D, Koegel S, Wiemken A, Boller T, Courty PE (2015) Plant phosphorus acquisition in a common mycorrhizal network: regulation of phosphate transporter genes of the Pht1 family in sorghum and flax. New Phytol 205(4):1632–1645. doi:10.1111/nph.13292
Yang SY, Grønlund M, Jakobsen I, Grotemeyer MS, Rentsch D, Miyaod A, Hirochikad H, Kumare CS, Sundaresane V, Salaminf N, Catausang S, Mattesg N, Heuerg S, Paszkowskia U (2012) Nonredundant regulation of rice arbuscular mycorrhizal symbiosis by two members of the PHOSPHATE TRANSPORTER1 gene family. Plant Cell 24(10):4236–4251. doi:10.1105/tpc.112.104901
Yosuke T, Yoshihiro K, Toyotaka M, Shingo H (2012) Identification and expression analysis of arbuscular mycorrhiza-inducible phosphate transporter genes of soybean. Biosci Biotechnol Biochem 76(2):309–313. doi:10.1271/bbb.110684
Zavišić A, Nassal P, Yang N, Heuck C, Spohn M, Marhan S, Pena R, Kandeler E, Polle A (2016) Phosphorus availabilities in beech (Fagus sylvatica L.) forests impose habitat filtering on ectomycorrhizal communities and impact tree nutrition. Soil Biol Biochem 98:127–137. doi:10.1016/j.soilbio.2016.04.006
Zhu LH, Wu XQ, Qu HY, Ji J, Ye JR (2010) Micropropagation of Pinus massoniana and mycorrhiza formation in vitro. Plant Cell Tiss Org 102(1):121–128. doi:10.1007/s11240-010-9711-y
Acknowledgements
The project is supported by grants from Special Core Program of Guizhou Province, P. R. China (20126011–1) and the 863 Program of China (2011AA10020301). All authors have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by J. Zwiazek.
Rights and permissions
About this article
Cite this article
Zhang, T., Wen, XP. & Ding, GJ. Ectomycorrhizal symbiosis enhances tolerance to low phosphorous through expression of phosphate transporter genes in masson pine (Pinus massoniana). Acta Physiol Plant 39, 101 (2017). https://doi.org/10.1007/s11738-017-2392-y
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11738-017-2392-y