Abstract
Genetic mosaicism, characterized by multiple genotypes within an individual, is considered an obstacle to CRISPR/Cas9 genome editing in animal models. Despite the various strategies for minimizing mosaic mutations, no definitive methods exist to eliminate them. This study aimed to enhance gene editing efficiency in porcine zygotes using CRISPR/Cas9, which targets specific genes through centrifugation and zona pellucida removal before electroporation. Centrifugation at 2000 × g did not adversely affect blastocyst formation rates in zygotes electroporated with gRNA targeting the GGTA1 gene; instead, it led to increased total and monoallelic mutation rates compared with control zygotes without centrifugation. However, the groups had no significant differences in biallelic mutation rates. In zygotes electroporated with gRNA targeting the CMAH gene, centrifugation treatments exceeding 1000 × g significantly increased both biallelic mutation rates and mutation efficiency. The combination of centrifugation and zona pellucida removal did not have a detrimental effect on blastocyst formation rates. It led to a higher rate of double biallelic mutations in embryos targeting both GGTA1 and CMAH compared to embryos without centrifugation treatment. In summary, our results demonstrate that pre-electroporation treatments, including centrifugation and zona pellucida removal, positively influenced the reduction of mosaic mutations, with the effectiveness of centrifugation depending on the specific gRNA used.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Genetic mosaicism, defined as the presence of more than one genotype within a single individual, can arise through various mechanisms. These include natural processes, such as chromosomal non-disjunction, anaphase lag, endoreplication, mutations occurring during development, and manipulation mechanisms, such as genome editing (Taylor et al. 2014; Mehravar et al. 2019). The method of choice for introducing genomic alterations into animal models is currently the CRISPR/Cas9 system (Mehravar et al. 2019). A typical approach for generating knockout and transgenic animal models involves directly introducing CRISPR/Cas9 components, including DNA, RNA, or protein molecules, into fertilized zygotes using microinjection (Li et al. 2013c; Wang et al. 2013; Yang et al. 2013), electroporation (Hirata et al. 2019; Namula et al. 2019), or transfection (Le et al. 2022) methods. However, one of the outcomes of employing CRISPR-mediated gene editing in embryos is the occurrence of genetic mosaicism in the founders. Various degrees of mosaicism have been observed in mice (Shen et al. 2013), rats (Li et al. 2013a), cynomolgus monkeys (Niu et al. 2014), and zebrafish (Ablain et al. 2015). To overcome mosaicism, generating a founder animal with the desired modifications is essential, followed by developing new mutant strains by outcrossing the mosaic founders. Although rodents can undergo this procedure within a few months, its completion requires several years in other species, such as non-human primates (Niu et al. 2014; Mehravar et al. 2019; Wang et al. 2024). Therefore, overcoming mosaicism at the founder stage is advantageous because it reduces the time required.
Various strategies (such as accelerating the editing process, shortening Cas9 longevity through embryo splitting, implementing germline modifications, and utilizing precise genome editing with the CRISPR/Cas9 system) can effectively minimize mosaic mutations (Mehravar et al. 2019). However, it should be noted that there are no definitive and guaranteed strategies to eliminate mosaic mutations arising from CRISPR/Cas9 genome editing. Previously, we successfully generated mutant blastocysts by introducing the CRISPR/Cas9 system into zygotes via electroporation (Hirata et al. 2019; Namula et al. 2019), a technique commonly used to introduce foreign genetic material, such as DNA or RNA, into cells by applying an electric field. Therefore, our objective was to enhance gene editing efficiency by mitigating the mosaicism associated with electroporation. Given that lipid droplets in cells play crucial roles in lipid metabolism, energy storage, and signaling (Wang 2016), modulating the polarization of lipid droplets could potentially influence the efficiency of cellular uptake during electroporation, thereby facilitating the targeted introduction of genetic material into specific cellular compartments. One method for polarizing intracellular lipid droplets in putative zygotes is high-speed centrifugation (Kato and Nagao 2009; Wirtu et al. 2013). Moreover, the zona pellucida surrounding the plasma membrane of the zygote may present a barrier preventing access of CRISPR/Cas9 components to the zygote by electroporation. Therefore, in the present study, we examined the effect of centrifugation treatment, with or without zona pellucida removal before electroporation, on the development, mutation, and mutation efficiency of putative porcine zygotes edited with gRNA targeting the α1,3-Galactosyltransferase (GGTA1) and CMP-Neu5Ac hydroxylase (CMAH) genes.
Materials and methods
Ethical approval
All animal experiments were approved by the Institutional Animal Care and Use Committee of Tokushima University (approval number: T2019-11).
Oocyte collection, in vitro maturation, fertilization, and embryo culture
Oocyte collection, in vitro maturation, fertilization, and embryo culture were performed as previously described (Lin et al. 2022). Briefly, pig ovaries were obtained from prepubertal crossbred gilts (Landrace × Large White × Duroc) at a local slaughterhouse. Cumulus-oocyte complexes (COCs) with a uniform ooplasm and compact cumulus cell mass were cultured in a maturation medium at 39 °C in a humidified incubator containing 5% CO2. The maturation medium consisted of 25 mM HEPES tissue culture medium 199 with Earle’s salts (TCM 199; Invitrogen Co., Carlsbad, CA) supplemented with 10% (v/v) porcine follicular fluid, 0.6 mM of cysteine (Sigma-Aldrich, St. Louis, MO), 50 µM of sodium pyruvate (Sigma-Aldrich), 2 mg/mL of D-sorbitol (Wako Pure Chemical Industries Ltd., Osaka, Japan), 1 µg/mL of 17 β-estradiol (Sigma-Aldrich), 10 IU/mL of equine chorionic gonadotropin (Kyoritu Seiyaku, Tokyo, Japan), 10 IU/mL of human chorionic gonadotropin (Kyoritu Seiyaku), and 50 µg/mL of gentamicin (Sigma-Aldrich). After 20–22 h of maturation, the COCs were cultured for an additional 24 h in a maturation medium without hormones under the same atmosphere.
For in vitro fertilization (IVF), frozen-thawed spermatozoa were transferred to 6 mL of porcine fertilization medium (PFM; Research Institute for the Functional Peptides Co., Yamagata, Japan) and washed by centrifugation at 500 × g for 5 min. Pelleted spermatozoa were resuspended in PFM and adjusted to a concentration of 5 × 106 cells/mL. The matured oocytes were then transferred to sperm-containing PFM and co-incubated for 5 h at 39 °C under 5% CO2, 5% O2, and 90% N2.
After IVF, the putative zygotes were denuded from the cumulus cells and attached spermatozoa by mechanical pipetting, transferred to porcine zygote medium (PZM-5; Research Institute for the Functional Peptides Co.), and cultured continuously in vitro at 39 °C in a humidified incubator containing 5% CO2, 5% O2, and 90% N2. All cleaved embryos were transferred into 100 μL droplets of porcine blastocyst medium (PBM; Research Institute for the Functional Peptides Co.) 72 h after insemination. The embryos were subsequently cultured for an additional 4 d to evaluate their ability to develop to the blastocyst stage and to examine the efficiency of genome editing in blastocysts.
Centrifugation treatment
The putative zygotes were collected 10 h after initiating IVF. Before electroporation treatment, the zygotes were transferred into a 2 mL Sarstedt screw cap tube containing 500 µL of PZM-5, and then the tubes were centrifuged at various gravities for 10 min at 35.0 °C by a high-speed refrigerated centrifuge (MX-307; Tomy Seiko, Tokyo, Japan) to polarize the intracellular lipid droplets (Fig. 1A and B).
Electroporation
The gRNAs targeting the GGTA1 or CMAH genes were prepared as previously described (Le et al. 2022). The target sequences were GGTA1 (5′- AGACGCTATAGGCAACGAAA-3′) and CMAH (5′-GAAGCTGCCAATCTCAAGGA-3′). Electroporation was performed as described previously (Tanihara et al. 2016). An electrode (LF501PT1-20; BEX, Tokyo, Japan) was connected to a CUY21EDIT II electroporator (BEX) and placed under a stereoscopic microscope. The zygotes with or without centrifugation treatment (approximately 30–40 zygotes) were placed in a line in the electrode gap in a chamber slide filled with 10 μL of Opti-MEM I solution with 100 ng/μL of gRNA and 100 ng/µL of Cas9 protein, and then electroporated by five 1 ms pulses at 25 V. After electroporation, the embryos were cultured in PZM-5 and PBM as described above.
Analysis of targeted gene sequence in a single blastocyst
We analyzed the frequencies of base insertions or deletions (indels) in the target regions of individual blastocysts to compare the efficiency of introducing target mutations into the embryos. The genomic DNA of individual blastocysts was extracted by heat treatment with 50 mM NaOH to investigate the CRISPR/Cas9-mediated mutations. After neutralization, the DNA was subjected to polymerase chain reaction (PCR) using the KOD One PCR Master Mix (Toyobo, Osaka, Japan) according to the manufacturer’s instructions. Gene-specific primers used for amplifications were as follows: GGTA1:5′-AAAAGGGGAGCACTGAACCT-3′ (forward) and 5′-ATCCGGACCCTGTTTTAAGG-3′ (reverse), and CMAH: 5′- GCTGTCAATGCTCAGGGATT-3′ (forward) and 5′- TGCCAAACCTAATTGGGAGA-3′ (reverse). After purification of the PCR products with a Fast Gene Gel/PCR Extraction Kit (Nippon Genetics, Tokyo, Japan), the sequences of the target regions were analyzed by Sanger sequencing using a BigDye Terminator Cycle Sequencing Kit version 3.1 (Thermo Fisher Scientific K.K., Tokyo, Japan) on an ABI 3500 genetic analyzer (Applied Biosystems, CA). The TIDE (tracking of indels by decomposition; https://tide.deskgen.com/) bioinformatics package was used to quantify the frequency of indel mutations in blastocysts electroporated with GGTA1 or CMAH gRNAs (Brinkman et al. 2014). According to the sequences of the target regions, blastocyst genotypes were classified as biallelic mutants (carrying no WT sequence), monoallelic mutants (carrying a mutation and the WT sequence), or WT (carrying only the WT sequence). The mutation rate was defined as the ratio of the number of gene-edited blastocysts to the total number of sequenced blastocysts. The mutation efficiency was defined as the proportion of indel mutation events in the mutant blastocysts.
Experimental design
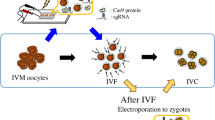
In the first experiment, we examined the effects of gravity centrifugation before electroporation in putative zygotes on embryonic development and target mutations in the resulting blastocysts. Zygotes collected 10 h after the start of IVF were centrifuged at various gravities (1000, 2000, 5000, and 10,000 × g) for 10 min. After centrifugation, the zygotes were electroporated with gRNAs targeting either GGTA1 or CMAH (Fig. 2) and then cultured with PZM-5 and PBM for 7 d, as described above. As a control, the zygotes were electroporated without centrifugation.
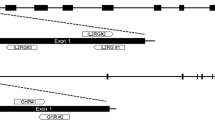
Relative positions of gRNA target sequences and primer pairs (flanking the gRNA target site). Fragments with 449 bp and 333 bp were generated for sequence analysis of GGTA1 and CMAH, respectively. Nucleotides in blue and red represent the target sequences and PAM sequences of each gRNA, respectively.
In the second experiment, we compared the effects of removing the zona pellucida (ZP) after centrifugation in putative zygotes on embryonic development and target mutations in the resulting blastocysts because ZP removal may increase mutation rates (Namula et al. 2022). Some zygotes, with or without centrifugation at 1000 × g, were exposed to 0.5% (w/v) actinase-E (Kaken-Seiyaku Corp., Tokyo, Japan) in Dulbecco’s PBS (Nissui Pharmaceutical, Tokyo, Japan) for 20–30 s, transferred to PZM-5 without actinase-E, and freed completely from their ZP by gentle pipetting (Fig. 1C and D). For the one-step generation of multiple gene modifications in pigs, after removing the ZP, ZP-free zygotes were electroporated with two gRNAs targeting GGTA1 and CMAH to generate embryos with both GGTA1 and CMAH mutations. Electroporated zygotes were then cultured with PZM-5 and PBM in four-well dishes for 7 d, as described above. Centrifugation at 1000 × g was the most suitable for embryonic development and target mutation in the first experiment was used in the second experiment. As a control, the ZP-intact zygotes with or without centrifugation were electroporated.
Statistical analysis
The percentage data for embryos that developed to the blastocyst stage were compared by analysis of variance (ANOVA) using the general linear model procedure in SAS for Windows, version 9.1 (SAS Institute, Cary, NC). The percentage of gene-edited blastocysts was analyzed using the chi-square test with Yates correction. In the second experiment, the statistical model included the ZP, centrifugation, and two-way interactions. If the interactions were not significant, they were excluded from the model but retained to test the effects of ZP or centrifugation treatment. Differences with a p-value (P) of 0.05 or less were considered statistically significant.
Results
As shown in Tables 1 and 2, the centrifugation treatment did not affect the blastocyst formation rates of zygotes electroporated with gRNA targeting GGTA1 and CMAH genes, irrespective of the centrifugation gravity before electroporation treatment. Assessing the target sequence of the resulting blastocysts revealed that centrifugation at 2000 × g increased the total and monoallelic mutation rates of zygotes electroporated with gRNA targeting the GGTA1 gene compared to those of control zygotes without centrifugation treatment. However, the groups had no significant differences in biallelic mutation rates. In contrast, centrifugation at more than 1000 × g significantly increased the biallelic mutation rates and mutation efficiency of zygotes electroporated with gRNA targeting CMAH (P < 0.05).
When comparing the effects of centrifugation of zygotes before electroporation in combination with ZP removal on the embryonic development of zygotes electroporated simultaneously with two gRNAs targeting GGTA1 and CMAH genes, there were no significant differences in the rates of blastocyst formation (Fig. 3). When mutation rates were assessed after individual sequencing of the target sites of each gene in the resulting blastocysts (Fig. 4), the rate of double biallelic mutations was significantly higher (P < 0.05) in embryos with centrifugation treatment combined with ZP removal than in embryos without centrifugation treatment. Moreover, all embryos had mutations in the GGTA1 and CMAH genes due to ZP removal. However, there were differences in the rates of double biallelic mutations between ZP-free embryos with and without centrifugation.
Effects of removing the zona pellucida (ZP) after centrifugation in putative zygotes on embryonic development and target mutations. After centrifugation at 1000 × g, ZP-intact (ZP +) and ZP-free (ZP-) zygotes were electroporated. As a control, both zygotes without centrifugation treatment were electroporated. Electroporation treatment introduced two gRNAs targeting GGTA1 and CMAH genes simultaneously into zygotes. (A) Blastocyst formation rates of cultured embryos. Blastocyst development was assessed after 7 d. (B) The genotype of blastocysts was determined using Sanger sequencing and TIDE analysis. Each mutation rate was defined as the ratio of the number of edited blastocysts to the total number of sequenced blastocysts. Single mutation, blastocysts carrying a mutation in GGTA1 or CMAH; Double-mutation, blastocysts carrying mutations in GGTA1 and CMAH; Double biallelic mutation, blastocysts carrying biallelic mutations in GGTA1 and CMAH. Five replicate trials were performed. Data are expressed as mean ± SEM. The numbers within parentheses indicate the total number of examined oocytes or blastocysts. Superscript letters, A-B and a-b, show a significant difference in the rate of “Double biallelic mutation” and “Double mutation,” respectively (P < 0.05).
Representative results of Sanger sequencing of blastocysts formed after electroporation treatment with Cas9 protein, GGTA1 (left), and CMAH (right) gRNAs. The TIDE bioinformatics package was used to determine the genotype of each blastocyst. Single mutation: GGTA1 indicates a mixture of mosaic mutation with multiple indel mutations and wild type (WT), and CMAH indicates WT. Double mutation: GGTA1 and CMAH indicate a mixture of mosaic mutation with multiple indel mutations and WT. Double biallelic mutation: GGTA1 indicates biallelic mutation with only 10 base deletions, and CMAH indicates biallelic mutation with no WT sequence and two types of indels (14 base deletions and 5 base insertions).
Discussion
Mosaicism reduction can generally be achieved using in vitro transcription (IVT) of sgRNAs and Cas9 instead of CRISPR/Cas9 plasmids (Horii et al. 2014), and for further improvement, utilizing the Cas9 protein in the Cas9 protein/sgRNA format, rather than Cas9 RNA, has been demonstrated to be effective (Aida et al. 2015; Hendel et al. 2015; Burger et al. 2016). However, these two approaches have decreased mosaicism without complete elimination (Mehravar et al. 2019). In the present study, we investigated additional methods to enhance system efficacy and mitigate the persistent challenges of mosaicism.
In the first experiment, we examined the effects of gravitational centrifugation before electroporation in putative zygotes on embryonic development and target mutations in the resulting blastocysts. When employing gRNA targeting either the GGTA1 or CMAH genes, centrifugation of putative zygotes before electroporation yielded superior results for certain parameters, such as total mutation, monoallelic mutation, biallelic mutation, or mutation efficiency in the resulting blastocysts, compared to the control condition. Porcine embryos may exhibit greater tolerance for the uptake of foreign genetic material because of their higher cytoplasmic lipid content than embryos of other domestic animals (Tominaga et al. 2000). This factor is potentially attributed to the crucial role of lipids in signaling, membrane trafficking, and cell polarization (Rajendran and Simons 2005). This suggests that lipid droplet polarization during high-speed centrifugation could potentially influence electroporation efficiency without adversely affecting embryo development, as it triggers cellular responses, such as alterations in membrane dynamics, endocytosis (Rajendran and Simons 2005), or other processes related to the uptake of foreign genetic material. However, the effects of centrifugation gravity on the target mutations in the resulting blastocysts appear to be dependent on the gRNA used; the gRNA targeting the GGTA1 gene yielded favorable results for total and monoallelic mutation rates at 2000 × g, whereas the gRNA targeting the CMAH gene produced favorable results for biallelic mutation rates and mutation efficiency at levels exceeding 1000 × g. This difference may be associated with the intrinsic properties of the target locus, where the resulting double-strand breaks (DSBs) and repair products could influence the nature of genetic mutations (Mehravar et al. 2019). Despite these observations, the exact underlying mechanism remains to be conclusively identified and understood.
In the second experiment, the effects of ZP removal after centrifugation in putative zygotes before simultaneous electroporation with two gRNAs targeting the GGTA1 and CMAH genes on both embryonic development and the induction of target mutations in the resulting blastocysts were compared, as increased mutation rates may be associated with ZP removal. The ZP serves as a protective coating for mammalian oocytes and embryos, preserving the integrity of the preimplantation embryo, ensuring a stable microenvironment, and shielding the embryo from bacteria, fungi, immune cells, and mechanical injury while traversing the oviduct (Wassarman and Litscher 2008; Vajta et al. 2010; Li et al. 2013b). In vitro procedures can induce hardening of the ZP due to the influence of various chemical agents (Cohen et al. 1990), resulting in alterations to the ZP structure (Michelmann et al. 2007), which could pose an obstacle during electroporation. Our results indicate that ZP removal had no significant adverse effects on embryonic development. This finding aligns with the findings of Li et al. (2013b), who reported that ZP removal might enhance embryonic quality by accelerating embryonic development and reducing the number of apoptotic cells in blastocysts. This suggests that combining ZP removal with centrifugation is viable for preparing porcine zygotes before electroporation, as it demonstrates no negative impact on embryo development.
When evaluating the mutation rates, all embryos showed mutations in GGTA1 and CMAH attributable to ZP removal, regardless of centrifugation. We used a group culture system of ZP-free zygotes and embryos after electroporation, which may lead to the aggregation of two or more embryos in some embryos, resulting in chimeric blastocysts. This was a serious limitation of our group culture system as a method for evaluating the development and mutation of edited zygotes. Therefore, individual culture systems for ZP-free embryos should be optimized, e.g., using individual micro-well cultures, to achieve efficient embryo development and thus address the limiting factor of embryo aggregation leading to chimerism. However, our observation supports the hypothesis that the presence of ZP impedes electroporation. Generally, the electrical pulse applied during electroporation induces the reversible breakdown of the ZP and the cellular membrane, leading to pore formation that facilitates the introduction of DNA/RNA into embryos (Hirata et al. 2019). Consequently, without the ZP, the electrical pulse would only be needed to induce pore formation in the cellular membrane, simplifying the introduction of foreign genetic material into the embryos. This suggests that, in the present study, the removal of the ZP was crucial for delivering all CRISPR/Cas9 components into the embryos, including the two gRNAs targeting the GGTA1 and CMAH genes. Although the limited amount of genomic DNA in each blastocyst did not allow for analysis of the large deletions generated during double-strand break repair after Cas9 cleavage, the analysis of targeted gene sequences in a single blastocyst revealed that the rates of double biallelic mutations in ZP-free embryos without centrifugation were significantly lower than those observed after centrifugation. This finding indicates that centrifugation may be pivotal in improving double biallelic mutation rates and reducing mosaicism, possibly by influencing cellular responses (Rajendran and Simons 2005) or other processes associated with introducing CRISPR/Cas9 components.
Conclusions
In conclusion, our findings indicated that pre-electroporation treatments involving centrifugation and ZP removal positively affect the reduction of mosaic mutations. Notably, the effect of centrifugation varied depending on the gRNA used. Therefore, the combined application of centrifugation and ZP removal has emerged as a potential strategy to mitigate mosaicism in the CRISPR system, particularly in porcine embryos.
Data availability
The data used to support the findings of this study have been included in this article.
References
Ablain J, Durand EM, Yang S, Zhou Y, Zon LI (2015) A CRISPR/Cas9 vector system for tissue-specific gene disruption in zebrafish. Dev Cell 32:756–764. https://doi.org/10.1016/j.devcel.2015.01.032
Aida T, Chiyo K, Usami T, Ishikubo H, Imahashi R, Wada Y, Tanaka KF, Sakuma T, Yamamoto T, Tanaka K (2015) Cloning-free CRISPR/Cas system facilitates functional cassette knock-in mice. Genome Biol 16:87. https://doi.org/10.1186/s13059-015-0653-x
Brinkman EK, Chen T, Amendola M, van Steensel B (2014) Easy quantitative assessment of genome editing by sequence trace decomposition. Nucleic Acids Res 42:e168. https://doi.org/10.1093/nar/gku936
Burger A, Lindsay H, Felker A, Hess C, Anders C, Chiavacci E, Zaugg J, Weber LM, Catena R, Jinek M, Robinson MD, Mosimann C (2016) Maximizing mutagenesis with solubilized CRISPR-Cas9 ribonucleoprotein complexes. Development 143:2025–2037. https://doi.org/10.1242/dev.134809
Cohen J, Elsner C, Kort H, Malter H, Massey J, Mayer MP, Wiemer K (1990) Impairment of the hatching process following IVF in the human and improvement of implantation by assisting hatching using micromanipulation. Hum Reprod 5:7–13. https://doi.org/10.1093/oxfordjournals.humrep.a137044
Hendel A, Bak RO, Clark JT, Kennedy AB, Ryan DE, Roy S, Steinfeld I, Lunstad BD, Kaiser RJ, Wilkens AB, Bacchetta R, Tsalenko A, Dellinger D, Bruhn L, Porteus MH (2015) Chemically modified guide RNAs enhance CRISPR-Cas genome editing in human primary cells. Nat Biotechnol 33:985–989. https://doi.org/10.1038/nbt.3290
Hirata M, Tanihara F, Wittayarat M, Hirano T, Nguyen NT, Le QA, Namula Z, Nii M, Otoi T (2019) Genome mutation after introduction of the gene editing by electroporation of Cas9 protein (GEEP) system in matured oocytes and putative zygotes. In Vitro Cell Dev Biol Anim 55:237–242. https://doi.org/10.1007/s11626-019-00338-3
Horii T, Arai Y, Yamazaki M, Morita S, Kimura M, Itoh M, Abe Y, Hatada I (2014) Validation of microinjection methods for generating knockout mice by CRISPR/Cas-mediated genome engineering. Sci Rep 4:4513. https://doi.org/10.1038/srep04513
Kato Y, Nagao Y (2009) Effect of PVP on sperm capacitation status and embryonic development in cattle. Theriogenology 72:624–635. https://doi.org/10.1016/j.theriogenology.2009.04.018
Le QA, Wittayarat M, Namula Z, Lin Q, Takebayashi K, Hirata M, Tanihara F, Do LTK, Otoi T (2022) Multiple gene editing in porcine embryos using a combination of microinjection, electroporation, and transfection methods. Vet World 15:2210–2216. https://doi.org/10.14202/vetworld.2022.2210-2216
Li D, Qiu Z, Shao Y, Chen Y, Guan Y, Liu M, Li Y, Gao N, Wang L, Lu X, Zhao Y, Liu M (2013a) Heritable gene targeting in the mouse and rat using a CRISPR-Cas system. Nat Biotechnol 31:681–683. https://doi.org/10.1038/nbt.2661
Li R, Liu Y, Pedersen HS, Kragh PM, Callesen H (2013b) Development and quality of porcine parthenogenetically activated embryos after removal of zona pellucida. Theriogenology 80:58–64. https://doi.org/10.1016/j.theriogenology.2013.03.009
Li W, Teng F, Li T, Zhou Q (2013c) Simultaneous generation and germline transmission of multiple gene mutations in rat using CRISPR-Cas systems. Nat Biotechnol 31:684–686. https://doi.org/10.1038/nbt.2652
Lin Q, Le QA, Takebayashi K, Hirata M, Tanihara F, Thongkittidilok C, Sawamoto O, Kikuchi T, Otoi T (2022) Viability and developmental potential of porcine blastocysts preserved for short term in a chemically defined medium at ambient temperature. Reprod Domest Anim 57:556–563. https://doi.org/10.1111/rda.14095
Mehravar M, Shirazi A, Nazari M, Banan M (2019) Mosaicism in CRISPR/Cas9-mediated genome editing. Dev Biol 445:156–162. https://doi.org/10.1016/j.ydbio.2018.10.008
Michelmann HW, Rath D, Töpfer-Petersen E, Schwartz P (2007) Structural and functional events on the porcine zona pellucida during maturation, fertilization and embryonic development: a scanning electron microscopy analysis. Reprod Domest Anim 42:594–602. https://doi.org/10.1111/j.1439-0531.2006.00829.x
Namula Z, Hirata M, Le QA, Lin Q, Takebayashi K, Yoshimura N, Tanihara F, Thongkittidilok C, Otoi T (2022) Zona pellucida treatment before CRISPR/Cas9-mediated genome editing of porcine zygotes. Vet Med Sci 8:164–169. https://doi.org/10.1002/vms3.659
Namula Z, Wittayarat M, Hirata M, Hirano T, Nguyen NT, Le QA, Fahrudin M, Tanihara F, Otoi T (2019) Genome mutation after the introduction of the gene editing by electroporation of Cas9 protein (GEEP) system into bovine putative zygotes. In Vitro Cell Dev Biol Anim 55:598–603. https://doi.org/10.1007/s11626-019-00385-w
Niu Y, Shen B, Cui Y, Chen Y, Wang J, Wang L, Kang Y, Zhao X, Si W, Li W, Xiang AP, Zhou J, Guo X, Bi Y, Si C, Hu B, Dong G, Wang H, Zhou Z, Li T, Tan T, Pu X, Wang F, Ji S, Zhou Q, Huang X, Ji W, Sha J (2014) Generation of gene-modified cynomolgus monkey via Cas9/RNA-mediated gene targeting in one-cell embryos. Cell 156:836–843. https://doi.org/10.1016/j.cell.2014.01.027
Rajendran L, Simons K (2005) Lipid rafts and membrane dynamics. J Cell Sci 118:1099–1102. https://doi.org/10.1242/jcs.01681
Shen B, Zhang J, Wu H, Wang J, Ma K, Li Z, Zhang X, Zhang P, Huang X (2013) Generation of gene-modified mice via Cas9/RNA-mediated gene targeting. Cell Res 23:720–723. https://doi.org/10.1038/cr.2013.46
Tanihara F, Takemoto T, Kitagawa E, Rao S, Do LT, Onishi A, Yamashita Y, Kosugi C, Suzuki H, Sembon S, Suzuki S, Nakai M, Hashimoto M, Yasue A, Matsuhisa M, Noji S, Fujimura T, Fuchimoto D, Otoi T (2016) Somatic cell reprogramming-free generation of genetically modified pigs. Sci Adv 2:e1600803. https://doi.org/10.1126/sciadv.1600803
Taylor TH, Gitlin SA, Patrick JL, Crain JL, Wilson JM, Griffin DK (2014) The origin, mechanisms, incidence and clinical consequences of chromosomal mosaicism in humans. Hum Reprod Update 20:571–581. https://doi.org/10.1093/humupd/dmu016
Tominaga K, Shimizu M, Ooyama S, Izaike Y (2000) Effect of lipid polarization by centrifugation at different developmental stages on post-thaw survival of bovine in vitro produced 16-cell embryos. Theriogenology 53:1669–1680. https://doi.org/10.1016/S0093-691X(00)00306-X
Vajta G, Rienzi L, Bavister BD (2010) Zona-free embryo culture: is it a viable option to improve pregnancy rates? Reprod Biomed Online 21:17–25. https://doi.org/10.1016/j.rbmo.2010.03.014
Wang CW (2016) Lipid droplets, lipophagy, and beyond. Biochim Biophys Acta 1861:793–805. https://doi.org/10.1016/j.bbalip.2015.12.010
Wang H, Yang H, Shivalila CS, Dawlaty MM, Cheng AW, Zhang F, Jaenisch R (2013) One-step generation of mice carrying mutations in multiple genes by CRISPR/Cas-mediated genome engineering. Cell 153:910–918. https://doi.org/10.1016/j.cell.2013.04.025
Wang J, Torres IM, Shang M, Al-Armanazi J, Dilawar H, Hettiarachchi DU, Paladines-Parrales A, Chambers B, Pottle K, Soman M, Su B, Dunham RA (2024) One-step knock-in of two antimicrobial peptide transgenes at multiple loci of catfish by CRISPR/Cas9-mediated multiplex genome engineering. Int J Biol Macromol:129384. https://doi.org/10.1016/j.ijbiomac.2024.129384
Wassarman PM, Litscher ES (2008) Mammalian fertilization: the egg’s multifunctional zona pellucida. Int J Dev Biol 52:665–676. https://doi.org/10.1387/ijdb.072524pw
Wirtu G, McGill J, Crawford L, Reddy G, Bergen W, Simon L (2013) Targeting lipid metabolism to improve oocyte cryopreservation (OCP) in domestic animals. Trans Clin Bio 1:15–20
Yang H, Wang H, Shivalila CS, Cheng AW, Shi L, Jaenisch R (2013) One-step generation of mice carrying reporter and conditional alleles by CRISPR/Cas-mediated genome engineering. Cell 154:1370–1379. https://doi.org/10.1016/j.cell.2013.08.022
Acknowledgements
We thank Nippon Food Packer and K. K. Shikoku (Tokushima, Japan) for supplying the pig ovaries.
Funding
This study was partly supported by the JSPS KAKENHI (grant numbers JP22H02499 and JP22K19896). We also received financial support from the Uzushio Program of Tokushima University.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, B., Wittayarat, M., Takebayashi, K. et al. Effects of centrifugation treatment before electroporation on gene editing in pig embryos. In Vitro Cell.Dev.Biol.-Animal 60, 732–739 (2024). https://doi.org/10.1007/s11626-024-00926-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11626-024-00926-y