Abstract
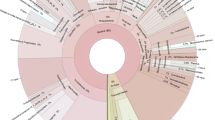
By combining low nutrient enrichments and molecular methods, a high diversity of new amylase genes was detected in a neutral sulphide-rich hot spring in Iceland. Enrichments based on hot spring water and low concentrations of starch were used to select slow-growing, starch-degrading microorganisms. Six enrichments had in total 17 bacterial types detected by 16S rRNA analysis, mostly related to the Thermus-Deinococcus group, green non-sulphur bacteria, gram positives, and uncultivated new candidate divisions. No Archaea were found. The apparent 16S rRNA species composition of the enrichments was very different from that of the microbial mat in the same hot spring. DNA samples obtained from 4 enrichments and from hot spring biomass were screened by PCR for amylase genes in glycoside-hydrolase family 13. Degenerate primers, based on conserved amino acid sequences from multiple␣alignments of family 13, enabled the detection of 18 amylase sequence types in the enrichments, including α-amylases, α-glucosidases, 1,4-α-glucan branching enzymes, cyclomaltodextrin hydrolases, maltogenic amylases and neopullulanases, and unspecified family 13 glycoside-hydrolases. Only one unique neopullulanase sequence, also found in most of the enrichments, was detected in the hot spring biomass DNA. The results suggest that the enrichment method combined with sequence-based screening is an efficient way to access the silent, i.e. not detectable, gene diversity in natural environments.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Altschul, S.F., Madden, T.L., Schaffer, A.A., Zhang, J.H., Zhang, Z., Miller, W. & Lipman, D.J. 1997 Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Research 25, 3389–3402.
Amann, R.I., Ludwig, W. & Schleifer, K.-H. 1995 Phylogenetic identification and in situ detection of individual microbial cells without cultivation. Microbiological Reviews 59, 143–169.
Chrisostomos, S., Patel, B.K.C., Dwivedi, P.P. & Denman, S.E. 1996 Caloramator indicus gen. nov., sp. nov., a new thermophilic anaerobic bacterium isolated from a non-volcanically heated waters of an Indian artesian aquifer. International Journal of Systematic Bacteriology 46, 497–501.
Cole, J.R., Chai, B., Marsh, T.L., Farris, R.J., Wang, Q., Kulam, S.A., Chandra, S., McGarrell, D.M., Schmidt, T.M., Garrity, G.M. & Tiedje, J.M. 2003 The Ribosomal Database Project (RDP-II): previewing a new autoaligner that allows regular updates and the new prokaryotic taxonomy. Nucleic Acids Research 31, 442–443.
Collins, M.D., Lawson, P.A., Willems, A., Cordoba, J.J., Fernandez-Garayzabal, J., Garcia, P., Cai, J., Hippe, H. & Farrow, J.E.A. 1994 The phylogeny of the genus Clostridium: Proposal of five new genera and eleven species combinations. International Journal of Systematic Bacteriology 44, 812–826.
Cote, R.J. & Gherna, R.L. 1994 Nutrition and Media. In Methods for General and Molecular Bacteriology, eds. Gerhardt, P., Murray, R.G.E., Wood, W.A. & Krieg, N.R. pp. 155–178. Washington DC, USA: American Society of Microbiology. ISBN: 1-55581-048-9.
Coutinho, P.M. & Henrissat, B. 1999a Carbohydrate-active enzymes: an integrated database approach. In Recent Advances in Carbohydrate Bioengineering, eds. Gilbert, H.J., Davies, G., Henrissat, B. & Svensson, B. pp. 3–12. Cambridge, UK: The Royal Society of Chemistry. ISBN 0-85404774-3.
Coutinho, P.M. & Henrissat, B. 1999b The modular structure of cellulases and other carbohydrate-active enzymes: an integrated database approach. In Genetics;Biochemistry and Ecology of Cellulose Degradation, eds. Ohmiya, K., Hayashi, K., Sakka, K., Kobayashi, Y., Karita, S. & Kimura, T. pp. 15–23. Tokyo, Japan: Uni Publishers Co. ISBN 0-85404774-3.
Dahllof, I. 2002 Molecular community analysis of microbial diversity. Current Opinion in Biotechnology 13, 213–217.
Hall, T.A. 1999 BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symposium Series 41, 95–98.
Henikoff, S., Henikoff, J.G., Alford, W.J. & Pietrokovski, S. 1995 Automated construction and graphical presentation of protein blocks from unaligned sequences. Gene 163, GC17–GC26.
Henrissat, B. & Davies, G. 1997 Structural and sequence-based classification of glycoside-hydrolases. Current Opinion in Structural Biology 7, 637–644.
Hjorleifsdottir, S., Skirnisdottir, S., Hreggvidsson, G.O., Holst, O. & Kristjansson, J.K. 2001 Species composition of cultivated and noncultivated bacteria from short filaments in an Icelandic hot spring at 88_C. Microbial Ecology 42, 117–125.
Horvathova, V., Janecek, S. & Sturdik, E. 2001 Amylolytic enzymes: molecular aspects of their properties. General Physiology and Biophysics 20, 7–32.
Hugenholtz, P., Pitulle, C., Hershberger, K.L. & Pace, N.R. 1998 Novel division level bacterial diversity in a Yellowstone hot spring. Journal of Bacteriology 180, 366–376.
Jackson, C.R., Roden, E.E. & Churchill, P.F. 1998 Changes in bacterial species composition in enrichment cultures with various dilutions of inoculum as monitored by denaturing gradient gel electrophoresis. Applied and Environmental Microbiology 64, 5046–5048.
Kristjansson, J.K., Hjorleifsdottir, S., Marteinsson, V.T. & Alfredsson, G.A. 1994 Thermus scotoductus, sp. nov., a pigment-producing thermophilic bacterium from hot tap water in Iceland and including Thermus sp. X-1. Systematic and Applied Microbiology 17, 44–50.
Loginova, L.G., Egorova, L.A., Golovacheva, R.S. & Seregina, L.M. 1984 Thermus ruber sp. nov. nom rev. International Journal of Systematic Bacteriology 34, 498–499.
Marteinsson, V.T., Kristjansson, J.K., Kristmannsdottir, H., Dahlkvist, M., Saemundsson, K., Hannington, M., Petursdottir, S.K., Geptner, A. & Stoffers P. 2001 Discovery and description of giant submarine smectite cones on the seafloor in Eyjafjordur, northern Iceland, and a novel thermal microbial habitat. Applied and Environmental Microbiology 67, 827–833.
Niehaus, F., Bertoldo, C., Kahler, M. & Antranikian, G. 1999 Extremophiles as a source of novel enzymes for industrial applications. Applied Microbiology and Biotechnology 51, 711–729.
Pikuta, E., Lysenko, A., Chuvilskaya, N., Mendrock, U., Hippe, H., Suzina, N., Nikitin, D., Osipov, G. & Laurinavichius, K. 2000 Anoxybacillus pushchinensis gen. nov., sp. nov., a novel anaerobic, alkaliphilic, moderately thermophilic bacterium from manure, and description of Anoxybacillus flavithermus comb. nov. International Journal of Systematic and Evolutionary Microbiology 50, 2109–2117.
Rainey, F.A., Fritze, D. & Stackebrandt, E. 1994 The phylogenetic diversity of thermophilic members of the genus Bacillus as revealed by 16S rDNA analysis. FEMS Microbiology Letters 115, 205–211.
Rose, T.M., Schultz, E.R., Henikoff, J.G., Pietrokovski, S., McCallum, C.M. & Henikoff, S. 1998 Consensus-degenerate hybrid oligonucleotide primers for amplification of distantly-related sequences. Nucleic Acids Research 26, 1628–1635.
Santegoeds, C.M., Nold, S.C. & Ward, D.M. 1996 Denaturing gradient gel electrophoresis used to monitor the enrichment culture of aerobic chemoorganotrophic bacteria from a hot spring cyanobacterial mat. Applied and Environmental Microbiology 62, 3922–3928.
Skirnisdottir, S., Hreggvidsson, G.O., Hjorleifsdottir, S., Marteinsson, V.T., Petursdottir, S.K., Holst, O. & Kristjansson, J.K. 2000 Influence of sulfide and temperature on species composition and community structure of hot spring microbial mats. Applied and Environmental Microbiology 66, 2835–2841.
Skirnisdottir, S., Hreggvidsson, G.O., Holst, O. & Kristjansson, J.K. 2001 Isolation and characterization of a mixotrophic sulphuroxidizing Thermus scotoductus. Extremophiles 5, 45–51.
Slobodkin, A., Reysenbach, A.-L., Mayer, F. & Wiegel, J. 1997a Isolation and characterization of the homoacetogenic thermophilic bacterium Moorella glycerini sp. nov. International Journal of Systematic Bacteriology 47, 969–974.
Slobodkin, A., Reysenbach, A.-L., Strutz, N., Dreier, M. & Wiegel, J. 1997b Thermoterrabacterium ferrireducens gen. nov., sp. nov., a thermophilic anaerobic dissimilatory Fe(III)-reducing bacterium from a continental hot spring. International Journal of Systematic Bacteriology 47, 541–547.
Sonnhammer, E.L.L., Eddy, S.R. & Durbin, R. 1997 Pfam: a comprehensive database of protein domain families based on seed alignments. Proteins 28, 405–420.
Stackebrandt, E. & Goebel, B.M. 1994 Taxonomic note; A place for DNA-DNA reassociation and 16S rRNA sequence analysis in the present species definition in bacteriology. International Journal of Systematic Bacteriology 44, 846–849.
Suzuki, M.T. & Giovannoni, S.J. 1996 Bias caused by template annealing in the amplification of mixtures of 16S rRNA genes by PCR. Applied and Environmental Microbiology 62, 625–630.
Thompson, J.D., Gibson, T.G., Plewniak, F., Jeanmougin, F. & Higgins, D.G. 1997 The ClustalX windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Research 24, 4876–4882.
Von Wintzingerode, F., Gobel, U.B. & Stackebrandt, E. 1997 Determination of microbial diversity in environmental samples: pitfalls of PCR-based rRNA analysis. FEMS Microbiology Reviews 21, 213–229.
Ward, D.M., Ferris, M.J., Nold, S.C. & Bateson, M.M. 1998 A natural view of microbial biodiversity within hot spring cyanobacterial mat communities. Microbiology and Molecular Biology Reviews 62, 1353–1370.
Williams, R.A.D., Smith, K.E., Welch, S.G. & Micallef, J. 1996 Thermus oshimai sp. nov., isolated from hot springs in Portugal, Iceland, and the Azores, and comment on the concept of a limited geographical distribution of Thermus species. International Journal of Systematic Bacteriology 46, 403–408.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Hobel, C.F., Marteinsson, V.T., Hauksdóttir, S. et al. Use of low nutrient enrichments to access novel amylase genes in silent diversity of thermophiles. World Journal of Microbiology and Biotechnology 20, 801–809 (2004). https://doi.org/10.1007/s11274-004-2623-4
Issue Date:
DOI: https://doi.org/10.1007/s11274-004-2623-4