Abstract
Purpose
Dientamoeba fragilis (D. fragilis) is a common intestinal protozoan with a global distribution. In the present study, we aimed to determine genetic diversity of D. fragilis isolates with multilocus sequence typing (MLST) in the southwest of Turkey and analyse the clinical findings.
Materials and Methods
The study included faecal samples from 200 individuals in Aydin, Turkey. The positivity of D. fragilis was determined with 18 S rRNA gene-based PCR assay. Six nested-PCR reactions were set to amplify partial D. fragilis housekeeping genes in the positive samples. The sequences were aligned with the references from GenBank to detect nucleotide polymorphisms and haplotypes. Additionally, the clinical findings and demographic characteristics of patients were statistically analysed between D. fragilis-infected and non-infected cases.
Results
The positivity of D. fragilis was 16% (32 out of 200 cases) with 18 S rRNA based-PCR, and all were classified as “genotype 1”. The analysis of six MLST loci revealed different haplotypes only at one locus; the remaining five loci exhibited no polymorphisms. The haplotypes in the present study were identical to at least one previously reported reference, except the locus “large subunit of RNA polymerase II” locus. There were no significant differences in any of the clinical findings or demographic characteristics between the infected and non-infected groups.
Conclusions
Our study revealed a low genetic diversity of D. fragilis isolates from Turkey, like other countries including Italy, Denmark, the UK, Australia, and Brazil. The high degree of sequence similarity in housekeeping genes indicated the clonal distribution of D. fragilis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Dientamoeba fragilis (D. fragilis) is an anaerobic intestinal protozoan commonly detected in human faecal samples. It is closely related to trichomonads, the phylum Parabasalia, despite its name “Dientamoeba” [1]. A few mammalian and avian species other than humans can serve as natural hosts for D. fragilis. However, their potential role in human infections still requires investigation [2,3,4]. The trophozoite stage is binucleate in high percentage and nuclei can be observed in permanently stained faecal smears [1, 5]. No cyst stage has been identified in the life cycle of D. fragilis, and the vector-like role of pinworm eggs has been proposed for a long time [6]. Putative pre-cyst or cyst stages have been described in some animal models and human clinical samples [7,8,9]. However, these stages are rarely observed in human samples and potentially contribute to waterborne transmission [10].
A high percentage of D. fragilis infected people remain asymptomatic, but it may also detected in humans with gastrointestinal symptoms, particularly abdominal pain, and diarrhoea [11]. Some studies reported an improvement in symptoms following the eradication of D. fragilis [12]. The pathogenicity of D. fragilis has been a subject of debate for a long time, as well as describing it as a non-pathogen, opportunistic pathogen, or commensal [13]. Additionally, the number of studies related to gut microbiota, clinical findings, and D. fragilis infections has been increasing [14]. The primary method for the detection of intestinal parasites is direct microscopy of fresh smears in most routine laboratories. However, detecting D. fragilis is not possible without additional permanent staining methods, such as trichrome or molecular techniques like real-time or conventional PCR [15]. The prevalence of D. fragilis varies significantly between studies based on diagnostic method and target population. Unlike other intestinal parasites, higher frequencies have been reported in developed countries because of the use of molecular methods [5, 16]. In addition, it was more prevelant in paediatric age group [13].
Studies on genetic diversity of D. fragilis are important for many aspects such as clinical presentation, diagnosis, treatment, and host-parasite interaction. The genetic variation among the isolates might influence the clinical presentation of the infection, leading to different clinical pictures [1]. Another significance of genetic studies is to investigate the zoonotic potential of the infection and its hosts. Phylogenetically related genotypes were identified in both humans and pigs, leading to a conclusion that pigs might act as a natural host for D. fragilis [4]. Research on the genetic diversity of D. fragilis isolates has involved the analysis of various genes, including 18 S rRNA, actin, ITS, and EF-1α [10, 17, 18]. Two genetic variants of D. fragilis (genotype 1 and 2) have been identified; however, it is unclear whether these genotypes exhibit distinct pathogenicity. “Genotype 1” is globally predominant in both humans and animal hosts [5, 16, 19]. Multilocus sequence typing (MLST) is a molecular technique that enables the genetic characterization of distinct isolates of the same microorganism. The method relies on the analysis of fragment sequences from usually 6 or 7 housekeeping genes [20]. It offers a high distinctiveness, sequence typing, and the ability to reveal its clonal distribution and evolutionary relationship [21]. The method has been used for genotyping of common parasites including Entamoeba histolytica, Toxoplasma gondii, Leishmania spp., and D. fragilis [4, 22,23,24]. A method for genotyping D. fragilis isolates using a multilocus sequencing approach was introduced in 2016 [25]. Following the metagenomic analysis of two D. fragilis positive samples, many candidate markers were identified based on their sequence similarity to Trichomonas vaginalis. Despite having a worldwide distribution, understanding the molecular epidemiology and clinical importance of D. fragilis is challenging, because of the limited genotype data from most of the countries. The aims of the present study were to detect D. fragilis positivity in the Southwest of Turkey, investigate the genetic diversity of the isolates with multilocus sequence typing (MLST), and analyse the clinical findings.
Materials and Methods
Faecal Samples and Ethical Approval
The sample size (n = 200) for the study was calculated with a post hoc power of 98.5%, a sampling error of 5%, and a 95% confidence interval [26]. Faecal samples of 200 cases were randomly selected from Aydin Adnan Menderes University Hospital Parasitology Laboratory with a simple random selection method. The city is located in the southwest of Turkey (37° 50′ 53″ N, 27° 50′ 43″ E). The native-lugol, formol-ethyl acetate sedimentation, and cellophane tape methods were used as routine parasitological examinations of samples in the laboratory. The faecal samples that were positive for intestinal parasites or commensals were not included in the study, e.g., Entamoeba coli, Blastocystis, Giardia intestinalis, Enterobius vermicularis. Information about the gender, residence and age of the cases was obtained from the hospital data system. In addition, the cases were questioned about clinical findings including abdominal pain, diarrhoea, constipation, nausea vomiting, lack of appetite, weight loss, and itching.
This study was approved by Aydin Adnan Menderes University Faculty of Medicine Non-Interventional Research Ethics Committee (Prot. No. 2020/224).
Detection of D. fragilis Positivity
Faecal samples were stored at -20 °C for a maximum of one week before DNA isolation. Genomic DNA isolation was performed using the QIAamp Fast DNA Stool Mini Kit (Qiagen) following the manufacturer’s instructions. D. fragilis positivity was investigated with 18 S rRNA-based PCR with DF400 and DF1250 primers [27]. The reaction was set in 30 μl: 1 μl template DNA, 3 μl Buffer (with MgCl2), 1.5 μl each primer (10 pmol), 1.8 μl dNTP (2.5 mM), and 0.24 μl Taq DNA Polymerase (Applied Biological Materials). The amplification cycle was as follows: initial denaturation of 94 °C for 3 min, 30 cycles (94 °C for 1 min, 57 °C for 1.5 min, and 72 °C for 2 min), and final extension at 72 °C for 10 min. The amplicons were separated on a 1.5% agarose gel and visualized under ultraviolet light (340 nm) using a gel documentation system (Vilber Lourmat). The amplicons with the expected length (863 bp) were purified and sequenced using the Applied Biosystems 377 DNA Sequencer (Medsantek, Istanbul).
Multilocus Sequence Typing (MLST) of D. fragilis
The positive samples, D. fragilis housekeeping genes were partially amplified with nested-PCR for MLST analysis as previously reported, except for the locus “large subunit of RNA polymerase II”, for this locus different set of primers was used [25] Primer pairs used to amplify these genes, expected product sizes and different annealing temperatures are given in Table 1. Reaction in 30 μl was set as follows: 1 μl template DNA or PCR product, 3 μl Buffer (with MgCl2), 1.5 μl forward primer (10 pmol), 1.5 μl reverse primer (10 pmol), 1.8 μl dNTP (2.5 mM), 0.24 μl Taq DNA Polymerase (ABM) and 21 μl dH2O. The PCR cycle was set as initial denaturation at 95 °C for 2 min, 30 cycles (95 °C 30 s denaturation, 52–59 °C 45 s annealing, 72 °C 30 s extension) and 72 °C 5 min final extension. The positive samples with the expected length were purified and sequenced.
Genetic Analyses
The editing and alignment of the DNA sequences were performed with BioEdit 7.7.1. The partial 18 S rRNA gene sequences of isolates were compared with reported D. fragilis 18 S rRNA sequences in Genbank using the Basic Local Alignment Search Tool (BLAST). The Neighbor-Joining method was used to construct the tree, and the robustness of the tree topology was validated with bootstrap tests (1000 repetitions) [28]. The sequences included in the phylogenetic analysis of D. fragilis 18 S rRNA gene were as follows: Genotype 1 (Acc. No: AY730405, MN914083, AB92772, JQ677147, MN560149, OQ345680, OP375682, FJ649228, AB692771, AY30405) and “genotype 2” (Acc. No: U37461).
The housekeeping gene sequences were aligned with BioEdit 7.7.1 and compared to the references in Genbank with BLAST. There were 16 haplotype sequences representing six housekeeping genes of D. fragilis (Acc. No. KX669659-KX669671, MZ405080-4) [25, 29]. The haplotype number, haplotype diversity, nucleotide diversity (Nei, 1972) and polymorphic region number were calculated using the DNAsp 4.50 software for polymorphic loci [30]. A phylogenetic tree was created using The Neighbor-Joining method.
Statistical Analyses of Demographic Characteristics and Clinical Findings
In the present study, we evaluated three demographic parameters: gender, residence, and gender of the cases. In addition, the clinical findings including abdominal pain, diarrhoea, constipation, nausea vomiting, lack of appetite, weight loss and itching were compared between D. fragilis positive and negative cases. The categorical variables were tested with the chi-square test and the quantitative variables were tested with the Student’s t-test. We used IBM SPSS Statistics 26 for the analysis.
Results
The Positivity of D. fragilis in Faecal Samples
The molecular analysis of 200 faecal samples by PCR identified 16% (n = 32) positivity for D. fragilis. The sequences of 29 isolates can be used in genotype determination because three exhibited crowded or noisy backgrounds in the chromatograms. The partial 18 S rRNA gene sequences were identical for all 29 D. fragilis isolates. Therefore, we deposited the sequence of a single isolate (DF10) to the GenBank as a sample. The accession number for this sequence was OM250406.
The Analysis of Housekeeping Genes
The six housekeeping genes of D. fragilis (large subunit of RNA polymerase II, laminin A, TKL family-protein kinase, cathepsin L-like cysteine peptidase, clan Sc-family S9-serine peptidase, and clan MH-family M20 metallopeptidase) were analysed separately for each locus. The summary of isolate numbers, haplotype count, and references can be found in Table 2.
For the locus “large subunit of RNA polymerase II” the samples in our study share the same sequence (Accession No. OM287401). In GenBank, there were two haplotypes (Accession Nos. KX669660 and KX669659) for this locus. The haplotype in our study differs by a single base substitution from each of the reference sequences, a novel haplotype (Supplementary file).
For the loci “laminin A” and “Clan Sc-family S9-serine peptidase”, all the isolates in our study share the same sequence (Accession No. OM287402 and OM287404, respectively). In GenBank, a single haplotype was reported for each of these loci (Acc. Nos. KX669671 and KX669670, respectively). The haplotypes in our study and the references were identical.
For the locus “TKL family-protein kinase,” the samples in our study share the same sequence (Acc. No. OM287403). In GenBank, two haplotypes have been reported for this locus (Acc. Nos. KX669661 and KX669662), both from the UK. The haplotype in our study was identical to the first sequence. (Supplementary file)
For the locus “Cathepsin L-like cysteine peptidase,” the samples in our study exhibit two different sequences (Acc. Nos. OM977006 and OM977007). In GenBank, four haplotypes have been reported for this locus (Acc. Nos. KX669666-KX669669). The first haplotype in our study was identical to KX669666 (a human isolate from the UK), and the second was identical to KX669667 (a human isolate from Italy). In addition, the sequences were also identical to the ones reported from Turkey (Supplementary file).
For the locus “Clan MH-family M20 metallo-peptidase”, the samples in our study have the same sequence (Acc. No. OM287405). For this locus, three haplotypes have been reported in Genbank. The haplotype in our study was identical to the reported sequences from both the UK and Turkey (Supplementary file).
Genetic Analyses
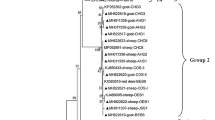
The comparison of the 18 S rRNA sequence with the references from GenBank using BLAST analysis revealed that it corresponds to “genotype 1”, the dominant genotype in the world. In the phylogenetic tree constructed using the Neighbor-Joining method, the present haplotype (Acc. No. OM250406) was in the “genotype 1” clade and stands with the sequences reported from Iran, Italy, the UK, Australia, and Germany (Fig. 1). In addition, other common trichomonads from different hosts such as T. vaginalis (human), Histomonas meleagridis (avian), and Tritrichomonas foetus (cattle) were located distinctly from “genotype 1”.
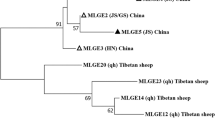
The partial sequences of housekeeping genes of our isolates were identical for the following five loci “large subunit of RNA polymerase II”, “laminin A”, “TKL family-protein kinase”, “clan Sc-family S9-serine peptidase”, and “clan MH-family M20 metallo-peptidase”, presenting a single haplotype. Only for the locus “cathepsin L-like cystein peptidase”, we defined two haplotypes (Acc. Nos. OM977006 and OM977007). The haplotype number was 2, haplotype diversity was 0.513, nucleotide diversity: was 0.0190 and the number of polymorphic regions was 1. We did not calculate these values for the other five loci because there was no polymorphism. The estimated evolutionary distance of isolates for “cathepsin L-like cystein peptidase”, the only polymorphic locus, was presented in Fig. 2. Haplotype 1 was grouped with references from the UK and Turkey, while haplotype 2 was grouped with references from Italy and Turkey.
The Analysis of Demographic Parameters and Symptoms
Of the participants, 108 (54%) were male and 92 (46%) were female. The majority (%53.5) were living in the city centre. The age of cases ranged from 18 to 77, with a mean age of 35.9 ± 10.4 for positive cases and 34.2 ± 11.9 for negative cases. All the cases had at least one symptom. Abdominal pain was the most common (28.1%) symptom among D. fragilis-positive cases, followed by lack of appetite (25%). Neither the demographic characteristics nor the clinical findings of cases were significantly different between the groups. The details of these parameters were presented in Tables 3 and 4.
Discussion
Dientamoeba fragilis has been classified among anaerobic flagellated organisms, in the subkingdom Metamonada and the phylum Parabasalia [1]. The conventional microscopic method faces challenges in detecting fragile trophozoites, often leading to misdiagnoses of infections. Therefore, extended observation periods and experienced staff are necessary [6]. In the last decade, advancements in nucleic acid amplification techniques for diagnosis have led to a rise in detecting D. fragilis in clinical samples. In addition, these techniques have contributed to the understanding of the faecal-oral transmission of D. fragilis [31]. Both conventional and real-time polymerase chain reactions (PCR) have significantly shown higher sensitivity and specificity than microscopy-based methods [5, 32, 33]. For example, a study from Egypt detected D. fragilis in 13%, 17%, and 41% of the stool samples using wet mount smears, trichrome stain, and PCR, respectively [34]. In the present study, 32 out of 200 (16%) stool samples tested positive for D. fragilis via PCR. In Turkey, a few studies have used molecular methods for the detection of D. fragilis. The frequency of D. fragilis was reported as 28.4%, 10.7%, 12%, and 6.5% among symptomatic cases in studies from different cities in Turkey [35,36,37,38]. The recent PCR-based studies in the following countries: France, Gabon, Italy, Czech Republic, Cuba, and Egypt reported the frequencies of 2.3%, 4%, 9.1%, 12%, 24%, and 41% for D. fragilis, respectively [34, 39,40,41,42,43]. However, making a reasonable comparison of D. fragilis positivity between countries is challenging due to significant variations in the studied populations and amplification methods.
The genetic variation among D. fragilis isolates was first studied with restriction enzyme patterns of the SSU-rRNA gene [44]. Sequencing of a partial segment of the SSU-rRNA gene revealed minimal variation, with two D. fragilis genotypes being identified [27, 45]. Globally, “genotype 1” is reported as the sole or most prevalent genotype in both human and animal studies. However, “genotype 2” has been rarely reported [19, 39, 40, 44, 46]. Consistent with other studies, all D. fragilis isolates in our study have the same partial SSU rRNA sequence and were classified as “genotype 1”. There is limited research on D. fragilis genetic diversity in our country. To date, no study in Turkey has reported the presence of “genotype 2”; all documented human isolates, including four from Central Anatolia and 26 from Aydin, were identified as “genotype 1” [29, 36]. In addition, D. fragilis isolates from cattle and budgerigars also found only “genotype 1” [2, 3]. Intra- and inter-diversity were limited in the 18S rRNA coding gene; the similarity between the two genotypes is almost 97%. Therefore, reliable markers are required for a deeper investigation of D. fragilis genotype [45].
Multilocus sequence typing (MLST) is a method that allows detailed genotyping of specific organisms. The method relies on sequence polymorphisms in specific housekeeping genes (e.g., cathepsin-like cysteine peptidase and RNA polymerase II) among isolates of an organism. The cysteine peptidase coding genes may serve as useful targets for studying the genetic diversity of D. fragilis isolates [1]. Proteolytic digestion of host proteins by cysteine peptidases is a vital source of nutrition for many parasites, including Plasmodium spp., Trypanosoma spp., and Entamoeba histolytica [47, 48]. These peptidases have key functions in the pathogenesis of protozoan parasites including cell/tissue invasion, autophagy, and modulation of host immune response [49, 50]. Among the proteases or peptidases, cathepsin L-like cysteine peptidase was detected as the most abundant virulence factor in D. fragilis. Most of the cysteine protease transcripts in D. fragilis had similar homology to cytotoxic and proteolytic cysteine proteases from T. vaginalis [1]. Additionally, the large subunit of RNA polymerase is commonly used as a housekeeping gene in MLST analysis, and its regulatory role in the virulence of malaria parasites has been previously reported [51]. The cathepsin L-like peptidases have been reported as potential targets for the development of novel chemotherapy options for various parasitic protozoa [50]. Caccio et al. (2016) developed six genetic markers for the genotyping of D. fragilis isolates from different countries including Denmark, Italy, the UK Australia, and Brazil. They reported low genetic variability among isolates and supposed a clonal genetic structure of D. fragilis population [25]. They studied the genetic diversity among 111 human D. fragilis isolates, except for one “genotype 2” isolate, all were “genotype 1”. In our study, we used the same markers for typing 32 isolates from Turkey. Only for the locus “cathepsin L-like cysteine peptidase” we identified two haplotypes. For the other loci, all our isolates shared the same sequences. For the locus “large subunit of RNA polymerase II”, we identified a new haplotype that differed by a single base substitution from the sequences reported from Italy. Following the present results, previous studies have demonstrated the existence of a major D. fragilis clone with a widespread geographic distribution [25]. Another study studied five previously reported loci in a limited number of samples (six patients), but most isolates failed to amplify [29]. In addition, D. fragilis genotype 1 specific housekeeping genes such as actin and elongation factor 1 alpha (EF-1α) were reported to have limited epidemiological value in detecting genetic resolution of D. fragilis isolates [18]. The study of alternative housekeeping genes may provide better genetic resolution, as previously reported for Giardia intestinalis, Trichomonas vaginalis, and Blastocystis [52,53,54].
The clinical significance or role of D. fragilis in human gastrointestinal diseases is one of the most debated topics in the literature [11]. In our study, we compared common gastrointestinal symptoms between the D. fragilis-infected and non-infected groups, and none of these symptoms showed significant differences. The most frequently reported symptoms in D. fragilis-infected patients are diarrhoea and abdominal pain [55, 56]. Today we are still far from concluding whether D. fragilis causes specific symptoms or not [12, 46]. It was difficult in our study to differentiate the specific symptoms of D. fragilis, as they are non-specific [11]. European countries including, the Netherlands, Denmark, and Belgium have reported higher D. fragilis colonization in asymptomatic control groups compared to the patients with gastrointestinal symptoms [57,58,59]. However, large-scale studies, mostly in Europe, have reported a correlation between D. fragilis infection and clinical symptoms, supporting the idea of its pathogenic nature [11]. The frequency of D. fragilis was 8.2% in patients with gastrointestinal symptoms and 26.9% in the control group [60]. Recently, microbiota-based studies also support the asymptomatic beneficial effect or relation of D. fragilis in the human gut [61]. The genetic diversity among D. fragilis isolates may lead to the variations in the clinical outcomes of the infection [18]. Therefore, the genetic analysis of cysteine protease-coding genes and other virulence factors has the potential to explain the clinical differences [18, 62]. It was reported that a specific cysteine proteinase (CP5) coding gene was greatly differed between pathogenic and non-pathogenic human Entamoeba species. The CP5 coding gene was highly degenerated, with many insertions and deletions in non-pathogenic E. dispar [63]. Currently, we have very limited knowledge about the possible outcomes of cathepsin like peptidase polymorphisms in the pathogenesis of D. fragilis. Therefore, additional studies both in vitro and in vivo are needed. We observed that gender and age were not significantly associated with D. fragilis positivity. In contrast to our finding, higher frequencies were reported in females than in males [1, 62, 64]. However, the predisposing role of gender and age in the infection remains unclear [16, 55]. We excluded faecal samples containing parasitic agents at the beginning of the study, and these samples were not included in the analysis. A limitation of our study to analyse the clinical findings is that, since we collected samples from a routine parasitology laboratory, we did not have the opportunity to include microbiological examination results of the samples. Therefore, the lack of microbial examination hindered our ability to analyse whether the symptoms were related to D. fragilis or any microbiological agents including bacteria, viruses, and fungi.
In conclusion, the D. fragilis isolates in the southwest of Turkey exhibited minimal genetic variation in both SSU-rRNA and six housekeeping gene sequences, supporting the clonal distribution of D. fragilis. In the study, we gave a detailed methodology for the amplification of nested-PCR protocol for D. fragilis six housekeeping genes. Clinical findings indicated asymptomatic or non-pathogenic colonization. However, additional molecular studies are needed for a more comprehensive understanding of clinical characteristics and the molecular epidemiology of D. fragilis.
References
Stark D, Barratt J, Chan D et al (2016) Dientamoeba fragilis, the neglected trichomonad of the human bowel. Clin Microbiol Rev 29(3):553–580. https://doi.org/10.1128/CMR.00076-15
Yildiz I, Erdem Aynur Z (2022) First detection and molecular characterization of Dientamoeba fragilis in cattle. Zoonoses Public Health 69(8):897–903. https://doi.org/10.1111/zph.12986
Yetismis G, Yildirim A, Pekmezci D et al (2022) First report and genotyping of Dientamoeba fragilis in pet budgerigars (Melopsittacus undulatus), with zoonotic importance. Zoonoses Public Health 69(5):572–578. https://doi.org/10.1111/zph.12949
Caccio SM, Sannella AR, Manuali E et al (2012) Pigs as natural hosts of Dientamoeba fragilis genotypes found in humans. Emerg Infect Dis 18(5):838–841. https://doi.org/10.3201/eid1805.111093
Johnson EH, Windsor JJ, Clark CG (2004) Emerging from obscurity: biological, clinical, and diagnostic aspects of Dientamoeba fragilis. Clin Microbiol Rev 17(3):553–570. https://doi.org/10.1128/CMR.17.3.553-570.2004
Ogren J, Dienus O, Lofgren S et al (2013) Dientamoeba fragilis DNA detection in Enterobius vermicularis eggs. Pathog Dis 69(2):157–158. https://doi.org/10.1111/2049-632X.12071
Stark D, Garcia LS, Barratt JL et al (2014) Description of Dientamoeba fragilis cyst and precystic forms from human samples. J Clin Microbiol 52(7):2680–2683. https://doi.org/10.1128/JCM.00813-14
Abdo SM, El-Adawy H, Farag HF et al (2021) Detection and molecular identification of Blastocystis isolates from humans and cattle in northern Egypt. J Parasitic Dis 2021:1–8
Munasinghe VS, Vella NG, Ellis JT et al (2013) Cyst formation and faecal-oral transmission of Dientamoeba fragilis-the missing link in the life cycle of an emerging pathogen. Int J Parasitol 43(11):879–883. https://doi.org/10.1016/j.ijpara.2013.06.003
Garcia LS (2016) Dientamoeba fragilis, one of the neglected intestinal Protozoa. J Clin Microbiol 54(9):2243–2250. https://doi.org/10.1128/JCM.00400-16
van Gestel RS, Kusters JG, Monkelbaan JF (2019) A clinical guideline on Dientamoeba fragilis infections. Parasitology 146(9):1131–1139. https://doi.org/10.1017/S0031182018001385
Miguel L, Salvador F, Sulleiro E et al (2018) Clinical and epidemiological characteristics of patients with Dientamoeba fragilis infection. Am J Trop Med Hyg 99(5):1170–1173. https://doi.org/10.4269/ajtmh.18-0433
Jokelainen P, Hebbelstrup Jensen B, Andreassen BU et al (2017) Dientamoeba fragilis, a commensal in children in Danish Day Care centers. J Clin Microbiol 55(6):1707–1713. https://doi.org/10.1128/JCM.00037-17
van Kalleveen MW, Budding AE, Benninga MA et al (2021) Intestinal microbiota in children with symptomatic Dientamoeba fragilis infection: a case-control study. Pediatr Infect Dis J 40(4):279–283. https://doi.org/10.1097/INF.0000000000002975
Gough R, Ellis J, Stark D (2019) Comparison and recommendations for use of Dientamoeba fragilis real-time PCR assays. J Clin Microbiol 57(5). https://doi.org/10.1128/JCM.01466-18
Barratt JL, Harkness J, Marriott D et al (2011) The ambiguous life of Dientamoeba fragilis: the need to investigate current hypotheses on transmission. Parasitology 138(5):557–572. https://doi.org/10.1017/S0031182010001733
Windsor JJ, Macfarlane L, Clark CG (2006) Internal transcribed spacer dimorphism and diversity in Dientamoeba fragilis. J Eukaryot Microbiol 53(3):188–192. https://doi.org/10.1111/j.1550-7408.2006.00092.x
Stensvold CR, Clark CG, Roser D (2013) Limited intra-genetic diversity in Dientamoeba fragilis housekeeping genes. Infect Genet Evol 18:284–286. https://doi.org/10.1016/j.meegid.2013.05.003
Windsor JJ, Clark CG, Macfarlane L (2004) Molecular typing of Dientamoeba fragilis. Br J Biomed Sci 61(3):153–155. https://doi.org/10.1080/09674845.2004.11978138
Ibarz Pavon AB, Maiden MC (2009) Multilocus sequence typing. Methods Mol Biol 551:129–140. https://doi.org/10.1007/978-1-60327-999-4_11
Bastardo A, Ravelo C, Romalde JL (2012) Multilocus sequence typing reveals high genetic diversity and epidemic population structure for the fish pathogen Yersinia ruckeri. Environ Microbiol 14(8):1888–1897. https://doi.org/10.1111/j.1462-2920.2012.02735.x
Frazao-Teixeira E, Sundar N, Dubey JP et al (2011) Multi-locus DNA sequencing of Toxoplasma Gondii isolated from Brazilian pigs identifies genetically divergent strains. Vet Parasitol 175(1–2):33–39. https://doi.org/10.1016/j.vetpar.2010.09.030
Gilchrist CA, Ali IK, Kabir M et al (2012) A Multilocus sequence typing system (MLST) reveals a high level of diversity and a genetic component to Entamoeba histolytica virulence. BMC Microbiol 12:151. https://doi.org/10.1186/1471-2180-12-151
Herrera G, Hernandez C, Ayala MS et al (2017) Evaluation of a multilocus sequence typing (MLST) scheme for Leishmania (Viannia) braziliensis and Leishmania (Viannia) panamensis in Colombia. Parasit Vectors 10(1):236. https://doi.org/10.1186/s13071-017-2175-8
Caccio SM, Sannella AR, Bruno A et al (2016) Multilocus sequence typing of Dientamoeba fragilis identified a major clone with widespread geographical distribution. Int J Parasitol 46(12):793–798. https://doi.org/10.1016/j.ijpara.2016.07.002
Rosner B, Glynn RJ (2011) Power and sample size estimation for the clustered wilcoxon test. Biometrics 67(2):646–653. https://doi.org/10.1111/j.1541-0420.2010.01488.x
Peek R, Reedeker FR, van Gool T (2004) Direct amplification and genotyping of Dientamoeba fragilis from human stool specimens. J Clin Microbiol 42(2):631–635. https://doi.org/10.1128/JCM.42.2.631-635.2004
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4(4):406–425. https://doi.org/10.1093/oxfordjournals.molbev.a040454
Unalan-Altintop T, Vahabov C, Ergunay K et al (2022) Investigation of Dientamoeba fragilis and Blastocystis in patients from Turkey with ulcerative colitis and irritable bowel syndrome: any relation with genotypes? Acta Trop 231:106451. https://doi.org/10.1016/j.actatropica.2022.106451
Rozas J, Sanchez-DelBarrio JC, Messeguer X et al (2003) DnaSP, DNA polymorphism analyses by the coalescent and other methods. Bioinformatics 19(18):2496–2497. https://doi.org/10.1093/bioinformatics/btg359
Roser D, Nejsum P, Carlsgart AJ et al (2013) DNA of Dientamoeba fragilis detected within surface-sterilized eggs of Enterobius vermicularis. Exp Parasitol 133(1):57–61. https://doi.org/10.1016/j.exppara.2012.10.009
Stark D, Beebe N, Marriott D et al (2005) Detection of Dientamoeba fragilis in fresh stool specimens using PCR. Int J Parasitol 35(1):57–62. https://doi.org/10.1016/j.ijpara.2004.09.003
Stark D, Beebe N, Marriott D et al (2006) Evaluation of three diagnostic methods, including real-time PCR, for detection of Dientamoeba fragilis in stool specimens. J Clin Microbiol 44(1):232–235. https://doi.org/10.1128/JCM.44.1.232-235.2006
Tolba MM, Allam AF, Khalil SS et al (2022) Evaluation of microscopy and PCR for detection of Dientamoeba fragilis. Trop Parasitol 12(2):87–93. https://doi.org/10.4103/tp.tp_93_21
Aykur M, Caliskan Kurt C, Dirim Erdogan D et al (2019) Investigation of Dientamoeba fragilis prevalence and evaluation of Sociodemographic and clinical features in patients with gastrointestinal symptoms. Acta Parasitol 64(1):162–170. https://doi.org/10.2478/s11686-018-00017-5
Yildiz I, Ertug S, Tileklioglu E et al (2021) Investigation of Dientamoeba fragilis frequency in faecal samples of patients with Enterobius vermicularis infection by polymerase chain reaction. Turkiye Parazitol Derg 45(3):195–200. https://doi.org/10.4274/tpd.galenos.2021.7257
Sivcan E, Charyyeva A, Ceylan SS et al (2018) Dientamoeba fragilis infection in patients with gastrointestinal system complaints. Mikrobiyol Bul 52(2):166–179. https://doi.org/10.5578/mb.66468
Gureser AS, Karasartova D, Sarzhanov F et al (2023) Prevalence of Blastocystis and Dientamoeba fragilis in diarrheal patients in Corum, Turkiye. Parasitol Res 122(12):2977–2987. https://doi.org/10.1007/s00436-023-07987-0
Jirku M, Kasparova A, Lhotska Z et al (2022) A cross-sectional study on the occurrence of the intestinal protist, Dientamoeba fragilis, in the gut-healthy volunteers and their animals. Int J Mol Sci 23(23). https://doi.org/10.3390/ijms232315407
Guadano-Procesi I, Berrilli F, Di Montalbano M et al (2024) Detection and genotyping of Dientamoeba fragilis from symptomatic patients: new insights from Italy into a little-known gastrointestinal protozoan. Parasitol Int 98:102816. https://doi.org/10.1016/j.parint.2023.102816
Jerez Puebla LE, Nunez-Fernandez FA, Fraga Nodarse J et al (2020) Diagnosis of intestinal protozoan infections in patients in Cuba by microscopy and molecular methods: advantages and disadvantages. J Microbiol Methods 179:106102. https://doi.org/10.1016/j.mimet.2020.106102
Oyegue-Liabagui SL, Ndjangangoye NK, Kouna LC et al (2020) Molecular prevalence of intestinal parasites infections in children with diarrhea in Franceville, Southeast of Gabon. BMC Infect Dis 20(1):350. https://doi.org/10.1186/s12879-020-05071-x
Menu E, Mary C, Toga I et al (2019) A hospital qPCR-based survey of 10 gastrointestinal parasites in routine diagnostic screening. Marseille France Epidemiol Infect 147:e100. https://doi.org/10.1017/S0950268819000165
Johnson JA, Clark CG (2000) Cryptic genetic diversity in Dientamoeba fragilis. J Clin Microbiol 38(12):4653–4654. https://doi.org/10.1128/JCM.38.12.4653-4654.2000
Caccio SM (2018) Molecular epidemiology of Dientamoeba fragilis. Acta Trop 184:73–77. https://doi.org/10.1016/j.actatropica.2017.06.029
Oliveira-Arbex AP, David EB, Caccio SM et al (2021) Prevalence and genetic characterization of Dientamoeba fragilis in asymptomatic children attending daycare centers. Rev Inst Med Trop Sao Paulo 63:e39. https://doi.org/10.1590/S1678-9946202163039
McKerrow JH, Sun E, Rosenthal PJ et al (1993) The proteases and pathogenicity of parasitic protozoa. Annu Rev Microbiol 47:821–853. https://doi.org/10.1146/annurev.mi.47.100193.004133
Na BK, Kang JM, Sohn WM (2008) CsCF-6, a novel cathepsin F-like cysteine protease for nutrient uptake of Clonorchis sinensis. Int J Parasitol 38(5):493–502. https://doi.org/10.1016/j.ijpara.2007.09.001
Yang N, Matthew MA, Yao C (2023) Roles of cysteine proteases in biology and pathogenesis of parasites. Microorganisms 11(6):1397. https://doi.org/10.3390/microorganisms11061397
Siqueira-Neto JL, Debnath A, McCall LI et al (2018) Cysteine proteases in protozoan parasites. PLoS Negl Trop Dis 12(8):e0006512. https://doi.org/10.1371/journal.pntd.0006512
Ukaegbu UE, Deitsch KW (2015) The emerging role for RNA polymerase II in regulating virulence gene expression in malaria parasites. PLoS Pathog 11(7):e1004926. https://doi.org/10.1371/journal.ppat.1004926
Chang Y, Li J, Zhang L (2023) Genetic diversity and molecular diagnosis of Giardia. Infect Genet Evol 113:105482. https://doi.org/10.1016/j.meegid.2023.105482
Cornelius DC, Robinson DA, Muzny CA et al (2012) Genetic characterization of Trichomonas Vaginalis isolates by use of multilocus sequence typing. J Clin Microbiol 50(10):3293–3300. https://doi.org/10.1128/JCM.00643-12
Ertug S, Malatyali E, Ertabaklar H et al (2016) Multilocus sequence typing of Blastocystis isolates in Aydin. Turk Parasitol Int 65(6 Pt B):760–762. https://doi.org/10.1016/j.parint.2016.06.006
Stark D, Barratt J, Roberts T et al (2010) A review of the clinical presentation of dientamoebiasis. Am J Trop Med Hyg 82(4):614–619. https://doi.org/10.4269/ajtmh.2010.09-0478
Vandenberg O, Peek R, Souayah H et al (2006) Clinical and microbiological features of dientamoebiasis in patients suspected of suffering from a parasitic gastrointestinal illness: a comparison of Dientamoeba fragilis and Giardia lamblia infections. Int J Infect Dis 10(3):255–261. https://doi.org/10.1016/j.ijid.2005.05.011
Brands MR, Van de Vijver E, Haisma SM et al (2019) No association between abdominal pain and Dientamoeba in Dutch and Belgian children. Arch Dis Child 104(7):686–689. https://doi.org/10.1136/archdischild-2018-316383
van Bruijnesteijn LE, Dullaert-de Boer M, Ruijs GJ et al (2015) Case-control comparison of bacterial and protozoan microorganisms associated with gastroenteritis: application of molecular detection. Clin Microbiol Infect 21(6):592 e9-19. https://doi.org/10.1016/j.cmi.2015.02.007
Krogsgaard LR, Engsbro AL, Stensvold CR et al (2015) The prevalence of intestinal parasites is not greater among individuals with irritable bowel syndrome: a population-based case-control study. Clin Gastroenterol Hepatol 13(3):507–513e2. https://doi.org/10.1016/j.cgh.2014.07.065
Kurt O, Oktem Okullu S, Ozen NM et al (2020) Is Dientamoeba fragilis friend or foe? A study from Istanbul on patients and healthy individuals from different cohorts. Turk Mikrobiyol Cem Derg 50:244–249. https://doi.org/10.5222/TMCD.2020.244
Gotfred-Rasmussen H, Stensvold CR, Ingham AC et al (2021) Impact of metronidazole treatment and Dientamoeba fragilis colonization on gut microbiota diversity. J Pediatr Gastroenterol Nutr 73(1):23–29. https://doi.org/10.1097/MPG.0000000000003096
Barratt JL, Harkness J, Marriott D et al (2011) A review of Dientamoeba fragilis carriage in humans: several reasons why this organism should be considered in the diagnosis of gastrointestinal illness. Gut Microbes 2(1):3–12. https://doi.org/10.4161/gmic.2.1.14755
Willhoeft U, Hamann L, Tannich E (1999) A DNA sequence corresponding to the gene encoding cysteine proteinase 5 in Entamoeba histolytica is present and positionally conserved but highly degenerated in Entamoeba dispar. Infect Immun 7(11):5925–5929. https://doi.org/10.1128/IAI.67.11.5925-5929.1999
Ganci M, Butt H, Tyrrell J et al (2023) The effect of Blastocystis sp. and Dientamoeba fragilis on psychological symptom severity in a sample of clinically diverse males and females. Curr Psychol 42:4017–4030. https://doi.org/10.1007/s12144-021-01700-z
Acknowledgements
This work was supported by the Scientific and Technological Research Council of Turkey (TUBITAK), Project no: 220S421.
Author information
Authors and Affiliations
Contributions
Study conception and design: E.M., IY., and S.E.; data collection: I.Y., and H.E. Analysis and interpretation of results: O.G., and EM., and manuscript preparation: E.M., OG., S.E., and H.E.
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Malatyali, E., Guclu, O., Yildiz, I. et al. The Low Genetic Diversity of Dientamoeba fragilis Isolates in Southwest Turkey and Analysis of Clinical Findings. Acta Parasit. 69, 1576–1586 (2024). https://doi.org/10.1007/s11686-024-00896-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11686-024-00896-x