Abstract
En1, Nr2f1, Gpc4, Sfrp2, Shox2, Tbx15 and Thbd are among the genes involved in development process of an organism in a number of tissues, in particular adipose tissue. Considering the involvement of isobutyl-methyl-xanthine (IBMX), indomethacin, dexamethasone (DEX), triiodothyronine (T3), and insulin in adipocyte differentiation, we propose that these differentiation-inducing agents may regulate differentiation in brown adipose tissue through a developmental process. Stromavascular cells isolated from interscapular brown fat of mice were cultured in DMEM-LG medium. Proliferating brown preadipocytes were differentiated in the presence of IBMX, indomethacin, DEX, T3 and insulin. Pref1 (marker of proliferation stage) and uncoupling protein 1 (UCP1, marker for differentiation stage) were upregulated during proliferation and differentiation, respectively. Relative expression of Pref1, UCP1 and developmental genes was determined in different stages of adipogenesis. En1, Gpc4, Nr2f1, Sfrp2 and Shox2 were upregulated during differentiation. Differentiation of preadipocytes in the absence of IBMX, indomethacin, and DEX resulted in drastic reduction in fat accumulation in differentiated adipocytes with simultaneous decrease in En1, Gpc4, Nr2f1, Sfrp2, Shox2 and Tbx15 gene expression. T3 upregulated the expression of En1, Gpc4, Sfrp2 and Tbx15 genes during differentiation and downregulated Shox2 expression as compared to proliferated state. Insulin upregulated the expression of Shox2.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Brown as well as white adipose tissues (WAT) store fat during energy excess and release triglycerides under low-energy condition. Brown adipose tissue (BAT) also generates heat by oxidizing lipids during non-shivering thermogenesis where uncoupling protein 1 (UCP1), unique protein present in inner mitochondrial membrane, prohibits formation of ATP (Cannon and Nedergaard 2010). As opposed to WAT, BAT is not considered as unhealthy fat. Mesenchymal stem cells present in adipose tissue and committed to become preadipocytes are first converted to preadipocytes which finally differentiate into mature adipocytes. This occurs both in vitro and in vivo (Dani et al. 1997). Proliferation and differentiation stages are morphologically distinct stages which appear during adipogenesis. The former stage results in fibroblast-like cells while the latter is identified by accumulation of fat. Both the stages can be easily ascertained microscopically. The marker Pref1 predominantly appears when preadipocytes are proliferating while UCP1 is abundantly noted when brown preadipocytes are differentiated into mature adipocytes (Villena et al. 2002). The fat synthesis in mature cells is regulated by transcription factor PPARγ, its co-activator PGC1alpha and other molecules, viz. CCAAT/enhancer-binding proteins, specifically C/EBPα/β/δ, sirtuin 1 and early B cell factor 2 (Tontonoz et al. 1994; Wu et al. 1999; Qiang et al. 2012; Rajakumari et al. 2013).
En1 (mouse homolog of a drosophila patterning gene), Gpc4 (gene involved in cell division and growth regulation), Nr2f1 (gene involved in organogenesis), Sfrp2 (modulator of Wnt signaling), Shox2 (transcription factor expressed during embryonic development), Tbx15 (transcription factor involved in craniofacial and limb development) and Thbd (a surface glycoprotein of endothelial and placental cells) are differentially expressed in subcutaneous and intra-abdominal white fat depots (Hanks et al. 1998; de Cat and David 2001; Singh et al. 2005; Gesta et al. 2006; Nakajima et al. 2009; Schneider et al. 2011; Li et al. 2012; Bosch et al. 2014). Ever since, developmental genes are reported in adipose tissue; precise role of developmental genes is still difficult to understand (Cowherd et al. 1997; Cantile et al. 2002; Gesta et al. 2006; Gesta et al. 2007; Yamamoto et al. 2009; Waldén et al. 2012). The expression of developmental genes varies between white and brown adipose tissues (Gesta et al. 2006; Gesta et al. 2007; Yamamoto et al. 2009; Waldén et al. 2012). Although preadipocytes and adipocytes constitutes more than 80% of adipose tissue, the presence of other cells such as stem cells, fibroblasts, smooth muscle cells, endothelial cells, macrophages, and pericytes can also account for expression of developmental genes (Katz 2002). In vitro culture system allows preadipocytes and differentiated adipocytes in purified form and therefore, expression of developmental genes at these stages can be studied to understand the role of developmental genes in adipogenesis. The adipogenesis is regulated by isobutyl-methyl-xanthine (IBMX), indomethacin, dexamethasone (DEX), triiodothyronine (T3) and insulin. However, whether these differentiation-inducing agents influence expression of developmental genes is not known. In this paper, the effect of these agents on expression of En1, Gpc4, Nr2f1, Sfrp2, Shox2, Tbx15 and Thbd is reported.
Materials and Methods
Subjects.
Swiss albino mice in random fed groups were allowed ad libitum access to water and food and were maintained in a temperature-controlled room (25 ± 3°C) for 12 h light–dark cycle at small animal house facilities of the National Dairy Research Institute, Karnal. Experiments were performed on 4–6-wk-old male mice in compliance with the Institute Animal Ethics Committee (IAEC) guidelines.
Isolation of stromavascular fraction and proliferation of preadipocytes.
BAT (200–300 mg) was collected aseptically from the interscapular region of mice. The methodology described by Fasshauer et al. (2000) was used for isolation of stromavascular fraction (SVF). BAT was excised from three animals and treated with a digestion medium containing 2 ml 0.2% collagenase (type II) in HEPES buffer (100 mM HEPES, 123 mM NaCl, 5 mM KCl, 1.3 mM CaCl2, 5 mM glucose, 1.5% BSA, pH 7.4) for 10–15 min at 37°C on an orbital shaker (90 rpm). The digested contents were filtered through a 100-μm nylon filter and centrifuged at 450×g for 10 min. The pellet comprising SVF was suspended in Dulbecco’s modified Eagle’s medium-low glucose (DMEM-LG) proliferation medium containing 15% fetal bovine serum (FBS), 10 mM HEPES, 50 IU penicillin/ml, 50 μg streptomycin/ml, 3.7 g/l NaHCO3, 3 nM insulin and 15 μM ascorbic acid and then plated in a 25cm2 tissue culture flask and cultured in 5% CO2 atmosphere at 37°C. After 24 h of incubation, non-adherent cells were removed by repeatedly washing the cells with proliferation medium without FBS. Adherent cells were further maintained in DMEM-LG proliferation medium, and spent medium was replenished with fresh medium until the cells reached 70–80% confluency.
Adipocyte differentiation.
Differentiation of brown preadipocytes was achieved as per method described by Klein et al. (1999) with some modifications. Proliferative preadipocytes were cultured in an induction medium (DMEM-LG medium (pH 7.4) containing 10% FBS, 500 μM IBMX, 125 μM indomethacin, 2 μg/ml DEX, 18 nM insulin and 1 nM T3) for 2 d and then cultured in a differentiation medium (induction medium devoid of IBMX, indomethacin and DEX) for 5 d. The medium was changed in every alternate day.
Differentiation in absence of induction medium.
Proliferative preadipocytes were also cultured in a differentiation medium for 7 d without pre-culturing in an induction medium.
Effect of T3.
Effect of three different levels of T3, viz. 1, 2 and 4 nM on expression pattern of developmental genes during differentiation of brown adipocytes was studied, and the scheme of culture is shown in Table 1 (Cousin et al. 1992). SVF was isolated from three Swiss albino mice and were cultured in a proliferation medium containing 15% charcoal–dextran-treated FBS and 15 μM ascorbic acid for 7 d to induce preadipocytes in proliferation stage. At day 8, the spent medium was replaced with DMEM-LG medium containing 10% charcoal–dextran-treated FBS (devoid of IBMX, indomethacin, DEX, T3 and insulin) for 48 h. Further, cells were treated with different concentration of T3 for 2 d when cultured in DMEM-LG medium containing 10% charcoal–dextran-treated FBS and 1 nM/2 nM/4 nM concentration of T3.
Effect of insulin concentration.
Effect of insulin was studied similar to T3 except insulin at three different levels, viz. 0.5, 3 and 18 nM was added in cultured medium at day 10 instead of T3 (Table 1).
RNA extraction and gene expression analysis.
Using TRI reagent (Sigma Chemicals Co., St. Louis, MO) total RNA from SVF, proliferated adipocytes, and differentiated adipocytes was isolated and quantified by NanoPhotometer (IMPLEN, Munich, Germany). RNA was treated with DNase (Fermentas Life Sciences, Ontario, Canada). Subsequently, RNA (1 μg) was reverse transcribed with RevertAid™ first-strand complementary DNA (cDNA) synthesis kit (Fermentas Life Sciences, Canada). Quantitative PCR was done on Mx3005 apparatus (Stratagene, La Jolla, CA) using SYBR Green, double-stranded DNA-specific fluorescent dye (Roche Diagnostics, Mannheim, Germany) using comparative quantitation method and β-actin normalization. cDNA samples were prepared in three biological replicates from SVF and proliferated and differentiated adipocytes. Primers of genes Pref1, UCP1 and β-actin were designed using the web-based software PRIMER-3. Mouse sequences for genes Pref1 and UCP1 were obtained from NCBI (www.ncbi.nlm.nih.gov) and β-actin from ENSEMBL genome (www.ensembl.org) browser. The primers for genes En1, Gpc4, Nr2f1, Sfrp2, Shox2, Tbx15 and Thbd were used as reported by Gesta et al. (2006). Details of primers are given in Table 2.
Critical threshold (CT) of SVF value was considered as a calibrator except in case of hormonal studies where CT value in the absence of hormone was used as a calibrator. Relative mRNA fold change was calculated in comparison to β-actin. The data were analyzed using 2−ΔΔCT method (Livak and Schmittgen 2001).
Statistical analysis.
The gene expression was studied in three biological replicates, and each value was determined in triplicates. Comparison between groups was performed using one-way analysis of variance (ANOVA) followed by Tukey: Compare all pairs of columns test. P value of <0.05 was considered as significant. Statistical analysis was performed using GraphPad Prism version 5.01 (GraphPad Software Inc., San Diego, CA).
Results
Adipogenesis.
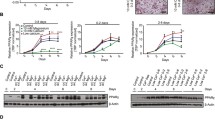
When stromavascular cells were plated in DMEM-LG proliferation medium, fibroblast-like cells were visible after 24 h of plating. The fibroblast-like brown fat cells multiplied rapidly and at day 6, 70–80% confluency is usually reached. At day 7, proliferated cells (70–80% confluency) were treated with induction medium containing IBMX, DEX, indomethacin, T3 and insulin for 2 d. At day 9, distinct changes in cell morphology were noted. Cells acquired almost a round shape. The cells were then cultured in differentiation medium containing T3 and insulin for 6 d. At day 10, cytoplasm of these cells became enlarged and cells contained numerous small-sized lipid droplets. At day 14 of cell culture, lipid incorporation increased and small-sized multilocular lipid droplets were observed in differentiated cells. Differentiated cells on staining with Oil Red O: [1-{2, 5-dimethyl-4-(2-5-dimethylphenyl) phenyldiazene}azonapthalen-2-ol] dye, appeared red (Fig. 1A ) (Peng et al. 2013).
Proliferated and differentiated brown adipocytes and expression of stage-specific marker and developmental genes (A); brown adipocytes at different days of culture (B); brown adipocytes cultured without induction medium at day 14 (C); expression pattern of marker genes and developmental genes. SVF stroma vascular fraction, P proliferation stage, D differentiation stage, w/o IM without induction medium. Bars represent relative mRNA expression fold change and error bars represent standard error of the mean. The number of technical and biological replicates was three each. *P < 0.05; **P < 0.01; ***P < 0.001.
Pref1 and UCP1 levels changed as expected, according to their specificity in proliferation and differentiation stage, respectively (Fig. 1C ). The expression of seven developmental genes, viz. En1, Gpc4, Nr2f1, Sfrp2, Shox2, Tbx15 and Thbd in SVF and proliferated and differentiated brown adipocytes indicated (Table 3) that there was about 16.4-fold, 2.5-fold, 18-fold, 3.1-fold and 67.9-fold higher expression of En1, Gpc4, Nr2f1, Sfrp2 and Shox2 in differentiated over proliferated adipocytes (Fig. 1C ), respectively. However, expression of Tbx15 and Thbd was not changed significantly on differentiation of proliferated cells. Also, the expression of UCP1, Shox2 and Thbd decreased in proliferated cells over SVF while that of Pref1 and Gpc4 increased.
Effect of induction medium on differentiation.
Cells grown in the absence of induction medium components do not fully differentiate, and few tiny droplets of fat were visible at day 14 of culture (Fig. 1B ). Expression of UCP1, En1, Gpc4, Nr2f1, Sfrp2, Shox2 and Tbx15 were downregulated approximately 62, 116, 81, 38, 42, 294 and 10 times (Fig. 1C , Table 3) respectively compared to differentiation achieved in the presence of induction medium components. However, Thbd expression increases three times (Fig. 1C ).
Effect of T3 on differentiation.
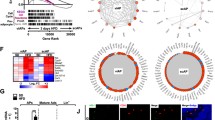
Cells grown in the presence of T3 (at all the concentrations) but in the absence of insulin had swollen cytoplasm in comparison to cell grown in the absence of T3. Lipid droplets were absent both in the presence and absence of T3 in medium (Fig. 2A ). Significantly higher expression of UCP1 was noted at 2 and 4 nM T3 (Fig. 2B ). Expression of Pref1 was not altered at all the three levels of T3 (Fig. 2B ).
Effect of T3 on differentiation of brown adipocytes and expression of stage-specific marker and developmental genes. Proliferative adipocytes were cultured in the absence (C control) or in the presence of T3 at indicated concentrations. (A) Cells at day 12; (B) expression pattern of marker genes and developmental genes. Bars represent relative mRNA expression fold change and error bars represent standard error of the mean. The number of technical and biological replicates was three each. *P < 0.05; **P < 0.01; ***P < 0.001.
Effect of different levels of T3 on expression En1, Gpc4, Nr2f1, Sfrp2, Shox2, Tbx15 and Thbd is shown in Fig. 2 and Table 4. The expression of En1, Gpc4, Nr2f1, Sfrp2, Shox2 and Tbx15 was dependent on concentration of T3. Gpc4, Sfrp2 and Tbx15 were maximally produced at 2 nM T3 and difference in expression was statistically significant when compared with control. At other concentrations of T3 (1 and 4 nM), the expression was lower. The En1 expression was significantly higher at 1 nM T3 in comparison to the control group. At 2 or 4 nM T3, there was no significance difference in expression of En1 between the treatment group and control group. Although the expression of Nr2f1 was significantly higher (P < 0.05) at 2 nM T3 in comparison to the control group, the fold difference in expression was less than 2 and therefore, the results should be taken as indicative in nature. On the other hand, expression of Shox2 was significantly lower at 4 nM in comparison to the control group. There was no effect on the expression of Thbd at all the three levels of T3. At 1 nM T3, the expression of En1 and Sfrp2 enhanced while other genes remained unaffected.
Effect of insulin on differentiation.
Cells grown in the presence of insulin (at all the concentrations) but in the absence of T3 in the differentiation medium had a distinct morphology. Differentiated cells were spindle in shape with a swollen cytoplasm and little lipid droplets (Fig. 3A ). Insulin did not alter expression of En1, Nr2f1, Sfrp2, Tbx15 and Thbd (Fig. 3B ; Table 4). Expression of Shox2 was significantly higher at 0.5 nM, whereas Gpc4 decreased at 3 nM (Fig. 3B ). At all the concentrations of insulin, Pref1 was expressed at a reduced level. UCP1 expressed was enhanced only at 0.5 nM insulin (Fig. 3B ).
Effect of insulin on differentiation of brown adipocytes and expression of stage-specific marker and developmental genes. Proliferative adipocytes were cultured in the absence (C control) or in the presence of insulin at indicated concentrations. (A) Cells at day 12; (B) expression pattern of marker genes and developmental genes. Bars represent relative mRNA expression fold change and error bars represent standard error of the mean. The number of technical and biological replicates was three each. *P < 0.05; **P < 0.01; ***P < 0.001.
Discussion
En1, Gpc4, Nr2f1, Sfrp2, Shox2, Tbx15, Thbd and a number of Hox genes including HoxA1, HoxA5, HoxC4, HoxC8, HoxC9 and HoxA10 are present in adipose tissue or preadipocytes and mature adipocytes derived from adipose tissue (Cowherd et al. 1997; Cantile et al. 2002; Gesta et al. 2006; Gesta et al. 2007; Yamamoto et al. 2009; Waldén et al. 2012). Adipose tissue comprises stem cells, preadipocytes and mature adipocytes and these cells types accounts for >80% (Katz 2002). The other cells such as macrophages, pericytes, fibroblast and endothelial cells are also present in adipose tissue and the possibility exists that these can also account of developmental genes identified so far (Katz 2002). In vitro culture system provides purified preparation of preadipocytes which on appropriate culture condition can be differentiated. Large fat globule(s) are present in differentiated white adipocyte, whereas several fat globules of small size are observed in differentiated brown adipocytes (Seale et al. 2009). Our studies on differentiation of brown adipocytes indicate that either insulin or T3 alone in the absence of differentiation-inducing agents (IBMX, indomethacin and DEX) cannot lead to accumulation of fat in brown adipocytes in a significant way. In the presence of T3 alone, even proliferative preadipocytes were not healthy.
The involvement of developmental genes in adipogenesis is further supported from the present study. Treating preadipocytes with induction medium results in the upregulation of En1, Gpc4, Nr2f1, Sfrp2 and Shox2, whereas these genes are downregulated in the absence of an induction medium. Further, significant lipid accumulation is observed only when preadipocytes are pretreated with a cocktail of IBMX, indomethacin and DEX during differentiation. The results therefore support that role of IBMX, indomethacin and DEX in lipid accumulation is linked with regulation of developmental genes. Tbx15 and Thbd expressions remains unaltered during differentiation in the presence of induction medium suggesting that these genes are not of much consequence in differentiation (Fig. 1C ).
Effect of T3 on UCP1 (marker for brown adipocytes differentiation) was dose dependent and therefore, T3 affects differentiation. However, except Thbd, the effects of T3 on other developmental genes were dose specific (Fig. 2). The results presented in the paper suggest that Nr2f1, Sfrp2, Tbx15 and Thbd are not regulated through insulin (Fig. 3).
References
Bosch DG, Boonstra FN, Gonzaga-Jauregui C, Xu M, de Ligt J, Jhangiani S et al (2014) NR2F1 mutations cause optic atrophy with intellectual disability. Am J Hum Genet 94:303–309
Cannon B, Nedergaard J (2010) Metabolic consequences of the presence or absence of the thermogenic capacity of brown adipose tissue in mice (and probably in humans). Int J Obesity 34:S7–S16
Cantile M, Procino A, D’Armiento M, Cindolo L, Cillo C (2002) HOX gene network is involved in the transcriptional regulation of in vivo human adipogenesis. J Cell Physiol 194:225–236
Cousin B, Cinti S, Morroni M (1992) Occurrence of brown adipocytes in rat white adipose tissue: molecular and morphological characterization. J Cell Sci 103:931–942
Cowherd RM, Lyle RE, Miller CP, Mcgehee RE (1997) Developmental profile of homeobox gene expression during 3T3-L1 adipogenesis. Biochem Biophys Res Commun 237:470–475
Dani C, Smith AG, Dessolin S, Leroy P, Staccini L, Villageois P et al (1997) Differentiation of embryonic stem cells into adipocytes in vitro. J Cell Sci 110:1279–1285
De Cat B, David G (2001) Developmental roles of the glypicans. Semin Cell Dev Biol 12:117–125
Fasshauer M, Klein J, Ueki K, Kriauciunas KM, Benito M, White MF et al (2000) Essential role of insulin receptor substrate-2 in insulin stimulation of Glut4 translocation and glucose uptake in brown adipocytes. J Biol Chem 275:25494–25501
Gesta S, Bluher M, Yamamoto Y, Norris AW, Berndt J, Kralisch S et al (2006) Evidences for a role of developmental genes in the origin of obesity and body fat distribution. Proc Natl Acad Sci U S A 103:6676–6681
Gesta S, Tseng YH, Kahn CR (2007) Developmental origin of fat: tracking obesity to its source. Cell 131:242–256
Hanks MC, Loomis CA, Harris E, Tong CX, Anson-Cartwright L, Auerbach A et al (1998) Drosophila engrailed can substitute for mouse Engrailed1 function in mid-hindbrain, but not limb development. Development 125:4521–4530
Katz AJ (2002) Mesenchymal cell culture: adipose tissue. In: Methods of tissue engineering; Atala, A., Lanza, R. eds. California, pp 277–286
Klein J, Fasshauer M, Ito M, Lowell BB, Benito M, Kahn CR (1999) β3-adrenergic stimulation differentially inhibits insulin signaling and decreases insulin induced glucose uptake in brown adipocytes. J Biol Chem 274:34795–34802
Li YH, Kuo CH, Shi GY, Wu HL (2012) The role of thrombomodulin lectin-like domain in inflammation. J Biomed Sci 19:34
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(−delta delta C (T)). method. Methods 25:402–408
Nakajima H, Ito M, Morikawa Y, Komori T, Fukuchi Y, Shibata F et al (2009) Wnt modulators, SFRP-1, and SFRP-2 are expressed in osteoblasts and differentially regulate hematopoietic stem cells. Biochem Bioph Res Co 390:65–70
Peng Y, Xiang H, Chen C, Zheng R, Chai J, Peng J et al (2013) MiR-224 impairs adipocyte early differentiation and regulates fatty acid metabolism. Int J Biochem Cell Biol 45:1585–1593
Qiang L, Wang L, Kon N, Zhao W, Lee S, Zhang Y et al (2012) Brown remodeling of white adipose tissue by SirT1-dependent deacetylation of Pparγ. Cell 150:620–632
Rajakumari S, Wu J, Ishibashi J, Lim HW, Giang AH, Won KJ (2013) EBF2 determines and maintains brown adipocyte identity. Cell Metab 17:562–574
Schneider KU, Dietrich D, Fleischhacker M, Leschber G, Merk J, Schäper F et al (2011) Correlation of SHOX2 gene amplification and DNA methylation in lung cancer tumors. BMC Cancer 11:102
Seale P, Kajimura S, Spiegelman BM (2009) Transcriptional control of brown adipocyte development and physiological function of mice and men. Genes Dev 23:788–797
Singh M, Petry M, Haenig B, Lescher B, Leitges M, Kispert A (2005) The T-box transcription factor Tbx15 is required for skeletal development. Mech Develop 122:131–144
Tontonoz P, Hu E, Spiegelman BM (1994) Stimulation of adipogenesis in fibroblasts by PPAR γ 2, a lipid-activated transcription factor. Cell 79:1147–1156
Villena JA, Kim KH, Sul HS (2002) Pref-1 and ADSF/resistin: two secreted factors inhibiting adipose tissue development. Horm Metab Res 34:664–670
Waldén TB, Hansen IR, Timmons JA (2012) Recruited vs. non recruited molecular signatures of brown, “brite” and white adipose tissues. Am J Physiol – Endocrinol and Metab 302:E19–E31
Wu Z, Puigserver P, Spiegelman BM (1999) Transcriptional activation of adipogenesis. Curr Opin Cell Biol 11:689–694
Yamamoto Y, Gesta S, Lee KY, Tran TT, Saadatirad P, Kahn CR (2009) Adipose tissue possess unique developmental gene signature. Obesity 8:872–878
Acknowledgments
The authors would like to acknowledge the National Agricultural Innovation Project and National Initiative on Climate Resilient Agriculture for the financial support.
Authors’ contributions
Y.S.R. developed the concept. S.S. and A.K.B. performed the experiments, and collected and analyzed the data. Y.S.R., S.S., and A.K.B. wrote the manuscript. R.S. and S.G. gave the conceptual advice.
Conflict of interest
The authors declare that they have no competing interests.
Author information
Authors and Affiliations
Corresponding author
Additional information
Editor: Tetsu Okamoto
Rights and permissions
About this article
Cite this article
Singh, S., Rajput, Y.S., Barui, A.K. et al. Expression of developmental genes in brown fat cells grown in vitro is linked with lipid accumulation. In Vitro Cell.Dev.Biol.-Animal 51, 1003–1011 (2015). https://doi.org/10.1007/s11626-015-9930-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11626-015-9930-y