Abstract
Purpose
The purpose of this study was to determine the first-order rate constants and half-lives of aerobic and anaerobic biomineralization of atrazine in soil samples from an agricultural farm site that had been previously used for mixing pesticide formulations and washing application equipment. Atrazine catabolic genes and atrazine-degrading bacteria in the soil samples were analyzed by molecular methods.
Materials and methods
Biomineralization of atrazine was measured in soil samples with a [U-ring-14C]-atrazine biometer technique in soil samples. Enrichment cultures growing with atrazine were derived from soil samples and they were analyzed for bacterial diversity by constructing 16S rDNA clone libraries and sequencing. Bacterial isolates were also obtained and they were screened for atrazine catabolic genes.
Results and discussion
The soils contained active atrazine-metabolizing microbial communities and both aerobic and anaerobic biomineralization of [U-ring-14C]-atrazine to 14CO2 was demonstrated. In contrast to aerobic incubations, anaerobic biometers displayed considerable differences in the kinetics of atrazine mineralization between duplicates. Sequence analysis of 16S rDNA clone libraries constructed from the enrichment cultures revealed a preponderance of Variovorax spp. (51 %) and Schlesneria (16 %). Analysis of 16S rRNA gene sequences from pure cultures (n = 12) isolated from enrichment cultures yielded almost exclusively Arthrobacter spp. (83 %; 10/12 isolates). PCR screening of pure culture isolates for atrazine catabolic genes detected atzB, atzC, trzD, trzN, and possibly atzA. The presence of a complete metabolic pathway was not demonstrated by the amplification of catabolic genes among these isolates.
Conclusions
The soils contained active atrazine-metabolizing microbial communities. The anaerobic biometer data showed variable response of atrazine biomineralization to external electron acceptor conditions. Partial pathways are inevitable in soil microbial communities, with metabolites linking into other catabolic and assimilative pathways of carbon and nitrogen. There was no evidence for the complete set of functional genes of the known pathways of atrazine biomineralization among the isolates.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Atrazine is in common use as an agricultural herbicide in the U.S.A. As a consequence, atrazine residuals and metabolites and atrazine-degrading microorganisms are widely found in surface soils, rivers and wetlands in agricultural watersheds (Lazorko-Connon and Achari 2009; Mudhoo and Garg 2011). Half-lives of atrazine biodegradation can be shorter than 10 days in agricultural soils that have decades of annual atrazine applications (Pussemier et al. 1997). Soil microbial communities vary greatly in atrazine biodegradation potential. Biodegradation and biomineralization of atrazine involve multiple enzymatic steps, most hydrolytic, and some alternatively oxidative, which vary in pathway sequence, metabolites and genes in soil and sediment bacterial communities (Krutz et al. 2010; Lian et al. 2012; Udiković-Kolić et al. 2012; Fenner et al. 2013; Fan and Song 2014; Singh and Singh 2016). The best known biodegradative pathway is the Atz, which was originally elucidated in Pseudomonas ADP and involves six proteins (AtzA, AtzB, AtzC, AtzD, AtzE, and AtzF) (de Souza et al. 1995; Boundy-Mills et al. 1997; Sadowsky et al. 1998; Martinez et al. 2001; Cheng et al. 2005). Other degradative enzymes have been characterized, such as TrzN (triazine chlorohydrolase) in Nocardioides sp. strain C190 (Mulbry et al. 2002) and Arthrobacter aurescens TC1 (Sajjaphan et al. 2004), and TrzD (cyanuric acid amidohydrolase) in Pseudomonas sp. strain NRRLB-12227 (Eaton and Karns 1991) and Klebsiella pneumonia strain 99 (Karns and Eaton 1997). To date, analyses of catabolic genes in soil bacterial isolates suggest that the complete Atz pathway is relatively rare and various combinations of atz and trz genes are more common in environmental isolates (Devers et al. 2007; Vaishampayan et al. 2007; El Sebaï et al. 2011). It is likely that pathway intermediates feed into other degradative pathways in microbial communities and the complete mineralization through the Atz pathway may not be dominant. Examples of cooperative pathways could include the hydrolytic release of the ring side chains ethanolamine and isopropyl amine, both biodegradable by many microorganisms that cannot degrade the parent molecule atrazine, or the ring cleavage metabolites such as urea, an excellent N source for soil microorganisms. Neither the Atz nor Trz pathway requires molecular O2 and thus they can function in both aerobic and anaerobic environments.
Cooperative pathways and synergistic metabolism of intermediates are central in the attenuation of atrazine by soil microorganisms. Several eukaryotic soil microorganisms in genera such as Cryptococcus, Phanerochaete, Pichia, and Pleurotus have also been described that can mediate atrazine biodegradation but these pathways do not proceed beyond cyanuric acid formation (Donnelly et al. 1993; Mougin et al. 1994; Masaphy et al. 1996; Bastos and Magan 2009; Abigail et al. 2012, 2013; Romero et al. 2014). Laccases may be involved, but specific catabolic genes associated with atrazine biodegradation have yet to be characterized in eukaryotic microorganisms.
For this study, the biomineralization potential of atrazine in an agricultural farm setting in central Ohio was examined. Two sites at this farm were of specific interest: (1) a pesticide mixing area that had been abandoned by late 1990s, about 10 years prior to this study, and (2) an adjacent (about 30 m) area that has received spills from washings of pesticide formulations from application equipment in a shed. Previous studies have shown that agricultural farm and agrochemical dealer mixing and spill areas have microbial populations that can actively degrade residual pesticide mixtures (Arthur et al. 2000; Chirnside et al. 2007, 2009; Udiković et al. 2003; Udiković-Kolić et al. 2007, 2010). Potential problems of point source pesticide contamination in agricultural farm sites have been mitigated with systems whereby the spray or washing area is covered with rich organic material which supports pesticide retention and microbial degradation. Several configurations and performance data have been discussed in the literature (Grigg et al. 1997; Arthur et al. 2000; de Wilde et al. 2007; del Pilar Castillo et al. 2008; Spanoghe et al. 2009). However, soil-bound residues of atrazine from historic applications or spills can be persist for decades (Liu et al. 2016).
At the farm site in this study, atrazine and other pesticides have been used on and off depending on crop rotation and integrated pesticide management practices. No special remediation was undertaken over the years of practicing pesticide disposal until the late 1990s, at which time the containment and storage of liquid pesticide residues was initiated. A previous study of enrichment of atrazine degraders from the farm site yielded a soil bacterial isolate M91-3 (Radosevich et al. 1995), subsequently identified as Ralstonia basilensis (Stamper et al. 2002), obtained originally from samples collected from the pesticide mixing and spill site. The purpose of this study was to determine the half-lives of aerobic and anaerobic biomineralization of atrazine in soil samples from the past pesticide mixing and spill and the machinery washing areas. Biomineralization was measured in biometers using soil samples and [U-ring-14C]-atrazine. Bio-Sep beads were also tested for trapping atrazine degraders from soil. Soil-borne atrazine degraders have been previously probed with porous Bio-Sep beads (Ghosh et al. 2009; Omotayo et al. 2011). Atrazine enrichment cultures from the soil samples and field-incubated Bio-Sep beads yielded atrazine-degrading mixed cultures and isolates of bacteria. Phylogenetic analyses based on 16S rRNA gene sequences and the screening of known catabolic genes of atrazine metabolism were included in the scope of this study.
2 Materials and methods
2.1 Sampling
The Western Agricultural Research Station of the Ohio Agricultural Research and Development Center is located 5.6 km northwest of South Charleston, Ohio (39°86′38″N, 83°67′22″W). The farm (established in 1958) is a 173 ha agricultural research facility that supports agronomic and specialty crop programs, including corn and soy, in the western part of Ohio. As part of the weed control programs, the management practice has involved the application of atrazine as a pre-emergence weed killer in corn fields. The soil at the site is primarily Strawn–Crosby complex (0 to 2 % slopes) (http://websoilsurvey.nrcs.usda.gov/app/WebSoilSurvey.aspx).
The farm site has a shed building where herbicide formulations are stocked and prepared for transportation to the field. Next to the shed was a grassy area (Fig. 1) where pesticides were mixed (site 1), and this area received numerous pesticide spills over the years. A second adjacent grassy area was at the base of the ramp to the shed where pesticide application machinery was rinsed (site 2). About 10 years prior to this study, the practice of allowing spills and rinsing outdoors was discontinued and liquid waste was contained to a special holding tank for further treatment. Both sites are now grass-covered.
Soil samples were collected from site 1 (the former pesticide mixing site) and site 2 (machinery washing area) in mid-May. Several grab samples of top soil (0–7 cm) were removed with a sterile trowel to yield about 500 g composite and stored in sterile plastic bags on ice until processed in the laboratory. The sampling avoided grass roots. Previous studies generally indicate that maximum activity of atrazine biodegradation is usually associated with the soil top layer (e.g., Tuovinen et al. 2015). No effort was made to sample the subsurface.
Bio-Sep beads were equilibrated in solution of 2 mg atrazine/l or distilled water for 3 days before they were closed in sterile nylon mesh bags (~30 beads per bag) and sealed with plastic clips. Bead bags were buried in the top 10-cm layer of soil in site 1 at the former pesticide mixing area and retrieved after 33 days of contact time with soil (mid-May to mid-June). Only site 1 was sampled with Bio-Sep beads because the thrust of the study was on soil sampling directly for biometers and enrichments of atrazine degraders.
2.2 Biomineralization of [U-ring-14C]-atrazine in biometers
Aerobic and anaerobic mineralization of atrazine in soil samples and Bio-Sep beads was measured with the biometer technique that has been previously described (Douglass et al. 2014, 2015). The biometers (60-ml volumetric capacity) contained 5 g soil samples or 6–10 diced Bio-Sep beads and 0.064 μmol [U-ring-14C]-atrazine ± glucose (11.1 mM). In order to prevent possible microelement limitation, biometers with Bio-Sep beads received 5 ml mineral salts (MS) solution, which contained (per liter): K2HPO4, 0.5 g; MgSO4·7H2O, 0.5 g; FeCl3·H2O, 0.1 mg; CaCl2·H2O, 1 mg; MnCl2, 0.01 mg; ZnCl2, 1 μg; pH 6.8. Anaerobic incubations under N2 head space involved 25 μmol nitrate as KNO3, 125 μmol Fe(III) as ferrihydrite, or 15 μmol sulfate as Na2SO4 per biometer as external electron acceptors or distilled H2O as a blank. These additions were intended to approximate e−-equimolar acceptor conditions. Controls of sterile soil were included in biometer incubations.
The biometers were incubated at 22 ± 2 °C in the laboratory. Biometer sampling and measurement of 14CO2 have been previously described (Douglass et al. 2014). 14CO2 evolution was calculated as a percentage of the total amount of radioactivity of the [U-ring-14C]-atrazine added to the biometers at the beginning of the time course. Standard error bars were included for the replicates. Rate constants (k, days−1) and half-lives (t 1/2, days) of atrazine mineralization were determined with a pseudo first-order rate expression (Guerin and Boyd 1992).
2.3 Enrichment of atrazine degraders
Aerobic enrichment cultures were set up with 2.5 g soil or 6–10 Bio-Sep beads in MS media containing 30 mg/l atrazine ± glucose amendment (2 g glucose/l) and were incubated in shake flasks (150 rpm) at 22 ± 2 °C. Aliquots of enrichment cultures were transferred to the corresponding fresh media at intervals of 2–4 weeks. After at least six transfers to fresh MS media with 30 mg/l atrazine, enrichment cultures were screened for residual atrazine concentration using HPLC with a UV detector at 220 nm as previously described (Radosevich et al. 1996). Samples of cultures were mixed with mobile phase (65 % HPLC-grade acetonitrile in HPLC-grade distilled water) followed by centrifugation at 16,000×g for 15 min. The supernatants were transferred to HPLC vials in an autosampler and pumped through an Adsorbosphere C18 HPLC column at a flow rate of 1.0 mL/min.
Reference samples containing uninoculated sterile MS + atrazine media and standards with varying concentrations of atrazine dissolved in mobile phase were included with every HPLC run. Enrichments that exhibited at least 80 % decrease in atrazine concentration were deemed to contain atrazine-degrading mixed cultures.
2.4 Isolation of atrazine-degrading pure cultures
Prospective enrichment cultures of atrazine degraders were inoculated to solid MS + atrazine media. Dilutions were spread-plated in MS + atrazine media solidified with 1.5 % Noble agar. All plates were incubated at 22 ± 2 °C for up to 2 weeks. Discreet colonies from the spread plates were subcultured twice on streak plates before transferring them into 5 ml of corresponding liquid MS media. Changes in atrazine concentration were monitored with HPLC during subsequent incubation in liquid media. Pure cultures were stored at 4 °C. Liquid cultures were stored as 20 % glycerol stocks at −55 °C.
2.5 DNA isolation, 16S rRNA gene amplification, and denaturing gradient gel electrophoresis (DDGE)
Genomic DNA was isolated from pure and mixed cultures with MoBio UltraClean Microbial DNA Isolation Kits per manufacturer’s instructions (MoBio Laboratories, Carlsbad, CA). The V3 region of the 16S rRNA gene from atrazine-degrading mixed cultures was amplified in 50 μl total volume, consisting of 1X TaKaRa Premix Taq (1.25 U Taq; 0.2 mM dATP; 0.2 mM dTTP; 0.2 mM dCTP; 0.2 mM dGTP; 10 mM Tris-HCl [pH 8.3], 50 mM KCl, 1.5 mM MgCl2), 1 μM primers 357fGC and 518r (Muyzer et al. 1993), and 20 ng of template DNA. The PCR amplification followed a previously established protocol (Douglass et al. 2015). PCR products were resolved with DGGE using the DCode Universal Mutation Detection System (Bio-Rad, Hercules, CA). Gels consisted of 8 % acrylamide and a 40–60 % denaturant gradient. Gels were electrophoresed at 83 V for 16 h in 0.5X TAE buffer solution at 60 °C. After electrophoresis, gels were stained in 1X SYBR Gold (Molecular Probes, Eugene, OR) for 30 min on a rocking platform and visualized on a UV transilluminator. Mixed cultures corresponding to unique DGGE banding patterns were selected for further analysis via 16S rDNA clone library sequencing.
2.6 16S rDNA clone library formation and analysis
Genomic DNA from atrazine-degrading mixed cultures and isolates was amplified for the approximately 1,500-bp region of the 16S rDNA using 1 μM primers 27f and 1525r (Lane 1991) and 20 ng of template DNA. The PCR conditions followed previously published protocols (Douglass et al. 2015). PCR products were used to construct 16S rDNA clone libraries. The clones were made using the Promega pGEM-T vector system and JM109 high-efficiency competent cells (Promega, Madison, WI). Overnight cultures of clones were plated onto 96-well plates and sent to the Clemson University Genomics Institute for sequencing. For each pure culture, three clones were sequenced; for each mixed culture, 25 clones were sequenced. Sequences were edited in Geneious (http://www.geneious.com/). A consensus sequence was generated for each set of three clones resulting from pure cultures when possible, and these consensus sequences were entered into the NCBI-BLAST and Ribosomal Database Project (RDP) databases. If a consensus sequence of three clones was not available, individual edited clone sequences or consensus sequences constructed from two clones were entered as queries into the aforementioned databases. Each individual edited clone sequence resulting from mixed cultures was searched in these databases.
The 16S rRNA gene sequences were deposited with GenBank. The accession numbers for aerobic mixed culture clones are KP779907-KP779962. The accession numbers for aerobic isolates are KP779963-KP779997.
2.7 Atrazine catabolic gene detection in atrazine-degrading bacterial isolates
Genomic DNA from atrazine-degrading isolates was amplified for known atrazine catabolic genes atzA, atzB, atzC, atzD, atzE, atzF, trzD, and trzN. Escherichia coli genomic DNA was also amplified as a negative control for all catabolic genes. Pseudomonas ADP was used as a positive control for atzA, atzB, atzC, atzD, atzE, and atzF, and as a negative control for trzD and trzN. Arthrobacter aurescens TCI was used as a positive control for trzN as well as an additional positive control for atzB and atzC and a negative control for atzA, atzD, atzE, and atzF. Klebsiella pneumoniae 99 was used as a positive control for trzD.
PCR reactions were conducted with 1 μM forward primer and 1 μM reverse primer as specified in Table 1 and 20-ng template DNA as described by Douglass et al. (2015). PCR products were loaded onto agarose gel, and after electrophoresis the gel was stained with ethidium bromide and visualized with a UV gel imager. Detection of a band of anticipated size was treated as preliminary evidence of the presence of the given catabolic gene, and a random assortment of these PCR reactions were used to create clone libraries as outlined in section 2.6 . For each PCR reaction, three clones were sequenced. When possible, a consensus sequence for these three clone sequences was generated and used for bioinformatics analysis; otherwise, a consensus sequence of two clones was generated, or a single edited clone sequence was used. Query sequences were entered into NCBI-BLAST and aligned with their corresponding catabolic gene sequences. Catabolic gene sequences were deposited with GenBank, and the accession numbers are KP863790-KP863798.
3 Results and discussion
3.1 Measurement of atrazine mineralization in biometers with [U-ring-14C]-atrazine
Under aerobic conditions, mineralization of atrazine in both sites reached ~20–25 % in biometers (Fig. 2). Glucose amendment had no discernible effect on mineralization. Enhancement of glucose amendment could indicate that atrazine-mineralizing bacteria are limited by carbon rather than other nutrients. Whether glucose is co-metabolized with atrazine or only increases the amount of active biomass is not known. In this case, the lack of glucose effect suggests that bacteria were not limited for carbon and glucose did not specifically increase the biomass or activity of atrazine-mineralizing bacteria.
Aerobic mineralization of [U-ring-14 C]-atrazine in biometers in the absence (a) and presence (b) of glucose amendment. Symbols: (black circle), site 1, soil from the former machinery washing area; (white circle), site 2, soil from the former pesticide mixing area; (black down-pointing triangle), sterile soil control; (white diamond), water-equilibrated Bio-Sep beads retrieved from site 1; (black diamond), atrazine-equilibrated Bio-Sep beads retrieved from site 2. Bio-Sep beads were only used to inoculate biometers without glucose. Data points represent mean cumulative percentages obtained from duplicate biometers, with the exception of those representing Bio-Sep biometers, which represent results from single biometers. Bars indicate the standard error for the replicates, and in some cases are smaller than the data point symbols
Mineralization in biometers inoculated with Bio-Sep beads was comparable to sterile controls (Fig. 2). The lack of 14CO2 evolution from Bio-Sep bead biometers was attributed to negligible size of atrazine-degrading microbial community associated with the beads used in the assay. This assertion is supported by the fact that after an enrichment phase, mixed cultures derived from Bio-Sep beads degraded atrazine as confirmed via HPLC analysis. Subsequently, atrazine-degrading pure cultures were successfully isolated from mixed cultures enriched from Bio-Sep beads.
Under anaerobic conditions, pesticide mixing area soil (site 1) outperformed machinery washing area soil (site 2) in biometers with Fe(III) amendment (~20 vs. 6 %) and no electron acceptor amendment (~18 vs. 7 %). The opposite trend was true in the nitrate-amended biometers (~26 vs. 22 %). Microbial communities in both soils mineralized atrazine to a similar extent (~20 %) under sulfate amendment (Fig. 3). The soil sample from the washing area (site 2) mineralized less than 10 % of the added atrazine in the absence of external electron acceptors as well as in ferrihydrite-amended biometers. The reduction of the external electron acceptors nitrate, Fe(III), and sulfate was not tested in the 14C-labeled atrazine biometer experiments. In general, biometers constructed with samples from the former pesticide mixing area (site 1) had large variation between duplicate anaerobic biometers. The variation was due to different lag times and extent of mineralization between the duplicates. In general, the variation reflects spatial heterogeneity in microscale in soil microbiology that has been reported numerous times in terms of microbial numbers and activity as well as community structure and composition (e.g., Ettema and Wardle 2002; O’Donnell et al. 2007).
Anaerobic mineralization of [U-ring-14 C]-atrazine in biometers. The biometers were amended with a no electron acceptor, b K-nitrate, c ferrihydrite, and d Na-sulfate. Symbols: (white circle), site 1, soil from the former pesticide mixing area; (black circle), site 2, soil from the former machinery washing area; (black down-pointing triangle), sterile soil control. Data points represent mean cumulative percentages obtained from duplicates. Bars indicate the standard error for the replicates, and in some cases are smaller than the data point symbols
Thus the anaerobic biometer results indicated that nitrate and sulfate enhance the biomineralization of atrazine. Atrazine and nitrate especially are common pollutants in surface runoff from agricultural fields. If the runoff is trapped in a riparian zone or a wetland system, nitrate flux may promote the concurrent biodegradation of atrazine. Whether it is due to denitrification is not clear because atrazine can also be metabolized via hydrolytic steps. However, metabolites such as the ethyl and isopropyl side chains and the lower pathway intermediates from atrazine biodegradation may link into other degradative pathways involving respiratory activities in microbial consortia.
The half-lives of aerobic mineralization of atrazine calculated from the first-order rate constants indicated relatively fast atrazine attenuation, about 3 days in the soil samples from the washing area (site 2) and 2 days in the pesticide mixing area (site 1) (Table 2). These half-lives clearly indicate that soil microorganisms at these sites have previously acclimated to atrazine. The half-lives of anaerobic mineralization in site 2 soil were 4 days under nitrate amended conditions, but were extended to 11–34 days in the absence of external electron acceptors or in the presence of Fe(III) or sulfate. In contrast, the half-lives of anaerobic mineralization in soil samples from site 1 were in the range of 3 to 5 days (Table 2). For reference, the half-lives of 2–3 days for aerobic biomineralization estimated in this study are in the general range of atrazine attenuation determined for agricultural soils with histories of atrazine application (Krutz et al. 2010, 2012; Mueller et al. 2010). The half-life of 1.7 days (k = 0.397 days−1) is among the shortest ever reported for a soil sample in the literature. If the soil from this site were used for crop production, atrazine application would have little impact on weed control because its concentration would attenuate in less than a week to non-herbicidal levels based on the estimated half-life. As a point of reference, given the wide use of atrazine over several decades, verifiable information on rate constants or half-lives of atrazine biomineralization in U.S. soils with no previous environmental exposure to atrazine is not available in the literature.
3.2 16S rRNA gene sequence based analysis of atrazine-degrading enrichment cultures and bacterial isolates
Aerobic enrichments of atrazine degraders from the soil samples of sites 1 and 2 and Bio-Sep bead samples from site 1 yielded mixed cultures which consistently depleted atrazine in liquid media upon successive subcultures. The HPLC chromatograms indicated the presence of multiple metabolites that varied in time course samples, but metabolite analysis was outside the scope of the study.
Three mixed cultures were selected for further analysis based upon the 16S rDNA gene banding patterns in DGGE profiles: two enrichments created with soil from site 1, the former pesticide mixing area (one amended with glucose, one not amended with glucose) and an enrichment created with soil from site 2, the former farm machinery wash area. Amplification of 16S rRNA genes from the three mixed cultures yielded 49 clones for sequencing; the remainder of the 75 clones failed to produce usable sequence data. Sequence data from 16S rDNA clones was queried in the RDP database for nearest matches. As shown in Fig. 4, most clones (≥90 % similarity match) were consistent with Variovorax (51 %) and Schlesneria (16 %) spp. Other clusters included Acidovorax and Legionella spp. (8 % each). Some clones could only be matched at a family (Phyllobacteriaceae, 2 %) or class (Alphaproteobacteria, 6 %) level, due to the lack of cultured representatives in the RDP database matching the query sequences. Variovorax spp. have been previously isolated from atrazine-degrading mixed cultures constructed from agricultural soils (Smith et al. 2005; Zhang et al. 2012), whereas Schlesneria spp. have been previously associated with N-cycling (Kulichevskaya et al. 2007). It is conceivable that some clones were incidental and not relevant in the biodegradation of atrazine. Bacteria closely related to the clones were not recovered by cultivation in atrazine media, signaling bias in methods and media of isolation.
Identification of 16S rDNA clones from atrazine-degrading aerobic mixed cultures. Forty-nine clones in total are represented. All sequence queries returned similarity scores of 0.90 or greater. All identifications are made at the genus level except for Alphaproteobacteria (one asterisk) at the class level and Phyllobacteriaceae (two asterisks) at the family level
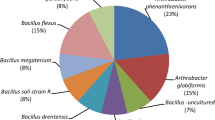
Atrazine-degrading mixed cultures were also plated onto solidified MS media. Two mixed cultures, one from the soil sampled at site 1 and the other from site 2, were selected for constructing clone libraries. Additionally, clone libraries were also created from two glucose-amended enrichments of Bio-Sep beads that had been originally incubated in the soil at site 1. One Bio-Sep contained atrazine-soaked beads and the other water-soaked beads. A total of 12 pure cultures were isolated from the plates. DNA was extracted and amplified for 16S rRNA genes; amplification products were cloned and sequenced. Figure 5 summarizes the RDP nearest matches (≥96 % similarity) of the resulting 16S rDNA consensus sequences. Arthrobacter spp. dominated the pure culture collection (80 %; 8/10), with isolates similar to Arthrobacter sp. B3051 constituting the largest number (25 %; 3/12). A single match each was also made with Hydrogenophaga sp. Gsoil 1545 and Sinorhizobium sp. TJ170. Similar to the findings in this study, previous efforts to isolate bacteria from atrazine-degrading soil microbial communities have yielded numerous Arthrobacter strains in spite of the lack of selective culture-based techniques specific for these Actinobacteria. Arthrobacter spp. have been isolated from around the world from soils that have been exposed to atrazine (Aislabie et al. 2005; Vaishampayan et al. 2007; Vibber et al. 2007; Zhang et al. 2011). The reasons for the cultural bias for Arthrobacter remain unclear. This bias is particularly noteworthy because Arthrobacter was not detected through cloning. Conversely, clones detected in enrichment cultures were notably absent from the isolations.
3.3 Detection of atz and trz catabolic genes
The presence of atrazine catabolic genes was investigated in seven of the 10 pure culture isolates by PCR amplification and agarose gel visualization (Table 3). All isolates were shown to contain atzBC. Additionally, an isolate consistent with Arthrobacter sp. M2012083 was demonstrated to contain atzBC-trzN, a pattern which has been previously reported in the literature (Devers et al. 2004; Sajjaphan et al. 2010; Zhang et al. 2011).
An isolate matching with Hydrogenophaga sp. Gsoil 1545 was shown to contain atzBC-trzD, and potentially also atzA and/or trzN. Hydrogenophaga spp. have previously been isolated from river sediment samples containing atrazine degraders (Vargha et al. 2005). A trzN–atzABC–trzD pattern of catabolic genes has also been reported for Hydrogenophaga spp. isolated from soil sampled in an agrochemical factory (Udiković-Kolić et al. 2010).
None of the isolates contained the atzDEF combination. Three of the successfully amplified catabolic genes were chosen for subsequent cloning and sequencing (Table 4). BLAST searches against respective database catabolic gene sequences yielded similarity scores in the range of 99–100 % and query coverage values of 100 %.
4 Conclusions
The biomineralization of atrazine was comparable in soil samples from both sites, demonstrating that the soils contained active atrazine-mineralizing microbial communities. While the reduction of external electron acceptors was not confirmed in 14C-atrazine-containing soil biometers, the data showed atrazine biomineralization anaerobic conditions. Some replicate anaerobic biometers showed considerable variation in the time course of atrazine biomineralization. This indicated that the response to external electron acceptors was variable. Some anaerobic electron acceptors may not necessarily stimulate the activity. Variovorax spp. in enrichment cultures and isolates of Arthrobacter spp. were particularly dominant. There was no evidence for the functional genes of the complete Atz pathway among the isolates. Partial pathways are inevitable in soil microbial communities, with metabolites linking into other catabolic and assimilative pathways of carbon and nitrogen. In contrast to the identification of isolates predominantly as Arthrobacter spp. in the phylum Actinobacteria, the 16S rDNA sequences from the clones from enrichment cultures matched the phylum Planctomycetes and classes Alpha-, Beta-, and Gammaproteobacteria.
References
Abigail MEA, Lakshmi V, Das N (2012) Biodegradation of atrazine by Cryptococcus laurentii isolated from contaminated agricultural soil. J Microbiol Biotechnol Res 2:450–457
Abigail MEA, Salam JA, Das N (2013) Atrazine degradation in liquid culture and soil by a novel yeast Pichia kudriavzevii strain Atz-EN-01 and its potential application for bioremediation. J Appl Pharmaceut Sci 3(6):35–40
Arthur EL, Perkovich BS, Anderson TA, Coats JR (2000) Degradation of an atrazine and metolachlor herbicide mixture in pesticide-contaminated soils from two agrochemical dealerships in Iowa. Water Air Soil Pollut 119:75–90
Aislabie J, Bej AK, Ryburn J, Lloyd N, Wilkins A (2005) Characterization of Arthrobacter nicotinovorans HIM, an atrazine-degrading bacterium, from agricultural soil New Zealand. FEMS Microbiol Ecol 52:279–286
Bastos AC, Magan N (2009) Trametes versicolor: potential for atrazine bioremediation in calcareous clay soil, under low water availability conditions. Int Biodeter Biodegr 63:389–394
Boundy-Mills KL, de Souza ML, Mandelbaum RT, Wackett LP, Sadowsky MJ (1997) The atzB gene of Pseudomonas sp. strain ADP encodes the second enzyme of a novel atrazine degradation pathway. Appl Environ Microbiol 63:916–923
Cheng G, Shapir N, Sadowsky MJ, Wackett LP (2005) Allophanate hydrolase, not urease, functions in bacterial cyanuric acid metabolism. Appl Environ Microbiol 71:4437–4445
Chirnside AEM, Ritter WF, Radosevich M (2007) Isolation of a selected microbial consortium from a pesticide-contaminated mix-load site soil capable of degrading the herbicides atrazine and alachlor. Soil Biol Biochem 39:3056–3065
Chirnside AEM, Ritter WF, Radosevich M (2009) Biodegradation of aged residues of atrazine and alachlor in a mixed-load site soil. Soil Biol Biochem 41:2484–2492
de Souza ML, Wackett LP, Boundy-Mills KL, Mandelbaum RT, Sadowsky MJ (1995) Cloning, characterization, and expression of a gene region from Pseudomonas sp. strain ADP involved in the dechlorination of atrazine. Appl Environ Microbiol 61:3373–3378
de Souza ML, Seffernick J, Martinez B, Sadowsky MJ, Wackett LP (1998) The atrazine catabolism genes atzABC are widespread and highly conserved. J Bacteriol 180:1951–1954
de Wilde T, Spanoghe P, Debaer C, Ryckeboer J, Springael D, Jaeken P (2007) Overview of on-farm bioremediation systems to reduce the occurrence of point source contamination. Pest Manag Sci 63:111–128
del Pilar Castillo M, Torstensson L, Stenström J (2008) Biobeds for environmental protection from pesticide use—a review. J Agric Food Chem 56:6206–6219
Devers M, Soulas G, Martin-Laurent F (2004) Real-time reverse transcription PCR analysis of expression of atrazine catabolism genes in two bacterial strains isolated from soil. J Microbiol Meth 56:3–15
Devers M, El Azhari N, Kolic N-U, Martin-Laurent F (2007) Detection and organization of atrazine-degrading genetic potential of seventeen bacterial isolates belonging to divergent taxa indicate a recent common origin of their catabolic functions. FEMS Microbiol Lett 273:78–86
Donnelly PK, Entry JA, Crawford DL (1993) Degradation of atrazine and 2,4-dichlorophenoxyacetic acid by mycorrhizal fungi at three nitrogen concentrations in vitro. Appl Environ Microbiol 59:2642–2647
Douglass JF, Radosevich M, Tuovinen OH (2014) Mineralization of atrazine in the river water intake and sediments of a constructed flow-through wetland. Ecol Eng 72:35–39
Douglass JF, Radosevich M, Tuovinen OH (2015) Molecular analysis of atrazine-degrading bacteria and catabolic genes in the water column and sediment of a created wetland in an agricultural/urban watershed. Ecol Eng 83:405–412
Eaton RW, Karns JS (1991) Cloning and analysis of s-triazine catabolic genes from Pseudomonas sp. strain NRRLB-12227. J Bacteriol 173:1215–1222
El Sebaï T, Devers-Lamrani M, Changey F, Rouard N, Martin-Laurent F (2011) Evidence of atrazine mineralization in a soil from the Nile Delta: isolation of Arthrobacter sp. TES6, an atrazine-degrading strain. Int Biodeter Biodegrad 65:1249–1255
Ettema CH, Wardle DA (2002) Spatial soil ecology. Trends Ecol Evol 17:177–183
Fan X, Song F (2014) Bioremediation of atrazine: recent advances and promises. J Soils Sedim 14:1727–1737
Fenner K, Canonica S, Wackett LP, Elsner M (2013) Evaluating pesticide degradation in the environment: blind spots and emerging opportunities. Science 341:752–758
Ghosh D, Roy K, Srinivasan V, Mueller T, Tuovinen OH, Sublette K, Peacock A, Radosevich M (2009) In-situ enrichment and analysis of atrazine-degrading microbial communities using atrazine-containing porous beads. Soil Biol Biochem 41:1331–1334
Grigg BC, Assaf NA, Turco RF (1997) Removal of atrazine contamination in soil and liquid systems using bioaugmentation. Pestic Sci 50:211–220
Guerin WF, Boyd SA (1992) Differential bioavailability of soil-sorbed naphthalene to two bacterial species. Appl Environ Microbiol 58:1142–1152
Karns JS, Eaton RW (1997) Genes encoding s-triazine degradation are plasmid-borne in Klebsiella pneumoniae strain 99. J Agric Food Chem 45:1017–1022
Krutz LJ, Shaner DL, Weaver MA, Webb RMT, Zablotowicz RM, Reddy KN, Huang Y, Thomson SJ (2010) Agronomic and environmental implications of enhanced s-triazine degradation. Pest Manag Sci 66:461–481
Krutz LJ, Zablotowicz RM, Reddy KN (2012) Selection pressure, cropping system, and rhizosphere proximity affect atrazine degrader populations and activity in s-triazine-adapted soil. Weed Sci 60:516–524
Kulichevskaya IS, Ivanova AO, Belova SE, Baulina OI, Bodelier PLE, Rijpstra WIC, Damsté JSS, Zavarzin GA, Dedysh SN (2007) Schlesneria paludicola gen. nov., sp. nov., the first acidophilic member of the order Planctomycetales, from Sphagnum-dominated boreal wetlands. Int J System Evol Microbiol 57:2680–2687
Lane DJ (1991) 16S/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M (eds) Nucleic Acid Techniques in Bacterial Systematics. Wiley, Chichester, pp 115–175
Lazorko-Connon S, Achari G (2009) Atrazine: its occurrence and treatment in water. Environ Rev 17:199–214
Lian B, Jiang J, Zhang J, Zhao Y, Li S (2012) Horizontal transfer of dehalogenase genes involved in the catalysis of chlorinated compounds: evidence and ecological role. Crit Rev Microbiol 38:95–110
Liu X, Hui C, Bi L, Romantschuk M, Kontro M, Strömmer R, Hui N (2016) Bacterial community structure in atrazine treated reforested farmland in Wuying China. Appl Soil Ecol 98:39–46
Martinez B, Tomkins J, Wackett LP, Wing R, Sadowsky MJ (2001) Complete nucleotide sequence and organization of the atrazine catabolic plasmid pADP-1 from Pseudomonas sp. strain ADP. J Bacteriol 183:5684–5697
Masaphy S, Levanon D, Henis Y (1996) Degradation of atrazine by the lignocellulolytic fungus Pleurotus pulmonarius during solid-state fermentation. Bioresour Technol 56:207–214
Mougin C, Laugero C, Asther M, Dubroca J, Frasse P, Asther M (1994) Biotransformation of the herbicide atrazine by the white rot fungus Phanerochaete chrysosporium. Appl Environ Microbiol 60:705–708
Mudhoo A, Garg VK (2011) Sorption, transport and transformation of atrazine in soils, minerals and composts: a review. Pedosphere 21:11–25
Mueller TC, Steckel LE, Radosevich M (2010) Effect of soil pH and previous atrazine use history on atrazine degradation in a Tennessee field soil. Weed Sci 58:478–483
Mulbry WW, Zhu H, Nour SM, Topp E (2002) The triazine hydrolase gene trzN from Nocardioides sp. strain C190: cloning and construction of gene-specific primers. FEMS Microbiol Lett 206:75–79
Muyzer G, de Waal EC, Uitterlinden AG (1993) Profiling of complex microbial populations by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes coding for 16S rRNA. Appl Environ Microbiol 59:695–700
O’Donnell AG, Young IM, Rushton SP, Shirley MD, Crawford JW (2007) Visualization, modelling and prediction in soil microbiology. Nature Rev Microbiol 5:689–699
Omotayo AE, Ilori MO, Amund OO, Ghosh D, Roy K, Radosevich M (2011) Establishment and characterization of atrazine degrading cultures from Nigerian agricultural soil using traditional and Bio-Sep bead enrichment techniques. Appl Soil Ecol 48:63–70
Pussemier L, Goux S, Vanderheyden V, Debongnie P, Tresinie I, Foucart G (1997) Rapid dissipation of atrazine in soils taken from various maize fields. Weed Res 37:171–179
Radosevich M, Traina SJ, Hao Y-L, Tuovinen OH (1995) Degradation and mineralization of atrazine by a soil bacterial isolate. Appl Environ Microbiol 61:297–302
Radosevich M, Traina SJ, Tuovinen OH (1996) Biodegradation of atrazine in surface soils and subsurface sediments collected from an agricultural research farm. Biodegradation 7:137–149
Romero MC, Urrutia MI, Reinoso EH, Vedoval RD, Reynaldi FJ (2014) Atrazine degradation by wild filamentous fungi. Global Res J Microbiol 4:10–16
Rousseaux S, Hartmann A, Soulas G (2001) Isolation and characterisation of new Gram-negative and Gram-positive atrazine degrading bacteria from different French soils. FEMS Microbiol Ecol 36:211–222
Sadowsky MJ, Tong Z, de Souza M, Wackett LP (1998) AtzC is a new member of the amidohydrolase protein superfamily and is homologous to other atrazine-metabolizing enzymes. J Bacteriol 180:152–158
Sajjaphan K, Shapir N, Wackett LP, Palmer M, Blackmon B, Tomkins J, Sadowsky MJ (2004) Arthrobacter aurescens TC1 atrazine catabolism genes trzN, atzB, and atzC are linked on a 160-kilobase region and are functional in Escherichia coli. Appl Environ Microbiol 70:4402–4407
Sajjaphan K, Heepngoen P, Sadowsky MJ, Boonkerd N (2010) Arthrobacter sp. strain KU001 isolated from a Thai soil degrades atrazine in the presence of inorganic nitrogen sources. J Microbiol Biotechnol 20:602–608
Singh B, Singh K (2016) Microbial degradation of herbicides. Crit Rev Microbiol 42:245–261
Smith D, Alvey S, Crowley DE (2005) Cooperative catabolic pathways within an atrazine-degrading enrichment culture isolated from soil. FEMS Microbiol Ecol 53:265–273
Spanoghe P, Maes A, Steurbaut W (2009) Limitation of point source pesticide pollution: results of bioremediation system. Comm Appl Biol Sci Ghent Univ 74(2):1–14
Stamper DM, Radosevich M, Hallberg KB, Traina SJ, Tuovinen OH (2002) Ralstonia basilensis M91-3, a denitrifying soil bacterium capable of using s-triazines as nitrogen sources. Can J Microbiol 48:1089–1098
Tuovinen OH, Desmukh V, Özkaya B, Radosevich M (2015) Kinetics of aerobic and anaerobic biomineralization of atrazine in surface and subsurface agricultural soils in Ohio. J Environ Sci Health B50:718–726
Udiković N, Hršak D, Mendaš G, Filipčić D (2003) Enrichment and characterization of atrazine degrading microbial communities. Food Technol Biotechnol 41:211–217
Udiković Kolić N, Hršak D, Kolar AB, Petrić I, Stipičevic S, Soulas G, Martin-Laurent F (2007) Combined metabolic activity within an atrazine-mineralizing community enriched from agrochemical factory soil. Int Biodeter Biodegrad 60:299–307
Udiković-Kolić N, Hršak D, Devers M, Klepac-Ceraj V, Petrič I, Martin-Laurent F (2010) Taxonomic and functional diversity of atrazine-degrading bacterial communities enriched from agrochemical factory soil. J Appl Microbiol 109:355–367
Udiković-Kolić N, Scott C, Martin-Laurent F (2012) Evolution of atrazine-degrading capabilities in the environment. Appl Microbiol Biotechnol 96:1175–1189
Vargha M, Takátsc Z, Márialigeti K (2005) Degradation of atrazine in a laboratory scale model system with Danube river sediment. Water Res 39:1560–1568
Vaishampayan PA, Kanekar PP, Dhakephalkar PK (2007) Isolation and characterization of Arthrobacter sp. strain MCM B-436, an atrazine-degrading bacterium, from rhizospheric soil. Int Biodeter Biodegr 60:273–278
Vibber LL, Pressler MJ, Colores GM (2007) Isolation and characterization of novel atrazine-degrading microorganisms from an agricultural soil. Appl Microbiol Biotechnol 75:921–928
Zhang Y, Jiang Z, Cao B, Hu M, Wang Z, Dong X (2011) Metabolic ability and gene characteristics of Arthrobacter sp. strain DNS10, the sole atrazine-degrading strain in a consortium isolated from black soil. Int Biodeter Biodegrad 65:1140–1144
Zhang Y, Cao B, Jiang Z, Dong X, Hu M, Wang Z (2012) Metabolic ability and individual characteristics of an atrazine-degrading consortium DNC5. J Hazard Mater 237–238:376–381
Acknowledgments
The work was funded through the USDA National Research Initiative, grant no. 2004-35107-14884. JFD gratefully acknowledges a Fellowship from the Robert H. Edgerley Environmental Toxicology Fund, Division of Natural and Mathematical Sciences of the College of Arts and Sciences, Ohio State University. We thank two anonymous reviewers for insightful suggestions that were helpful in improving the manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible editor: Jizheng He
Rights and permissions
About this article
Cite this article
Douglass, J.F., Radosevich, M. & Tuovinen, O.H. Biomineralization of atrazine and analysis of 16S rRNA and catabolic genes of atrazine degraders in a former pesticide mixing site and a machinery washing area. J Soils Sediments 16, 2263–2274 (2016). https://doi.org/10.1007/s11368-016-1416-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11368-016-1416-3