Abstract
In neurosurgery, cranial incisions during craniotomy can be recovered by cranioplasty—a surgical operation using cranial implants to repair skull defects. However, surgeons often encounter difficulties when grafting prefabricated cranial plates into defective areas, since a perfect match to the cranial incision is difficult to achieve. Previous studies using mirroring technique, surface interpolation, or deformed template had limitations in skull reconstruction to match the patient’s original appearance. For this study, we utilized low-resolution and high-resolution computed tomography images from the patient to repair skull defects, whilst preserving the original shape. Since the accuracy of skull reconstruction was associated with the partial volume effects in the low-resolution images and the percentage of the skull defect in the high-resolution images, the low-resolution images with intact skull were resampled and thresholded followed by active contour model to suppress partial volume artifacts. The resulting low-resolution images were registered with the high-resolution ones, which exhibited different percentages of cranial defect, to extract the incised cranial part. Finally, mesh smoothing refined the three-dimensional model of the cranial defect. Simulation results indicate that the reconstruction was 93.94% accurate for a 20% skull material removal, and 97.76% accurate for 40% skull material removal. Experimental results demonstrate that the proposed algorithm effectively creates a customized implant, which can readily be used in cranioplasty.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
In clinical neurosurgery, neurological surgeons commonly perform a craniotomy to control elevated intracranial pressure, caused by brain trauma or stroke [10, 18, 24, 26]. Once the acute cause of raised intracranial pressure is resolved, cranial incisions made during the craniotomy may be repaired by cranioplasty, a surgical operation using cranial implants to replace the removed bone section, and repair the skull defect. Cranioplasty uses material from autograft, allograft, xenograft, or uses synthetic material, not only to protect intracranial structures, but also serves an esthetic purpose. To best fit the hole on the skull and recover pre-operative head shape, the customized metal, and plastic bone substitutes need to match the incision as closely as possible. Patients will feel more comfortable if post-operative shape resembles the original appearance.
Previous literature reports fall into three method categories addressing the issue of modeling skull reconstruction. The most popular one is the mirroring technique that duplicates the healthy counterpart section from the opposite side of the skull, as a model for the mold. Although this method has many applications [13, 14, 19, 28], it uses the assumption of the ideal bilateral symmetry of the human skull [25]. Besides, mirroring is suitable only for the unilateral damage, and is unable to deal with bilateral defects. The second class of method uses surface interpolation, such as a Bézier surface [5], and radial basis function [4], to approximate the cranial hole and reconstruct a fitting surface. The major limitation of these two methods is that only a mathematical approximation is applied, instead of making use of the actual anatomical structure. To improve upon the previous two categories, a third approach involves the construction of a template and applying deformation to approximate an appropriate profile. The template (or reference model) can be either selected from a database [27], or constructed from the mirroring or an average skull [7]. A warping or deformation algorithm is employed based on anatomical landmarks on the patient’s, and template images [7, 27]. However, this method requires manually selected landmarks and the deformation process proves to be computationally expensive.
We propose an alternative approach to recover the impaired skull material based on the subject’s own anatomical structure of the incised section, rather than using mirroring duplication, mathematical models, or artificial template deformation. The proposed method uses information from the subject’s intact, and defective cranial anatomy. The former is acquired from clinical, diagnostic low-resolution computed tomography (CT) images, and the latter acquired from a high-resolution CT image, generally used for surgical planning purposes. The slice thickness of the clinical diagnostic images was about 5 mm, limited by in-plane resolution due to scattered radiation. The slice thickness of the high-resolution images was about 0.5 mm for the cranioplasty. The low-resolution CT image for the intact skull was used to provide the general appearance of the incised cranial section, whereas the high-resolution defective image was used to register with the low-resolution image, so that the incised part could be localized and extracted for skull reconstruction. We used the low-resolution diagnostic images to extract the anatomical structure of the cranial defect for skull reconstruction, for the following two reasons. Firstly, patients with skull defects may not receive a prior high-resolution CT image scan, due to an accidental cranial trauma or emergency craniotomy, and only the low-resolution diagnostic image would be available for skull reconstruction, if the patient had ever a medical examination by brain CT images before. Secondly, the radiation dose for high-resolution brain CT images is significant, and patients receiving two such scans in a short period of time (immediately before and after craniotomy) increase their risk of induced cancers caused by the radiation exposure.
The low-resolution image exhibited partial volume effects that require suppression using interpolation processes, thresholding and active contour methods before skull reconstruction can begin. Specifically, the interpolation process along the slice-selection direction is applied to match the voxel size of the high-resolution image. We performed image thresholding to extract skull content, whilst ignoring the soft tissue detail. The active contour method, also known as “snakes”, was in turn used to eliminate the penumbra on the low-resolution skull surface. Snakes were first introduced by Kass et al. [11], and modified by Cohen [6]; the method minimizes energy curves with piecewise smoothing properties on the edge of the target. Many related applications of snakes are reported in the literature, such as Kozerke et al. [12] using snakes to define vessel regions, and Yushkevich et al. [29] segmenting out the subcortical structure in brain images. The high-resolution defective skull images, which were much less affected by the partial volume effect, were directly created from the thresholding.
The low- and high-resolution skull images were registered to extract the incised cranial portion. Image registration is a process of bringing images into alignment, for comparison or integration of information by intensity- or feature-based metrics [9]. Compared to an intensity-based method, such as using mutual information [17], or correlation ratio [23], as the cost function computed from whole volume content, the feature-based method is more computationally efficient from using points [2], edges [3], or surfaces [21]. In this study, registration used the binarized, extracted skulls as the registration feature. The use of skull-based image registration has the advantage that soft tissue, which can be deformed due to the change of cranial pressure following craniotomy [8], is excluded, thus ensuring the validity of rigid-body transformation from the diagnostic, to defective images. Therefore, only six parameters are required to produce a computationally efficient registration.
2 Methods
The proposed algorithm for reconstruction of the cranial defect consisted of the following steps: resampling, thresholding, active contour model, registration, difference operations, three-dimensional (3D) model reconstruction, and mesh smoothing. Figure 1 displays a flow chart of the proposed algorithm. First, the low-resolution diagnostic CT image is resampled so that the resolutions of the resampled image and the high-resolution defective image are the same. Second, both low- and high-resolution images are thresholded to extract the skull. Third, we apply the snake algorithm on the diagnostic skull image to remove artifacts on the skull contour. Fourth, the two images are registered followed by a difference operation to extract the defective area. Sixth, the marching cube algorithm is applied to construct a 3D model of the cranial defect. Finally, Laplacian smoothing is applied to the mesh to refine the model. A series of computer simulations is conducted, to validate the proposed algorithm for defective skull reconstruction.
2.1 Image acquisition
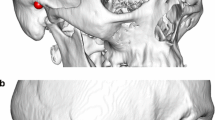
A 52-year-old male was informed of radiation issues, and completed a consent form before participating in this study. The first diagnostic scan was performed with a LightSpeed 16 CT scanner (GE Healthcare, Inc.) at National Taiwan University Hospital, Taipei, Taiwan. Axial images were acquired for diagnosis, with a matrix size of 512 × 512, in-plane resolution of 0.47 mm × 0.47 mm, and 5-mm-thick contiguous slices (Fig. 2a). After 6 weeks, the subject underwent craniotomy on the right side of skull and was scanned a second time. Axial data were acquired for cranioplasty using the same model CT scanner as the initial scans, at Chang Gung Memorial Hospital, Taoyuan, Taiwan. The high-resolution volumetric image, i.e., defective image, consisted of 512 × 512 × 409 voxels with the voxel dimension 0.49 mm × 0.49 mm × 0.63 mm (Fig. 2b).
The second data set was collected from a 52-year-old female with a skull defect on the right side. She filled out a consent form after being informed of radiation issues. Data were acquired with an Aquilion 64 CT scanner (Toshiba Medical Systems, Inc.) at Chang Gung Memorial Hospital, Taoyuan, Taiwan, and the field of view was 246 mm, with a 512 × 512 matrix and 0.5-mm-thick contiguous slices. The raw data were reconstructed into low-resolution images with 5-mm slice thickness (Fig. 2c) and high-resolution images with 0.6-mm slice thickness (Fig. 2d), which had similar resolution to clinical data (Table 1). Since the subject’s skull defect was on the right hemisphere, we first created intact low- and high-resolution skull images by flipping the left-hemisphere skull to the right for subsequent simulations.
2.2 Image pre-processing
Trilinear interpolation was applied to the diagnostic image to produce a resampled image with the same resolution as that of the defective image. In order to reduce computation for subsequent registration processes, we cropped both image volumes to limit their field of view, covering only areas above the orbitomeatal plane. Since the skull exhibited much higher intensity in brain CT images, we segmented it out by thresholding and used the binarized images (1 for skull and 0 for background) as the matching feature. The threshold was empirically determined to a value close to the lower bound of the CT number (Hounsfield units) for bone.
2.3 Active contour model
The penumbra of skull content in the resampled and binarized low-resolution image presented the partial volume effect near the top of volume images (white arrows in Fig. 2a, c). To remove these artifacts, the active contour model was used to smooth the inner and outer edges of skull for each slice along the coronal and sagittal directions, referred to as ‘inner–outer active contour’ hereafter. Before we applied the active contour method, a closed profile was generated by flipping the upper half portion of the image to facilitate initialization of active contours (Fig. 3a, b). A contour within the middle of the white area was initialized to extract the inner edge of the skull (the blue curve in Fig. 3c). The initial contour was stretched and smoothed piecewise, until it stopped at the concave corners of skull boundary (the blue curve in Fig. 3d). Similarly, we placed the initial contour outside the white area to extract the outer edge of the skull (the red curve in Fig. 3c). In order to encapsulate the recessed profile, rather than the bumped profile (the artifacts) for the outer edge of the skull, image erosion was first executed, followed by applying the active contour method (the red curve in Fig. 3d). Once the inner and outer curves were identified, the regions outside these two curves were considered as artifacts, and removed accordingly (see Fig. 3e). Finally, we replaced the upper portion of original image, with the upper portion of the resulting image (see Fig. 3f).
The steps of the active contour method to remove the artifacts on the top half of the skull images. a Original skull, b a closed profile by flipping the upper half portion of the image, c inner (blue) and outer (red) contours are initialized, d resulting contours after active contour method, e regions outside these two contours are removed, f resulting skull after replacing the upper portion of original skull. (Color figure online)
2.4 Image registration
The alignment of the low-resolution intact skull and high-resolution defective skull images provides a model of the skull structure of the removed cranial portion. We defined the reference image, R, as the image that remained unchanged during the registration process, and the floating image, F, as the transformed image based on the reference image. The defective image was assigned as the reference, and the diagnostic image was the floating image in our implementation. All skull voxels used in the computation of the cost function, which we expressed in the form of the sum of squared differences (SSD). This SSD is reliable for aligning two images with different resolutions and content removal [15]. The binary maps of floating and reference images were denoted as F B and R B, respectively. We express the cost function as:
where,
More specifically, we transformed each skull voxel at position x on F B into R B via the transformation T, and computed the intensity difference, d, between the two binary maps of floating and reference images. The trilinear interpolation was adopted to obtain the values on the non-integer positions T(x) in R B. Note that once the transformed position is outside the reference image domain ΩR, the calculation of differences is invalid and an additional penalty value p is assigned to the cost function to prevent exaggerated transformations. The penalty p is set to a larger value when the skull voxels of the floating image are anticipated to be within the domain of the reference image following transformation. In this study, the penalty value p was set to 100 and the six rigid-body transformation parameters were optimized by Powell’s direction set method combined with Brent’s line search [22].
2.5 Difference between complete and defective images
We designed a difference operation to extract the difference between the intact and defective skull images for reconstructing the repaired skull portion. The difference operation consisted of four steps. Firstly, the defective images were subtracted from the complete (diagnostic) images. Secondly, one-step erosion and dilation processes removed isles in the image. Thirdly, 3D region growing from a seed point on the repaired skull portion removed the unlinked portions not relevant to the target [1]. Finally, manual intervention refined the image.
2.6 Mesh smoothing
We constructed a 3D model of the repair from the volumetric image, using the marching cubes algorithm [16], a classic method for surface extraction, where each surface element is described by a triangle. Since the 3D mesh constructed from the volume grid is not sufficiently smooth, we applied a mesh smoothing method to rectify this problem. The Laplacian filter is a conventional method used to obtain a smooth polygonal surface [20]. The positions of all vertices P in the model were updated according to the formula:
where Q 0, Q 1, …, Q n−1 are the 1-ring neighbors of P, and λ is the step-size parameter controlling the magnitude of displacement.
2.7 Simulation design
We conducted a series of computer simulations based on a pair of intact low- and high-resolution skull images created from mirroring, to validate the proposed algorithm for defective skull reconstruction. The low-resolution image, with 5.00 mm slice thickness, was resampled using trilinear interpolation so that the resolution of the resampled image and the high-resolution image were the same, i.e., the voxel size was adjusted to 0.48 mm × 0.48 mm × 0.60 mm (Table 1). Once the two image resolutions matched, the skull portions were segmented by thresholding, and the active contour method applied to eliminate superfluous artifacts. Since the accuracy of reconstruction depended on performance of the active contour and registration, we assessed the accuracies for these operations in addition to the skull reconstruction.
We evaluated performance of the active contour in eliminating the artifacts on the low-resolution images. Since the low- and high-resolution images in the simulation were reconstructed from the same raw data set, they were inherently aligned before we applied the active contour method on the low-resolution image. Therefore, the difference in skull content between the low- and high-resolution images was determined both before and after artifact elimination.
We evaluated the accuracy of registration under different ratios of the hypothetically removed skull content. We randomly placed a ball-shaped eraser on the high-resolution skull image to simulate the effect of craniotomy, and generated two levels of content-removal images, namely 20 and 40% removal ratios for both unilateral and bilateral skull, respectively. The floating images were randomly translated and rotated, using hypothetical parameters consisting of six uniformly distributed random offsets for all translational ([−20 mm, 20 mm]) and rotational ([−20°, 20°]) parameters. The rigid-body registration was performed repeatedly (ten times) to transform the floating image.
After the registration process, the difference operation was applied to extract the repaired skull portion under different simulation conditions. The accuracies for skull reconstruction were calculated from the differences between the reconstructed skulls and the hypothetically removed portion on the high-resolution image.
3 Results
3.1 Simulation results
We evaluated performance of the active contour by comparing similarities between the low- and high-resolution images before, and after artifact elimination. We denote the number of overlapping voxels between the low- and high-resolution skulls as ‘O’, and the number of voxels in the union of the low- and high-resolution skulls as ‘U’. The similarities before and after artifact elimination, S b and S a, were defined by O b/U b and O a/U a, respectively. The results for the simulation data were S b = 0.748 and S a = 0.842, which represent the active contour effectiveness.
Registration errors for the simulation data can be quantified using the root mean squared error (RMSE) for the translation RMSEt and rotation RMSEr.
where (t x , t y , t z , r x , r y , r z ) are estimated parameters and \( (t_{x}^{\prime } ,t_{y}^{\prime } ,t_{z}^{\prime } ,r_{x}^{\prime } ,r_{y}^{\prime } ,r_{z}^{\prime } ) \) are true parameters. Figure 4 shows the registration accuracy results, evaluated using the RMSE’s. The results demonstrated that the median RMSEt for unilateral 20%, unilateral 40%, bilateral 20%, and bilateral 40% removal ratios were 0.25, 0.17, 0.19, and 0.22 mm, respectively. Regarding the median RMSEr for four different removal ratios were 0.01°, 0.08°, 0.02°, and 0.02°, respectively. The image registration was accurate for unilateral and bilateral cases at 20 and 40% removal ratios alike, in which the median RMSEt and RMSEr were all less than 0.25 mm and 0.09°, respectively.
To estimate the accuracy of the reconstruction of the removed skull portion, we compared the reconstructed and hypothetically removed skulls. By denoting the number of overlapping voxels between the reconstructed skull and the hypothetically removed skulls as N and the number of voxels in the hypothetically removed skulls as M, then, the reconstructed accuracy is (N/M) × 100%. In the simulation data, the bilateral skull removal resulted in the better reconstructed accuracy: 97.76 ± 0.55% (mean ± standard deviation) and 97.23 ± 0.75% for 20 and 40% removal ratios, respectively, whereas the lower accuracy was presented in the cases of unilateral skull removal: 97.21 ± 0.75 and 93.94 ± 2.33% for 20 and 40% removal ratios, respectively.
3.2 Experimental results
Three selected sagittal slices and their corresponding 3D surface models (Fig. 5) show the result of the removal of artifacts from the low-resolution images. Figure 5a and d displays the selected sagittal slice of the binarized 5.00-mm slice images, and its reconstructed 3D surface model, respectively. The “stair-shape” surface caused by the low resolution along the slice-selection direction was not acceptable for cranioplasty. Tri-linear interpolation was applied so that the voxel size of low- and high-resolution images were identical (0.49 mm × 0.49 mm × 0.63 mm) as shown in Fig. 5b. However, a skull penumbra caused by partial volume effects, was evident as ripples on the reconstructed surface (Fig. 5e). Elimination by the inner–outer snakes (Fig. 5c) significantly suppressed this effect (Fig. 5f).
The registration results are displayed in Fig. 6, where diagnostic skull images (in green) were overlaid and transparently fused with the defective image (in Indian red). The registration process transformed the position of diagnostic images into that of the defective images, which allowed the difference operation to extract the removed skull portion correctly.
The registration results of the experimental data. a The originally defective image, b the originally intact image, c the transformed intact image. d Intact (green) and defective (Indian red) skulls are initially mis-aligned. e After registration, most skull voxels are successfully matched. (Color figure online)
A 3D mesh smoothing technique was applied to the reconstructed skull portion to increase the target surface smoothness. Figure 7a and b shows the reconstructed 3D mesh, and the surface of the reconstructed skull (colored in pink) on the defective volume (colored in white), respectively. The results demonstrate that the boundary between the repaired skull and the incised part of the craniotomy match each other well. Accordingly, our method faithfully reconstructed the removed skull portion to its original appearance, and the 3D mesh coordinates could be used to sculpt a real artificial implant for the cranioplasty.
4 Discussion
In this paper, we propose an alternative method to reconstruct cranial defects. The use of the patients’ diagnostic and defective images with the aid of active contour model, and image registration techniques, were effective in modeling the skull’s original shape. This approach differs from mirroring techniques, which assume that contralateral skull information is available [13, 14, 19, 28]. However, patients with defects crossing the midsagittal plane of the brain, or those with a bilateral craniotomy do not fulfill this condition. Although surface interpolation [4, 5] can be used to reconstruct the dissected skull for cases with bilateral defects, such reconstructions are coarse approximations because a mathematical model, rather than anatomical structure, is applied. Compared to deformation-based methods [7, 27], which warp a template according to anatomical landmarks to fill the hole, our method has two main advantages. Firstly, we use the patient’s own anatomical data, which is more natural than an artificial template to reconstruct the preoperative shape. Secondly, manual selection of landmark locations and computationally expensive surface deformation were substituted by a simple image thresholding and rigid-body registration, achieving a large reduction in computer processing time.
The artifacts induced by the partial volume effect on the diagnostic images introduce inaccuracies during the rigid-body registration process between the diagnostic and defective images. Once the artifacts were suppressed using the active contour method, the registration results were satisfactory, as shown in the simulation data (Fig. 4), and in a real example (Fig. 6). Since the severity of partial volume effects depends upon slice thickness, the amount of shrinkage in cranial surfaces is adjusted based on different slice thicknesses. Specifically, the two-, four-, six-, and seven-pixel image erosions were used empirically for the 3-, 5-, 10-, and 15-mm slice thickness, respectively. As Fig. 5f shows, the artifact-removed surface model resemble to the real cranial contour more closely, compared to that in Fig. 5e.
The results of the simulation data indicated that the median RMSEt and RMSEr were all less than 0.25 mm and 0.09°, respectively, for all 20 and 40% removal ratios. In the worst case, although the unilateral 40% removal ratio was almost twice the usual clinical craniotomy, the overall reconstruction accuracy was greater than 93.94 ± 2.33%. Simulation results suggested that not only could the unilateral skull defect be repaired accurately, but that the bilateral craniotomy could also be recovered satisfactorily. For the same ratio of skull material removal, the overall reconstructed accuracies of the bilateral reconstructions were always higher in mean values and lower in standard deviations, than the unilateral reconstructions. This was because more balanced errors were derived in the cost function by removal in bilateral skull operations, than by unilateral instances, hence producing more robust results in the registration process.
Although there were one and two outliers in the RMSEr boxplots of unilateral 40% and bilateral 40% skull removal, respectively, their results appeared to be comparable with other cases. These outliers were caused by larger local minima at the low-resolution stage in the multi-resolution scheme that propagated to the higher-resolution levels. In the worst case, the RMSEt and RMSEr were 0.30 mm and 0.33°, respectively. Nevertheless, subvoxel accuracy was achieved (the voxel size was 0.48 mm × 0.48 mm × 0.60 mm) for all of our registration simulations. The reconstructed accuracy was as good as 90.09%.
In conclusion, we proposed an alternative method to faithfully reconstruct removed skull material based on the patient’s own intact diagnostic and defective high-resolution CT images. The results for the simulations and the real case demonstrated that the inner-outer snake could effectively eliminate artifacts induced by the partial volume effect, which facilitate the subsequent rigid-body registration to achieve satisfactory results. Our method was able to produce a repaired skull model matching the border of the incised section, and recover the original cranial appearance.
References
Adams R, Bischof L (1994) Seeded region growing. IEEE Trans Pattern Anal Mach Intell 16:641–647
Besl PJ, Mckay ND (1992) A method for registration of 3-D shapes. IEEE Trans Pattern Anal Mach Intell 14:239–256
Borgefors G (1988) Hierarchical chamfer matching: a parametric edge matching algorithm. IEEE Trans Pattern Anal Mach Intell 10:849–865
Carr JC, Fright WR, Beatson RK (1997) Surface interpolation with radial basis functions for medical imaging. IEEE Trans Med Imaging 16:96–107
Chong CS, Lee HP, Kumar AS (2006) Automatic hole repairing for cranioplasty using Bezier surface approximation. J Craniofac Surg 17:344–352
Cohen LD (1991) On active contour models and balloons. CVGIP: Image Underst 53:211–218
Dean D, Min K-J (2003) Deformable templates for preoperative computer-aided design and fabrication of large cranial implants. Int Congr Ser 1256:710–715
Hartkens T, Hill DLG, Castellano-Smith AD, Hawkes DJ, Maurer CR, Martin AJ, Hall WA, Liu H, Truwit CL (2003) Measurement and analysis of brain deformation during neurosurgery. IEEE Trans Med Imaging 22:82–92
Hill DLG, Batchelor PG, Holden M, Hawkes DJ (2001) Medical image registration. Phys Med Biol 46:R1–R45
Hutchinson P, Timofeev I, Kirkpatrick P (2007) Surgery for brain edema. Neurosurg Focus 22:E14
Kass M, Witkin A, Terzopoulos D (1987) Snakes: active contour models. Int J Comput Vis 1:321–331
Kozerke S, Botnar R, Oyre S, Scheidegger MB, Pedersen EM, Boesiger P (1999) Automatic vessel segmentation using active contours in cine phase contrast flow measurements. J Magn Reson Imaging 10:41–51
Lee M-Y, Chang C-C, Lin C-C, Lo L-J, Chen Y-R (2002) Custom implant design for patients with cranial defects. IEEE Eng Med Biol 21:38–44
Lee S-C, Wu C-T, Lee S-T, Chen P-J (2009) Cranioplasty using polymethyl methacrylate prostheses. J Clin Neurosci 16:56–63
Liao Y-L, Sun Y-N, Lu C-F, Wu Y-T, Wu C-T, Lee S-T, Lee J-D (2010) Skull-based registration of intra-subject CT images: the effects of different resolutions and partial contents. In: Mahadevan V, Zhou J (eds) Proceeding of the 2nd International Conference on Bioinformatics and Biomedical Technology (ICBBT), Research Publishing Services, Singapore, pp 269–272
Lorensen WE, Cline HE (1987) Marching cubes: a high resolution 3D surface construction algorithm. ACM SIGGRAPH Comput Graph 21:163–169
Maes F, Collignon A, Vandermeulen D, Marchal G, Suetens P (1997) Multimodality image registration by maximization of mutual information. IEEE Trans Med Imaging 16:187–198
Manawadu D, Quateen A, Findlay JM (2008) Hemicraniectomy for massive middle cerebral artery infarction: a review. Can J Neurol Sci 35:544–550
Maravelakis E, David K, Antoniadis A, Manios A, Bilalis N, Papaharilaou Y (2008) Reverse engineering techniques for cranioplasty: a case study. J Med Eng Technol 32:115–121
Ohtake Y, Belyaev A, Bogaevski I (2001) Mesh regularization and adaptive smoothing. Comput Aided Design 33:789–800
Pelizzari CA, Chen GTY, Spelbring DR, Weichselbaum RR, Chen CT (1989) Accurate three-dimensional registration of CT, PET, and/or MR images of the brain. J Comput Assist Tomogr 13:20–26
Press WH (1992) Numerical recipes in C: the art of scientific computing. Cambridge University Press, Cambridge/New York
Roche A, Malandain G, Pennec X, Ayache N (1998) The correlation ratio as a new similarity measure for multimodal image registration. In: Wells WM, Colchester A, Delp S (eds) 1st International Conference on Medical Image Computing and Computer Assisted Intervention (MICCAI), LNCS 1496, Springer, Berlin/Heidelberg, pp 1115–1124
Schirmer CM, Ackil AA, Malek AM (2008) Decompressive craniectomy. Neurocrit Care 8:456–470
Sommer HJ, Eckhardt RB, Shiang TY (2006) Superquadric modeling of cranial and cerebral shape and asymmetry. Am J Phys Anthropol 129:189–195
Subramaniam S, Hill MD (2009) Decompressive hemicraniectomy for malignant middle cerebral artery infarction: an update. Neurologist 15:178–184
Wu T, Engelhardt M, Fieten L, Popovic A, Radermacher K (2006) Anatomically constrained deformation for design of cranial implant: methodology and validation. In: Larsen R, Nielsen M, Sporring J (eds) 9th International Conference on Medical Image Computing and Computer Assisted Intervention (MICCAI), LNCS 4190, Springer, Berlin/Heidelberg, pp 9–16
Wu WZ, Zhang Y, Li H, Wang WS (2009) Fabrication of repairing skull bone defects based on the rapid prototyping. J Bioact Compat Polym 24:125–136
Yushkevich PA, Piven J, Hazlett HC, Smith RG, Ho S, Gee JC, Gerig G (2006) User-guided 3D active contour segmentation of anatomical structures: significantly improved efficiency and reliability. Neuroimage 31:1116–1128
Acknowledgments
The authors express appreciation to Kevin Fortune for his assistance in English language editing. This work is funded by the Department of Industrial Technology of Ministry of Economic Affairs, Taiwan, under the contract numbers 97-EC-17-A-19-S1-035 and 98-EC-17-A-19-S1-035.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Liao, YL., Lu, CF., Sun, YN. et al. Three-dimensional reconstruction of cranial defect using active contour model and image registration. Med Biol Eng Comput 49, 203–211 (2011). https://doi.org/10.1007/s11517-010-0720-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11517-010-0720-0