Summary
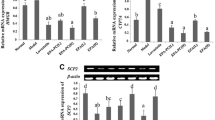
Abnormal cholesterol metabolism is associated with an elevated risk of developing atherosclerosis, hypertension, and diabetes etc. Na+/K+-ATPase was found to regulate cholesterol synthesis, distribution and trafficking. This study aimed to examine the effect of high-fat diet on cholesterol metabolism in rats and the role of Na+/K+-ATPase/Src/ERK signaling pathway in the process. Forty male SD rats were evenly divided into high-fat diet group and control group at random. Animals in the former group were fed on high-fat diet for 12 weeks, and those fed on basic diet served as control. Blood lipids, including total cholesterol (TC), triglyceride (TG), high density lipoprotein-cholesterol (HDL-C), and low density lipoprotein-cholesteral (LDL-C) levels, were detected at 3, 6 and 12 weeks. The ratio of cholesterol content in cytoplasm to that in cell membrane was detected in liver tissues. RT-PCR and Western blotting were used to measure the expression of lipid metabolism-associated genes (HMG-CoA reductase and SREBP-2) after 12-week high-fat diet. Na+/K+-ATPase/Src/ERK signaling pathway-related components (Na+/K+-ATPase α1, Src-PY418 and pERK1/2) were also measured by Western blotting. The results showed that the serum TC, TG, and LDL-C levels were significantly higher in high-fat diet group than those in control group, while the HDL-C level was significantly lower in high-fat diet group at 6 weeks (P<0.01). High-fat diet led to an increase in the cholesterol content in the cytoplasm and cell membrane. The ratio of cholesterol content in cytoplasm to that in cell membrane was elevated over time. The expression of HMG-CoA reductase and SREBP-2 was significantly suppressed at mRNA and protein levels after 12-week high-fat diet (P<0.05). Moreover, high-fat diet promoted the expression of Na+/K+-ATPase α1 but suppressed the phosphorylation of Src-PY418 and ERK1/2 at 12 weeks (P<0.05). It was concluded that high-fat diet regulates cholesterol metabolism, and Na+/K+-ATPase signaling pathway is involved in the process possibly by regulating the expression of lipid metabolism-associated proteins HMG-CoA reductase and SREBP-2.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Ikonen E. Cellular cholesterol trafficking and compartmentalization. Nat Rev Mol Cell Biol, 2008,9(2):125–138
Ouimet M, Franklin V, Mak E, et al. Autophagy regulates cholesterol efflux from macrophage foam cells via lysosomal acid lipase. Cell Metab, 2011,13(6):655–167
Glynn IM. A hundred years of sodium pumping. Annu Rev Physiol, 2002,64:1–18
Li ZC, Zhang ZB, Xie JX, et al. Na+/K+-ATPase mimetic pNaKtide peptide inhibits the growth of human cancer cells. J Biol Chem, 2011,286:32394–32403
Adamian L, Naveed H, Liang J. Lipid-binding surfaces of membrane proteins: evidence from evolutionary and structural analysis. Biochim Biophys Acta, 2011,1808(4):1092–1102
Briand N, Dugail I, Le Lay S. Cavin proteins: New players in the caveolae field. Biochimie, 2011,93(1):71–77
Chidlow JH Jr, Sessa WC. Caveolae, caveolins, and cavins: complex control of cellular signalling and inflammation. Cardiovasc Res, 2010,86(2):219–225
Hansen CG, Nichols BJ. Exploring the caves: cavins, caveolins and caveolae. Trends Cell Biol, 2010,20(4):177–186
Cai T, Wang H, Chen Y, et al. Regulation of caveolin-1 membrane trafficking by the Na+/K+-ATPase. J Cell Biol, 2008,182(6):1153–1169
Ye Q, Lai F, Banerjee M, et al. Expression of mutant a1 Na+/K+-ATPase defective in conformational transition attenuates Src-mediated signal transduction. J Biol Chem, 2013,288(8):5803–5814
Ness GC. Physiological feedback regulation of cholesterol biosynthesis: Role of translational control of hepatic HMG-CoA reductase and possible involvement of oxylanosterols. Biochim Biophys Acta, 2015,1851(5):667–673
Brouwer IA, Wanders AJ, Katan MB. Effect of animal and industrial trans fatty acids on HDL and LDL cholesterol levels in humans—a quantitative review. PLoS One, 2010,5(3):9434
Yokoyama C, Wang X, Briggs MR, et al. SREBP-1, a basic-helix-loop-helix-leucine zipper protein that controls transcription of the low density lipoprotein receptor gene. Cell, 1993,75:187–197
Theodore L, Lange SY. Cell cholesterol homeostasis: mediation by active cholesterol. Trends Cell Biol, 2010,20(11):680–687
Fukumitsu S, Villareal MO, Onaga S, et al. a-Linolenic acid suppresses cholesterol and tricylglycerol biosynthesis pathway by suppressing SREBP-2, SREBP-1a and-c expression. Cytotechnology, 2013,65(6):899–907
Aguilar D, Fernandez ML. Hypercholesterolemia induces adipose dysfunction in conditions of obesity and nonobesity. Adv Nutr, 2014,5(5):497–502
Chen Y, Cai T, Wang H, et al. Regulation of intracellular cholesterol distribution by Na+/K+-ATPas. J Biol Chem, 2009,284(22):14881–14890
Xie JX, Li X, Xie Z. Regulation of renal function and structure by the signaling Na+/K+-ATPase. IUBMB Life, 2013,65(12):991–998
Robinson JG. 2013 ACC/AHA cholesterol guideline for reducing cardiovascular risk: what is so controversial? Curr Atheroscler Rep, 2014,16(6):413
Chen M, Mason RP, Tulenko TN. Biochim Biophys Acta, 1995,1272(2):101–112
Author information
Authors and Affiliations
Corresponding author
Additional information
This study was supported by a grant from the National Natural Science Foundation of China (No. 81200637).
Rights and permissions
About this article
Cite this article
Wang, L., Xu, F., Zhang, Xj. et al. Effect of high-fat diet on cholesterol metabolism in rats and its association with Na+/K+-ATPase/Src/pERK signaling pathway. J. Huazhong Univ. Sci. Technol. [Med. Sci.] 35, 490–494 (2015). https://doi.org/10.1007/s11596-015-1458-6
Received:
Revised:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11596-015-1458-6