Abstract
Diversity of the microbial community in the Zharkent geothermal hot spring, located in the southeastern region of Kazakhstan, was assessed using both culture-dependent and -independent approaches. Shotgun metagenomic sequencing of DNA extracted from the spring water yielded 11,061,725 high-quality sequence reads, totaling >1,67 Gb of nucleotide sequences. Furthermore, water samples were enriched in nutrient broth at varying high temperatures, and colonies isolated by being streaked onto nutrient agar. Finally, DNA extraction and amplification, as well as sequencing and phylogenetic analysis, were conducted. Bacteria constituted more than 99.97% of the total prokaryotic abundance, with Archaea contributing only an extremely small component; Firmicutes, Proteobacteria, and Actinobacteria dominated the community. At genus level, Firmicutes reads affiliated with Desmospora, Parageobacillus, Paenibacillus, and Brevibacillus, accounting for more than 60% of total prokaryotic abundance. Eight morphologically distinct, aerobic, endospore-forming thermophilic bacteria were recovered; isolates differed significantly in substrate utilization patterns, as well as their production of thermophilic, extracellular, hydrolytic enzymes for degradation of starch, lipids, cellulose, and protein. Five strains could degrade all four macromolecular types at temperatures ranging from 55 to 75 °C. Phylogenetic analyses based on 16S rRNA gene sequences placed all isolates into the genus Geobacillus with some of them possibly representing novel species. The results indicate that this hot spring represents a rich source of novel thermophilic bacteria and potentially useful thermostable enzymes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Geothermal environments, such as hot springs, solfatara fields, hydrothermal vents, and deep subsurface habitats, host a rich diversity of thermophiles that synthesize unique thermophilic and thermostable enzymes (extremozymes) with the potential for novel application in high-temperature biocatalytic processes [1,2,3]. Thermophilic enzymes are adapted to function at the growth temperature of their host. Therefore, the diapason of extremes at which life is detected determines the range of conditions under which enzyme activity might occur [4, 5]. Isolation and characterization of thermophiles has been performed for many geothermal places in various parts of the world, including the USA [6,7,8,9,10], Turkey [11], Italy [12], Bulgaria [13], Greece [14], India [15], and New Zealand [16]. Kazakhstan is home to many geothermal sites of which the microbial communities remain unexplored, likely representing potential sources of novel extremophilic microbes with biotechnological application potential.
Metagenomics analysis permits identification of microbes and potentially useful novel enzymes in thermal springs, using culture-independent methods [17]. At present, high-throughput metagenomic sequence analysis supplements traditional culture-based methods, allowing for comprehensive understanding of complex microbial communities [18, 19]. Nevertheless, to understand the physiology and potential applications of microorganisms in a specific ecological niche, it remains important to isolate pure cultures. Using a combination of culture-dependent and culture-independent molecular methods has largely increased our comprehension of the functional and structural diversity of microbial communities [20]. Here, we present results of both culture-dependent and -independent analyses, in the first study of the microbial diversity of a hot spring in Kazakhstan.
Materials and Methods
Study Site and Collection of Samples
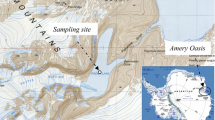
The Zharkent geothermal hot spring is located at 43°97′14.93″ N, 79°66′12.09″ E. Due to its remote location, this hot spring is less influenced by human interference and believed to possess rich microbial wealth. The temperature of the hot spring during the sampling period was 90 °C. The pH level was approximately 7.5, indicating a slightly alkaline environment. Sampling was performed on January 19, 2019. Sample was collected in 50-ml sterile tubes and maintained at 4 °C during transportation to Almaty. Temperature and pH were determined in situ using a portable combination pH/EC/TDS meter (BEM802, Milwaukee, Wisconsin, US).
Water sample was filtered through 0.2-µm filter and stored at 7 °C in plastic tube, until chemical analyses could be performed at the ICP Laboratory of the University of Bergen (https://www.uib.no/en/geo/111639/icp-laboratory). Major and minor elements—Al, As, Ba, B, Ca, Co, Cr, Cu, Eu, Fe, Pb, La, Li, Mg, Mn, Ni, Na, Sr, S, Si, Ti, K, V, Y, and Zn—were assessed using an iCAP™ 7600 ICP-OES Analyzer. Anions—Cl−, SO42−, NO3−, and Br−—were determined by ion chromatography (IC, Metrohm). The major cations to be detected (>1200 ppm) were sodium (227.4 ppm), calcium (4.3 ppm), and potassium (3.2 ppm) (Supplementary Table S1). Major anions to be detected were chloride (138.7 ppm) and sulfate (89.4 ppm). Nitrate, iron, cobalt, chromium, copper, europium, lanthanum, titanium, yttrium, and zinc could not be detected.
Isolation and Morphological Characterization
Water samples were enriched in nutrient broth (NB) containing 2.0% bacteriological peptone, 2.0% D-glucose, 1.0% yeast extract, and 0.6% NaCl (w/v) at 65, 70, and 75 °C, for three days. The enriched cultures were streaked onto nutrient agar to obtain separate colonies. Selected colonies were purified by streaking onto the same medium at least four times, using the quadrant streak technique. The isolates were considered pure upon microscopic observation of a single morphological cell type per culture. Plastic bags were used to prevent evaporation during the incubation. Observations were recorded for color, shape, margin, elevation, light transmission, and pigmentation after 12, 24, and 48 h of incubation (Supplementary Fig. S1). Cell morphology of the isolates was determined by phase-contrast microscopy (Nikon Eclipse E400 biological microscope (1,001,360, Nikon, Minato City, Japan)) using freshly prepared wet mounts. Scanning electron microscopy (SEM) was performed using a Jeol JSM-7400F scanning electron microscope at the Molecular Imaging Center of the University of Bergen (https://www.uib.no/en/rg/mic). For SEM, 1 ml of dense culture was fixed with 80 µl of 25% glutaraldehyde (G6257, Sigma-Aldrich) for 1 h at 25 °C. After 3 washes with 0.1 M phosphate buffer (pH 7.2) (P5244, Sigma-Aldrich), post-fixation was performed using 1% osmium tetroxide (201,030, Sigma-Aldrich) for 1 h. Dehydration was performed successively in 30%, 50%, 90%, and twice in 100% ethanol, followed by filtration of the cells onto a 0.2-µm black polycarbonate filter (WHA70632502, Cyclopore). Pieces of the filter were critical point dried for at least 1 h, mounted onto an SEM specimen stub, and sputter coated with 10 nm gold/palladium particles.
Metabolic and Biochemical Characterization
Most biochemical characteristics were tested after 1–2 days of incubation at 65–75 °C. Catalase activity was determined based on the formation of bubbles after the addition of a few drops of 3% H2O2 (v/v) to the prepared samples [21]. The activity of oxidase was detected based on the oxidation of tetramethyl-p-phenylenediamine (TMPD) [22]. Isolates were examined for the assimilation and fermentation of 49 carbohydrates using API® 50 CH strips (SKU:50,300, bioMérieux, Inc.) as recommended by the manufacturer. Following transfer of 100 µl aliquots of suspended bacteria to the strips, they were incubated at 50–55 °C for 48 h and then visually evaluated according to the manufacturer’s protocols. To determine the impact of temperature on growth, the isolates were grown at temperatures in the range of 40–95 °C at regular 5 °C increments, for 24 h. In addition, the effect of pH on the growth of the isolated microorganisms was studied by growing the organisms for 24 h at 65–75 °C in NB adjusted for pH values ranging from 4–10 [5]. Growth was measured by determining the optical density at 620 nm using a PD-303 spectrophotometer (Apel, Kawaguchi City, Saitama Prefecture, Japan).
Degradation of starch, carboxymethyl cellulose (CMC), polysorbate (Tween 60, Tween 80), and casein was determined qualitatively using agar plates with different basal media. The degradation of casein was examined on plates containing (w/v) skimmed milk powder (20.00%), glucose (0.50%), and agarose (1.5%). Positive results were indicated by clear halo zones developing around the colonies, as described [23]. CMC degradation was investigated on plates containing (w/v) CMC (1.00%), NaCl (0.01%), NaNO3 (0.02%), K2HPO4 (0.01%), MgSO4 (0.003%), KCl (0.003%), peptone (0.01%), and agar (1.50%), as described [24]. Degradation of starch was tested on plates containing (w/v) soluble starch (2.00%), tryptone (1.00%), yeast extract (0.10%), and agarose (1.50%). The presence of cellulase and amylase activity was confirmed by observing a clear zone around the colony, formed after staining with Lugol solution (1.00% KI and 0.50% I2 (w/v) in Milli-Q water) (L6146, Sigma-Aldrich), as described [21, 25]. Lipolytic enzyme production was tested by streaking strains onto Spirit blue agar [23] medium (S4306, Sigma-Aldrich) supplemented with 1% (w/v) Tween 60 or Tween 80 (P1754, Sigma-Aldrich). Formation of visible precipitates of fatty acid calcium salts, indicated the presence of lipase activity, as described [23, 26]. Additional enzymatic activities were assessed using the API® ZYM system (SKU:25,200, bioMérieux, Inc.) according to the manufacturer’s instructions.
Extraction of DNA and PCR Amplification
Cultures grown in NB were centrifuged at 4 °C for 15 min at 7500 rpm, and DNA was isolated from the pelleted cells using a GenElute™ Bacterial Genomic DNA Extraction Kit (NA2100, Sigma-Aldrich, Darmstadt, Hesse, Germany), as per the manufacturer’s protocol. DNA quality and quantity were determined using a NanoDrop™ One/OneC Microvolume UV–Vis spectrophotometer (ND-ONE/ONEC-W4, Thermo Fisher Scientific, Waltham, MA, USA). Thereafter, samples were electrophoresed on 0.8% agarose gel to assess DNA quality, and bands were compared to the GeneRuler DNA Ladder Mix (SM0333, Thermo Fischer Scientific, Waltham, MA, USA). Universal primer pairs 27f (5′-GAG TTT GAT CCT GGC TCA -3′) and 1525r (5′-GAA AGG AGG AGA TCC AGC C-3′) [27, 28] were used to amplify the 16S rRNA genes. Composition of the polymerase chain reaction (PCR) mixtures was as follows: 10 ng template DNA, 5 µl 10 × PCR buffer for Taq DNA polymerase (M0480S, New England BioLabs Inc., Ipswich, MA, USA), 5 µl of a stock solution with 10 mM of each deoxynucleoside triphosphate (dNTP), 0.5 µl of each primer (100 mM stock solution), 0.125 µl Taq DNA polymerase (0.625 U) (M0273S, New England BioLabs Inc., Ipswich, MA, USA), and sterile water to a final volume of 25 µl. PCR amplification was conducted per the following process: initial denaturation of templates for 3 min at 96 °C, in 30 cycles consisting of the following steps: denaturation for 30 s at 96 °C, annealing for 30 s at 55 °C, and extension for 2.5 min at 72 °C, followed by a final extension for 10 min at 72 °C. GenElute™ PCR Cleanup Kit (NA1020, Sigma-Aldrich, Darmstadt, Hesse, Germany) was used to purify PCR products, prior to Sanger sequencing.
For extraction of environmental DNA, the sample was filtered through a 1.2-µm pore size membrane (RAWP04700, Whatman) to eliminate any traces of sand and large particles. The filtrate was then passed through a 0.2-µm pore size membrane (WHA7402004, Whatman), which was cut into small pieces and used for DNA isolation using a GenElute™ Bacterial Genomic DNA Extraction Kit, as per the manufacturer’s instructions. DNA quality and quantity were determined using the NanoDrop™ spectrophotometer as above, along with a Life Technologies Qubit 2.0 fluorometer (Q33238, Thermo Fischer Scientific, Waltham, MA, USA) and a Qubit dsDNA HS Assay Kit (Q32851, Thermo Fischer Scientific, Waltham, MA, USA) for quantification.
Sequence analysis of the metagenomic DNA was performed by Eurofins Gen omics (Constance, Germany) using Illumina HiSeq paired-end technology, in NovaSeq 6000 S2 PE150 XP sequencing mode (Illumina, San Diego, CA, USA). The sequence data were trimmed with the Microbial Metagenomics Module implemented in the CLC Genomic Workbench 20.0.01 software (QIAGEN Bioinformatics, Aarhus, Denmark), using the “Trim Reads” tool. Taxonomic profiling to calculate the relative abundance of taxa, was performed using the “Taxonomic Profiling” tool.
Accession Numbers
The 16S rRNA gene sequences reported in this study have been deposited in GenBank under accession numbers MW485510-MW485517. The metagenomic raw sequence data are deposited in GenBank under accession number SPP14001913.
Results
Metagenomic Analysis
Illumina shotgun sequencing yielded a total of 11,061,725 reads after trimming, totaling 1,670,320,475 bp. Taxonomic profiling revealed that 99.97% of the prokaryotes belonged to the domain Bacteria, with only an extremely small fraction from the domain Archaea. The phylum Firmicutes was a highly dominant group, with a relative abundance of 89.00%. The next most abundant phylum was Proteobacteria representing 5.30% of total sequences obtained, and Actinobacteria constituting 3.90%. Minor components—each representing ≤1.00% of total sequences obtained—were also detected; these included Chloroflexi, Cyanobacteria, Bacteroidetes, Deinococcus–Thermus, and Aquificae. Among the Firmicutes, the Bacilli class constituted the vast taxonomic majority, making up 84.00% of the total class level distinction, of which Thermoactinomycetaceae was a dominant group with 40.00% relative abundance at family level (Fig. 1). Members of the Thermoactinomycetaceae family were represented by four main genera, namely, Desmospora (34.0%), Thermoactinomyces (4.00%), Laceyella (1.90%), and Shimazuella (0.30%); respective relative abundances at genus level are given in brackets (Fig. 1). This high abundance of Thermoactinomycetaceae, and in particular the dominance of genus Desmospora, is interesting as this group contains mesophilic or moderately thermophilic bacteria [29]. Other major members of Bacilli included Parageobacillus, Paenibacillus, Brevibacillus, Bacillus, and Alicyclobacillus and Geobacillus. Proteobacteria was made up of Alphaproteobacteria (2.00%), Gammaproteobacteria (1.90%), Betaproteobacteria (0.72%), and Deltaproteobacteria (0.56%); respective relative abundances at class level are given in brackets (Fig. 1). Pseudomonas was the most prominent genus at 0.40% abundance at genus level, while a total of 134 taxa were predicted at species level.
Morphological and Biochemical Characteristics of Isolates
Eight different bacterial isolates were obtained after enrichment in NB and being streaked onto nutrient agar. Enrichments and isolation were performed at three different temperatures, namely, 65 °C, 70 °C, and 75 °C. The isolates were named 3WAk1, 3WAk2, 3WAk3, 3SAk1, 3SAk2, 3SAk3, 3SAk4, and 3SAk5. Colonies developed after 24 h of incubation. The colonies varied in terms of shape, color, texture, and margin, with diameters between 0.9 and 8.0 mm (Table 1). All colonies were translucent, with consistencies varying from mucoid to smooth, and with irregular or circular shapes (Table 1). All the isolates were catalase positive, motile and spore forming, but negative for H2S production. They grew in a pH range of 5–10 and in a temperature range of 40–85 °C, with optima around pH 7.4 and 65–75 °C (Table 1). Strains 3WAk3 and 3SAk4 were extremely thermophilic, with 75 °C as optimal temperature and a maximum tolerated temperature of 85 °C. All isolates except one—strain 3SAk2—were oxidase positive. None of the strains grew under anaerobic conditions.
The API® 50 CH and API® ZYM profiles for the eight strains indicated major phenotypic diversity, as none shared the same phenotypic profile (Supplementary Tables S2, S3). Furthermore, the fermentation of carbohydrates varied between isolates; for example, strains 3SAk1 and 3WAk1 could utilize 16–20 substrates, as opposed to 3WAk3 for which it was only 7. Only strain 3WAk2 yielded a positive reaction with inositol. All strains were positive for esterase (C4), alkaline phosphatase, esterase lipase (C8), α-glucosidase, naphthol-AS-BI-phosphohydrolase, and acid phosphatase, but negative for β-glucuronidase and β-galactosidase, when tested using the API® ZYM kit. The other enzyme tests showed variable results (Supplementary Table S3).
Cells of all the isolates were Gram positive, rod shaped, and endospore forming, as evaluated by light microscopy. Six of the isolates were subjected to SEM, which revealed short and long cells, sometimes slightly curved and with bulbously shaped ends (Fig. 2), consistent with the morphologies typical of the Bacillaceae family.
Each isolate produced at least two of the four extracellular hydrolytic enzymes that were tested for—protease, cellulase, amylase, and lipase—while all produced lipase (Table 2). Five isolates (3WAk1, 3SAk1, 3SAk2, 3SAk4, and 3SAk5) produced a combination of all four relevant extracellular enzymes. The clear zones around the colonies, representing the amounts and activity levels of secreted enzymes, varied with incubation temperatures but generally appeared maximal at 65 °C (Table 2).
Phylogenetic Identification
Phylogenetic analysis of the strains entailed amplification and sequencing of their 16S rRNA genes. When analyzed through BLASTn searches, all fell into the Geobacillus genus with 16S rRNA gene sequence identities in the range 99.4–99.8% (Supplementary Table S4). Strain 3WAk1 showed a particularly close correlation to Geobacillus lituanicus strain N-3T.
A phylogenetic tree including representative strains of Geobacillus species, was constructed (Fig. 3). The new strains fell into five clusters, one of which contained four highly similar strains (3SAk3, 3SAk4, and 3WAk2) clustering tightly around Geobacillus stearothermophilus, with high bootstrap support. Strains 3SAk2 and 3WAk1 also affiliated with G. stearothermophilus, leading us to believe that these five strains most likely belong to this species. Strain 3SAk1 grouped closely with Geobacillus lituanicus, while strains 3SAk5 and 3WAk3 possibly represent novel Geobacillus species.
Phylogenetic maximum-likelihood tree based on 16S rRNA gene sequences of isolates (in bold) and representative type strains Geobacillus species. Accession numbers are shown in brackets. Bootstrap values (>70%) are shown as percentages at the nodes. Bar: 0.010 changes per nucleotide position. Parageobacillus thermantarcticus R-35644T (NR_116986.1) was used as outgroup
Discussion
Metagenomics is now broadly applied in the discovery of novel and potentially significant biocatalysts in various environments [30], including hot springs [17, 31, 32]. Geothermal areas are the selective habitats of thermophiles that are useful sources of unique extremozymes and biomolecules. Thermozymes are adapted to remain stable and active at high temperatures, allowing them to contribute significantly to biocatalytic manufacturing operations. This is due to their robustness, ability to enable higher substrate solubility at elevated temperatures, reduced risk for causing system contamination, as well as lower viscosity and increased miscibility [4, 5]. Nowadays, more than 500 products are made using enzyme technology and approximately 150 manufacturing processes are based on catalysts derived from microbes [20]. Additionally, more than 3000 known enzymes are used in manufacturing processes; roughly 65% of these are hydrolases used in the starch detergent, paper, and pulp industries, and 25% are applied in food processing [32]. Studies have shown that the diversity of extremophilic microorganisms may be greater than previously estimated [33]. To our knowledge, this study is the first to reveal the diversity of the bacterial community inhabiting the Zharkent geothermal hot spring, located in the southeastern region of Kazakhstan, through a multidisciplinary study implementing both classic culture-based and molecular methods. Most of the isolates obtained from samples, belonged to the bacterial genus, Geobacillus. Metagenomic analysis revealed many bacterial taxa and a strong dominance of Bacteria. An extremely small group of representatives of the Archaea domain, were found in the hot spring water samples (~0.03%). These results are surprising and inconsistent with those of other hot spring studies, as Archaea tend to dominate in extremely hot geothermal environments. For example, phylogenetic analysis of ten hot springs in Tibet revealed the presence of >50% Archaea in relevant samples [34]. Another study [35] analyzed the thermophilic microbial community in the hydrothermal water reservoirs of the Tattapani geothermal area in the Surguja district, Chhattisgarh, India. Here, water temperature ranges from 55 to 98 °C and study results revealed the presence of >4.8% Archaea.
Extremely high-temperature environments are commonly inhabited by chemolithoautotrophic thermophilic species, with the majority comprising Archaea and Aquificae. However, in the Zharkent hot spring, the most abundant species were of the Paenibacillus, Geobacillus, and Bacillus genera (Fig. 1). The high abundance of Bacillus species in this environment was anticipated due to the ability of most extremophilic Bacillales to form spores [36]. However, these microbes are not merely surviving but rather thriving, which means that the Zharkent spring is a unique source of these microbial communities.
In the present study, eight isolates were recovered using culture-dependent methods. For the identification and characterization of these isolates, both phenotypic and genotypic characteristics were determined. Based on growth and biochemical characteristics, such as tolerance to high temperatures and low-nutrient environments, spore-forming ability, and fermentative activity, all eight strains appeared to be members of the genus Geobacillus. One of the most interesting features was that majority of the strains could grow at higher temperatures than other, previously described Geobacillus species [36]. As Geobacillus species show poor identity values based on the 16S rRNA gene sequence [37], it is possible that some strains from this study—3WAk1, 3WAk2, 3WAk3, 3SAk1, 3SAk2, 3SAk4, and 3SAk5—could represent new species. However, this would need to be confirmed through comparative genome sequence analysis. Positive results regarding thermophilic protease, cellulase, amylase, and lipase activity indicate potential for application of these isolates in biotechnology. Of the thermophiles, Geobacillus species have received considerable attention due to their production of industrially important extremozymes [36]. Strain 3SAk4 is particularly interesting in this respect as it produces all four types of hydrolytic enzymes that were tested for in this study.
While attempts are being made to develop an understanding of the role of unculturable microorganisms in modern biotechnology, culturing of relevant microorganisms remains important. The combination of culture-dependent and -independent approaches results in a powerful tool for understanding microbial communities in hot springs and assists in the identification of new biomolecules and extremozymes.
Conclusion
The present study provides important information regarding the microbial diversity thriving in the Zharkent geothermal spring. As genomic and metagenomic databases become more expansive, particularly with regard to extremophiles and unculturable microbes, the taxonomic classification of currently unresolved sequences will strengthen the comparative analysis ability of the combined approach presented in this study, allowing for a more complete understanding of these unique environments.
Comparative genome sequencing analysis of the strains isolated during this study could provide further insight into the potential novelty of one or more of these strains.
Data Availability
The datasets generated during and/or analyzed during the current study are available in the GenBank repository, https://www.ncbi.nlm.nih.gov/genbank/. Any data generated or analyzed during this study that are not included in the published article will be available from the corresponding author on reasonable request.
References
Raddadi N, Cherif A, Daffonchio D, Neifar M, Fava F (2015) Biotechnological applications of extremophiles, extremozymes and extremolytes. Appl Microbiol Biotechnol 99(19):7907–7913. https://doi.org/10.1007/s00253-015-6874-9
DeCastro ME, Rodriguez-Belmonte E, Gonzalez-Siso MI (2016) Metagenomics of thermophiles with a focus on discovery of novel thermozymes. Front in Microbiol 7:1521. https://doi.org/10.3389/fmicb.2016.01521
Khalil A (2011) Screening and characterization of thermophilic bacteria (lipase, cellulase and amylase producers) from hot springs in Saudi Arabia. J Food Agric Environ 9(2):672–675. https://doi.org/10.1234/4.2011.2187
Khalil A (2011) Isolation and characterization of three thermophilic bacterial strains (lipase, cellulose and amylase producers) from hot springs in Saudi Arabia. Afr J of Biotechnol 10(44):8834–8839. https://doi.org/10.5897/AJB10.1907
Panda MK, Sahu MK, Tayung K (2013) Isolation and characterization of a thermophilic Bacillus sp. with protease activity isolated from hot spring of Tarabalo, Odisha, India. Iran J Microbiol 5(2):159–165
Reysenbach AL, Wickham GS, Pace NR (1994) Phylogenetic analysis of the hyperthermophilic pink filament community in Octopus Spring, Yellowstone National Park. Appl Environ Microbiol 60(6):2113–2119. https://doi.org/10.1128/AEM.60.6.2113-2119.1994
Slobodkin A, Reysenbach AL, Mayer F, Wiegel J (1997) Isolation and characterization of the homoacetogenic thermophilic bacterium Moorella glycerini sp. nov. Int J Syst Bacteriol 47(4):969–974. https://doi.org/10.1099/00207713-47-4-969
Slobodkin A, Reysenbach AL, Strutz N, Dreier M, Wiegel J (1997) Thermoterrabacterium ferrireducens gen. nov., sp. nov., a thermophilic anaerobic dissimilatory Fe(III)-reducing bacterium from a continental hot spring. Int J Syst Bacteriol 47(2):541–547. https://doi.org/10.1099/00207713-47-2-541
Huber R, Eder W, Heldwein S, Wanner G, Huber H, Rachel R et al (1998) Thermocrinis ruber gen. nov., sp. nov., a pink-filament-forming hyperthermophilic bacterium isolated from Yellowstone National Park. Appl Environ Microbiol 64(10):3576–3583. https://doi.org/10.1128/AEM.64.10.3576-3583.1998
Reysenbach AL, Ehringer H, Hershberger K (2000) Microbial diversity at 83 degrees C in Calcite springs, Yellowstone national park: another environment where the aquificales and “Korarchaeota” coexist. Extremophiles 4(1):61–67. https://doi.org/10.1007/s007920050008
Adiguzel A, Ozkan H, Baris O, Inan K, Gulluce M, Sahin F (2009) Identification and characterization of thermophilic bacteria isolated from hot springs in Turkey. J Microbiol Methods 79(3):321–328. https://doi.org/10.1016/j.mimet.2009.09.026
Maugeri TL, Gugliandolo C, Caccamo D, Stackebrandt E (2001) A polyphasic taxonomic study of thermophilic bacilli from shallow, marine vents. Syst Appl Microbiol 24(4):572–587. https://doi.org/10.1078/0723-2020-00054
Derekova A, Mandeva R, Kambourova M (2008) Phylogenetic diversity of thermophilic carbohydrate degrading bacilli from Bulgarian hot springs. World J Microbiol Biotechnol 24:1697–1702. https://doi.org/10.1007/s11274-008-9663-0
Sievert SM, Ziebis W, Kuever J, Sahm K (2000) Relative abundance of archaea and bacteria along a thermal gradient of a shallow-water hydrothermal vent quantified by rRNA slot-blot hybridization. MicroSoc 146(6):1287–1293. https://doi.org/10.1099/00221287-146-6-1287
Verma A, Gupta M, Shirkot P (2014) Isolation and characterization of thermophilic bacteria in natural hot water springs of Himachal Pradesh (India). Bioscan 9:947–952. https://doi.org/10.1007/s13213-014-0984-y
Niederberger TD, Ronimus RS, Morgan HW (2008) The microbial ecology of a high-temperature near-neutral spring situated in Rotorua New Zealand. Microbiol Res 163(5):594–603. https://doi.org/10.1016/j.micres.2006.09.001
López-López O, Cerdán ME, González-Siso MI (2013) Hot spring metagenomics. Life 3(2):308–320. https://doi.org/10.3390/life3020308
Giampaoli S, Berti A, Di Maggio RM, Pilli E, Valentini A, Valeriani F et al (2014) The environmental biological signature: NGS profiling for forensic comparison of soils. Forensic Sci Int 240:41–47. https://doi.org/10.1016/j.forsciint.2014.02.028
White RA, Chan AM, Gavelis GS, Leander BS, Brady AL, Slater GF et al (2016) Metagenomic analysis suggests modern freshwater microbialites harbor a distinct core microbial community. Front Microbiol 6:1531. https://doi.org/10.3389/fmicb.2015.01531
Antranikian G, Egorova K (2007) Extremophiles, a unique resource of biocatalysts for industrial biotechnol. Physiology and biochemistry of extremophiles. American Society of Microbiology, Washington, pp 361–406. https://doi.org/10.1128/9781555815813.ch27
Cowan DA (1991) Industrial enzymes in biotechnol. The science and the business. Harwood Academic, Amsterdam, pp 311–340
Kovacs N (1956) Identification of Pseudomonas pyocyanea by the oxidase reaction. Nature 178(4535):703. https://doi.org/10.1038/178703a0
Wehr HM, Frank JF, Association APH (2004) Standard methods for the examination of dairy products. American Public Health Association, Washington
Shaikh NM, Patel A, Mehta S, Patel N (2013) Isolation and screening of cellulolytic bacteria inhabiting different environment and optimization of cellulase production. Univers J Environ Res Technol 3(1):39–49
Kasana RC, Salwan R, Dhar H, Dutt S, Gulati A (2008) A rapid and easy method for the detection of microbial cellulases on agar plates using gram’s iodine. Curr Microbiol 57(5):503–507. https://doi.org/10.1007/s00284-008-9276-8
Gonzalez C, Gutierrez C, Ramirez C (1978) Halobacterium vallismortis sp. nov. An amylolytic and carbohydrate-metabolizing, extremely halophilic bacterium. Can J Microbiol 24(6):710–715. https://doi.org/10.1139/m78-119
Woese CR, Gutell R, Gupta R, Noller HF (1983) Detailed analysis of the higher-order structure of 16S-like ribosomal ribonucleic acids. Microbiol Rev 47(4):621–669. https://doi.org/10.1128/MMBR.47.4.621-669.1983
Peake I (1989) The polymerase chain reaction. J Clin Pathol 42(7):673–676. https://doi.org/10.1136/jcp.42.7.673
Yassin AF, Hupfer H, Klenk HP, Siering C (2009) Desmospora activa gen. nov., sp. nov., a thermoactinomycete isolated from sputum of a patient with suspected pulmonary tuberculosis, and emended description of the family Thermoactinomycetaceae Matsuo et al. 2006. Int J Syst Evol Microbiol 59(3):454–459. https://doi.org/10.1099/ijs.0.001362-0
Lorenz P, Eck J (2005) Metagenomics and industrial applications. Nat Rev Microbiol 3(6):510–516. https://doi.org/10.1038/nrmicro1161
Stamatakis A (2014) RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30(9):1312–1313. https://doi.org/10.1093/bioinformatics/btu033
Adrio JL, Demain AL (2014) Microbial enzymes: tools for biotechnological processes. Biomolecule 4(1):117–139. https://doi.org/10.3390/biom4010117
Kumar L, Awasthi G, Singh BJB (2011) Extremophiles: a novel source of industrially important enzymes. Biotechnology 10(2):121–135. https://doi.org/10.3923/biotech.2011.121.135
Huang Q, Dong CZ, Dong RM, Jiang H, Wang S, Wang G et al (2011) Archaeal and bacterial diversity in hot springs on the Tibetan Plateau, China. Extremophiles 15(5):549–563. https://doi.org/10.1007/s00792-011-0386-z
Kaushal G, Kumar J, Sangwan RS, Singh SP (2018) Metagenomic analysis of geothermal water reservoir sites exploring carbohydrate-related thermozymes. Int J Biol Macromol 119:882–895. https://doi.org/10.1016/j.ijbiomac.2018.07.196
Schleifer K-H (2009) Phylum XIII. Firmicutes Gibbons and Murray, 5 (Firmacutes [sic] Gibbons and Murray 1978, 5). Bergey’s manual® of syst bacter. Springer, Berlin, pp 19–1317
Zeigler DR (2005) Application of a recN sequence similarity analysis to the identification of species within the bacterial genus Geobacillus. Int J Syst Evol Microbiol 55(Pt 3):1171–1179. https://doi.org/10.1099/ijs.0.63452-0
Acknowledgements
The authors are grateful to the Molecular Imaging Center and the ICP-Laboratory at the University of Bergen for their excellent service in performing electron microscopy and chemical analyses for this study, respectively.
Funding
The research was supported by the Eurasia program of the Norwegian Agency for International Cooperation and Quality Enhancement in Higher Education (Diku) under Project CPEA-LT-2017/10061.
Author information
Authors and Affiliations
Contributions
AM, NKB, AK contributed to conceptualization and methodology; AM, RJL, NKB, and AK were involved in formal analysis and investigation; AM, NKB, AK, IS were involved in writing––original draft preparation; AM and NKB were involved in writing––review and editing; AK and NKB contributed to funding acquisition; AK, IS, and NKB performed supervision.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Consent for Publication
All the authors consent to publication of this study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Mashzhan, A., Javier-López, R., Kistaubayeva, A. et al. Metagenomics and Culture-Based Diversity Analysis of the Bacterial Community in the Zharkent Geothermal Spring in Kazakhstan. Curr Microbiol 78, 2926–2934 (2021). https://doi.org/10.1007/s00284-021-02545-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-021-02545-2