Abstract
Polyporus sensu lato is understood to be a polyphyletic genus, including two narrowly defined genera (Favolus and Neofavolus), two subclades (Melanopus and Polyporellus), and some scattered species. In this study, Polyporus hapalopus is described and illustrated as a new species on the basis of morphological and molecular evidence. This species is characterized by laterally stipitate, imbricate and large basidiocarps (up to 40 cm diam.), grapefruit odor when fresh, angular pores with lacerate dissepiments, soft (when fresh) to tough (when dry) context, a dimitic hyphal system in context and stipe with variable wide skeleto-binding hyphae, a monomitic hyphal system in trama, and cylindrical basidiospores bearing one or two guttules. Phylogenetically, on the basis of the combined internal transcribed spacer and nuclear large subunit rDNA sequences, P. hapalopus is nested within the Polyporus sensu lato clade, but is separated from four previously established subclades and from other known species.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Polyporus P. Micheli ex Adans. is a widespread polypore genus. It includes species with stipitate to substipitate basidiocarps, a dimitic hyphal system, thick-walled and arboriform skeleto-binding hyphae, thin-walled, smooth and cylindrical to subellipsoid basidiospores with negative reaction in Melzer’s reagent, and causing a white rot (Gilbertson and Ryvarden 1987; Núñez and Ryvarden 1995).
Núñez and Ryvarden (1995) treated 32 species of Polyporus worldwide, arranged into six morphological groups. Subsequently, several Polyporus species have been reinstated (Zheng and Liu 2005), newly described (Buchanan and Ryvarden 1998; Dai et al. 2003, 2007d, 2009; Sotome et al. 2007; Xue and Zhou 2012) and emended (Dai 2012a). With the aid of molecular phylogeny from various genetic regions, Polyporus has been consistently characterized as a polyphyletic genus (Ko and Jung 2002; Krüger et al. 2006; Sotome et al. 2008). Recently, from both morphological and phylogenetic perspectives, Sotome et al. (2013) studied the taxonomy of morphological group Favolus as circumscribed in Núñez and Ryvarden (1995), and placed the species of this group in two genera Favolus Fr. and Neofavolus Sotome & T. Hatt., both belonging to Polyporus sensu lato. Meanwhile, Dai et al. (2014) recovered four clades (Favolus, Neofavolus, Melanopus and Polyporellus) and some scattered species in Polyporus sensu lato.
China has a high diversity of wood-decay fungi (Dai 2011, 2012b). Species of Polyporus have been repeatedly collected and documented from different regions (Dai et al. 2003, 2004, 2007a, b, c, d, 2009; Dai and Penttilä 2006; Cui et al. 2008; Li et al. 2008; Yuan and Dai 2008; Wang et al. 2011), and, to date, a total of 37 species of Polyporus have been recorded in China (Dai 2012b; Xue and Zhou 2012; Dai et al. 2014).
During a field trip in the Guangxi Autonomous Region of southern China in 2011, an unknown Polyporus specimen of giant size was collected. It was identified as a new species according to both morphological and phylogenetic analyses. We describe and illustrate it as Polyporus hapalopus in the present paper.
Materials and methods
Morphological study
The studied specimen, Yuan 5809, is deposited in the herbarium of the Institute of Applied Ecology, Chinese Academy of Sciences (IFP). The microscopic procedure follows Dai (2010). In this text, CB stands for Cotton Blue, CB+ for cyanophilous, CB– for acyanophilous, IKI for Melzer’s reagent, IKI– for negative reaction in Melzer’s reagent, and KOH for 5 % potassium hydroxide. The section was studied at magnification up to 1,000× using a Nikon Eclipse 80i microscope and phase contrast illumination. Measurements are made from sections stained with CB. In describing the variation in size of basidiospores, 5 % of measurements were excluded from each end of the range and given in parentheses. The meanings of abbreviations are as follows: L = mean basidiospore length (arithmetic average of all basidiospores), W = mean basidiospore width (arithmetic average of all basidiospores), Q = the ratio of L/W, and n = number of basidiospores measured from given number of specimens. Drawings are made with the aid of a drawing tube. Special color terms follow Petersen (1996).
Molecular procedures and phylogenetic analysis
Internal transcribed spacer (ITS) and nuclear large subunit rDNA (nLSU) sequences were obtained from the herbarium specimen using Phire® Plant Direct PCR Kit (Finnzymes Oy, Finland). The PCR procedure was conducted following Zhou and Xue (2012). For the genomic DNA extraction, a small piece of specimen Yuan 5809 was incubated in 20 μl Dilution Buffer for at least 3 min at room temperature. Then, 0.9 μl of the supernatant was used as template for a 30 μl PCR reaction. The ITS region was amplified and sequenced with the primer pair ITS5 and ITS4 (White et al. 1990), while nLSU sequence was amplified with the primers LR0R and LR7, and sequenced with the primers LR0R, LR3R, LR3, and LR7 (http://biology.duke.edu/fungi/mycolab/primers.htm). DNA sequencing was performed at Beijing Genomics Institute, China.
The newly generated sequences were submitted to GenBank (http://www.ncbi.nlm.nih.gov; Table 1). Other ITS and nLSU sequences for phylogenetic analysis (Table 1) were mainly adapted from Krüger et al. (2006), Sotome et al. (2009, 2011, 2013) and Dai et al. (2014). Pycnoporus cinnabarinus (Jacq.) P. Karst. and Trametes orientalis (Yasuda) Imazeki were selected as outgroup (Dai et al. 2014; Sotome et al. 2013). Following Dai et al. (2014), ITS and nLSU sequences were combined for phylogenetic analysis. The combined sequences were aligned by ClustalX 2.0 (Larkin et al. 2007) with default parameters. The resulting alignment was deposited in TreeBASE (http://www.treebase.org; accession number S14664). The best-fit evolutionary model was estimated by jModelTest (Guindon & Gascuel 2003; Posada 2008) based on corrected Akaike information criterion. Maximum likelihood (ML) analysis and Bayesian inference (BI) were performed using raxmlGUI 1.2 (Stamatakis 2006; Silvestro and Michalak 2012) and MrBayes 3.2 (Ronquist et al. 2012), respectively. The ML tree was constructed with the GTR model and auto FC option (Pattengale et al. 2010) in bootstrap (BS) replicates. For BI, two independent runs were launched. Each run performed a Metropolis-coupled Markov chain Monte Carlo analysis with four chains for 500,000 generations. Trees were saved every 1,000th generation. Chain convergence was determined using Tracer v1.5 (http://tree.bio.ed.ac.uk/software/tracer/). The value of burn-in was set to discard the first 25 % trees, and the remaining trees were used to compute a 50 % consensus tree and for calculating Bayesian posterior probabilities (BPPs).
Results
Molecular phylogeny
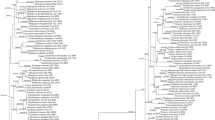
The combined ITS and nLSU dataset, resulting in an alignment with 1,492 characters, includes 23 ITS and 22 nLSU sequences from 23 collections representing 19 species. The best-fit evolutionary model of this alignment for BI was estimated as HKY. The ML search stopped after 200 BS replicates. All chains for BI converged after 500,000 generations, which was indicated by the effective sample sizes of all parameters above 250 and the potential scale reduction factors close to 1.000. ML and BI analyses generated nearly congruent topologies, so both BS values and BPPs were shown at the nodes of the topology from ML (Fig. 1).
The current phylogeny (Fig. 1) is similar to that in Dai et al. (2014). Four subclades, Favolus, Neofavolus, Melanopus and Polyporellus, were recovered within the Polyporus sensu lato clade. The newly sequenced specimen, Yuan 5809, was nested within the Polyporus sensu lato clade, and formed a single branch separate from other sampled species and the four subclades.
Taxonomy
Polyporus hapalopus H.J. Xue & L.W. Zhou, sp. nov. (Figs. 2, 3)
MycoBank no.: MB 802658
Basidiocarps annual, laterally stipitate, imbricate. Pileus fan-shaped, up to 40 cm diam. Pileal surface cinnamon-buff to clay-buff, with more or less radial stripes. Pore surface white to cream; pores angular, 4–6 per mm. Stipe concolorous with the pileal surface. Hyphal system dimitic; generative hyphae with clamp connections. Basidiospores cylindrical, hyaline, IKI–, CB–, 6.1–6.9 × 2.2–2.4 μm.
Type. China, Guangxi Autonomous Region, Laibin, Jinxiu County, Dayaoshan Nature Reserve, on fallen angiosperm trunk, 23.VIII.2011, Yuan 5809 (Holotype in IFP 019116).
Etymology: hapalopus (Lat.): referring to the soft context when fresh.
Fruitbody. Basidiocarps annual, laterally stipitate, imbricate, fleshy and with grapefruit odor when fresh, becoming corky and light in weight upon drying. Pileus fan-shaped, up to 40 cm diam. when fresh. Pileal surface cinnamon-buff to clay-buff, bearing a cuticle, glabrous, with more or less radial stripes, wrinkled when dry; margin sharp. Pore surface white to cream when fresh, becoming cinnamon-buff to honey-yellow upon drying; pores angular, 4–6 per mm; dissepiments thin, lacerate. Context cream to straw-yellow, soft (when fresh) to tough (when dry), up to 4 mm thick. Tubes less than 1 mm long, decurrent along one side of the stipe, similar to pore surface in color. Stipe concolorous with the pileal surface, glabrous, corky, up to 2 cm long, up to 2.5 cm diam., bearing a concolorous cuticle on one side of the stipe.
Hyphal structure. Hyphal system dimitic in context and stipe, monomitic in trama; generative hyphae with clamp connections; skeleto-binding hyphae frequently branched, with straight proximal and central parts and tapering ends, rarely with arboriform branches, IKI–, CB+; tissue unchanged in KOH.
Context. Generative hyphae infrequent, hyaline, thin-walled, 4–5 μm diam.; skeleto-binding hyphae dominant, thick-walled with a distinct wide lumen, loosely interwoven, 1.5–7.5 μm diam. Hyphae in cuticle of pileal surface parallel; generative hyphae thin-walled to slightly thick-walled, sometimes branched, 2.5–9 μm diam.; skeleto-binding hyphae thick-walled with a distinct wide lumen, 3–8 μm diam.
Tubes. Generative hyphae hyaline, thin-walled, scarcely branched, subparallel along tubes, 2.6–4 μm diam. Cystidia and cystidioles absent; basidia clavate, with a basal clamp and four sterigmata, 15–17 × 5.5–6.5 μm; basidioles similar to basidia, but smaller.
Stipe. Hyphal structure similar to context; skeleto-binding hyphae dominant, thick-walled with a distinct wide lumen, 1.5–6 μm diam. Hyphae in cuticle gelatinized, interwoven; generative hyphae frequent, thin-walled, 2–4 μm diam.; skeleto-binding hyphae dominant, thick-walled, 1.5–6 μm diam.
Spores. Basidiospores cylindrical, hyaline, thin-walled, smooth, bearing one or two guttules, IKI–, CB–, (5.5–)6.1–6.9(–7.6) × (2–)2.2–2.4(–2.7) μm, L = 6.32 μm, W = 2.29 μm, Q = 2.76 (n = 30/1).
Discussion
Polyporus hapalopus is characterized by its laterally stipitate, imbricate and large basidiocarps (up to 40 cm diam.) with grapefruit odor when fresh, angular pores, lacerate dissepiments, soft (when fresh) to tough (when dry) context, a dimitic hyphal system in context and stipe with variable wide skeleto-binding hyphae, a monomitic hyphal system in trama, and cylindrical basidiospores with one or two guttules. Its laterally stipitate basidiocarps, frequently branched skeleto-binding hyphae, cylindrical and IKI– basidiospores, and causing a white rot are features that fit the generic concept of Polyporus.
Polyporus hapalopus and P. udus Jungh. are both distributed in pantropical to warm-temperate areas (Núñez and Ryvarden 1995). Polyporus udus, belongs to the morphological group Polyporus, resembles P. hapalopus in sharing glabrous pilei and when dry a wrinkled pileal surface, angular pores, and generative hyphae dominant (or exclusive) in trama, but P. udus has much larger pores (1–2 per mm), smooth and grayish-brown cuticle with pink tints when fresh, and cylindrical to broadly ellipsoid basidiospores (10–15 × 4–6 μm, Núñez and Ryvarden 1995).
In the ITS plus nLSU based phylogenetic analysis (Fig. 1), Polyporus hapalopus nested within the Polyporus sensu lato clade separated from the subclade Melanopus and the genera Favolus and Neofavolus. Polyporus tuberaster (Jacq.) Fr., the generic type of Polyporus, seems close to P. hapalopus in phylogeny but without reliable statistical supports; however, its larger pores (< 2 per mm) and basidiospores (> 10 μm long) distinguish it from P. hapalopus (Núñez and Ryvarden 1995). In morphology, species within the subclade Melanopus often bear coriaceous basidiocarps and black cuticle on the stipe, which differs from Polyporus hapalopus (Dai et al. 2014). Neofavolus is characterized by the solitary basidiocarp (Sotome et al. 2013), which makes it different from P. hapalopus with imbricate basidiocarps. In addition, P. hapalopus has a radially striate pileal surface. Favolus also has a pileal surface with radial striate, but differs from P. hapalopus in its indistinct pileal cuticle (Sotome et al. 2013). In addition, P. hapalopus has a monomitic hyphal system in the trama, while Favolus and Neofavolus have a dimitic hyphal system in both context and trama (Sotome et al. 2013). Therefore, based on the evidence from morphological and phylogenetic perspectives, Polyporus hapalopus is a new member of the genus Polyporus, but its phylogenetic relationship with other species of Polyporus sensu lato is not resolved.
References
Buchanan PK, Ryvarden L (1998) New Zealand polypore fungi (Aphyllophorales): three new species and a new record. N Z J Bot 36:219–231
Cui BK, Yuan HS, Dai YC (2008) Wood-rotting fungi in eastern China 1. Polypores from Wuyi Mountains, Fujian Province. Sydowia 60:25–40
Dai YC (2010) Hymenochaetaceae (Basidiomycota) in China. Fungal Divers 45:131–343
Dai YC (2011) A revised checklist of corticioid and hydnoid fungi in China for 2010. Mycoscience 52:69–79
Dai YC (2012a) Two new polypores from tropical China, and renaming two species of Polyporus and Phellinus. Mycoscience 53:40–44
Dai YC (2012b) Polypore diversity in China with an annotated checklist of Chinese polypores. Mycoscience 53:49–80
Dai YC, Penttilä R (2006) Polypore diversity of Fenglin Nature Reserve, northeastern China. Ann Bot Fenn 43:81–96
Dai YC, Härköene M, Niemelä T (2003) Wood-inhabiting fungi in southern China 1. Polypores from Hunan Province. Ann Bot Fenn 40:381–393
Dai YC, Wei YL, Wang Z (2004) Wood-inhabiting fungi in southern China 2. Polypores from Sichuan Province. Ann Bot Fenn 41:319–329
Dai YC, Cui BK, Huang MY (2007a) Polypores from eastern Inner Mongolia, northeastern China. Nova Hedwigia 84:513–520
Dai YC, Cui BK, Yuan HS (2007b) Notes on polypores from Gansu and Qinghai Province, Northwest China. Cryptogam Mycol 28:177–187
Dai YC, Wei YL, Yuan HS, Huang MY, Penzina T (2007c) Polypores from Altay and Tian Mts. in Xinjiang, northwest China. Cryptogam Mycol 28:269–279
Dai YC, Yu CJ, Wang HC (2007d) Polypores from eastern Xizang (Tibet), western China. Ann Bot Fenn 44:135–145
Dai YC, Yuan HS, Wang HC, Yang F, Wei YL (2009) Polypores (Basidiomycota) from Qin Mts. in Shaanxi Province, Central China. Ann Bot Fenn 46:54–61
Dai YC, Xue HJ, Vlasák J, Rajchenberg M, Wang B, Zhou LW (2014) Phylogeny and global diversity of Polyporus group Melanopus (Polyporales, Basidiomycota). Fungal Divers. doi:10.1007/s13225-013-0248-3
Gilbertson RL, Ryvarden L (1987) North American polypores 2. Fungiflora, Oslo
Guindon S, Gascuel O (2003) A simple, fast and accurate method to estimate large phylogenies by maximum-likelihood. Syst Biol 52:696–704
Ko KS, Jung HS (2002) Phylogenetic evaluation of Polyporus s. str. based on molecular sequences. Mycotaxon 82:315–322
Krüger D, Petersen RH, Hughes KW (2006) Molecular phylogenies and mating study data in Polyporus with special emphasis on group “Melanopus” (Basidiomycota). Mycol Prog 5:185–206
Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ, Higgins DG (2007) Clustal W and Clustal X version 2.0. Bioinformatics 23:2947–2948
Li J, Xiong HX, Dai YC (2008) Polypores from Shennongjia Nature Reserve in Hubei Province, Central China. Cryptogam Mycol 29:267–277
Núñez M, Ryvarden L (1995) Polyporus (Basidiomycotina) and related genera. Synopsis Fungorum 10:1–85
Pattengale ND, Alipour M, Bininda-Emonds ORP, Moret BME, Stamatakis A (2010) How many bootstrap replicates are necessary? J Comput Biol 17:337–354
Petersen JH (1996) Farvekort. The Danish Mycological Society’s colour-chart. Foreningen til Svampekundskabens Fremme, Greve
Posada D (2008) jModelTest: Phylogenetic model averaging. Mol Biol Evol 25:1253–1256
Ronquist F, Teslenko M, van der Mark P, Ayres D, Darling A, Hōhna S, Larget B, Liu L, Suchard MA, Huelsenbeck JP (2012) MrBayes 3.2: Efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol 61:539–542
Silvestro D, Michalak I (2012) raxmlGUI: a graphical front-end for RAxML. Org Divers Evol 12:335–337
Sotome K, Hattori T, Kakishima M (2007) Polyporus phyllostachydis sp. nov. with notes on other rhizophilic species of Polyporus (Basidiomycota, Polyporaceae). Mycoscience 48:42–46
Sotome K, Hattori T, Ota Y, To-anun C, Salleh B, Kakishima M (2008) Phylogenetic relationships of Polyporus and morphologically allied genera. Mycologia 100:603–615
Sotome K, Hattori T, Ota Y, Kakishima M (2009) Second report of Polyporus longiporus and its phylogenetic position. Mycoscience 50:415–420
Sotome K, Hattori T, Ota Y (2011) Taxonomic study on a threatened polypore, Polyporus pseudobetulinus, and a morphologically similar species, P. subvarius. Mycoscience 52:319–326
Sotome K, Akagi Y, Lee SS, Ishikawa NK, Hattori T (2013) Taxonomic study of Favolus and Neofavolus gen. nov. segregated from Polyporus (Basidiomycota, Polyporales). Fungal Divers 58:245–266
Stamatakis A (2006) RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 22:2688–2690
Wang B, Cui BK, Li HJ, Du P, Jia BS (2011) Wood-rotting fungi in eastern China 5. Polypore diversity in Jiangxi Province. Ann Bot Fenn 48:237–246
White TJ, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ (eds) PCR protocols: a guide to methods and applications. Academic Press, San Diego, pp 315–322
Xue HJ, Zhou LW (2012) Polyporus submelanopus sp. nov. (Polyporales, Basidiomycota) from Northwest China. Mycotaxon 122:433–441
Yuan HS, Dai YC (2008) Polypores from northern and central Yunnan Province, Southwestern China. Sydowia 60:147–159
Zheng HD, Liu PG (2005) Type studies on Albatrellus henanensis and A. jianfenglingensis. Mycotaxon 93:257–263
Zhou LW, Xue HJ (2012) Fomitiporia pentaphylacis and F. tenuitubus spp. nov. (Hymenochaetales, Basidiomycota) from Guangxi, southern China. Mycol Prog 11:907–913
Acknowledgments
We express our gratitude to Prof. Yu-Cheng Dai (IFP, China) for his guidance in the taxonomic study and also for his recommendations for improving the manuscript. L.W.Z. thanks Drs. Hai-Sheng Yuan (IFP, China), Shuang-Hui He (BJFC, China) and Wei Li (HMAS, China) for their company in the field trip. Dr. Hai-Sheng Yuan is also thanked for providing the in situ photo of Polyporus hapalopus and preparing the line drawing. The research was financed by National Natural Science Foundation of China (Project Nos. 31200015 & 30910103907) and the Youth Fund for Creative Research Groups, Institute of Applied Ecology, Chinese Academy of Sciences.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Xue, HJ., Zhou, LW. Polyporus hapalopus sp. nov. (Polyporales, Basidiomycota) from China based on morphological and molecular data. Mycol Progress 13, 811–817 (2014). https://doi.org/10.1007/s11557-014-0964-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11557-014-0964-4