Abstract
Extensive and dynamic chromatin remodeling occurs after fertilization, including DNA methylation and histone modifications. These changes underlie the transition from gametic to embryonic chromatin and are thought to facilitate early embryonic development. Histone H3 lysine 4 methylation (H3K4me) is an important epigenetic mechanism that associates with gene-specific activation and functions in development. However, dynamic regulation of H3K4me during early embryonic development remains unclear. Herein, the authors examined the dynamic changes of H3K4me and its key regulators (Ash1l, Ash2l, Kmt2a, Kmt2b, Kmt2c, Setd1a, Setd7, Kdm1a, Kdm1b, Kdm5a, Kdm5b, Kdm5c, and Kdm5d) in mouse oocytes and preimplantation embryos. An increase in levels of H3K4me2 and me3 was observed at the one- to two-cell stages (P < 0.05), corresponding to the period of embryonic genome activation (EGA). Subsequently, the H3K4me2 level dramatically decreased at the four-cell stage and remained at low level until the blastocyst stage (P < 0.05), whereas the H3K4me3 level transiently decreased in the four-cell embryos but steadily increased to the peak in the blastocysts (P < 0.05). The high level of H3K4me2 during the EGA was coinciding with a peak expression of its methyltransferase, ASH2L, which may stabilize this methylation level during this period. Correspondingly, a concomitant decrease in levels of its demethylases, KDM5B and KDM1A, was observed. H3K4me3 was correlated to the expression of its methyltransferase (KMT2B) and demethylase (KDM5A). Thus, these enzymes may function for the EGA and the first lineage segregation in preimplantation mouse embryos.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Epigenetic modifications, such as DNA methylation, histone methylation, and acetylation, regulate the patterns of gene expression through the changes of chromatin structure and DNA accessibility (Beisel and Paro 2011). The methylation of lysine 4 of histone H3 (H3K4me) is one of these modifications, which is involved in gene activation (Eissenberg and Shilatifard 2010). Specifically, H3K4 di- and trimethylation (H3K4me2 and me3) residues mark the transcription start sites of actively transcribed genes (Bernstein et al. 2006; Barski et al. 2007; Mikkelsen et al. 2007), while high level of H3K4 monomethylation (H3K4me1) is associated with enhancer sequences (Heintzman et al. 2007). H3K4me2/3 has also been shown to regulate the expression of pluripotency genes (Pan et al. 2007; Wang et al. 2009a, b) and Hox genes (VerMilyea et al. 2009) in mouse preimplantation embryos and embryonic stem cells.
Epigenetic regulation of gene expression is mediated primarily by SET-domain- and JmjC-domain-containing proteins. The SET/mixed lineage leukemia (MLL) family of SET proteins is primarily responsible for the generation of H3K4me1/2/3 to induce gene activation, and it has seven major members as follows: lysine (K)-specific methyltransferase 2A (KMT2A)/MLL1, KMT2B/MLL2, KMT2C/MLL3, ASH1L, ASH2L, SETD1A, and SETD7 (Dou et al. 2006; Black et al. 2012). In contrast, the lysine (K)-specific demethylase 5 (KDM5) family of JmjC proteins (KDM5A/RBP2, KDM5B/PLU1, KDM5C/SMCX, and KDM5D/SMCY) has recently been identified as demethylases of H3K4me2/3 causing gene repression (Klose et al. 2006). In addition, KDM1A/LSD1 and KDM1B/LSD2 demethylate H3K4me1/2 and result in gene repression (Shi et al. 2004; Fang et al. 2010).
The families of MLL methyltransferases and KDM5 demethylases are essential for normal development in mammals. Loss of either Kmt2a or Kmt2b in oocytes results in anovulation and oocyte death (Terranova et al. 2006; Andreu-Vieyra et al. 2010). Knockout of Ash2l or Kdm5b causes early embryonic lethality (Stoller et al. 2010; Albert et al. 2013). In the absence of Kdm1b, germ cells do not undergo de novo DNA methylation during oogenesis (Ciccone et al. 2009), while Kdm1a deficiency causes embryonic lethality due to a progressive loss of genomic DNA methylation and therefore lack of cell differentiation (Wang et al. 2009a, b). Inhibition of KDM1A also results in early embryo arrest in our previous studies (Shao et al. 2008a). Recently, Kdm1a mutants display germline sterility and defects in ovary development (Di Stefano et al. 2007; Katz et al. 2009; Eliazer et al. 2011). More recently, antagonistic functions of Kdm5a and Ash2l regulate lifespan and control a set of genes involved in lifespan determination (Greer et al. 2010). These studies suggest that the dynamic regulation of the H3K4me state continues throughout early development. However, the embryonic expression of these epigenetic regulators has not been well investigated, especially in early developmental stages.
Here, we investigated the levels of H3K4me in preimplantation mouse embryos, and performed quantitative real-time polymerase chain reaction (PCR) analyses of the messenger RNA (mRNA) expression of Ash1l, Ash2l, Kmt2a, Kmt2b, Kmt2c, Setd1a, Setd7, Kdm1a, Kdm1b, Kdm5a, Kdm5b, Kdm5c, Kdm5d, H2afz, and Gapdh (Table 1). Furthermore, immunofluorescence was performed to analyze the protein expression of these regulators to find out their association with the levels of H3K4me.
Materials and Methods
Animals.
Kunming (KM) white female mice (3–4 wk old) and imprinting control region (ICR) male mice (12–16 wk old) were obtained from the Experimental Animal Breeding Center of Jiangsu University. All animals were maintained under controlled temperature (25 ± 2°C) and lighting (light from 0700 to 1900 h) for at least 1 wk prior to use. Animal care and experimental procedures were conducted in accordance with the Animal Research Committee Guidelines of Jiangsu University.
Reagents and media.
All reagents and media used in this study were from Sigma-Aldrich Inc. (St. Louis, MO) unless noted otherwise. M2 (M7167; Sigma-Aldrich) and KSOM (MR-020P-5F; Millipore, Billerica, MA) media were prepared for the handling and culture of mouse oocytes and embryos, respectively.
Oocyte collection.
To retrieve fully grown oocytes arrested at prophase I of meiosis, KM white mice were treated with 10 IU of pregnant mare serum gonadotrophin (PMSG; Ningbo Second Hormone Factory, Zhejiang, China). The ovaries were removed from the mice 48 h after PMSG treatment and transferred to M2 medium. The ovarian follicles were punctured with a 30-gauge needle to release the oocytes in the cumulus oophorus. The cumulus cells surrounding the oocytes were removed by gentle pipetting through a narrow glass pipette in the M2 medium. Oocytes showing germinal vesicles (GVs) were collected.
Mature oocytes were retrieved by treatment with 10 IU of PMSG followed by 10 IU of human chorionic gonadotropin (hCG; Ningbo Second Hormone Factory) 48 h later. Cumulus oocyte complexes (COCs) were collected from the ampulla of oviducts 15 h after hCG injection, placed in M2 medium, and treated with 0.1% hyaluronidase until the cumulus cells dispersion.
In vivo-derived embryo collection.
Female KM white mice were treated with 10 IU of PMSG followed by 10 IU of hCG 48 h later. Immediately after the hCG injection, the mice were mated overnight with ICR male mice. Embryos were harvested from pregnant mice at 18, 42, 50, 62, 76, and 90 h post-hCG injection to obtain the one-cell, two-cell, four-cell, eight-cell, morulae, and blastocysts stages, respectively. One-cell, two-cell, four-cell, eight-cell, and morulae stages were flushed from oviducts with prewarmed M2 medium and cultured in KSOM medium. Blastocysts were retrieved by flushing uterine horns. The recovered embryos were placed in RNA extraction buffer and stored at −80°C.
mRNA extraction and complementary DNA (cDNA) synthesis.
Polyadenylated mRNA was extracted from pools (30) of embryos of the respective developmental stage using PicoPure RNA Isolation Kit (Arcturus, Molecular Devices, Mountain View, CA). Polyadenylated mRNA was extracted from pools of embryos collected on a single day. Each mRNA extraction was replicated three times for each experiment. Oocytes and embryos were washed three times in PVA-PBS and then lysed in 20 μl of extraction buffer at 42°C for 30 min and stored at −80°C until mRNA isolation. The mRNAs were reverse transcribed following the manufacturer’s instructions using PrimeScript RT Reagent Kit (Takara, Shiga, Japan).
Real-time PCR.
The quantification of all gene transcripts was carried out in three replicate by quantitative real-time PCR on the Bio-Rad CFX96 system using SsoFast EvaGreen Supermix (Bio-Rad, Hercules, CA). The primer sequences for each gene are listed in Table 2. Primers for Kdm5a, Kdm5b, Kdm5c, Kdm1a, Kdm1b, Ash1l, and Kmt2a were designed using Primer Premier 5.0 based on available sequences from Database of GenBank (NCBI). Primers for Kdm5d, Ash2l, Kmt2b, Kmt2c, Setd1a, Setd7, H2afz, and Gapdh were referred to previous literatures (VerMilyea et al. 2009; Andreu-Vieyra et al. 2010; Wang et al. 2010; Albert et al. 2013). The reaction mixture of the total 20 μl volume consisted of 10 μl of 1× SsoFast EvaGreen Supermix, 1 μl of each primer (0.5 μM), 6 μl of RNase/DNase-free water, and 2 μl of cDNA template. The PCR was conducted in an initial step of 95°C for 30 s, 40 cycles of 95°C for 5 s, and 58–60°C for 5 s. The fluorescence values were determined after each elongation step. The dissociation curve was analyzed following the final cycle to determine the specificity of amplification. The melting protocol consisted of heating from 65°C to 95°C, holding at each temperature for 0.1 s while monitoring fluorescence. To identify specific amplification of a single PCR product, the product was confirmed by 2% agarose gel electrophoresis. Negative controls, comprised of the PCR reaction mix, without nucleic acid, were also run with each group of samples. The comparative cycle threshold method was used for quantification of expression levels. The abundance of each single gene was determined relative to the averages of two housekeeping genes (H2afz and Gapdh). Fluorescence was acquired in each cycle to determine the threshold cycle (C T ), the cycle during the log-linear phase of the reaction at which fluorescence was increased above background for each sample. The sample ∆C T value was calculated by the difference between the average C T value and the sample C T value. The GV stage was used as the calibrator and used subsequently to obtain ∆∆C T values. Fold differences of the gene expression were determined by using the formula 2−∆∆CT.
Immunofluorescence analysis.
Embryos were fixed with 3.7% paraformaldehyde in PBS for 30 min at room temperature. After three 30-min washings in PBS containing 0.1% Tween-20 (PBST), fixed embryos were permeabilized for 30 min at room temperature in PBS containing 0.5% Triton X-100 and then blocked with 1% bovine serum albumin in PBST for 2 h at room temperature. The embryos then were incubated with primary antibody overnight at 4°C using the following antibodies: polyclonal rabbit anti-H3K4me1 (1/500; 07-436; Upstate Biotechnology, Lake Placid, NY), anti-H3K4me3 (1/600; ab8580; Abcam, Cambridge, MA), anti-KDM5A (1/1,000; 3,876; Cell Signaling Technology, Danvers, MA), anti-KDM5B (1/1,000; 3,273; Cell Signaling Technology), anti-KDM1A (1/600; 2,184; Cell Signaling Technology), anti-ASH2L (1/600; 5,019; Cell Signaling Technology), anti-KMT2B (1/1,000; sc-292359; Santa Cruz Biotechnology, Inc., Santa Cruz, CA), or monoclonal mouse anti-H3K4me2 (1/500; 05-1338; Upstate Biotechnology). After three 30-min washings with PBST, the embryos were incubated with a secondary antibody conjugated with fluorescein isothiocyanate (FITC; 1/300; Jackson ImmunoResearch, West Grove, PA) for 1 h in the dark at room temperature. After washing three times in PBST, the embryos were stained with 4′,6-diamidino-2-phenylindole (DAPI; 1 μg/ml; D9542, Sigma) for 15 min, mounted in Vectashield Mounting Medium (H-1,000; Vector Laboratories, Inc., Burlingame, CA), and sealed with fingernail polish. The slides were examined using a Zeiss Axio Observer microscope equipped with epifluorescence (Carl Zeiss, Oberkochen, Germany). Exposure times from fluorescent light were kept constant for the respective channel (FITC or DAPI). Quantitation of the fluorescence intensity was determined using an image analyzer system, SigmaScan-pro V5.01 (SPSS Inc., Chicago, IL), by the ratios of H3K4me or its regulators to DAPI DNA signals (Shao et al. 2008b).
Statistical analysis.
The difference in fluorescence intensity in mouse preimplantation embryos was tested by Kruskal-Wallis nonparametric analysis of variance. The fold change of gene expression between different developmental stages was analyzed by one-way ANOVA using SPSS 11.5 software (SPSS Inc.). Data were presented as mean ± SEM of three independent experiments. P < 0.05 was considered statistically significant.
Results
Changes in H3K4me during preimplantation development.
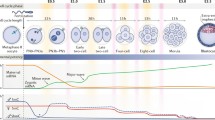
To evaluate the profile of H3K4me status during preimplantation development, H3K4me1, me2, and me3 signals were measured at various developmental stages in in vivo mouse embryos. H3K4me1/2/3 signals were observed in female pronuclei, whereas only weak H3K4me2 signal was detected in paternal chromatin (Fig. 1 a). Thereafter, intensity of H3K4me2 and me3 signals abruptly increased at the two-cell stage (time of embryonic genome activation, EGA; P < 0.05). Subsequently, H3K4me2 signal intensity dramatically decreased at the four-cell stage and remained at low level until the blastocyst stage (P < 0.05), whereas H3K4me3 signal intensity transiently decreased at the four-cell stage and then gradually increased to the peak at the blastocyst stage (P < 0.05; Fig. 1b ). Unlike H3K4me2 and me3, H3K4me1 signal intensity obviously increased at the two-cell stage (P < 0.05) but maintained at low level at other developmental stages (Fig. 1b ).
Intensity of H3K4me1, me2, and me3 in in vivo embryos during preimplantation development. (a) Microscopic images of H3K4me1, me2, and me3 at various developmental stages. Mouse one-cell, two-cell, four-cell, eight-cell, morula, and blastocyst embryos were collected at 18, 42, 50, 62, 76, and 90 h post-hCG injection, respectively, and subjected to immunocytochemistry with the antibody to methylated H3K4. The antibody was localized with FITC-conjugated secondary antibodies (green), and DNA was stained with DAPI (blue). The scale bar denotes 30 μm. (b) Patterns of H3K4me1, me2, and me3 in in vivo embryos. The numbers of embryos examined were 69, 65, 65, 68, 53, and 51 for H3K4me1; 67, 65, 68, 62, 51, and 52 for H3K4me2; and 63, 66, 62, 61, 54, and 56 for H3K4me3 in the one-cell, two-cell, four-cell, eight-cell, morula, and blastocyst embryos, respectively. Quantification of the fluorescence intensity was determined by the ratio of H3K4me to the DNA signal intensity. The data represent mean ± SEM of three independent experiments. Stages with different superscript letters represent statistically significant differ (P < 0.05). BL blastocyst, f female pronucleus, m male pronucleus.
mRNA expression of H3K4 methyltransferases and demethylases in preimplantation embryos.
Because enzymatic addition of methylated marks is a potential mechanism for the increase in H3K4me levels, we characterized the mRNA expression of H3K4 methyltransferases including Ash1l, Ash2l, Kmt2a, Kmt2b, Kmt2c, Setd1a, and Setd7, and of demethylases including Kdm1a, Kdm1b, Kdm5a, Kdm5b, Kdm5c, and Kdm5d in mouse oocytes and preimplantation embryos using quantitative reverse transcriptase (qRT)-PCR. Setd1a, Setd7, and Kmt2c exhibited a similar expression pattern. Their levels began to increase from the one-cell stage and were highest at the four-cell stage. In the subsequent cell divisions, the levels decreased gradually until the blastocyst stage (P < 0.05; Fig. 2). The transcript levels of Ash2l were highest in the two-cell embryos, and then showed a significant decrease in the four-cell embryos (P < 0.05). From the four- to eight-cell stages, no significant changes in the levels were noted (P > 0.05), but significant increases were detected in both morula and blastocyst stages (P < 0.05), which is consistent with the pattern of H3K4me2. The Kmt2b levels remained relatively low during the stages comprised between the oocytes and the eight-cell embryos, and a stable increase in the levels was observed from the eight-cell embryos until the blastocyst stage (P < 0.05; Fig. 2), which is a similar pattern with H3K4me3. Of all the methyltransferase genes, only Kmt2a and Ash1l displayed a constant level of transcripts at all stages examined except for a relative increase at the two-cell stage (P < 0.05).
Transcription patterns of H3K4 methyltransferases Setd1a, Setd7, Kmt2c, Ash2l, Kmt2b, Kmt2a, and Ash2l during mouse preimplantation development. The levels of the transcripts were normalized against H2afz and Gapdh. The transcript levels of GV oocytes were normalized to 1. The data represent mean ± SEM of three independent experiments. Thirty embryos were examined for each gene at various developmental stages. BL blastocyst, 1C one cell, 2C two cell, 4C four cell, 8C eight cell, GV germinal vesicle, MII metaphase II, M morula. For each mRNA within a panel, the values with different superscript letters represent statistically significant differ (P < 0.05).
On the other hand, the expression of Kdm1a and Kdm5b decreased to the lowest levels at the two-cell stage but increased significantly during subsequent cleavage stages (P < 0.05), which is opposite to the level of H3K4me2, whereas the expression of Kdm5a remained at low level at the two-cell stage, but abruptly increased at the four-cell stage and subsequently decreased to the lowest level at the blastocyst stage (P < 0.05; Fig. 3), which is opposite to the H3K4me3 level. Notably, Kdm5b was expressed at each stage but very high in the four-cell embryos suggesting a more active transcription state (∼180-fold; Fig. 3). In addition, Kdm5c, Kdm5d, and Kdm1b exhibited similar expression patterns—medium level in the one- and two-cell embryos, highest expression in the four-cell embryos, and low expression in the blastocysts (Fig. 3).
Transcription patterns of H3K4 demethylases Kdm1a, Kdm5b, Kdm5a, Kdm5c, Kdm5d, and Kdm1b during mouse preimplantation development. The levels of the transcripts were normalized against H2afz and Gapdh. The transcript levels of GV oocytes were normalized to 1. The data represent mean ± SEM of three independent experiments. Thirty embryos were examined for each gene at various developmental stages. For each mRNA within a panel, the values with different superscript letters represent statistically significant differ (P < 0.05).
Protein expression of H3K4 methyltransferases and demethylases in preimplantation embryos.
To specify the correlation between some methyltransferases and demethylases and H3K4me, we measured the signals of ASH2L, KMT2B, KDM5A, KDM5B, and KDM1A at the two-cell, four-cell, and blastocyst stages. The signals were observed at all stages (Fig. 4a ), but the quantitative profiles varied between these stages (Fig. 4b ). The ASH2L and KMT2B signals had their lowest intensity in the four-cell embryos and the highest intensity in the two-cell embryos for ASH2L and in the blastocysts for KMT2B. The intensity of KDM5A, KDM5B, and KDM1A signals significantly increased to the highest level at the four-cell stage and higher level at the blastocyst stage (P < 0.05; Fig. 4b ). However, the intensity of KDM5B was markedly higher than that of KDM5A and KDM1A at various stages (Fig. 4b ).
Intensity of ASH2L, KMT2B, KDM5A, KDM5B, and KDM1A in in vivo embryos during preimplantation development. (a) Microscopic images of H3K4 methyltransferases ASH2L and KMT2B, and of H3K4 demethylases KDM5A, KDM5B, and KDM1A at the two-cell, four-cell, and blastocyst stages. Mouse two-cell, four-cell, and blastocyst embryos were collected at 42, 50, and 90 h post-hCG injection, respectively. The antibody was localized with FITC-conjugated secondary antibodies (green), and DNA was stained with DAPI (blue). The scale bar denotes 30 μm. (b) Protein expression patterns of ASH2L, KMT2B, KDM5A, KDM5B, and KDM1A in in vivo embryos. The numbers of embryos examined were 62, 57, and 54 for ASH2L; 57, 53, and 51 for KMT2B; 56, 58, and 52 for KDM5A; 59, 54, and 52 for KDM5B; and 61, 56, and 51 for KDM1A in the two-cell, four-cell, and blastocyst embryos, respectively. Quantification of the fluorescence intensity was determined by the ratio of each regulator to the DNA signal intensity. The intensity of fluorescence in the two-cell embryos was normalized to 1. The data represent mean ± SEM of three independent experiments. The asterisk indicates significant differences between the indicated stages (P < 0.05). BL blastocyst.
Discussion
Histone methylation has emerged as a critical player in the regulation of gene expression (Black et al. 2012). H3K4me is particularly associated with gene activation (Santos-Rosa et al. 2002), although certain genes can be repressed due to H3K4me promoter occupancy (Wang et al. 2011). This epigenetic state is known to be involved in the regulation of key developmental genes in embryonic stem cells (Wang et al. 2009a, b; Ang et al. 2011) and is involved in the regulation of pluripotency (Pan et al. 2007). Previous studies have confirmed that H3K4me undergoes reprogramming during early embryonic development (Sarmento et al. 2004; Shao et al. 2008b). However, little information is available on this reprogramming and its regulators in preimplantation embryos. In this study, the profiles of H3K4me and its regulators were investigated during early development of in vivo mouse embryos. At the one-cell stage, H3K4me1/2/3 signals were present in the maternal genome, whereas only weak H3K4me2 signal was observed in the paternal chromatin. Our results are consistent with the previous findings of Sarmento et al. (2004), van der Heijden et al. (2005), and Black et al. (2012). The difference in H3K4me levels between female and male pronuclei may be responsible for distinctive formation of the respective chromatin. In addition, H3K4me2 and me3 exhibited the highest overall levels at the two-cell and blastocyst stages, respectively, in accordance with the previous reports (Santenard et al. 2010; Wu et al. 2012), indicating that H3K4me2 may play a role in early EGA, whereas H3K4me3 may be involved in initial lineage segregation in the blastocyst.
Histone methylation is catalyzed by a class of enzymes known as histone methyltransferases. We examined the expression of these enzymes during preimplantation development to elucidate which enzymes are involved in regulating H3K4me during this period. A previous study reported that Ash2l presented relatively high transcript levels during the EGA in in vitro porcine embryos from a microarray analysis (Whitworth et al. 2005). This result was confirmed in the present study by using a different animal species and the qRT-PCR and immunocytochemical staining. The high levels of Ash2l around the EGA were consistent with that of H3K4me2, which may stabilize this methylation levels during the EGA. A relative decrease in expression of Kmt2a was detected in another study (Gao et al. 2010). The peak in expression we observe at the two-cell stage is thus intriguing and should be investigated in further studies. Recently, Andreu-Vieyra et al. (2010) reported that Kmt2b transcripts exhibited higher level at the morula and blastocyst stages in in vitro mouse embryos. This result is consistent with our finding that the highest levels of Kmt2b mRNA were observed in in vivo mouse embryos at the same stages, suggesting that it may play a role in the initial cell lineage separation by means of methylating H3K4.
Removal of H3K4me by specific demethylases is crucial for gene repression to occur. Hence, it was expected that this histone methylation would be activated in order to allow for the EGA. This was to some degree substantiated, as the levels of H3K4me2 increased from the one- to two-cell stages. Correspondingly, a concomitant decrease in the mRNA and protein levels of its demethylases, Kdm5b and Kdm1a, was observed. This was in accordance with the recent results obtained by McGraw et al. (2007). Interestingly, the protein levels of KDM5B were the highest in the four-cell embryos, but its levels were only six times higher than that in the two-cell embryos. The mRNA levels of Kdm5b were at least 40 times higher in the four-cell embryos. It is possible that KDM5B causes a feedback inhibition of the splicing of the transcript of its own gene in the four-cell embryos. Another possibility is that post-translation modifications of KDM5B protein during the translation block the translation process or promote the degradation of the protein itself. In addition, McGraw et al. (2007) also found that a decrease in expression of Kdm5a was detected in the morula and blastocysts which reflects a similar pattern observed in our expression data which shows this reduced expression persists at the blastocyst stage. Noteworthily, the protein levels of KDM5A at the blastocyst stage was about two times higher than that at the two-cell stage, while the mRNA levels of Kdm5a were lower at the blastocyst stage. A possible explanation for this phenotype is that an increased availability of active KDM5A by unknown kinases may promote the translation of itself mRNA by virtue of upstream open reading frames (ORFS) in their 5′ untranslated regions (UTRs). Another possibility is that KDM5A itself causes a feedback inhibition of the transcription of its own gene.
Based on these findings, we suggest that H3K4me2 may play a role in the EGA. Hence, relatively high levels of this permissive histone modification coinciding with a peak expression of its methyltransferase, ASH2L, were noted at the two-cell stage, whereas a decrease in expression of its demethylases, KDM5B and KDM1A, was observed. H3K4me3 may play a major role in lineage segregation in the blastocyst when it occurs at relatively high levels that associate with the expression of KMT2B and KDM5A. For future studies, it will be interesting to investigate whether the depletions of these enzymes by RNAi change the H3K4me levels and therefore affect embryonic development.
References
Albert M, Schmitz SU, Kooistra SM, Malatesta M, Morales Torres C, Rekling JC, Johansen JV, Abarrategui I, Helin K (2013) The histone demethylase Jarid1b ensures faithful mouse development by protecting developmental genes from aberrant H3K4me3. PLoS Genet 9:e1003461
Andreu-Vieyra CV, Chen R, Agno JE, Glaser S, Anastassiadis K, Stewart AF, Matzuk MM (2010) MLL2 is required in oocytes for bulk histone 3 lysine 4 trimethylation and transcriptional silencing. PLoS Biol 8:e1000453
Ang YS, Tsai SY, Lee DF, Monk J, Su J, Ratnakumar K, Ding J, Ge Y, Darr H, Chang B, Wang J, Rendl M, Bernstein E, Schaniel C, Lemischka IR (2011) Wdr5 mediates self-renewal and reprogramming via the embryonic stem cell core transcriptional network. Cell 145:183–197
Barski A, Cuddapah S, Cui K, Roh TY, Schones DE, Wang Z, Wei G, Chepelev I, Zhao K (2007) High-resolution profiling of histone methylations in the human genome. Cell 129:823–837
Beisel C, Paro R (2011) Silencing chromatin: comparing modes and mechanisms. Nat Rev Genet 12:123–135
Bernstein BE, Mikkelsen TS, Xie X, Kamal M, Huebert DJ, Cuff J, Fry B, Meissner A, Wernig M, Plath K, Jaenisch R, Wagschal A, Feil R, Schreiber SL, Lander ES (2006) A bivalent chromatin structure marks key developmental genes in embryonic stem cells. Cell 125:315–326
Black JC, Van Rechem C, Whetstine JR (2012) Histone lysine methylation dynamics: establishment, regulation, and biological impact. Mol Cell 48:491–507
Chan SW, Hong W (2001) Retinoblastoma-binding protein 2 (Rbp2) potentiates nuclear hormone receptor-mediated transcription. J Biol Chem 276:28402–28412
Ciccone DN, Su H, Hevi S, Gay F, Lei H, Bajko J, Xu G, Li E, Chen T (2009) KDM1B is a histone H3K4 demethylase required to establish maternal genomic imprints. Nature 461:415–418
Di Stefano L, Ji JY, Moon NS, Herr A, Dyson N (2007) Mutation of Drosophila Lsd1 disrupts H3-K4 methylation, resulting in tissue-specific defects during development. Curr Biol 17:808–812
Dillon SC, Zhang X, Trievel RC, Cheng X (2005) The SET-domain protein superfamily: protein lysine methyltransferases. Genome Biol 6:227
Dou Y, Milne TA, Ruthenburg AJ, Lee S, Lee JW, Verdine GL, Allis CD, Roeder RG (2006) Regulation of MLL1 H3K4 methyltransferase activity by its core components. Nat Struct Mol Biol 13:713–719
Eissenberg JC, Shilatifard A (2010) Histone H3 lysine 4 (H3K4) methylation in development and differentiation. Dev Biol 339:240–249
Eliazer S, Shalaby NA, Buszczak M (2011) Loss of lysine-specific demethylase 1 nonautonomously causes stem cell tumors in the Drosophila ovary. Proc Natl Acad Sci U S A 108:7064–7069
Fang R, Barbera AJ, Xu Y, Rutenberg M, Leonor T, Bi Q, Lan F, Mei P, Yuan GC, Lian C, Peng J, Cheng D, Sui G, Kaiser UB, Shi Y, Shi YG (2010) Human LSD2/KDM1b/AOF1 regulates gene transcription by modulating intragenic H3K4me2 methylation. Mol Cell 39:222–233
Gao Y, Hyttel P, Hall VJ (2010) Regulation of H3K27me3 and H3K4me3 during early porcine embryonic development. Mol Reprod Dev 77:540–549
Greer EL, Maures TJ, Hauswirth AG, Green EM, Leeman DS, Maro GS, Han S, Banko MR, Gozani O, Brunet A (2010) Members of the H3K4 trimethylation complex regulate lifespan in a germline-dependent manner in C. elegans. Nature 466:383–387
Gregory GD, Vakoc CR, Rozovskaia T, Zheng X, Patel S, Nakamura T, Canaani E, Blobel GA (2007) Mammalian ASH1L is a histone methyltransferase that occupies the transcribed region of active genes. Mol Cell Biol 27:8466–8479
Heintzman ND, Stuart RK, Hon G, Fu Y, Ching CW, Hawkins RD, Barrera LO, Van Calcar S, Qu C, Ching KA, Wang W, Weng Z, Green RD, Crawford GE, Ren B (2007) Distinct and predictive chromatin signatures of transcriptional promoters and enhancers in the human genome. Nat Genet 39:311–318
Jensen LR, Amende M, Gurok U, Moser B, Gimmel V, Tzschach A, Janecke AR, Tariverdian G, Chelly J, Fryns JP, Van Esch H, Kleefstra T, Hamel B, Moraine C, Gecz J, Turner G, Reinhardt R, Kalscheuer VM, Ropers HH, Lenzner S (2005) Mutations in the JARID1C gene, which is involved in transcriptional regulation and chromatin remodeling, cause X-linked mental retardation. Am J Hum Genet 76:227–236
Katz DJ, Edwards TM, Reinke V, Kelly WG (2009) A C. elegans LSD1 demethylase contributes to germline immortality by reprogramming epigenetic memory. Cell 137:308–320
Klose RJ, Kallin EM, Zhang Y (2006) JmjC-domain-containing proteins and histone demethylation. Nat Rev Genet 7:715–727
Lee J, Saha PK, Yang QH, Lee S, Park JY, Suh Y, Lee SK, Chan L, Roeder RG, Lee JW (2008) Targeted inactivation of MLL3 histone H3-Lys-4 methyltransferase activity in the mouse reveals vital roles for MLL3 in adipogenesis. Proc Natl Acad Sci U S A 105:19229–19234
Lee JH, Skalnik DG (2008) Wdr82 is a C-terminal domain-binding protein that recruits the Setd1A histone H3-Lys4 methyltransferase complex to transcription start sites of transcribed human genes. Mol Cell Biol 28:609–618
Mamo S, Gal AB, Bodo S, Dinnyes A (2007) Quantitative evaluation and selection of reference genes in mouse oocytes and embryos cultured in vivo and in vitro. BMC Dev Biol 7:14
McGraw S, Vigneault C, Sirard MA (2007) Temporal expression of factors involved in chromatin remodeling and in gene regulation during early bovine in vitro embryo development. Reproduction 133:597–608
Mikkelsen TS, Ku M, Jaffe DB, Issac B, Lieberman E, Giannoukos G, Alvarez P, Brockman W, Kim TK, Koche RP, Lee W, Mendenhall E, O’Donovan A, Presser A, Russ C, Xie X, Meissner A, Wernig M, Jaenisch R, Nusbaum C, Lander ES, Bernstein BE (2007) Genome-wide maps of chromatin state in pluripotent and lineage-committed cells. Nature 448:553–560
Milne TA, Briggs SD, Brock HW, Martin ME, Gibbs D, Allis CD, Hess JL (2002) MLL targets SET domain methyltransferase activity to Hox gene promoters. Mol Cell 10:1107–1117
Pan G, Tian S, Nie J, Yang C, Ruotti V, Wei H, Jonsdottir GA, Stewart R, Thomson JA (2007) Whole-genome analysis of histone H3 lysine 4 and lysine 27 methylation in human embryonic stem cells. Cell Stem Cell 1:299–312
Pradhan S, Chin HG, Esteve PO, Jacobsen SE (2009) SET7/9 mediated methylation of non-histone proteins in mammalian cells. Epigenetics 4:383–387
Santenard A, Ziegler-Birling C, Koch M, Tora L, Bannister AJ, Torres-Padilla ME (2010) Heterochromatin formation in the mouse embryo requires critical residues of the histone variant H3.3. Nat Cell Biol 12:853–862
Santos-Rosa H, Schneider R, Bannister AJ, Sherriff J, Bernstein BE, Emre NC, Schreiber SL, Mellor J, Kouzarides T (2002) Active genes are tri-methylated at K4 of histone H3. Nature 419:407–411
Sarmento OF, Digilio LC, Wang Y, Perlin J, Herr JC, Allis CD, Coonrod SA (2004) Dynamic alterations of specific histone modifications during early murine development. J Cell Sci 117:4449–4459
Shao GB, Ding HM, Gong AH (2008a) Role of histone methylation in zygotic genome activation in the preimplantation mouse embryo. In Vitro Cell Dev Biol Anim 44:115–120
Shao GB, Ding HM, Gong AH, Xiao DS (2008b) Inheritance of histone H3 methylation in reprogramming of somatic nuclei following nuclear transfer. J Reprod Dev 54:233–238
Shi Y, Lan F, Matson C, Mulligan P, Whetstine JR, Cole PA, Casero RA, Shi Y (2004) Histone demethylation mediated by the nuclear amine oxidase homolog LSD1. Cell 119:941–953
Smith JA, White EA, Sowa ME, Powell ML, Ottinger M, Harper JW, Howley PM (2010) Genome-wide siRNA screen identifies SMCX, EP400, and Brd4 as E2-dependent regulators of human papillomavirus oncogene expression. Proc Natl Acad Sci U S A 107:3752–3757
Stoller JZ, Huang L, Tan CC, Huang F, Zhou DD, Yang J, Gelb BD, Epstein JA (2010) Ash2l interacts with Tbx1 and is required during early embryogenesis. Exp Biol Med (Maywood) 235:569–576
Terranova R, Agherbi H, Boned A, Meresse S, Djabali M (2006) Histone and DNA methylation defects at Hox genes in mice expressing a SET domain-truncated form of Mll. Proc Natl Acad Sci U S A 103:6629–6634
van der Heijden GW, Dieker JW, Derijck AA, Muller S, Berden JH, Braat DD, van der Vlag J, de Boer P (2005) Asymmetry in histone H3 variants and lysine methylation between paternal and maternal chromatin of the early mouse zygote. Mech Dev 122:1008–1022
VerMilyea MD, O’Neill LP, Turner BM (2009) Transcription-independent heritability of induced histone modifications in the mouse preimplantation embryo. PLoS One 4:e6086
Wan M, Liang J, Xiong Y, Shi F, Zhang Y, Lu W, He Q, Yang D, Chen R, Liu D, Barton M, Songyang Z (2013) The trithorax group protein Ash2l is essential for pluripotency and maintaining open chromatin in embryonic stem cells. J Biol Chem 288:5039–5048
Wang J, Hevi S, Kurash JK, Lei H, Gay F, Bajko J, Su H, Sun W, Chang H, Xu G, Gaudet F, Li E, Chen T (2009a) The lysine demethylase LSD1 (KDM1) is required for maintenance of global DNA methylation. Nat Genet 41:125–129
Wang J, Zhang M, Zhang Y, Kou Z, Han Z, Chen DY, Sun QY, Gao S (2010) The histone demethylase JMJD2C is stage-specifically expressed in preimplantation mouse embryos and is required for embryonic development. Biol Reprod 82:105–111
Wang K, Chen Y, Chang EA, Knott JG, Cibelli JB (2009b) Dynamic epigenetic regulation of the Oct4 and Nanog regulatory regions during neural differentiation in rhesus nuclear transfer embryonic stem cells. Cloning Stem Cells 11:483–496
Wang SS, Zhou BO, Zhou JQ (2011) Histone H3 lysine 4 hypermethylation prevents aberrant nucleosome remodeling at the PHO5 promoter. Mol Cell Biol 31:3171–3181
Whitworth KM, Agca C, Kim JG, Patel RV, Springer GK, Bivens NJ, Forrester LJ, Mathialagan N, Green JA, Prather RS (2005) Transcriptional profiling of pig embryogenesis by using a 15-K member unigene set specific for pig reproductive tissues and embryos. Biol Reprod 72:1437–1451
Wu FR, Liu Y, Shang MB, Yang XX, Ding B, Gao JG, Wang R, Li WY (2012) Differences in H3K4 trimethylation in in vivo and in vitro fertilization mouse preimplantation embryos. Genet Mol Res 11:1099–1108
Xiang Y, Zhu Z, Han G, Ye X, Xu B, Peng Z, Ma Y, Yu Y, Lin H, Chen AP, Chen CD (2007) JARID1B is a histone H3 lysine 4 demethylase up-regulated in prostate cancer. Proc Natl Acad Sci U S A 104:19226–19231
Yamane K, Tateishi K, Klose RJ, Fang J, Fabrizio LA, Erdjument-Bromage H, Taylor-Papadimitriou J, Tempst P, Zhang Y (2007) PLU-1 is an H3K4 demethylase involved in transcriptional repression and breast cancer cell proliferation. Mol Cell 25:801–812
Acknowledgments
This work was supported by grants from the National Natural Science Foundation of China (81170573) and the Open Research Fund Program of Jiangsu Provincial Key Laboratory for Science and Technology of Photon Manufacturing (Jiangsu University) (GZ200710) and the Natural Science Foundation of Jiangsu Province (BK2011485).
Author information
Authors and Affiliations
Corresponding author
Additional information
Editor: T. Okamoto
Rights and permissions
About this article
Cite this article
Shao, GB., Chen, JC., Zhang, LP. et al. Dynamic patterns of histone H3 lysine 4 methyltransferases and demethylases during mouse preimplantation development. In Vitro Cell.Dev.Biol.-Animal 50, 603–613 (2014). https://doi.org/10.1007/s11626-014-9741-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11626-014-9741-6