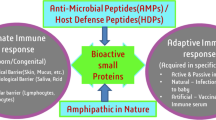
Abstract
In recent decades, antimicrobial resistance has been augmented as a global concern to public health owing to the global spread of multidrug-resistant strains from different ESKAPE pathogens. This alarming trend and the lack of new antibiotics with novel modes of action in the pipeline necessitate the development of non-antibiotic ways to treat illnesses caused by these isolates. In molecular biology, computational approaches have become crucial tools, particularly in one of the most challenging areas of multidrug resistance. The rapid advancements in bioinformatics have led to a plethora of computational approaches involving genomics, systems biology, and structural biology currently gaining momentum among molecular biologists since they can be useful and provide valuable information on the complex mechanisms of AMR research in ESKAPE pathogens. These computational approaches would be helpful in elucidating the AMR mechanisms, identifying important hub genes/proteins, and their promising targets together with their interactions with important drug targets, which is a crucial step in drug discovery. Therefore, the present review aims to provide holistic information on currently employed bioinformatic tools and their application in the discovery of multifunctional novel therapeutic drugs to combat the current problem of AMR in ESKAPE pathogens. The review also summarizes the recent advancement in the AMR research in ESKAPE pathogens utilizing the in silico approaches.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Hospital-acquired infections (HAI) caused by bacterial pathogens have emerged as a world health hazard and challenge to physicians due to their adaptation to antimicrobial resistance (AMR) (Chandler 2019). Conventional therapeutic drugs have largely improved the landscape of therapeutics. However, AMR in the microbial population remains a major problem in treating human infections, leading to the failure of conventional therapies. World Health Organisation (WHO) has recognized AMR as a global concern because of its recurrence in the emerging isolates predominantly caused by indiscriminate use of antibiotics. According to Global Antimicrobial Resistance and Use Surveillance System (GLASS), fluoroquinolones resistance has increased dramatically worldwide (70–80%) in Escherichia coli and Klebsiella pneumoniae, leaving carbapenems and colistin as the last resort. However, in Europe, Carbapenem-Resistant K. pneumoniae (CR-Kp) is the fastest-growing AMR threat, with a rising human mortality rate of 30–70% (Waddington et al. 2022). Recent reports have also suggested the acquisition of colistin resistance among extensive drug-resistant (XDR) enteric pathogens (El-Sayed Ahmed et al. 2020). According to GLASS, Methicillin-resistant Staphylococcus aureus (MRSA) and E. coli are developing resistance (12.11% and 36.0%, respectively) to third-generation cephalosporins antibiotics. Center for Disease Control and Prevention (CDC) has predicted that Acinetobacter baumannii and Enterobacteriaceae are the global threats, and cause pneumonia infection in the bloodstream (Enfield et al. 2014; Gupta et al. 2019).
Infectious Disease Society of America (IDSA) named these bacterial groups as ‘ESKAPE’ pathogens (Enterococcus faecium, Staphylococcus aureus, Klebsiella pneumoniae, Acinetobacter baumannii, Pseudomonas aeruginosa, and Enterobacter species) (Tiwari 2019). These pathogens become adaptable through time, and the overuse of antibiotics in clinical practices has resulted in resistance to antimicrobial drugs. These pathogens can transfer the AMR genes by gene mutation, extrachromosomal (plasmid-mediated), horizontal gene transfer (HGT) via conjugation or transformation, transposons, and integrons (Sun et al. 2019). Now-a-days, the synthesis of antimicrobial peptides (AMPs) gains an attention and has been considered as an alternative to the conventional therapies. They can easily interact with the microbial cell wall peptidoglycan through penetration or dissolving the biofilms (Bhattacharjya and Straus 2020; Dash and Bhattacharjya 2021; Bhattacharjya et al. 2022). Although AMPs may serve as promising leads against drug resistant microbes, only a few AMPs are in advanced clinical trials because of the development of a plethora of AMR in ESKAPE pathogens. This includes reduced cellular uptake of the drug, or enhanced efflux from cells, changed expression of drug targets, inactivating mutations of the targets, and others, resulting in fewer treatment options for patients and hence, increased morbidity and mortality (Goler-Baron and Assaraf 2011; Tolios et al. 2020).
To date, many research studies have unveiled countless putative AMR therapies. However, only a few of them have reached actual validation. In this context, bioinformatics research has immensely advanced over the last few years due to the high-throughput approaches introduced for promoting different aspects of contemporary biology. Currently, computational approaches serve different levels of micro-and macromolecular interactions responsible for the survival and development of numerous cellular systems (Vakser and Deeds 2019). With the dawn of genomics pipelines, specialized gene predictions, annotations, and variants calling for whole genomes and metagenomes, secondary analyses such as plasmid profiling have provided broad insights into the biomolecular framework of a living system (Su et al. 2019). The newly booming systems biology approach elucidates biomolecular cross-talks and their functional associations to shed light on the significant genes/proteins that can be interpreted as controlling hub biomolecules in the interaction networks (Sigurdsson et al. 2012). Tremendous improvements and immense applications of structural biology methods have encouraged scientists from all over the world to reach a new height in pharmaceutical research for solving a wide range of biomedical problems (Pandurangan et al. 2017; Ndagi et al. 2020). The hazardous complications concerning elevating levels of AMR in ESKAPE pathogens have compelled the bioinformaticians to scrutinize several possibilities for mitigating the dilemma.
The present review aims to provide up-to-date information on different computational approaches, encompassing genomics, systems biology, and structural biology, relevant to AMR research in ESKAPE pathogens. Further, in this study, we have compiled a comprehensive collection of various databases, servers, and tools to identify hub genes or proteins, the optimization of drug-target interactions, and, the use of these computational approaches to pave the way toward the discovery of novel drugs with multifunctional characteristics. This propels the finding of novel therapeutic possibilities against drug resistance in ESKAPE pathogens. This review could serve as a guide and provide insight to readers and medical researchers about the possibility and progression in designing potent therapeutic strategies for overcoming the current research gaps and problems associated with AMR in evolving ESKAPE pathogens.
Important ESKAPE pathogens
Enterococcus faecium
E. faecium is a Gram-positive coccus that causes severe nosocomial infections in humans and other immunocompromised patients (Bhatia et al. 2021). The resistance to antibiotics such as penicillin and vancomycin has been a significant concern worldwide. Glycopeptides are the most preferred choice for treating Enterococcal-mediated infections; however, their increasing resistance to antibiotics, viz., penicillin, tetracycline, vancomycin, and erythromycin has been a matter of global concern (Cauwerts et al. 2007; Pfaller et al. 2019).
Staphylococcus aureus
S. aureus is a Gram-positive coccus and is associated with different parts of the normal skin, especially the nose and perineum of humans and animals. This pathogen is typically harmful only in immunocompromised individuals. The infections caused by this species were treated with the well-known antibiotic penicillin; however, the repetitive use of this antibiotic could lead to the development of β-lactamase resistance. MRSA, an emerging strain of this pathogen, has evolved resistance against β-lactam antibiotics. Some other strains have also been identified as resistant to Vancomycin and are termed as Vancomycin-Resistant S. aureus (VRSA) (Loomba et al. 2010; McGuinness et al. 2017). The MRSA, coupled with the emergence of VRSA, has become a significant concern in the healthcare system.
Klebsiella pneumoniae
K. pneumoniae is an opportunistic Gram-negative bacterium that affects immunocompromised individuals and is associated with high morbidity and mortality due to the rare treatment options. K. Pneumonia reportedly involves extra-intestinal infections that includes urinary tract infections (UTIs), pneumonia, surgical wound infections and septicemia (Navon-Venezia et al. 2017; Ali et al. 2022). Besides this, K. pneumoniae shows intrinsic resistance toward ampicillin due to the presence of SHV-1 penicillinase in its chromosome. In contrast, resistance to other drugs may occur through chromosomal mutations or through the acquisition of AMR genes by HGT, especially through large conjugative plasmids (Wyres and Holt 2018).
Acinetobacter baumannii
A. baumannii is a Gram-negative pathogen and primary agent of nosocomial infections, including bacteremia, pneumonia, meningitis, and urinary tract infections in immune-compromised patients. This pathogen has been identified as one of the six most notorious microorganisms by the IDSA (Shafiee et al. 2021). Currently, owing to its unparalleled capacity to acquire foreign resistance mechanisms and prolonged survival in healthcare environments, large proportions of the A. baumannii strains are Multi Drug Resistant (MDR) and display high resistance to nearly all known antibiotics (Luo et al. 2015).
Pseudomonas aeruginosa
P. aeruginosa is a motile Gram-negative bacterium that affects patients, especially those who recovered from post-surgery or during their admission in Intensive Care Unit (Hilliam et al. 2020). In addition, a number of intrinsic (genetically encoded in the core genome) and acquired resistance (rises from the gain of resistance genes from other microorganisms) mechanisms exist within the P. aeruginosa population. Further, this pathogen possesses a whole armamentarium of virulence factors, including toxins, adhesins, siderophores, and others (Strateva and Mitov 2011). Sometimes the cooperative interaction between antibiotic resistance and virulence may lead to the development of ‘high-risk’ clones, which are widely disseminated and challenging to treat (Horna and Ruiz 2021).
Enterobacter species
Enterobacter spp. is motile Gram-negative bacilli belonging to the Enterobacterales order. Major nosocomial pathogens include E. cloacae, E. hormaechei, and E. aerogenes and they cause lung, soft tissue, intra-abdominal cavity, and urinary tract infection (Davin-Regli et al. 2019). The complication in treatment regimens has emerged due to increasing drug-resistant strains. E. aerogenes show resistance to third-generation cephalosporins by releasing β-lactamases and hydrolyzing the β-lactam ring. E. cloacae imparts resistance to aminoglycoside gentamicin by acquiring genetic elements, i.e., integrons conferring AMR genes (Davin-Regli and Pagès 2015).
Mechanism of Antibiotic Resistance attained by ESKAPE pathogens
The pathogens acquire resistance through various mechanisms, such as inhibition of the drug influx, modification of the drug target, enzymatic inactivation and alteration of the drug, activation of drug efflux pump resulting in rapid extrusion of drugs (Santajit and Indrawattana 2016). The well-known AMR mechanisms in ESKAPE pathogens have been depicted in Fig. 1.
A schematic representation of the AMR mechanisms in ESKAPE pathogens. Bacteria may resist the action of drugs by (a) Inhibition of the drug influx (cell membrane modification), (b) modification of the drug target site, (c) inactivation of drug-by-drug inactivating enzymes, (d) drug alteration, and (e) activation of drug efflux pump resulting in rapid extrusion of drugs
In silico approaches for the study of AMR in ESKAPE pathogens
Modern in silico approaches have made the drug target identification process more accessible than the conventional model (Basu et al. 2021b). There are various in silico approaches involved in genomics, systems biology and structural analysis and they are schematically represented in Fig. 2. These approaches play a significant role in identifying AMR genes, optimizing drug targets and screening lead compounds to identify those that could help to combat the resistance.
Understanding pathogenic transmission is essential to manage the outbreak that can be inferred through genotyping methods. Genome studies, including whole-genome analysis, transcriptomics studies, plasmid profiling, metagenomics and phylogenetic analysis, provide insights into the transmission, virulence and AMR (Quainoo et al. 2017). Systems biology helps to identify hub genes with their interacting partners that have a critical role at the molecular level in causing resistance, whereas structural biology plays a pivotal role in drug discoveries and could combat the resistance (Ndagi et al. 2020). Different in silico approaches to identify AMR genes and their function have been discussed below.
Genomics approaches to identify AMR genes in ESKAPE pathogens
The genomics approaches have revolutionized the identification of specific novel AMR genes in various ESKAPE pathogens (Hosen et al. 2014). Different bioinformatics databases and tools are shown in Fig. 3, which facilitates a comparative genomics approach to prioritize and identify specific AMR determinants. The genomics approach applies two criteria for drug target identification, i.e., “essentiality” and “selectivity” (Hossain et al. 2017). These AMR genes are involved in unique metabolic pathways which would help to design novel drugs and therapeutic components to combat the problem of resistance.
Whole Genome Sequencing (WGS) analysis is a high throughput technology that identifies AMR determinants and aids in developing novel antibiotics by elucidating various factors for resistance. It determines the genetic variants such as Single Nucleotide Polymorphism (SNPs), frameshift mutations, and copy number variation (CNVs) in the altered genome (Crofts et al. 2017; Hunt et al. 2017; Shi et al. 2019). WGS is used to acknowledge the infection and helps to diagnose the cause of resistance (Köser et al. 2014). In order to predict the factors responsible for phenotypic resistance in pathogens, different computational approaches, such as read-based (Genefinder), de Bruijn graph-based (Mykrobe), and BLAST-based (Typewriter) are used. The WGS study’s implication has helped AMR surveillance by consuming less time and cost. Wareth et al. (2021) have identified a broad range of AMR genes such as blaOXA−51-like, blaOXA−23, blaADC−25, blaADC−73, blaTEM−1 and blaNDM−1, which mediates resistance against β-lactam antibiotics. Another set of genes which shows resistance against aminoglycosides are ant(3”)-IIa, aph(3”)-Ib, armA, aph(6)-Id and aph(3’)-Ia. The gene catB8 has been identified to promote resistance against phenicol’s, tet.B and tet.39 against tetracyclines, and sul.1 and sul.2 against sulfonamides. Certain genes such as mphE, msrE and abaF have been reported to play a significant role in bacterial resistance to macrolides and lincosamide. The presence of tetM and tetL genes in Enterococcus strains show resistance against tetracycline, ermB against erythromycin, and vanA, vanB, and vanM shows resistance against vancomycin (Lee et al. 2020; Cui et al. 2020).
In S. aureus several genes, including blaZ against penicillin, mecA against methicillin, msrA against erythromycin, ermA, ermB, ermC, and ermT against erythromycin and clindamycin, tetK, tetL, and tetM against tetracycline, vanA against vancomycin; and fusA against fusidic acid have been reported to be responsible for the development of antimicrobial resistance (Gordon et al. 2014). Likewise, in K. pneumoniae, there are many genes (viz., mcr-1, blaCTX−M, blaOKP, blaOXA, blaSHV, blaTEM, blaLEN, aac(3′)-Ia, aac-(3′)-IIe, aac(6′)-Ib-cr, aac-(3′)-IId, aadA4, aadA5, aadA1a, aadA16, aph(6′)-Id, aph(3′)-Ia, qnrB, and qnrS) have been studied and reported to cause resistance against colistin (Stoesser et al. 2013; Schürch and van Schaik 2017). In A. baumannii, the co-existence of blaOXA−23, blaOXA−66, blaADC−25, and armA genes have been reported to be responsible for the development of multidrug resistance against carbapenems (Rao et al. 2020; Hwang et al. 2021); gyrA and parC show resistance against fluoroquinolones (Lean et al. 2015). EnvZ (osmolarity sensor histidine kinase) and phosphate regulon response regulator are associated with a two-component system by which E. coli show resistance against carbapenems. While, FtsZ gene is involved in the β-lactam resistance pathway in S. aureus and is sensitive to β-lactam antibiotics.
Transcriptomics analysis includes the RNA-sequencing of the bacterial gene expression that has the potential to fill the gap of antimicrobial resistance. The transcriptomic data links the genotypic and phenotypic results, which are important in understanding the resistance mechanism. This analysis also helps to reveal the putative gene effect that contributes to antimicrobial resistance and identifies cases where non-coding regulatory RNA promotes bacterial resistance. In P. aeruginosa, the transcriptome analysis identifies the key genes which show the resistance against aminoglycosides, β-lactam, and fluoroquinolone antibiotics (Khaledi et al. 2016). Another study states that the mutation in a two-component system (amgRS and pmrAB) shows colistin resistance (Schniederjans et al. 2017). In the case of K. pneumoniae, blaNDM-1 is mainly responsible for carbapenem resistance (Low et al. 2018). It has been reported that the downregulation of SprX might lead to increased resistance in S. aureus (Eyraud et al. 2014).
Metagenomics analysis provides insights and elucidates a relationship between AMR and the microbiome based on a microbial community’s structural and functional diversities. The advantage of metagenome analysis is that it covers a more significant number of species at a time and identifies the potential AMR determinants together with functional gene analysis. It employs two approaches for analysis: a function-driven approach that screens for an expressed trait and a sequence-driven approach for particular DNA sequences (Schloss and Handelsman 2003). Metagenomic data analysis starts from raw data sequencing, quality control, and obtaining high-quality reads. The reads are assembled using reference-based or de novo, for which different tools are available, represented in Fig. 3.
The contigs or scaffolds generated are further used for gene prediction. A list of gene sequences is collected and annotated, followed by putative gene function identification. Depending on the results, the taxonomical analysis is performed. The pipeline integrates many bioinformatics algorithms that address different aspects of complete metagenomic analysis. The downstream analysis identifies the antibiotic resistance genes and plasmids through gene network analysis. A study by (Matamoros-Recio et al. 2021) proved that the metagenome sequences from the sample and Comprehensive Antibiotic Resistance Database could be screened to find the antibiotic resistance gene. The analysis found that 259 ARGs are included in the database that contributes to antibiotic resistance. Yadav and Kapley (2019) carried out a metagenomic analysis study to monitor the AMR genes in E. coli and found a mobile genetic element, P-1 in plasmid pKJK5 that causes resistance. In addition, IncP-1beta (multi-resistance plasmid) and pB8 are important in spreading AMR through horizontal gene transfer.
Plasmid profiling is another important genomics approach that acknowledges the problem of resistance in bacterial pathogens by exchanging the AMR genes through HGT. The accessory genes responsible for the development of virulence and antibiotic resistance have clinically relevant traits. The plasmid profiling of these genes could be used to understand their molecular epidemiology. It has been illustrated that IncFIB plasmid is the most prevalent in E. coli and K. pneumoniae. Genes such as TEM-1B, aph, sul, tetA, and dfr are present in the plasmid showed resistance against penicillins, aminoglycosides, sulfonamides, tetracycline, and trimethoprim. In Norwegian isolates of E. coli and K. pneumoniae, ARGs viz., aph(3″)-Ib, aph(6)-Id, sul1, sul2, tet(D), and qnrS1 have been reported to be the most abundant plasmid-mediated resistance; plasmid with blaCTX−M gene and its interaction may be responsible for the development of resistance in these pathogens (Khezri et al. 2021). Plasmid types such as IncHI1B(pNDM-MAR), IncFIB(pQil), IncFIA, IncFII(K), IncR, IncFII(pRSB107), IncFIB(Mar), ColKP3 and ColpVC were found in K. pneumoniae strain from south India that were found to be involved in resistance. IncFIA, IncFII, IncFIB, Col (BS512), IncL1, IncX3 and IncH were mainly present in other isolates that showed resistance. In S. aureus, seven different plasmid groups viz., rep5, pSAS1 (rep7), pDLK1 (rep10), pUB110 (repUS12), Saa6159 (rep16), pKH12 (rep21), and pSA1308 (rep21) has been addressed for resistance. The E. coli plasmid IncHI1B(R27) with blaNDM−1 and blaCTX−M−15 sites and K. pneumoniae plasmids IncHI1B (pNDM-MAR) with blaCTX−M−15 site promotes resistance against β-lactamases. The blaTEM−1B shows a relationship between the genotypic and phenotypic expression of AMR that could further help to improve the strategies for better control of AMR dissemination (Ragupathi et al. 2019).
The genomics approaches adhere to profound surveillance to identify antimicrobial resistance with a greater fidelity that could refine the pathogenic risk. The amalgamation of the genomics approach with systems biology can help to deduce AMR networks and their functional analysis. The genomics approach shares a synergic area of interest with the systems biology field of network deduction and biological pathway analysis. Once the differentially expressed AMR genes are identified, their functional partners and mechanism of action can be better comprehended using the systems biology approach discussed below.
Systems Biology approach to identify hub gene/protein contributing towards AMR in ESKAPE pathogens
Systems biology-based networks are essential in pathological studies to decipher the disease-causing genes and their associated functional partners (Grimes et al. 2019). The approach mainly focuses on the coordinated systems-level interactions and they are crucial for the underlying mechanisms in various infectious diseases. The approach using gene interaction network (GIN) studies have efficiently explored the AMR mechanisms by identifying the potential target genes that can be targeted using a novel or repurposed drug (Debroy et al. 2020).
Systems biology approaches can be applied to decipher various insights about pathogenic genes (Miryala and Ramaiah 2019). The interaction data curated from various databases can be inferred to understand the direct functional interactors of pathogenic genes. Moreover, the indirect interactors essential for the progression of infection can be correlated using the interaction data. The constructed interactome can be better studied using various analyses such as clustering, topological parameter (TPA), functional enrichment (FEA) and pathway analyses (Naha et al. 2020). Clustering analysis aids in identifying stable gene clusters with a high probability of performing similar functions. Identifying such clusters is often important, as confining a set of genes from the interactome increases their specificity. Once the important pathogen-specific clusters are identified, their functional role can be validated using FEA. It consists of three main gene ontology terms, namely Biological Process (BP), Molecular Functions (MF) and Cellular Component (CC). The identification of enriched BP, MF and CC helps to easily comprehend various pathogen-associated genes’ functional status (Ashok et al. 2021).
Additionally, pathway analyses project the predominantly enhanced biological or signaling pathways critical for organism-induced infection. Another key insight that can be retrieved using TPA is the significance of each gene in the constructed interactome. This can be achieved by computing the various topological matrices and centrality parameters. Different parameters signify the genes importance and connectivity in the interactome. Network biology studies can be based on different biological entities depending on the biomolecule of interest such as genes, proteins, small molecules, and regulatory molecules. Therefore, different kinds of interactome can be constructed such as GIN, Protein-protein interaction (PPI), Gene Regulatory Network (GRN), Gene-co-expression Network (GCN), Drug-Gene interaction network (DGIN) and Host-Pathogen Interaction Network (HPIN) (Miryala et al. 2018).
PPI studies on antimicrobial-resistant genes have recognized different mechanisms in ESKAPE pathogens. Based on a PPI study, in S. aureus, the following genes have been identified to play a role in AMR resistance and they are NorA, aacA-aphD (aac6ie), aad9ib (ant), aadd (knt), baca (uppP), bl2a_pc (blaZ), ble, ermA, SAV0052 (ermb), ermc, fosB, mecA (mecI), mecR (mecr1), mepA, msrA1, qacA, vraR (str), tet38 and tetM (Anitha et al. 2016). A similar study in K. pneumoniae observed the role of SHV-11 gene in drug-resistance and functional partner genes gyrA, parC, glsA, osmE, yjhA, yhdT, rimL, pepB, KPN_00437, KPN_01875 (Miryala et al. 2020). In the case of A. baumannii, it has been reported that the presence of dfrA1 and dfrA10 genes confers resistance against trimethoprim (Anitha et al. 2014). The genes adeA, adeB and adeC are major components of adeABC efflux-pump and have been predicted to cause resistance in A. baumannii. The AMR genes associated with P. aeruginosa (arnA, cat, gyrA, str, fabG, parC, parE, phoP, phoQ, pmrApmrB) have also been identified to show resistance through gene network study. Genes gyrA and parC were also shown to be enriched through gene ontology terms that infer their role in P. aeruginosa (Miryala et al. 2019).
Intracellular pathogen requires an understanding of molecular-level interactions between the host and the pathogens, which are critical for their mechanism of infection. This interaction also indicates the probable drug-resistance mechanism in pathogens against antimicrobial therapeutics (Nourani et al. 2015; Basu et al. 2021a). The HPI network study in E. faecium has inscribed the role of hub genes asrR and vanA promoting AMR by associating with resistance to oxidative stress and membrane vesicle, which shows an increase in the survivability within the host (Lebreton et al. 2012; Wagner et al. 2018). In K. pneumoniae, mgrB mutation resists polymyxin by governing PhoPQ (Kidd et al. 2017), while AcrAB efflux pump is another virulence factor that resists the defense and promotes virulence in K. pneumoniae (Padilla et al. 2010). The antibiotic resistance genes specific to biofilm in P. aeruginosa are ndvB, PA1875–1877 and tssC1 (Zhang et al. 2013) and Esp protein in S. aureus (Sugimoto et al. 2013) have been studied, respectively. The interaction between host and bacterial protein could either lead to tolerance or resistance in the host. Thus, these interactions may lead to bacteria colonization, causing drug insensitivity (Casadevall and Pirofski 2000). The experimental methods to check the interaction between thousands of human proteins and pathogen proteins are expensive and time-consuming. So, systems biology approaches can be a better-suited means of predicting the putative interactions that can be further validated using experimental techniques. PPI also helps to address the important pathways that help bacteria survive and cause pathogenic virulence in the host system (Miryala and Ramaiah 2022). The various tools that can be utilized for the different analyses are depicted in Fig. 4.
An outline of tools and databases in Systems biology approach. Protein-Protein Interaction (PPI), Host-Pathogen Interaction (HPI), Gene-Regulatory Network (GRN), Drug-Gene/Protein Interaction (DGI), Gene-Gene Interaction (GGI), Gene Co-expression Interaction studies can aid to elucidate biomolecular level cross-talks that are involved in AMR mechanisms
Systems biology approaches to identify targets are easy, time-saving and cost-effective. Instead of the conventional trial and error method, which consumes time and money, systems biology approaches using network studies can efficiently identify functional targets playing a crucial role in infectious diseases. The identified target can be structurally validated using various structural biology approaches discussed below.
Structural Biology approach for target optimization and drug discovery to combat AMR in ESKAPE pathogens
Structural assessment and dynamicity of proteins, peptides and chemical compounds can be substantial tasks in computational biology that are being reliably dealt with the existing structural biology methods (Edwards and Cottage 2003). Preliminary data for structural evaluations can be collected from text-mining, literature-based databases and tools, and independent analyses in genomics and systems biology. The various servers and tools that are being commonly employed in the major aspects involved in the structural biology approaches have been represented in Fig. 5.
Important tools and databases commonly utilized in Structural biology approach. Protein Optimization and Validation, Mutational Study, Lead Identification and Optimization, Dynamics Simulation and Molecular Docking are vital aspects for designing therapeutic strategies against drug-resistant ESKAPE pathogens
Macromolecular structural predictions or modeling amalgamated with step-wise optimizations and validations have become vital sectors of contemporary bioinformatics. Protein optimization or refinement involves generating an improved 3D-structure of the protein with abundant Ramachandran favoured regions and a low number of Ramachandran outliers and poor rotamers (Sobolev et al. 2020). For this, energy minimization could be performed to obtain the protein conformation with the least internal energy, which corresponds to a stable protein configuration. Further, secondary structural predictions are being performed to determine the regions of α-helices, β-sheets and random coils in the protein (Ma et al. 2018). Secondary and tertiary structural validations include evaluation of the local and gross errors, number of misfolds, disorderness, solvent accessibility and thermal stability, and the backbone flexibility in the protein (Kesheri et al. 2015). In order to understand the functionality of a protein, active site identification and validation play an important role in studying the structure-function or structure-activity relationships (SAR) (Guha 2013). The stability analyses encompass the protein’s residual fluctuations and compactness that can be assessed through dynamics simulations (Justino et al. 2021). The prediction of drug-binding sites or pockets (druggability) in the protein structure must be executed to establish a protein as a therapeutic target.
The study of protein mutations is an important aspect of SAR studies, especially in the context of AMR evaluation in bacteria, since mutations can be responsible for transforming a non-AMR protein into an AMR protein and vice versa. Hence, a mutation’s functional outcome (gain or loss of function) is being assessed within the protein of interest in the emerging isolates (Singh et al. 2021; Shankar et al. 2021). The emergence of mutations in the conventional AMR proteins may result in the ineffective binding of standard drugs and has resulted in the investigation of alternative drug targets in many ESKAPE pathogens (Saha and Sarkar 2021).
On the other hand, ligand identification through virtual screening and prediction of its pharmacokinetic/ pharmacodynamics (PK/PD) profile has been widely performed as preliminary analyses in subsequent drug discovery (Gallo 2010). Ligands with favourable ‘Absorption, Distribution, Metabolism, Excretion and Toxicity’ (ADMET) profiles and high LD50 values need to be shortlisted for being considered safe for administration (Wu et al. 2020). Ligand optimizations, electrochemical stability, and reactivity profiles have greatly facilitated research revolving around drug design and repurposing.
The intermolecular interactions viz., protein-drug, protein-protein, and protein-nucleic acid accompanied by dynamics simulations have been proven tremendously beneficial in modern-day targeted therapy (Basu et al. 2020). Molecular docking has introspected the understanding of the interaction between the ligands and pathogen proteins by predicting their binding affinity through the estimated binding energy or cumulative docking scores (Takaya et al. 2008; Dar and Mir 2017). The intermolecular interaction profiles encompassing direct Hydrogen bonds, van der Waals forces and other non-canonical interactions in the protein-ligand complexes are being extensively studied (Varghese et al. 2022). Molecular dynamics simulations of the docked complexes indicate their stability profiles and strengthen the notion of the proposed therapeutic option (Miryala et al. 2021, 2022; Karthika et al. 2021). The protein-ligand complex studies have boosted the therapeutics development to combat the current problem (Naha et al. 2021).
Computer-aided drug design has been used to identify ZINC48942, which binds to the ‘Scm’ receptor that is involved in infection caused by E. faecium (Rasheed et al. 2021). Drugs such as ofloxacin, furazolidone and roflumilast affinity against the ‘FmtA’ active site is well studied in S. aureus and have a role in methicillin resistance (Dalal et al. 2021). Through virtual screening 1,3,4-oxadiazole (Naaz et al. 2021), 2-phenylquinolines (Sabatini et al. 2013), and CID 44,330,438 (Zárate et al. 2019) have been identified as potent inhibitors of NorA efflux pump that can combat AMR. It has been shown that BBN149 acts as the putative inhibitor of ArnT inhibitor in K. pneumoniae and P. aeruginosa (Ghirga et al. 2020). ZINC21811621, ZINC93091917 and ZINC19488569 have been identified as potent inhibitors against CTX-M-15 in K. pneumoniae (Farhadi et al. 2018). ZINC12670903, ZINC17465965, ZINC11681166 and ZINC13099024 against AmpC/ β-lactamases in P. aeruginosa (Farmer et al. 2010). In A. baumannii, BfmR is mainly involved in biofilm formation and, through structural analysis, compounds such as Calystegine B3,7,7 A-Diepialexine and Alpha-Methylnoradrenaline could bind to its active site (Lokhande et al. 2022). Another study pertaining to a similar approach has addressed ZINC4085364 as the potent inhibitor of blaVEB−1 in E. coli and P. aeruginosa (Messaoudi et al. 2013).
In order to combat multidrug resistance, the use of AMPs is one of the most efficient alternatives to the conventional approach. AMPs with therapeutic and antimicrobial activity against Gram-positive and Gram-negative bacteria are isolated from bacteria. (Mwangi et al. 2019). AMPs can be potentially useful in treating drug-resistant bacteria (Bhattacharjya and Straus 2020). Some AMPs such as Lactoferricin B and LL-37 belonging to both anionic and cationic peptide fragments have been reported to be effective against K. pneumoniae, S. aureus, and Enterobacter spp. Other sets of AMPs such as Parkerin, Ranalexin, BM Moricin, Melittin, and Circulin-A target P. aeruginosa and S. aureus. A study on Lactoferricin B-Mutant (M4) shows that this class of AMP have good binding high affinity towards SHV1, OXB48, NDM1, and AmpC, with the highest affinity for NDM1 (Basu et al. 2022). Polymyxin, another effective peptide with antimicrobial properties shows its effectiveness against P. aeruginosa, A. baumannii, and K. pneumoniae (Vaara 2019). Thanatin is another AMP that can be used against both Gram-positive and Gram-negative bacteria (Dash and Bhattacharjya 2021).
Depending on the structures, it has been reported that different AMPs having a different modes of action against bacterial pathogens depending on the structuere have been reported (Bhattacharjya et al. 2022). In silico approach can be implicated in designing and evaluating the antimicrobial activity of AMPs. Different databases such as Anti-microbial Peptide Database (APD3) and PhytAMP database have a collection of AMPs and are screened according to different classes such as anionic, cationic, cysteine-rich, and anionic/cationic peptide fragments. The efficacy of the AMPs can be enhanced by altering the template sequence (Duval et al. 2009). The enhancements of the altered/mutant AMPs can be assessed by iAMPred web tool, whereas HLP and CellPPD can be used to evaluate the physiochemical property of the parent AMPs to set a standard. For antigenic and toxicity prediction of AMPs, antigenic peptide tool and ToxinPred servers are being used. The 3D structures of the AMPs can be well optimised and modeled using servers like PEP-FOLD-3 and Swiss-Model servers, while the model can be validated by MolProbity tool. On the other hand, different protein structures of the ESKAPE pathogens can be retrieved and optimised by different tools mentioned in Fig. 5. The docking of AMPs with bacterial proteins can be performed through ClusPro2.0 and HPEP-DOCK servers. The simulation of the peptides-protein complex with superior binding profiles can be performed by CABS-dock server. Further, the stability of the complex can be determined using the MDWeb server (Basu et al. 2022).
Future directions
In silico approach not only facilitates the identification of novel resistance genes or proteins but also helps to enhance therapeutic options. As genome and protein sequencing are easily accessible and relentlessly performed in the present scenario, a staggering amount of high-throughput data has been generated and continues to date. The robust model of predicting the resistant genes helps to regulate the resistance at the local and global levels to monitor within countries. Determining precise resistant genes will help to facilitate the personalized approach to drive the current treatment regimens to the next level. Furthermore, Artificial Intelligence and Machine Learning algorithms can be applied to omics-based studies to establish advancements in medical informatics. The currently employed computational approaches emphasize enriching the therapeutic options and identifying the potential drug candidates that could combat the emerging and re-emerging AMR in the ESKAPE pathogens. Structural biology methods involving the prediction and optimization of therapeutic drug targets and potential lead molecules can be a promising approach to mitigate the dilemma of AMR. In antibiotic research, the early finding of prospective therapeutic targets and lead compounds is extremely desirable in order to reduce the research duration and experimental expenses. The review hopes for an enriching future of novel drug discovery against the ESKAPE pathogens and establishing a healthy collaboration between computational and experimental researchers.
Conclusions
The recent advancements in gaining knowledge of AMR in ESKAPE pathogens have been supported through different bioinformatics services in genomics, systems and structural biology. Numerous computational servers and tools have been designed and are currently being utilized globally to address the problem of creeping AMR in the recent ESKAPE isolates. The genomics approach provides a fast and affordable way of analyzing the outbreak of resistance and monitoring the AMR profiles in the emerging ESKAPE strains globally. Systems biology approaches have helped us identify the AMR genes and their interacting partners to better understand the AMR mechanisms in the pathogens. Structural biology methods have tremendously aided in investigating several alternative therapeutic options against the detected drug resistance. This review covers different aspects of the bioinformatics approaches which can be utilized to formulate effective research strategies for understanding the molecular mechanisms of drug resistance in ESKAPE pathogens. These approaches play a pivotal role in accelerating and enhancing drug discovery to combat bacterial drug resistance.
References
Ali S, Alam M, Hasan GM, Hassan MI (2022) Potential therapeutic targets of Klebsiella pneumoniae: a multi-omics review perspective. Brief Funct Genomics 21:63–77. https://doi.org/10.1093/bfgp/elab038
Anitha P, Anbarasu A, Ramaiah S (2016) Gene network analysis reveals the association of important functional partners involved in antibiotic resistance: A report on an important pathogenic bacterium Staphylococcus aureus. Gene 575:253–263. https://doi.org/10.1016/j.gene.2015.08.068
Anitha P, Anbarasu A, Ramaiah S (2014) Computational gene network study on antibiotic resistance genes of Acinetobacter baumannii. Comput Biol Med 48:17–27. https://doi.org/10.1016/j.compbiomed.2014.02.009
Ashok G, Miryala SK, Anbarasu A, Ramaiah S (2021) Integrated systems biology approach using gene network analysis to identify the important pathways and new potential drug targets for Neuroblastoma. Gene Rep 23:101101. https://doi.org/10.1016/j.genrep.2021.101101
Basu S, Joshi SM, Ramaiah S, Anbarasu A (2022) Designing Anti-Microbial Peptides Against Major β-Lactamase Enzymes in Clinically Important Gram-Negative Bacterial Pathogens: An In-Silico Study. Probiotics Antimicrob Proteins 14:263–276. https://doi.org/10.1007/s12602-022-09929-1
Basu S, Naha A, Veeraraghavan B et al (2021a) In silico structure evaluation of BAG3 and elucidating its association with bacterial infections through protein–protein and host-pathogen interaction analysis. J Cell Biochem jcb 29953. https://doi.org/10.1002/jcb.29953
Basu S, Ramaiah S, Anbarasu A (2021b) In-silico strategies to combat COVID-19: A comprehensive review. Biotechnol Genet Eng Rev 37:64–81. https://doi.org/10.1080/02648725.2021.1966920
Basu S, Veeraraghavan B, Ramaiah S, Anbarasu A (2020) Novel cyclohexanone compound as a potential ligand against SARS-CoV-2 main-protease. Microb Pathog 149:104546. https://doi.org/10.1016/j.micpath.2020.104546
Bhatia P, Sharma A, George AJ et al (2021) Antibacterial activity of medicinal plants against ESKAPE: An update. Heliyon 7:e06310. https://doi.org/10.1016/j.heliyon.2021.e06310
Bhattacharjya S, Mohid SA, Bhunia A (2022) Atomic-Resolution Structures and Mode of Action of Clinically Relevant Antimicrobial Peptides. Int J Mol Sci 23:4558. https://doi.org/10.3390/ijms23094558
Bhattacharjya S, Straus SK (2020) Design, Engineering and Discovery of Novel α-Helical and β-Boomerang Antimicrobial Peptides against Drug Resistant Bacteria. Int J Mol Sci 21:5773. https://doi.org/10.3390/ijms21165773
Casadevall A, Pirofski L (2000) Host-Pathogen Interactions: Basic Concepts of Microbial Commensalism, Colonization, Infection, and Disease. Infect Immun 68:6511–6518. https://doi.org/10.1128/IAI.68.12.6511-6518.2000
Cauwerts K, Decostere A, De Graef EM et al (2007) High prevalence of tetracycline resistance in Enterococcus isolates from broilers carrying the erm(B) gene. Avian Pathol 36:395–399. https://doi.org/10.1080/03079450701589167
Chandler CIR (2019) Current accounts of antimicrobial resistance: stabilisation, individualisation and antibiotics as infrastructure. Palgrave Commun 5:15–17. https://doi.org/10.1057/s41599-019-0263-4
Crofts TS, Gasparrini AJ, Dantas G (2017) Next-generation approaches to understand and combat the antibiotic resistome. Nat Rev Microbiol 15:422–434. https://doi.org/10.1038/nrmicro.2017.28
Cui P, Feng L, Zhang L et al (2020) Antimicrobial Resistance, Virulence Genes, and Biofilm Formation Capacity Among Enterococcus species From Yaks in Aba Tibetan Autonomous Prefecture, China. https://doi.org/10.3389/fmicb.2020.01250. Front Microbiol 11:
Dalal V, Dhankhar P, Singh V et al (2021) Structure-Based Identification of Potential Drugs Against FmtA of Staphylococcus aureus: Virtual Screening. MM-GBSA and QM/MM Protein J 40:148–165. https://doi.org/10.1007/s10930-020-09953-6. Molecular Dynamics
Dar AM, Mir S (2017) Molecular Docking: Approaches, Types, Applications and Basic Challenges. J Anal Bioanal Tech 08. https://doi.org/10.4172/2155-9872.1000356
Dash R, Bhattacharjya S (2021) Thanatin: An Emerging Host Defense Antimicrobial Peptide with Multiple Modes of Action. Int J Mol Sci 22:1522. https://doi.org/10.3390/ijms22041522
Davin-Regli A, Lavigne J-P, Pagès J-M (2019) Enterobacter spp.: Update on Taxonomy, Clinical Aspects, and Emerging Antimicrobial Resistance. Clin Microbiol Rev 32. https://doi.org/10.1128/CMR.00002-19
Davin-Regli A, Pagès J-M (2015) Enterobacter aerogenes and Enterobacter cloacae; versatile bacterial pathogens confronting antibiotic treatment. Front Microbiol 6. https://doi.org/10.3389/fmicb.2015.00392
Debroy R, Miryala SK, Naha A et al (2020) Gene interaction network studies to decipher the multi-drug resistance mechanism in Salmonella enterica serovar Typhi CT18 reveal potential drug targets. Microb Pathog 142:104096. https://doi.org/10.1016/j.micpath.2020.104096
Edwards YJK, Cottage A (2003) Bioinformatics Methods to Predict Protein Structure and Function: A Practical Approach. Mol Biotechnol 23:139–166. https://doi.org/10.1385/MB:23:2:139
El-Sayed Ahmed MAE-G, Zhong L-L, Shen C et al (2020) Colistin and its role in the Era of antibiotic resistance: an extended review (2000–2019). Emerg Microbes Infect 9:868–885. https://doi.org/10.1080/22221751.2020.1754133
Enfield KB, Huq NN, Gosseling MF et al (2014) Control of Simultaneous Outbreaks of Carbapenemase-Producing Enterobacteriaceae and Extensively Drug-Resistant Acinetobacter baumannii Infection in an Intensive Care Unit Using Interventions Promoted in the Centers for Disease Control and Prevention 2012. Infect Control Hosp Epidemiol 35:810–817. https://doi.org/10.1086/676857
Eyraud A, Tattevin P, Chabelskaya S, Felden B (2014) A small RNA controls a protein regulator involved in antibiotic resistance in Staphylococcus aureus. Nucleic Acids Res 42:4892–4905. https://doi.org/10.1093/nar/gku149
Farhadi T, Fakharian A, Ovchinnikov RS (2018) Virtual Screening for Potential Inhibitors of CTX-M-15 Protein of Klebsiella pneumoniae. Interdiscip Sci Comput Life Sci 10:694–703. https://doi.org/10.1007/s12539-017-0222-y
Farmer R, Gautam B, Singh S et al (2010) Virtual screening of AmpC/β-lactamase as target for antimicrobial resistance in Pseudomonas aeruginosa. Bioinformation 4:290–294. https://doi.org/10.6026/97320630004290
Gallo JM (2010) Pharmacokinetic/ Pharmacodynamic-Driven Drug Development. Mt Sinai J Med A J Transl Pers Med 77:381–388. https://doi.org/10.1002/msj.20193
Ghirga F, Stefanelli R, Cavinato L et al (2020) A novel colistin adjuvant identified by virtual screening for ArnT inhibitors. J Antimicrob Chemother 75:2564–2572. https://doi.org/10.1093/jac/dkaa200
Goler-Baron V, Assaraf YG (2011) Structure and Function of ABCG2-Rich Extracellular Vesicles Mediating Multidrug Resistance. PLoS ONE 6:e16007. https://doi.org/10.1371/journal.pone.0016007
Gordon NC, Price JR, Cole K et al (2014) Prediction of Staphylococcus aureus Antimicrobial Resistance by Whole-Genome Sequencing. J Clin Microbiol 52:1182–1191. https://doi.org/10.1128/JCM.03117-13
Grimes T, Potter SS, Datta S (2019) Integrating gene regulatory pathways into differential network analysis of gene expression data. Sci Rep 9:1–12. https://doi.org/10.1038/s41598-019-41918-3
Guha R (2013) On Exploring Structure–Activity Relationships. pp 81–94
Gupta V, Ye G, Olesky M et al (2019) Trends in resistant Enterobacteriaceae and Acinetobacter species in hospitalized patients in the United States: 2013–2017. BMC Infect Dis 19:742. https://doi.org/10.1186/s12879-019-4387-3
Hilliam Y, Kaye S, Winstanley C (2020) Pseudomonas aeruginosa and microbial keratitis. J Med Microbiol 69:3–13. https://doi.org/10.1099/jmm.0.001110
Horna G, Ruiz J (2021) Type 3 secretion system of Pseudomonas aeruginosa. Microbiol Res 246:126719. https://doi.org/10.1016/j.micres.2021.126719
Hosen MI, Tanmoy AM, Mahbuba D, Al et al (2014) Application of a subtractive genomics approach for in silico identification and characterization of novel drug targets in Mycobacterium tuberculosis F11. Interdiscip Sci Comput Life Sci 6:48–56. https://doi.org/10.1007/s12539-014-0188-y
Hossain T, Kamruzzaman M, Choudhury TZ et al (2017) Application of the Subtractive Genomics and Molecular Docking Analysis for the Identification of Novel Putative Drug Targets against Salmonella enterica subsp. enterica serovar Poona. Biomed Res Int 2017:1–9. https://doi.org/10.1155/2017/3783714
Hunt M, Mather AE, Sánchez-Busó L et al (2017) ARIBA: rapid antimicrobial resistance genotyping directly from sequencing reads. Microb Genomics 3. https://doi.org/10.1099/mgen.0.000131
Hwang SM, Cho HW, Kim TY et al (2021) Whole-Genome Sequencing for Investigating a Health Care-Associated Outbreak of Carbapenem-Resistant Acinetobacter baumannii. Diagnostics 11:201. https://doi.org/10.3390/diagnostics11020201
Justino GC, Nascimento CP, Justino MC (2021) Molecular dynamics simulations and analysis for bioinformatics undergraduate students. Biochem Mol Biol Educ 49:570–582. https://doi.org/10.1002/bmb.21512
Karthika A, Ramachandran B, Chitra J et al (2021) Molecular dynamics simulation of Toxin-Antitoxin (TA) system in Acinetobacter baumannii to explore the novel mechanism for inhibition of cell wall biosynthesis: Zeta Toxin as an effective therapeutic target. J Cell Biochem 122:1832–1847. https://doi.org/10.1002/jcb.30137
Kesheri M, Kanchan S, Chowdhury S, Sinha RP (2015) Secondary and Tertiary Structure Prediction of Proteins: A Bioinformatic Approach. pp 541–569
Khaledi A, Schniederjans M, Pohl S et al (2016) Transcriptome Profiling of Antimicrobial Resistance in Pseudomonas aeruginosa. Antimicrob Agents Chemother 60:4722–4733. https://doi.org/10.1128/AAC.00075-16
Khezri A, Avershina E, Ahmad R (2021) Plasmid identification and plasmid-mediated antimicrobial gene detection in norwegian isolates. Microorganisms 9:1–13. https://doi.org/10.3390/microorganisms9010052
Kidd TJ, Mills G, Sá-Pessoa J et al (2017) A Klebsiella pneumoniae antibiotic resistance mechanism that subdues host defences and promotes virulence. EMBO Mol Med 9:430–447. https://doi.org/10.15252/emmm.201607336
Köser CU, Ellington MJ, Peacock SJ (2014) Whole-genome sequencing to control antimicrobial resistance. Trends Genet 30:401–407. https://doi.org/10.1016/j.tig.2014.07.003
Lean S-S, Yeo CC, Suhaili Z, Thong K-L (2015) Whole-genome analysis of an extensively drug-resistant clinical isolate of Acinetobacter baumannii AC12: Insights into the mechanisms of resistance of an ST195 clone from Malaysia. Int J Antimicrob Agents 45:178–182. https://doi.org/10.1016/j.ijantimicag.2014.10.015
Lebreton F, van Schaik W, Sanguinetti M et al (2012) AsrR Is an Oxidative Stress Sensing Regulator Modulating Enterococcus faecium Opportunistic Traits, Antimicrobial Resistance, and Pathogenicity. PLoS Pathog 8:e1002834. https://doi.org/10.1371/journal.ppat.1002834
Lee T, Pang S, Stegger M et al (2020) A three-year whole genome sequencing perspective of Enterococcus faecium sepsis in Australia. PLoS ONE 15:e0228781. https://doi.org/10.1371/journal.pone.0228781
Lokhande KB, Pawar SV, Madkaiker S et al (2022) High throughput virtual screening and molecular dynamics simulation analysis of phytomolecules against BfmR of Acinetobacter baumannii : anti-virulent drug development campaign. J Biomol Struct Dyn 1–15. https://doi.org/10.1080/07391102.2022.2038271
Loomba P, Taneja J, Mishra B (2010) Methicillin and vancomycin resistant S. aureus in hospitalized patients. J Glob Infect Dis 2:275. https://doi.org/10.4103/0974-777X.68535
Low YM, Chong CW, Yap IKS et al (2018) Elucidating the survival and response of carbapenem resistant Klebsiella pneumoniae after exposure to imipenem at sub-lethal concentrations. Pathog Glob Health 112:378–386. https://doi.org/10.1080/20477724.2018.1538281
Luo L, Wu L, Xiao Y et al (2015) Enhancing pili assembly and biofilm formation in Acinetobacter baumannii ATCC19606 using non-native acyl-homoserine lactones. BMC Microbiol 15:62. https://doi.org/10.1186/s12866-015-0397-5
Ma Y, Liu Y, Cheng J (2018) Protein Secondary Structure Prediction Based on Data Partition and Semi-Random Subspace Method. Sci Rep 8:9856. https://doi.org/10.1038/s41598-018-28084-8
Matamoros-Recio A, Franco-Gonzalez JF, Forgione RE et al (2021) Understanding the Antibacterial Resistance: Computational Explorations in Bacterial Membranes. ACS Omega 6:6041–6054. https://doi.org/10.1021/acsomega.0c05590
McGuinness WA, Malachowa N, DeLeo FR (2017) Vancomycin Resistance in Staphylococcus aureus. Yale J Biol Med 90:269–281
Messaoudi A, Belguith H, Ben Hamida J (2013) Homology modeling and virtual screening approaches to identify potent inhibitors of VEB-1 β-lactamase. Theor Biol Med Model 10:22. https://doi.org/10.1186/1742-4682-10-22
Miryala SK, Anbarasu A, Ramaiah S (2018) Discerning molecular interactions: A comprehensive review on biomolecular interaction databases and network analysis tools. Gene 642:84–94. https://doi.org/10.1016/j.gene.2017.11.028
Miryala SK, Anbarasu A, Ramaiah S (2020) Role of SHV-11, a Class A β-Lactamase, Gene in Multidrug Resistance among Klebsiella pneumoniae Strains and Understanding Its Mechanism by Gene Network Analysis. Microb Drug Resist 26:900–908. https://doi.org/10.1089/mdr.2019.0430
Miryala SK, Anbarasu A, Ramaiah S (2019) Systems biology studies in Pseudomonas aeruginosa PA01 to understand their role in biofilm formation and multidrug efflux pumps. Microb Pathog 136:103668. https://doi.org/10.1016/j.micpath.2019.103668
Miryala SK, Basu S, Naha A et al (2022) Datasets comprising the quality validations of simulated protein-ligand complexes and SYBYL docking scores of bioactive natural compounds as inhibitors of Mycobacterium tuberculosis protein-targets. Data Br 42:108146. https://doi.org/10.1016/j.dib.2022.108146
Miryala SK, Basu S, Naha A et al (2021) Identification of bioactive natural compounds as efficient inhibitors against Mycobacterium tuberculosis protein-targets: A molecular docking and molecular dynamics simulation study. J Mol Liq 341:117340. https://doi.org/10.1016/j.molliq.2021.117340
Miryala SK, Ramaiah S (2019) Exploring the multi-drug resistance in Escherichia coli O157:H7 by gene interaction network: A systems biology approach. Genomics 111:958–965. https://doi.org/10.1016/j.ygeno.2018.06.002
Miryala SK, Ramaiah S (2022) Cellular and molecular level host-pathogen interactions in Francisella tularensis: A microbial gene network study. Comput Biol Chem 96:107601. https://doi.org/10.1016/j.compbiolchem.2021.107601
Mwangi J, Hao X, Lai R, Zhang Z-Y (2019) Antimicrobial peptides: new hope in the war against multidrug resistance. Zool Res 40:488–505. https://doi.org/10.24272/j.issn.2095-8137.2019.062
Naaz F, Khan A, Kumari A et al (2021) 1,3,4-oxadiazole conjugates of capsaicin as potent NorA efflux pump inhibitors of Staphylococcus aureus. Bioorg Chem 113:105031. https://doi.org/10.1016/j.bioorg.2021.105031
Naha A, Kumar Miryala S, Debroy R et al (2020) Elucidating the multi-drug resistance mechanism of Enterococcus faecalis V583: A gene interaction network analysis. Gene 748:144704. https://doi.org/10.1016/j.gene.2020.144704
Naha A, Vijayakumar S, Lal B et al (2021) Genome sequencing and molecular characterisation of XDR Acinetobacter baumannii reveal complexities in resistance: Novel combination of sulbactam–durlobactam holds promise for therapeutic intervention. J Cell Biochem 122:1946–1957. https://doi.org/10.1002/jcb.30156
Navon-Venezia S, Kondratyeva K, Carattoli A (2017) Klebsiella pneumoniae: A major worldwide source and shuttle for antibiotic resistance. FEMS Microbiol Rev 41:252–275. https://doi.org/10.1093/femsre/fux013
Ndagi U, Falaki AA, Abdullahi M et al (2020) Antibiotic resistance: bioinformatics-based understanding as a functional strategy for drug design. RSC Adv 10:18451–18468. https://doi.org/10.1039/D0RA01484B
Nourani E, Khunjush F, DurmuÅŸ S (2015) Computational approaches for prediction of pathogen-host protein-protein interactions. Front Microbiol 6. https://doi.org/10.3389/fmicb.2015.00094
Padilla E, Llobet E, Doménech-Sánchez A et al (2010) Klebsiella pneumoniae AcrAB Efflux Pump Contributes to Antimicrobial Resistance and Virulence. Antimicrob Agents Chemother 54:177–183. https://doi.org/10.1128/AAC.00715-09
Pandurangan AP, Ascher DB, Thomas SE, Blundell TL (2017) Genomes, structural biology and drug discovery: combating the impacts of mutations in genetic disease and antibiotic resistance. Biochem Soc Trans 45:303–311. https://doi.org/10.1042/BST20160422
Pfaller MA, Cormican M, Flamm RK et al (2019) Temporal and geographic variation in antimicrobial susceptibility and resistance patterns of Enterococci: Results from the SENTRY Antimicrobial Surveillance Program, 1997–2016. Open Forum Infect Dis 6:S54–S62. https://doi.org/10.1093/ofid/ofy344
Quainoo S, Coolen JPM, van Hijum SAFT et al (2017) Whole-Genome Sequencing of Bacterial Pathogens: the Future of Nosocomial Outbreak Analysis. Clin Microbiol Rev 30:1015–1063. https://doi.org/10.1128/CMR.00016-17
Ragupathi ND, Bakthavatchalam Y, Mathur P et al (2019) Plasmid profiles among some ESKAPE pathogens in a tertiary care centre in south India. Indian J Med Res 149:222. https://doi.org/10.4103/ijmr.IJMR_2098_17
Rao M, Rashid FA, Shukor S et al (2020) Detection of Antimicrobial Resistance Genes Associated with Carbapenem Resistance from the Whole-Genome Sequence of Acinetobacter baumannii Isolates from Malaysia. Can J Infect Dis Med Microbiol 2020:1–9. https://doi.org/10.1155/2020/5021064
Rasheed MA, Iqbal MN, Saddick S et al (2021) Identification of Lead Compounds against Scm (fms10) in Enterococcus faecium Using Computer Aided Drug Designing. Life 11:77. https://doi.org/10.3390/life11020077
Sabatini S, Gosetto F, Iraci N et al (2013) Re-evolution of the 2-Phenylquinolines: Ligand-Based Design, Synthesis, and Biological Evaluation of a Potent New Class of Staphylococcus aureus NorA Efflux Pump Inhibitors to Combat Antimicrobial Resistance. J Med Chem 56:4975–4989. https://doi.org/10.1021/jm400262a
Saha M, Sarkar A (2021) Review on Multiple Facets of Drug Resistance: A Rising Challenge in the 21st Century. J Xenobiotics 11:197–214. https://doi.org/10.3390/jox11040013
Santajit S, Indrawattana N (2016) Mechanisms of Antimicrobial Resistance in ESKAPE Pathogens. Biomed Res Int 2016:. https://doi.org/10.1155/2016/2475067
Schloss PD, Handelsman J (2003) Biotechnological prospects from metagenomics. Curr Opin Biotechnol 14:303–310. https://doi.org/10.1016/S0958-1669(03)00067-3
Schniederjans M, Koska M, Häussler S (2017) Transcriptional and Mutational Profiling of an Aminoglycoside-Resistant Pseudomonas aeruginosa Small-Colony Variant. Antimicrob Agents Chemother 61. https://doi.org/10.1128/AAC.01178-17
Schürch AC, van Schaik W (2017) Challenges and opportunities for whole-genome sequencing-based surveillance of antibiotic resistance. Ann N Y Acad Sci 1388:108–120. https://doi.org/10.1111/nyas.13310
Shafiee F, Naji Esfahani SS, Hakamifard A, Soltani R (2021) In vitro synergistic effect of colistin and ampicillin/sulbactam with several antibiotics against clinical strains of multi-drug resistant Acinetobacter baumannii. Indian J Med Microbiol 39:358–362. https://doi.org/10.1016/j.ijmmb.2021.04.006
Shankar C, Basu S, Lal B et al (2021) Aerobactin Seems To Be a Promising Marker Compared With Unstable RmpA2 for the Identification of Hypervirulent Carbapenem-Resistant Klebsiella pneumoniae: In Silico and In Vitro Evidence. Front Cell Infect Microbiol 11. https://doi.org/10.3389/fcimb.2021.709681
Shi J, Yan Y, Links MG et al (2019) Antimicrobial resistance genetic factor identification from whole-genome sequence data using deep feature selection. BMC Bioinformatics 20:535. https://doi.org/10.1186/s12859-019-3054-4
Sigurdsson G, Fleming RMT, Heinken A, Thiele I (2012) A Systems Biology Approach to Drug Targets in Pseudomonas aeruginosa Biofilm. PLoS ONE 7:e34337. https://doi.org/10.1371/journal.pone.0034337
Singh P, Jamal S, Ahmed F et al (2021) Computational modeling and bioinformatic analyses of functional mutations in drug target genes in Mycobacterium tuberculosis. Comput Struct Biotechnol J 19:2423–2446. https://doi.org/10.1016/j.csbj.2021.04.034
Sobolev OV, Afonine PV, Moriarty NW et al (2020) A Global Ramachandran Score Identifies Protein Structures with Unlikely Stereochemistry. Structure 28:1249–1258e2. https://doi.org/10.1016/j.str.2020.08.005
Stoesser N, Batty EM, Eyre DW et al (2013) Predicting antimicrobial susceptibilities for Escherichia coli and Klebsiella pneumoniae isolates using whole genomic sequence data. J Antimicrob Chemother 68:2234–2244. https://doi.org/10.1093/jac/dkt180
Strateva T, Mitov I (2011) Contribution of an arsenal of virulence factors to pathogenesis of Pseudomonas aeruginosa infections. Ann Microbiol 61:717–732. https://doi.org/10.1007/s13213-011-0273-y
Su M, Satola SW, Read TD (2019) Genome-Based Prediction of Bacterial Antibiotic Resistance. J Clin Microbiol 57. https://doi.org/10.1128/JCM.01405-18
Sugimoto S, Iwamoto T, Takada K et al (2013) Staphylococcus epidermidis Esp Degrades Specific Proteins Associated with Staphylococcus aureus Biofilm Formation and Host-Pathogen Interaction. J Bacteriol 195:1645–1655. https://doi.org/10.1128/JB.01672-12
Sun D, Jeannot K, Xiao Y, Knapp CW (2019) Editorial: Horizontal Gene Transfer Mediated Bacterial Antibiotic Resistance. Front Microbiol 10. https://doi.org/10.3389/fmicb.2019.01933
Takaya D, Takeda-Shitaka M, Terashi G et al (2008) Bioinformatics Based Ligand-Docking and in-Silico Screening. Chem Pharm Bull 56:742–744. https://doi.org/10.1248/cpb.56.742
Tiwari V (2019) Post-translational modification of ESKAPE pathogens as a potential target in drug discovery. Drug Discov Today 24:814–822. https://doi.org/10.1016/j.drudis.2018.12.005
Tolios A, De Las Rivas J, Hovig E et al (2020) Computational approaches in cancer multidrug resistance research: Identification of potential biomarkers, drug targets and drug-target interactions. Drug Resist Updat 48:100662. https://doi.org/10.1016/j.drup.2019.100662
Vaara M (2019) Polymyxins and Their Potential Next Generation as Therapeutic Antibiotics. Front Microbiol 10. https://doi.org/10.3389/fmicb.2019.01689
Vakser IA, Deeds EJ (2019) Computational approaches to macromolecular interactions in the cell. Curr Opin Struct Biol 55:59–65. https://doi.org/10.1016/j.sbi.2019.03.012
Varghese R, Basu S, Neeravi A et al (2022) Emergence of Meropenem Resistance Among Cefotaxime Non-susceptible Streptococcus pneumoniae: Evidence and Challenges. Front Microbiol 12. https://doi.org/10.3389/fmicb.2021.810414
Waddington C, Carey ME, Boinett CJ et al (2022) Exploiting genomics to mitigate the public health impact of antimicrobial resistance. Genome Med 14:15. https://doi.org/10.1186/s13073-022-01020-2
Wagner T, Joshi B, Janice J et al (2018) Enterococcus faecium produces membrane vesicles containing virulence factors and antimicrobial resistance related proteins. J Proteom 187:28–38. https://doi.org/10.1016/j.jprot.2018.05.017
Wu F, Zhou Y, Li L et al (2020) Computational Approaches in Preclinical Studies on Drug Discovery and Development. Front Chem 8. https://doi.org/10.3389/fchem.2020.00726
Wyres KL, Holt KE (2018) Klebsiella pneumoniae as a key trafficker of drug resistance genes from environmental to clinically important bacteria. Curr Opin Microbiol 45:131–139. https://doi.org/10.1016/j.mib.2018.04.004
Zárate S, Morales P, Świderek K et al (2019) A Molecular Modeling Approach to Identify Novel Inhibitors of the Major Facilitator Superfamily of Efflux Pump Transporters. Antibiotics 8:25. https://doi.org/10.3390/antibiotics8010025
Zhang L, Fritsch M, Hammond L et al (2013) Identification of Genes Involved in Pseudomonas aeruginosa Biofilm-Specific Resistance to Antibiotics. PLoS ONE 8:e61625. https://doi.org/10.1371/journal.pone.0061625
Acknowledgements
The authors gratefully acknowledge the Indian Council of Medical Research (ICMR), the Government of India agency, for the research grant (IRIS ID: 2020 − 0690). The authors would like to thank the Management of VIT, Vellore for the technical support. The authors would like to acknowledge Mr. Miryala SK (Post Doctoral Fellow, IIT Bombay, Mumbai), Mr. Soumya Basu, Mr. Aniket Naha and Mr. Hithesh Kumar C.K. (ICMR-Research Assistant) for genomics and structural biology inputs; and Ms. Gayathri Ashok (Teaching cum Research Assistant, VIT, Vellore) for the systems biology inputs. The authors are grateful to english language expert Ms. Lydia Ann Vinod (VIT, Vellore) for helping us in the language check. Reetika Debroy would like to thank ICMR, New Delhi for her Senior Research Fellowship (ID: 2021-10632).
Funding
The authors gratefully acknowledge the Indian Council of Medical Research (ICMR), the Government of India agency, for the research grant (IRIS ID: 2020 − 0690).
Author information
Authors and Affiliations
Contributions
Study conceptualization and funding requisition were performed by Sudha Ramaiah. Anand Anbarasu critically analysed and designed the study. Priyamvada and Reetika Debroy performed the literature search and drafted the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Priyamvada, P., Debroy, R., Anbarasu, A. et al. A comprehensive review on genomics, systems biology and structural biology approaches for combating antimicrobial resistance in ESKAPE pathogens: computational tools and recent advancements. World J Microbiol Biotechnol 38, 153 (2022). https://doi.org/10.1007/s11274-022-03343-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-022-03343-z