Abstract
Objectives
It is with a great prospect to develop an auxiliary diagnosis system for dental periapical radiographs based on deep convolutional neural networks (CNNs), and the indications and performances should be investigated. The aim of this study is to train CNNs for lesion detections on dental periapical radiographs, to evaluate performances across disease categories, severity levels, and train strategies.
Methods
Deep CNNs with region proposal techniques were constructed for disease detections on clinical dental periapical radiographs, including decay, periapical periodontitis, and periodontitis, leveled as mild, moderate, and severe. Four strategies were carried out to train corresponding networks with all disease and level categories (baseline), all disease categories (Net A), each disease category (Net B), and each level category (Net C) and validated by a fivefold cross-validation method afterward. Metrics, including intersection over union (IoU), precision, recall, and average precision (AP), were compared across diseases, severity levels, and train strategies by analysis of variance.
Results
Lesions were detected with precision and recall generally between 0.5 and 0.6 on each kind of disease. The influence of train strategy, disease category, and severity level were all statistically significant on performances (P < .001). Decay and periapical periodontitis lesions were detected with precision, recall, and AP values less than 0.25 for mild level, while 0.2–0.3 for moderate level and 0.5–0.6 for severe level. Net A performed similar to baseline (P > 0.05 for IoU, precision, and recall), while Net B and Net C performed slightly better than baseline under certain circumstances (P < 0.05), but Net C failed to predict mild decay.
Conclusions
The deep CNNs are able to detect diseases on clinical dental periapical radiographs. This study reveals that the CNNs prefer to detect lesions with severe levels, and it is better to train the CNNs with customized strategy for each disease.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The dental periapical radiography is a kind of radiology image that is essential for the diagnosis of dental hard tissue diseases, such as decay, periapical periodontitis, and periodontitis [1,2,3]. Based on the bisecting technique, a dental periapical radiography can shape several (usually 3–4) intact teeth and periodontal structures. However, the clarity of the radiography is strongly relied on the operator’s techniques, including the setting of project angle, radiation time, and doses [4]. Dental periapical radiographs are abundantly produced in daily clinical practice and read by a dentist to generate diagnosis reports, and provide guidance for treatment plans, or evaluate the outcomes. The reading and analyzing work consumes much mental effort and takes a substantial amount of time from the dentist, and radiographic interpretations tend to be with a variation between observers [3, 5, 6]. Moreover, there are misdiagnosis of non-endodontic lesions as periapical periodontitis lesions [7]. So, it is with a great prospect to develop an auxiliary diagnosis system for dental periapical radiographs.
There are kinds of algorithms applied in dental X-ray image processing, feature extraction, and segmentation, such as contour extraction [8, 9], adaptive threshold [10,11,12], iterative thresholding [13], level set [14, 15], mathematical morphology [16], Fourier descriptors [17], hierarchical contour matching [18], weighted Hausdorff distance [19], texture statistics techniques [20], local singularity analysis [21], semi-supervised fuzzy clustering algorithms [22, 23], neutrosophic orthogonal matrices [24], and so on. However, image features of target objects were always shallowly extracted in these algorithms and rely heavily upon manual definitions, which may raise issues of artificial errors.
With the development of artificial intelligence techniques, support vector machine (SVM) [25] and artificial neural networks [26,27,28,29] were introduced to detect lesions on dental images, which achieved better results compared to “rule-based” computer algorithms. Li et al. [25] used principal component analysis to extract pathological characteristics of the clinical images to train a SVM classifier, achieving automatic and fast clinical segmentations. Yu et al. [26] proposed a three-layer neural network in which normalized autocorrelation coefficients were treated as input features, and back-propagation algorithm was used to construct the weights of classifier to distinguish decayed teeth from normal. El-Bakry et al. [27] trained a neural network to classify sub-images which contain dental diseases or not and constructed a fast algorithm for dental disease detection by performing cross-correlation in the frequency domain between input image and the input weights of the neural networks. Tumbelaka et al. [28] used local image differentiation technique to extract edges as basis image features and then analyzed them by texture descriptors to obtain image entropy, which was further sent to artificial neural networks to detect the infected regions. However, in these researches, image features should be manually defined and pre-calculated before sent to classifiers like SVM or neural networks.
In recent years, deep convolution neural networks (CNNs) have been invented which can take the raw image data as input and did a good job on image classifications and object detections [30]. Srivastava et al. [29] constructed a deep fully convolutional neural network to mark caries on bitewing radiographs with precision reported to be 61.5 and recall 81.5. Al Kheraif et al. [31] performed teeth and bone segmentation work on panoramic radiographs using hybrid graph-cut technique and convolutional neural network. Lee et al. [32] implemented decay classification based on GoogLeNet Inception v3 CNN network, while the teeth for detection should be manually segmented before sent to neural networks, and the position of decay lesions was not precisely located in their research. Ekert et al. [33] applied deep convolutional neural networks (CNNs) to detect apical lesions on panoramic dental radiographs, but the CNN’s sensitivity needs to be improved before clinical application. Lately, regions with convolutional neural network (R-CNN) features [34] were developed to offer solutions for object detection tasks, where target objects (regions of interest) were automatically boxed out and annotated with labels. R-CNN was then upgraded to fast R-CNN [35] and furtherly to faster R-CNN [36], with higher efficiency and better performances. Although some researchers reported teeth segmentations on panoramic dental radiographs based on faster R-CNN [37], there were rarely researches applying R-CNNs in disease lesion detection on dental periapical radiographs.
In this research, faster R-CNN was utilized to detect decay, periapical periodontitis, and periodontitis in dental periapical radiographs. Influences of network train strategies, as well as disease categories and levels of severity, on the detection outcomes were invested, with intentions to figure out to what extent the faster R-CNN will perform in disease detection in dental X-ray, and to find out what kind of diseases and which levels will be detected with higher accuracy, i.e., the indications of this deep CNN auxiliary diagnosis methods.
Methods
Data collection and annotation
In total, 2900 digital dental periapical radiographs were collected. The inclusion criteria are (1) periapical radiographs with permanent teeth, (2) the radiation exposure is proper, and (3) the position and axial direction of teeth are proper. The exclusion criteria are (1) radiographs with deciduous teeth in it, (2) the image is too bright or too dark to distinguish the lesions, and (3) the teeth in the image is severely distorted. Each digital radiography was exported with a resolution of 96 dpi at size of approximately (300–500) × (300–400) pixels and saved as a “JPG” format image file with a unique identification code. These image files were collected anonymously to ensure that no private information, such as patient name, gender, or age, is revealed. Afterward, an expert dentist with more than 5 years of clinical experience draws minimum bounding boxes to frame each diseased area of decay, periapical periodontitis (labeled as periapi, for short), and periodontitis with bone resorptions (labeled as periodo, for short). Each type of disease was graded by three levels of severity, which are mild, moderate, and severe. Thus, a total of nine label names for bounding boxes were annotated: decay-mild, decay-moderate, decay-severe, periapi-mild, periapi-moderate, periapi-severe, periodo-mild, periodo-moderate, and periodo-severe. The criteria are as follows:
-
Decay-mild: the decay invasion depth less than 1/3 of the tooth sidewall or roof width;
-
Decay-moderate: the decay invasion depth between 1/3 and 1/2 of the tooth sidewall or roof width.
-
Decay-severe: the decay invasion depth larger than 1/2 of the tooth sidewall or roof width;
-
Periapi-mild: the width of the periapical periodontitis area (the miner axis) less than 1 mm;
-
Periapi-moderate: the width of the periapical periodontitis area (the miner axis) between 1 and 3 mm;
-
Periapi-severe: the width of the periapical periodontitis area (the minor axis) larger than 3 mm;
-
Periodo-mild: the bone resorption depth less than 1/3 of the tooth root length;
-
Periodo-moderate: the bone resorption depth between 1/3 and 1/2 of the tooth root length;
-
Periodo-severe: the bone resorption depth larger than 1/2 of the tooth root length;
The coordinates of points in the image were set as pixel distance from image’s left top corner, where the tooth bounding box could be recorded by its top left and bottom right corner points (xmin, ymin, and xmax, ymax).
Train and validation of faster R-CNNs
An object detection tool package [38] based on TensorFlow was utilized to construct faster R-CNN, which was one of the state-of-the-art object detectors for multiple categories. The training process was executed on a GPU (Quadro RTX 8000, NVIDIA, USA), with 48 GB memory and 4608 CUDA cores. The algorithms were running backend on TensorFlow version 1.13.1, and the operating system was Ubuntu 18.04. The training parameters were configured as: anchor scales [0.1, 0.2, 0.4, 0.8, 1.6], iterations 100,000, initial learning rate 0.003 and then reduced to 0.0003 after 30,000 iterations, and further to 0.00003 after 60,000 iterations. Also, a pre-trained model on the Coco dataset, version 2018-01-28, was loaded as a fine-tune checkpoint.
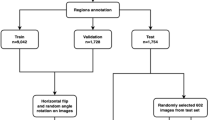
Faster R-CNN was trained and validated by several strategies of a different organization of annotated data (Table 1). Firstly, all annotated images with nine label names were treated as ground truth and were used to train and validate the faster R-CNN network as a baseline. Secondly, the level attribution of each bounding box (bbox) annotations was ignored, that is, decay-mild, decay-moderate, and decay-severe were all relabeled to be decay, same to periapi and periodo, to train and validate another fast R-CNN named Net A. After that, three faster R-CNNs (named Net B1, B2, and B3), one for each of the three disease name classifications, were trained, respectively. Similarly, C1, C2, and C3 were trained for each level of bboxes. Fivefold cross-validation was applied for every network listed in Table 1, where all included images were randomly and evenly divided to be five parts. The train and validation processes were run for five turns with every part used as validation dataset, and the rest four parts as train dataset. Before each run, the trained parameters were initialized or re-initialized to “forget” the trained memories and the networks were “renewed.”
Several metrics were calculated on validation dataset for each label name, including intersection over union (IoU), precision, recall, average precision (AP, also equal to area under curve, AUC). Metrics calculated from the fivefold cross-validation procedure were combined to calculate a mean value.
Firstly, the predicted bboxes were compared with ground truth bboxes, and IoU is defined as:
where \(\hbox{Area}_{\rm pred}\) and \(\hbox{Area}_{\rm gt}\) represent the areas of the predicted bbox and its corresponding ground truth bbox. The threshold of IoU was set to be 0.5, that is, if a predicted bbox whose IoU with corresponding ground truth bbox is larger than 0.5, it will be treated as true positive bboxes. Then precision and recall could be calculated:
where TP represents the count of true positive bboxes, while Pred is the count of predicted bboxes and GT is the count of ground truth bboxes. It can be easily inferred that precision defined here is equivalent to positive predictive value in clinical diagnosis, and recall is equivalent to sensitivity.
Receiver operating characteristic (ROC) curve is an important metric to evaluate diagnosis tools. However, the calculation of ROC curves relies on counts of true positive samples, true negative samples, false positive samples, and false negative samples. But negative samples are not applicable here in this kind of object detection tasks, because no bonding box has been drawn on negative targets (areas without disease lesions) by ground truth. Thus, ROC curves cannot be drawn. Instead, we calculated precision–recall curves in this study, which have deep connection with receiver operator characteristic curves; both can evaluate the accordance between test and reference [40]. The area under precision–recall curve was calculated as average precision (AP), which is widely used as an important metric of the performances of networks in object detection tasks. For each label name, the bboxes were predicted by faster R-CNN with confidence scores. A threshold of confidence score will be set to decide which of the predicted bboxes to finally output. If the confidence score of one predicted bbox was set as threshold, predicted bboxes whose confidence scores larger than the threshold will be finally output and matched with ground truth bboxes to produce a precision and a recall value. After every confidence score of predicted bboxes set as threshold, a series of precision and recall value pairs will be produced to draw a P–R curve. Thus, average precision (AP) [39] is defined as the area under smoothed P–R curve:
where \({p}_{{\rm interp}}(r)\) is the maximum precision for any recall values exceeding \(r\):
Statistical analysis
To evaluate the performance of each network on diseases and levels, analysis of variance (ANOVA) was used. Firstly, the performances of baseline were compared with Net A across all three diseases. Since there was no level attribution of bboxes output from Net A, the level attributions of predicted bboxes by baseline were ignored for comparison, that is, decay-mild, decay-moderate, and decay-severe were all treated as decay, also to periapi and periodo. Two-way ANOVA was applied with strategy names and disease names set as independent variables, and metrics calculated on validation dataset including IoU, precision, recall, and AP were set as dependent variables. Secondly, the performances of baseline were compared with Net B (composed of B1, B2, B3) and Net C (composed of C1, C2, C3). Multi-way ANOVA was applied, where strategy names, disease names, and level names were set as independent variables, and metrics calculated on validation dataset including IoU, precision, recall, and AP were set as dependent variables.
A statistical software program (IBM SPSS Statistics, v19.0; IBM Corp) was used for the statistical analysis. For those where the interactions of independent variables were significant, simple effects were analyzed with pairwise comparisons adjusted by the Bonferroni’s method (α = 0.05 for all tests).
Results
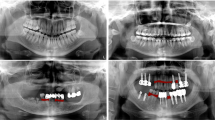
As shown in Fig. 1, although with some miss diagnosis, the diseases detected by faster R-CNNs were basically close to the ground truth. Performances of baseline and Net A were compared, and the metrics calculated on validation dataset are shown in Table 2. Two-way ANOVA results (Table 3) have shown that the strategy had no significant influence on all metrics, except AP, while the disease had a significant influence on all metrics. The interaction of strategy and disease had no significant influence on all metrics, except precision. Non-statistically significant interactions were removed from the analysis model, and \(F\) values as well as \(P\) values were re-estimated and shown in brackets. Otherwise, if interactions between factors were significant, simple effects were analyzed based on estimated marginal means. Factors with significant influence on according metrics were forwarded to pairwise comparisons adjusted by the Bonferroni’s method. Values of metrics and results of pairwise comparisons are illustrated in Fig. 2, where values with significant differences were annotated by different letters. As it can be seen, with comparison between different diseases, decay tends to be predicted with higher IoU, precision, and AP than periapi or periodo, while periodo tends to be predicted with higher recall (Fig. 2a). With comparison between different strategies (networks), Net A performed as good as baseline and even slightly better than baseline on AP value for prediction of periapi (Fig. 2b).
Samples of dental periapical radiographs with lesions detected by networks constructed and trained in this research. Ground truth was manual annotations of an expert dentist. The detections shown in baseline were output from neural network trained by all disease categories and all severity levels. The detections shown in Net A were output from neural network trained by all disease categories, ignoring severity levels. The detections shown in Net B were combination of detections from Net B1, Net B2, and Net B3, where decays were detected by Net B1, periapical periodontitis was detected by Net B2, and periodontitis was detected by Net B3. The detections shown in Net C were combination of detections from Net C1, Net C2, and Net C3, where mild-level diseases were detected by Net C1, moderate-level diseases were detected by Net C2, and severe-level diseases were detected by Net C3
Performances of baseline and Net B, C were also compared, and the metrics calculated on validation dataset are shown in Table 4. Multi-way ANOVA results (Table 5) show that all independent variables, as well as their interactions, had significant influences on all metrics. Simple effects were analyzed, and further pairwise comparisons were processed based on estimated marginal means and adjusted by the Bonferroni’s method. The values of metrics and the results of pairwise comparisons are illustrated in Fig. 3, 4, 5. As it can be seen, with comparison between different diseases on severe level, decay tends to be predicted with precision, recall, and AP values higher than periapi, and periapi tends to be higher than periodo, but the order was reversed on mild and moderate levels (Fig. 3). Mild decay tended to be predicted with lower IoU than mild periapi and mild periodo (Fig. 3). With comparison between different Levels, severe level tended to be predicted with precision, recall, and AP values higher than moderate level, and moderate level tended to be higher than mild level, particularly for decay and periapi (Fig. 4). With comparison between different strategies (networks), Net B and Net C performed better than baseline on certain circumstances, but Net C failed to predict mild decay (Fig. 5).
Discussion
Metrics included in this research have their clinical significances. As in clinical use, the overlapping rate between predicted disease areas and ground truth must reach a certain level that is beneficial for the dentist to position the potential disease. The IoU, which is defined as an overlapping area over the union area, can measure the precision of allocation of target diseases. The larger the IoU is, the more precise the location of target disease, and predicted target will completely overlap with target disease when IoU reaches 1. The IoU is always high. However, high IoU is not directly related to the diagnostic performance to predict either the presence/absence or severity of target diseases. Precision defined in this research is equivalent to positive predictive value in clinical diagnosis, and recall is equivalent to sensitivity. Thus, precision here represents the chance that a predicted disease bbox truly has the disease within it, and recall represents the probability that a prediction will indicate “disease” among those with the disease. AP, which is defined as the area under the precision–recall curve, can test the overall performances of the network, and the closer the AP reaches 1, the better the network model is. What’s more, precision–recall curves have a deep connection with receiver operating characteristic curves [40], both of which are able to evaluate the accordance between test and reference. However, when dealing with highly skewed datasets where the class distribution is not even, precision–recall (P–R) curves give a more informative picture of an algorithm's performance [40]. Precision, recall, and average precision did not show high performances, which implied the difficulty of correctly detection of dental lesions (Tables 2, 4).
As shown in the results, many values of precision, recall, and AP were less than 0.5, which is the random chance of two-category classifications. But the disease detection task here is not only to classify the multicategory disease lesions, but also to detect the actual position and size of the lesion, and the performances were the overall accuracy. Think of a small target square area like the mild decay in the dental radiography image, if determined its position and size randomly, the chance will be very small (about 0) to correctly match with the truth (IoU larger than 0.5), let alone the subsequent chance of multicategory classifications. Although the overall performance of CNNs has many values less than 0.5, they are still better than chance.
Different strategies and networks were designed in this research, and their influence on metrics was tested. Net A was designed to ignore levels for disease detection, which is reasonable for basic clinical applications, because usually we only need to know whether there are certain disease lesions or not on dental X-rays, and further manual examinations will be processed to determine the level of detected diseases. But we still want to figure out whether it will improve the recognition of disease names for the deep CNNs if we teach machine more details of the disease, such as disease levels here. However, the results of comparison between Net A and baseline show that, if only disease names were wanted (i.e., the levels of disease are not needed to output from network), baseline trained with extra disease level information performed no better than Net A with only disease name information. It can be inferred that there is no need to annotate objects with extra attributions other than what we need the deep CNNs to output.
As in baseline, we trained only one faster R-CNN network for all diseases and levels, but in Net B, we trained one faster R-CNN for each disease name, and in Net C, we trained one faster R-CNN for each level name. Thus, although Net B and Net C performed slightly better than baseline in certain circumstances, they were three times of baseline in scales of model parameters. Other than that, Net C failed to detect mild decay, which might because the differences of features between diseases within the same level were more obvious than differences of features between levels within the same disease. So, overall, Net B with trained faster R-CNN for each disease name performed better than baseline and Net C, but will cause more overheads in computation and memory than baseline.
Metrics were also compared among diseases and levels, with intention to find out the indications of faster R-CNNs in this research. The results turned out that periodontitis can be well detected among all levels, while decay and periapical periodontitis were better predicted with an increase in severity. Decay and periapical periodontitis with moderate or severe levels were in much larger scales and were more visually distinctive than mild level, and the small size objects are easy to be ignored after downsampling in faster R-CNN processes. Thus, disease lesions with too small sizes may not be indications for faster R-CNN. On the other hand, the distribution of lesions among different disease categories and severity levels is uneven in this study, which could also affect the performances of networks. There is also a trend of poorer performances with less train samples, because the networks need to be trained with enough amount of ground truth cases before they can correctly predict disease lesions. However, imbalance distribution of lesion counts across diseases and levels is the actual situation in real clinical dental radiographs. Thus, when it comes to the clinical application, radiographs should better be screened from the clinical images to form an evenly distributed train dataset.
The performances of current networks were poorer than Srivastava et al. [29] with precision reported to be 61.5 and recall 81.5 on bitewing images. This might because the ground truth used by Srivastava et al. for existence of caries was radiographic interpretation by dentists combined with clinical verification, and they did not specify the severity or size of the carious lesions. Although the performances were not sufficient to be used as diagnostic tools alone, it can prompt dentist with potential disease lesions, which is beneficial to improve the efficiency of clinical work. Furtherly, the structure of deep CNNs should be updated and some pre-image or post-image processing techniques should be carried out to improve the performances of disease detection on dental periapical radiographs, which will be our next work.
Conclusions
Some conclusions can be drawn with the constructed faster R-CNNs:
-
1.
The faster R-CNNs were able to detect diseases including decay, periapical periodontitis, and periodontitis in dental periapical radiographs.
-
2.
The network train strategy, disease category, and severity level all have significant influences on performances of faster R-CNNs.
-
3.
It is better to train one faster R-CNN for each disease classification, rather than training only one faster R-CNN for all disease classifications. However, training one R-CNN for each severity level is discouraged, because there tends to be a drawback of performances.
-
4.
Decays and periapical periodontitis with higher severity tend to be better predicted than lesions with lower severity.
-
5.
In mild and moderate levels, periodontitis can be better detected than periapical periodontitis, and periapical periodontitis better than decay, but the rank was reversed in severe levels.
References
American Dental Association Council on Scientific Affairs (2006) The use of dental radiographs: update and recommendations. J Am Dent Assoc 137(9):1304–1312
Rohlin M, Kullendorff B, Ahlqwist M, Henrikson CO, Hollender L, Stenström B (1989) Comparison between panoramic and periapical radiography in the diagnosis of periapical bone lesions. Dentomaxillofac Radiol 18(4):151–155
Douglass CW, Valachovic RW, Wijesinha A, Chauncey HH, Kapur KK, Mcneil BJ (1986) Clinical efficacy of dental radiography in the detection of dental caries and periodontal diseases. Oral Surg Oral Med Oral Pathol 62(3):330–339
Gupta A, Devi P, Srivastava R, Jyoti B (2014) Intra oral periapical radiography-basics yet intrigue: a review. Bangladesh J Dent Res Educ 4(2):83–87
Kaffe I, Gratt BM (1988) Variations in the radiographic interpretation of the periapical dental region. J Endod 14(7):330–335
Valachovic RW, Douglass CW, Berkey CS, Mcneil BJ, Chauncey HH (1986) Examiner reliability in dental radiography. J Dent Res 65(3):432–436
Sirotheau Corrêa Pontes F, Paiva Fonseca F, Souza De Jesus A, Garcia Alves AC, Marques Araújo L, Silva Do Nascimento L, Rebelo Pontes HA (2014) Nonendodontic lesions misdiagnosed as apical periodontitis lesions: series of case reports and review of literature. J Endod 40(1):16–27
Jain AK, Chen H (2004) Matching of dental X-ray images for human identification. Pattern Recogn 37(7):1519–1532
Shah S, Abaza A, Ross A, Ammar H (2006) Automatic tooth segmentation using active contour without edges. In: IEEE biometric consortium conference, biometrics symposium: special session on research, pp 1–6
Nomir O, Abdel-Mottaleb M (2005) A system for human identification from X-ray dental radiographs. Pattern Recogn 38(8):1295–1305
Zhou J, Abdel-Mottaleb M (2005) A content-based system for human identification based on bitewing dental X-ray images. Pattern Recogn 38(11):2132–2142
Razali MRM, Ismail W, Ahmad NS, Bahari M, Zaki ZM, Radman A (2017) An adaptive thresholding method for segmenting dental X-ray images. J Telecommun Electron Comput Eng (JTEC) 9(4):1–5
Lin PL, Lai YH, Huang PW (2010) An effective classification and numbering system for dental bitewing radiographs using teeth region and contour information. Pattern Recogn 43(4):1380–1392
Li S, Fevens T, Krzyżak A, Jin C, Li S (2007) Semi-automatic computer aided lesion detection in dental X-rays using variational level set. Pattern Recogn 40(10):2861–2873
Rad AEAR (2018) Automatic computer-aided caries detection from dental x-ray images using intelligent level set. Multimed Tools Appl 77(21):28843–28862
Said EH, Nassar DEM, Fahmy G, Ammar HH (2006) Teeth segmentation in digitized dental X-ray films using mathematical morphology. IEEE Trans Inf Forensics Secur 1(2):178–189
Nomir O, Abdel-Mottaleb M (2007) Human identification from dental X-ray images based on the shape and appearance of the teeth. IEEE Trans Inf Forensics Secur 2(2):188–197
Nomir O, Abdel-Mottaleb M (2008) Hierarchical contour matching for dental X-ray radiographs. Pattern Recogn 41(1):130–138
Lin P, Lai Y, Huang P (2012) Dental biometrics: human identification based on teeth and dental works in bitewing radiographs. Pattern Recogn 45(3):934–946
Rad AE, Shafry M, Rahim M, Norouzi A (2013) Digital dental X-Ray image segmentation and feature extraction. Telkomnika Indones J Electr Eng 11(6):3109–3114
Lin PL, Huang PY, Huang PW, Hsu HC, Chen CC (2014) Teeth segmentation of dental periapical radiographs based on local singularity analysis. Comput Methods Programs Biomed 113(2):433–445
Son LH, Tuan TM (2016) A cooperative semi-supervised fuzzy clustering framework for dental X-ray image segmentation. Expert Syst Appl 46(Supplement C):380–393
Tuan TM et al (2017) Dental segmentation from X-ray images using semi-supervised fuzzy clustering with spatial constraints. Eng Appl Artif Intell 59:186–195
Ali M, Khan M, Tung NT et al (2018) Segmentation of dental X-ray images in medical imaging using neutrosophic orthogonal matrices. Expert Syst Appl 91:434–441
Li S, Fevens T, Krzyżak A, Li S (2006) Automatic clinical image segmentation using pathological modeling, PCA and SVM. Eng Appl Artif Intell 19(4):403–410
Yu Y, Li Y, Li Y, Wang J, Lin D, Ye W (2006) Tooth decay diagnosis using back propagation neural network. In: IEEE international conference on machine learning and cybernetics, pp 3956–3959
El-Bakry HM, Mastorakis N (2008) An effective method for detecting dental diseases by using fast neural networks. WSEAS Trans Biol Biomed 11:293–301
Tumbelaka B, Oscandar F, Baihaki F, Sitam S, Rukmo M (2014) Identification of pulpitis at dental X-ray periapical radiography based on edge detection, texture description and artificial neural networks. Saudi Endod J 4(3):115–121
Srivastava MM, Kumar P, Pradhan L, Varadarajan S (2017) Detection of tooth caries in bitewing radiographs using deep learning. arXiv preprint arXiv:1711.07312.
Krizhevsky A, Sutskever I, Hinton GE (2012) ImageNet classification with deep convolutional neural networks. In: Advances in neural information processing systems, pp 1097–1105
Al Kheraif AA, Wahba AA, Fouad H (2019) Detection of dental diseases from radiographic 2d dental image using hybrid graph-cut technique and convolutional neural network. Measurement 146:333–342
Lee J, Kim D, Jeong S, Choi S (2018) Detection and diagnosis of dental caries using a deep learning-based convolutional neural network algorithm. J Dent 77:106–111
Ekert T, Krois J, Meinhold L, Elhennawy K, Emara R, Golla T, Schwendicke F (2019) Deep learning for the radiographic detection of apical lesions. J Endod 45(7):917–922
Girshick R, Donahue J, Darrell T, Malik J (2014) Rich feature hierarchies for accurate object detection and semantic segmentation. In: Proceedings of the IEEE conference on computer vision and pattern recognition, pp 580–587
Girshick R (2015) Fast r-cnn. In: Proceedings IEEE international conference on computer vision, pp 1440–1448
Ren S, He K, Girshick R, Sun J (2015) Faster r-cnn: towards real-time object detection with region proposal networks. In: Advances in neural information processing systems, pp 91–99
Laishram A, Thongam K (2020) Detection and classification of dental pathologies using faster-RCNN in orthopantomogram radiography image. Int Conf Signal Process Integr Netw (SPIN) 7:423–428
Huang J, Rathod V, Sun C, Zhu M, Korattikara A, Fathi A, Fischer I, Wojna Z, Song Y, Guadarrama S et al (2017) Speed/accuracy trade-offs for modern convolutional object detectors. In: Proceedings IEEE conference on computer vision and pattern recognition, pp 7310–7311
Everingham M, Van Gool L, Williams CKI, Winn J, Zisserman A (2010) The Pascal visual object classes (VOC) challenge. Int J Comput Vision 88(2):303–338
Davis J, Goadrich M (2006) The Relationship between precision–recall and ROC curves. In: Proceedings 23rd international conference on machine learning, pp 233–240
Funding
This study was funded by the National Natural Science Foundation of China (No. 51705006), Program for New Clinical Techniques and Therapies of Peking University School and Hospital of Stomatology (No. PKUSSNCT-19A08), and open fund of Shanxi Province Key Laboratory of Oral Diseases Prevention and New Materials (KF2020-04).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Declaration of Helsinki and its later amendments or comparable ethical standards.
Informed consent
For this type of study, formal consent is not required.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Chen, H., Li, H., Zhao, Y. et al. Dental disease detection on periapical radiographs based on deep convolutional neural networks. Int J CARS 16, 649–661 (2021). https://doi.org/10.1007/s11548-021-02319-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11548-021-02319-y