Abstract
In degradation of total petroleum hydrocarbon, 35 isolates belonging to 11 genera were sanitized and 3 isolates as well as their consortium were initiated to be able to raise in association with petroleum hydrocarbon as sole source of carbon under in vitro circumstances. The isolated strains were grounded on internal transcribed spacer (ITS) rDNA sequence analysis. The fungal strains with the utmost potentiality to reduce petroleum hydrocarbon without emerging antagonistic activities were Aspergillus niger, Penicillium ochrochloron, and Trichodema viride. For fungal growth on petroleum hydrocarbon, P. ochrocholon gained weight of 44%, A. niger 49%, and T. viride 39% within the first 30–40 days. As compared to the controls, these fungi accumulated significantly higher biomass, produced extracellular enzymes, and degraded total petroleum hydrocarbon and A. niger strongly degraded total petroleum hydrocarbon with a degradation of about 71.19%. These observations with GC-MS data confirm that these isolates displayed rapid total petroleum hydrocarbon biodegradation within a period of 60 days and the half-life showed that A. niger was the shortest with t1/2 = 21.280 day−1 corresponding to the highest percent degradation of 71.19% and first-order kinetic fitted into the present study. By multivariate analysis, five main factors were identified by factor analysis (FA). The first factor (F1) of the fungi species accounts for 20.0% which signifies that fungi species controls the degradation of petroleum variability and hierarchical cluster analysis (HCA) as a dendrogram with five observations and three variables shows two predominant clusters order cluster 1 > 2.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Soil pollution by hydrocarbons is one of the global environmental problems of recent time related to the petrochemical industry (Nilanjana and Preethy 2011) and contamination with hydrocarbons can have a profound effect on soil fauna (Anna et al. 2015) while Onojake et al. (2014) indicated in their study that hydrocarbon contamination of soils and the mangrove ecosystem has become a wide spread global environmental issue since the discovery of crude oil. Soil pollution with petroleum hydrocarbons has a high environmental impact, and oil spills cause serious environmental problems worldwide (David et al. 2017). The hydrocarbon contamination of the environment can be attributed to the transportation and utilization of petroleum oil and petroleum products all over the world and there are many ways of petroleum hydrocarbon contamination, some of which includes pipelines and oil well leakages, wrong methods of disposal of petroleum wastes, and accidental oil spills (Kriti and Subhash 2014). Petroleum hydrocarbons are composed of complicated mixtures of non-aqueous and hydrophobic components such as n-alkane, aromatics, resins, and asphaltenes. Due to the adverse impact of these chemicals on human health and the environment, they are classified as priority environmental pollutants by the US Environmental Protection Agency (1986). Over 84 million barrels of petroleum are expended globally per day, with over 50% of petroleum oil conveyed by ocean with high chances of oil spill over the environment due to the accidents that leads to water as well as soil pollutions (Hasan et al. 2010; Rhodes 2010; Eman et al. 2017). The physico-chemical processes such as incinerations, soil washing, the use of oil booms, soil vapor extraction, as well as solidification have been adopted for the purpose of remediating oil-polluted sites. Though, as expressed by Hasan (2014), these methods are labor-intensive, expensive, and relatively disruptive. These aforementioned disadvantages have led to the increasing interest in the use of bioremediation processes for the cleanup of petroleum-contaminated soil. The use of living organisms for remediation of contaminated sites, also known as bioremediation, has been studied by several researchers as an alternative to chemical treatment method over the last decade, as a sustainable and cost-effective method (Chikere et al. 2011; Kanaly and Harayama 2010). Biotreatment is cheap, ecofriendly, simple to execute, used over big areas, as well as high-mineralization efficiency and bioremediation of oil-polluted soil is efficient, economical, and a versatile alternative to physiochemical treatments (Barathi and Vasudevan 2003). Biodegradation of petroleum by fungi is well-known (Palanisamy et al. 2014), as many microorganisms are able to use hydrocarbons as both energy and carbon sources (Montagnolli et al. 2015). Through different extracellular and intracellular enzymatic activities reviewed by Fritsche and Hofrichter (2008), these microorganisms can mineralize petroleum hydrocarbons (PHCs) by degrading them to end product of CO2 but the rate and extent of mineralization depends on the metabolic abilities of the microorganisms (Dobler et al. 2000). To overcome slow innate PHCs biodegradation in soil, adjustments to optimum nutrient concentrations, pH, and/or temperature (biostimulation), along with microbial inoculation (bioaugmentation), may be used (Sayara et al. 2010; Tahhan et al. 2011) and in order to make bioaugmentation to be efficient in practice, inoculated microorganisms must tolerate contaminants, efficiently degrade compounds of interest, and thrive in the target environment (Yao et al. 2015). Polluted soil are of particular interest as sources for cultivation, since microbes in these soil are more likely to have developed multiple tolerance mechanisms, allowing them to survive and function effectively in the presence of PHCs (Caliz et al. 2012; Oriomah et al. 2014).
Numerous degrees of success have been recorded with bioremediation processes for the mineralization of soil contaminated with petroleum oil through inoculation of the affected soil with the exogenous hydrocarbon using microbial strains to increase the uptake of pollutants (Eman et al. 2017). Success in bioremediation processes is dependent on the activity and survival of the demeaning microbial strain once presented to the targeted environment. One of the strategies to expand the efficacy of bioremediation methods is the overview of exceedingly specialized microbes with high adaptive skill and ability to utilize petroleum as well as its derivatives as the only cause of carbon into the polluted environment (McGenity et al. 2012). In petroleum-contaminated soil, apart from carbon, other nutrients are deficient. Therefore, the addition of certain nutrients such as phosphate as well as nitrate will increase the development and efficiency of hydrocarbonoclastic bacteria.
Many isolated bacterial and fungal species have been reported to be capable of biodegrading petroleum hydrocarbons and even polynuclear aromatic hydrocarbons effectively (Marquez-Rocha et al. 2005). Laboratory studies provide greater control and manipulation in providing a basis to distinguish between biotic and abiotic processes and to determine the optimal conditions for biodegradation (Li et al. 2008). It is known that petroleum hydrocarbons can be removed by microorganisms such as fungi belonging to the genera Aspergillus, Penicillium, Fusarium, Amorphotheca, Neosartorya, Paecilomyces, Talaromyces, Graphium, and yeasts which includes Candida, Yarrowia, and Pichia and microalgae (Chaillan et al. 2004). There are insufficient confirmations which indicate that under few conditions, parasites could be a superior degrader of crude oil to customary bioremediation systems that enlist microorganisms (Saravanan and Sivakumar 2013). According to Kiran et al. (2009) and Passarini et al. (2011), fungi has the ability to utilize PAHs as benzo (a) pyrene to produce surfactants. Various fungi strains notably Aspergillus terreus MUT 271, Penicillium citreonigrum MUT 267, and Trichodema harzianum MUT 290 have been shown as good degraders of petroleum oil (Bovio et al. 2017). Cordyceps militaris is capable of metabolizing dichoro-trichlordibenzo-p-dioxin, trichlorodibenzo furan, and 3–3 ring polycyclic aromatic into their corresponding hydroxide (Toshio et al. 2015). Also, Hassan et al. (2018), Hadibarata et al. (2009), and Varjani 2017), stated that there are a large number of ligninolytic and non-ligninolytic fungi (such as Bjerkandera, Irpex, Lentinus, Polyporus, Penicillium, Aspergillus, and Candida) that have the capability to degrade petroleum and also the ability of fungi to secrete enzymes, especially those involved in the decomposition of lignin has a great effect on the degradation of petroleum hydrocarbons (Acevedoa et al. 2011). Though previous studies which have been done on the integration of biostimulation and bioaugmentation (Calvo et al. 2009; Zhao et al. 2011; Kauppi et al. 2011; Sheppard et al. 2011) in treatment of petroleum-contaminated soil worldwide and numerous studies have demonstrated the potential for petroleum hydrocarbon (PHC) bioremediation by fungi as seen in Table 1 (Sayara et al. 2011; Lee et al. 2015; Hadibarata et al. 2009; Eman et al. 2016; Hassan et al. 2018; Hung et al. 2008; Manli et al. 2016; Weiwei et al. 2017; Naga et al. 2017; Amechi and Chukwudi 2017; Naga et al. 2016) but very limited studies have been done on Libyan petroleum contamination of soil (Shaieb et al. 2015). Although researchers have not discussed studies on optimal conditions of isolation of fungi, this study examined the degradative ability of fungi isolation from olive oil mill effluent and its ability to remediate Libyan-contaminated soil with petroleum oil and half-life rate of degradation from first- and second-order kinetic.
2 Experimental Methods
2.1 Materials and Sample Preparation
Soil samples (silt-loam texture) were collected at depth of 0–15-cm layer from an olive oil mill effluent site in Lapta, Turkish Republic of North Cyprus (TRNC) and the samples were bagged and kept in ice coolers at 4 °C before transporting to the laboratory for further use majorly to maintain the quality of the soil. The collected soil samples were mixed to form composite soil sample followed by crushing with a mortar and a pestle. It was then sieved to evacuate enormous constituent part and plant leftover (Chouhan et al. 2014). Soil-petroleum oil concentrations (20:5 w/v) were prepared by first purifying the soil by autoclaving at 121 °C for 15 min followed by mixing the chosen amount of petroleum oil with 20 g of soil sample. The petroleum sample was removed with 100 mL of dichloromethane (CH2Cl2) by utilizing a separating funnel. Surplus water in the mineral medium after withdrawal was removed by adding anhydrous sodium sulfate (Na2SO4). The extract concerted in a rotary evaporator and 1 mL of the extract was used for GC-MS.
2.2 Total Petroleum Hydrocarbons and Physicochemical Analysis of the Soil Samples
The degradative ability of the fungi on the respective total petroleum hydrocarbons (TPHs) in the extracts was determined by using gas chromatography mass spectrophotometry (GCMS-QP2010 plus, Shimadzu, Japan) equipped with a TRB-1 capillary column (30 m × 0.25 mm × 0.25 μm). An analysis was conducted to determine the water-holding capacity (WHC), pH, moisture content, and soil texture of the soil samples. Hydrometer method was employed as described in ASTM D422-63 for soil particle size analysis. The pH of soil was carried out in water in a 1:1 soil solution ratio (Chima et al. 2016). The concentrations of inorganic elements (Zn, P, Ca, S, K, Fe, and Mg) were analyzed by using x-ray fluorescence spectroscopy (XRF-1800). The percentage of the concentration of organic carbon and nitrogen in the soil was determined by using a TOC-Vcph machine (Shimadzu mandel, Canada) and the results are shown in Table 2.
2.3 Fungal Growth and Identification
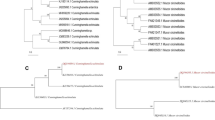
For the isolation of lipase-secreting fungi from the collected soil samples of the olive oil mill effluent site, serial dilution approach was adopted and 1 g of the soil sample was diluted in 9-mL sterile distilled water. The dilution was made up to 10−5 and 0.1 mL of the diluted aliquot was transferred onto 9-cm-diameter petri dishes containing 20 mL of potato dextrose agar (PDA) added with antibiotic (streptomycin 0.015 g L−1, chloromphenicol 0.05 g L−1). Plates were incubated at 24 °C with constant monitoring for slow-growing colonies for a period of 15 days. Each strain was isolated in pure culture for the taxonomic identification. Fungi were classified according to their macroscopic features and also by their molecular characterization. The isolated strains were identified on the basis of internal transcribed spacer (ITS) rDNA sequence analysis. For sequence analysis, the ITS1–5.8SITS4 rDNA gene of the fungi was amplified by using PCR with primer set pITS1 (5′ TCCGTAGGTGAACCTGCCG-3′) and pITS4 (5′ TCCTCCGCTTATTGATATGC-3′) (Al-Nasrawi 2012). The 550-bp amplicon were obtained, cloned, and sequenced. DNA sequence of the positive clones with 18S rDNA gene fragment was compared to the gene banks database. All identified strains were deposited in the biotechnology laboratory of Cyprus International University.
2.4 Fungi Growth on Basal Salt Medium
The three selected fungal strain notably: Aspergilus niger, Penicillium ochrochloron, and Trichoderma viride, were pre-grown in a basal salt medium to stimulate petroleum hydrocarbon degradation conditions based on compositions of Olga et al. (2015) and Toshio et al. (2015) with some modifications. The composition of the basal salt medium (BSM) was 0.8 K2HPO4, 0.8 KH2PO4, 0.8 (NH4) SO4, 0.6 NaCl, and 5 mL of magnesium/calcium solution (L−1): 0.8 g MgSO4.7H2O + 0.8 g CaCl2.2H2O. Thereafter, it was diluted to 200 mL with distilled water. The calcium/magnesium solution was sterilized, filtered (0.22 μm, Millipore), and added to the BSM after autoclaving. All selected fungi were cultured differently on 9-cm-diameter potato dextrose agar plates at 25 °C. The mycelia mats (two agar blocks) were transferred from agar plates to a sterilized conical flask containing 50-mL sterilized water and then homogenized for 30 s. Then, 1 mL of the homogenate was inoculated into 50-mL BSM (pH 6.9) containing 1.0% (w/v) glucose as the sole carbon source in a 100-mL Erlenmeyer flask. The cultures were pre-incubated statically at 25 °C for 6 days (Toshio et al. 2015; Hwanhwi et al. 2015; Zuzanna et al. 2015).
2.5 Enzyme Assay
For the determination of fungi, lipase, catalase, laccase, and manganese activities were assayed after filtration of the different fungi mycelium and spores by using a syringe filter (0.45 μm). The enzyme was measured by adding 200 μL of the supernatant crude in 0.1 mM tris-HCl buffer (pH 7.8). Then, 2.48 mL containing 0.15 M NaCl and 0.5% Triton X-100 carried out at a temperature condition of 40 °C for 5 min followed by addition of 50 mL MP-nitrophenyl palmitate. The sample was assayed with a UV-Vis spectrophotometer at a wavelength of 420 nm (Hwanhwi et al. 2015; Saowakon et al. 2017).
2.6 Culture Conditions and Artificial Contamination of Soil
Each 100-mL flask contained 20 g of sterilized petroleum oil contaminated soil. All the fungi were subculture for 7 days in 30-mL BSM (containing glucose as a source of carbon). The culture was directly added to the contaminated soil and then mixed thoroughly. The values were nurtured on a revolving shaker at 150 rpm (27 °C). The fluid as well as dense portions of the fungal values were removed in every 10-day interval for a total of 60 days. Every sample was removed three times with 100 mL dichloromethane (CH2Cl2) by utilizing a separating funnel. Surplus water in the mineral medium after removals was removed by adding anhydrous sodium sulfate (Na2SO4) and the extract concentrated in a rotary evaporator (Hwanhwi et al. 2015; Zuzanna et al. 2015).
2.7 Multivariate Analysis
Successful multivariate analysis by XLSTAT 2018 software relies greatly on the optimal selection of variables, particularly their independence and comparable relevance (Saowakon et al. 2017). Multiple correspondence analysis (MCA) was used for nominal categorical data to detect and represent underlying structures in data set of the result of this study. This is done by representing data as points in a low-dimensional Euclidean space and MCA can be viewed as an extension of simple correspondence analysis (CA). Additionally, CA was performed in order to validate the groups, similarities, or differences. Agglomerative hierarchical clustering (AHC) is a method of cluster analysis which seeks to build a hierarchy of clusters. Strategies for hierarchical clustering in this study fall into Agglomerative:
This is a “bottom up” approach: each observation starts in its own cluster, and pairs of clusters are merged as one moves up the hierarchy. Hierarchical cluster analysis (HCA) is widely applied in which clusters are sequentially formed, starting stepwise through repetition of forming and joining clusters by the largest similarity between a pair of objects to higher cluster formation until a single cluster is obtained. A dendrogram clarifying the clustering process and clusters proximity is then produced (de Amorim et al. 2016).
2.8 Kinetic Modeling
The kinetic was studied at a time interval of 10 days for equilibrium time of 60 days and the rate of total petroleum hydrocarbon by microorganisms is determined by the equation as described by (Agarry et al. 2013; Agarry and Oghenejoboh 2015) for differential rate law and integral rate law for first- and second-order rate kinetics.
Differential rate
Integral rate
Rearranging
t1/2 is a timescale by which the initial population is decreased by half of its original value. This can be represented by Eqs. (5) to (9) according to Amechi and Chukwudi (2017).
After a period of one half-life t = t1/2
Taking log of both sides In ex = x
For the second-order reaction, the differential rate equation are according to the study of Agarry et al. 2015 and Maletic et al. 2013 (Eqs. 8–16).
Rearranging
However, at half-life
where K = the biodegradation constant (day−1), where d = day, C0 = the initial concentration of total petroleum hydrocarbon, Ct = the residual concentration of total petroleum hydrocarbon in the soil at time (t), and time = days.
3 Results and Discussions
3.1 Isolation and Evaluation of the Fungal Communities
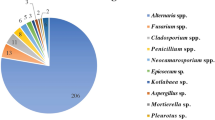
Thirty-five fungi, which were isolated with distinct morphological and physiogenetic characteristics, were derived from the soil samples from the olive oil mill effluent site. These isolated species include: two species of Candida, six species of Aspergillus, five species of Penicillium, and nine individual species of Trichordema, Emericella, Armonium, Paecilomyces, Eurotium, Cladosporium, Alternaria, and Geosmithia. Three isolates A. niger, T. viride, and P. ochrochloron (Fig. 1) were selected for further evaluation of biodegradation on the basis of higher growth and fungi biomass buildup in the course of co-cultivation with petroleum hydrocarbon and physiogenetic identification based on their internal spacer (ITS) region (Table 3).
3.2 Fungal Growth on Petroleum Oil
All the three selected fungi species were verified for their capability to grow in the presence of petroleum hydrocarbon as their only carbon source. The three selected fungi were able to grow on it with dissimilar metabolic efficacy when likened to the controls. The investigated inocula grew within 10–60 days to form colonies. Profuse growth of P. ochrochloron, A. niger, and T. viride. The P. ochrochloron had gained weight of 44% within 40 days followed by A. niger (49%) and T. viride (39%) within the first 30 days (Table 4). Figure 2a–c demonstrates gained treatment (g/L) which indicates the optimum weight to be within 30 days. Previous studies have demonstrated biodegradation ability of fungi biomass accumulation under similar conditions. A study by Abdulkadir et al. (2007) reported a maximum mycelia biomass of A. terreus, A. niger, and Penicillium chrysogenum growing on n-alkanes and crude oil as 50 (mg), 45 (mg), and 75 (mg), respectively, and A. niger demonstrated the highest weight loss (8.6%) (Al-Nasrawi 2012).
3.3 Enzymatic Activity of the Fungi Isolates
Lipase, catalase, and laccase enzymes were recorded in the treatment cultures of the three fungi (Table 5). A. niger secreted the highest amount of lipase (8.085 ± 0.069 U/mL) (Fig. 3b) followed by P. ochrochloron (8.025 ± 0.068 U/mL) and T. viride (8.039 ± 0.096 U/mL) which is in line with biomass increase in the same order while T. viride secreted the highest amount for catalase (8.912 ± 0.056 U/mL) followed by A. niger (8.146 ± 0.069 U/mL) and P. ochrochloron (7.815 ± 0.026 U/mL) (Fig. 3a) and that of laccase shows T. viride (8.05 ± 0.096 U/mL) to secrete the highest while A. niger secrete (7.83 ± 0.015 U/mL) and then P. ochrochloron (7.692 ± 0.026 U/mL) (Fig. 3c) (Table 5). There is a higher enzymatic activity compared to the controls. Bioremediation of petroleum hydrocarbons is mostly facilitated by secretion of extracellular enzymes by the acting microorganisms leading to the transformation of the oily substances into less toxic compounds. Hence, elevated enzyme levels in our treatment broth indicates a higher co-cultivation activity of T. viride with catalase. A study by Behnood et al. (2014) on biodegradation of PHCs by Phanerochaete chrysosporium reported lipase enzyme activity as 0.0426 (U/ml) during the fungal co-culture with petroleum hydrocarbon.
3.4 GC-MS Analysis for Total Petroleum Hydrocarbon Degradation
The GC-MS results of the total petroleum hydrocarbon substrate retrieved from the inoculum medium after interval of 10 days for up to 60 days were shown in Table 6. It was found that the selected fungi strains were efficient petroleum oil degraders as the number of PHCs concentration continue to reduce with increased degradation time. Treated total petroleum hydrocarbon substrate showed decrease in area of some peaks over time (days) which shows its breakdown while the new peaks represents breakdown of products or presumed metabolites (Fig. 4a–d). Also, the result shows that A. niger strongly degraded total petroleum hydrocarbon with a degradation of about 71.19% at the end of the 60-day treatment. The results also showed that the P. ochrochloron and T. viride reported degradation of about 66.56% and 35.80%, respectively, at the end of the 60-day treatment (Fig. 5).
3.5 Factor Analyses
Five main factors were identified by factor analyses, which accounted for the variances degradation percent of fungi species sample data (Tables 6). The spatial distribution of the variables in the spaces defined by F1 and F2 is shown in Fig. 6. The first factor (F1) of the fungi species accounts for 20.0% of the variance and has high to moderate positive loadings with P. ochrochloron, T. viride, and A. niger at 2.205, F1 signifies that fungi species controls the degradation of petroleum variability and suggests that this process has most likely been effective in the degradation. F2–F5, which accounts for another 20.0% of the variance and 1.0 of the eigenvalue, is characterized with negative loadings which reflects the infectiveness of degradation.
3.6 Hierarchical Cluster Analysis
Figure 6 shows the contributions of fungi species of petroleum degradation biplots on the loading space of primary factor (F1) versus second primary factor (F2) and Fig. 7 shows the results of hierarchical cluster analysis (HCA) presented as a dendrogram. A total of five observations and three variables were used for the analysis. The similarity of the HCA to the factor analysis confirms the interpretations made using the factor analysis. Distinctive classes of degradation time and fungi along with their spatial distribution are shown on Fig. 6. Two predominant clusters were identified in the fungi species data and their locations were found to be primarily controlled by time as described below. The fungi species is a major factor for increasing all major percentage of degradation in the order cluster 1 < 2 (Fig. 7).
Cluster 1 is the largest, containing three fungi species that are functional in the degradation of petroleum characterized by T. viride smallest degradation percentage (35.80%) followed by P. ochrochloron (66.56%) and A. niger (71.19%) classified into fungi species C2 = 4, 5; C1 = 1, 2; and C1 = 2, 3 in decreasing order while cluster 2 is grouped into only two fungi species and is characterized by low to high degradation.
3.7 Kinetic Study
In this study, total petroleum hydrocarbon biodegradation kinetics using first- and second-order rates by using linear plots of InC0/Ct and 1/Ct obtained from Eqs. (7) and (16), respectively, and then evaluated as seen in Fig. 8a, b. The half-life defined as time taken for half the contaminant/total petroleum hydrocarbon concentration to be consumed was shortest in the A. niger (t1/2 = 21.280 days). The order of half-lives for the three studied fungi could, however, be represented as thus; A. niger < P. ochrochloron < T. viride in both first- and second-order rates (Table 7). The highest r2 value and degradation constant were obtained in A. niger (r2 = 0.991, K1 = 2.243 × 10−2 day) for the first order. In the second-order kinetics, all the half-lives of the three fungi were significantly lower than those of first order which indicates a delay in biodegradation in term of first order for A. niger (30.903 days) and P. ochrochloron (32.496 days) while T. viride was higher in second order than first order with 108.710 and 98.598 days, respectively (Table 7). Meanwhile, the first-order kinetic fitted into the present study due to its higher r2. Biodegradation constant (Table 7) produced by first-order rate were significantly higher than those of second order in all treatments at P < 0.05.
4 Conclusion
The study has shown the efficiency of P. ochrochloron, A. niger, and T. viride isolated from olive oil mill effluent site in the biological treatment of contaminated Libyan soil. The pollutant biodegradation ability of the three isolated fungi as discovered in the study is attributed to their metabolic activities and secretion of catalase, lipase, and laccase enzymes during co-culture with Libyan petroleum oil-contaminated soil as sole source of carbon indicating a higher degradation activity of T. viride with catalase. Five main factors were identified which accounted for the variances in degradation percent of fungi species. The first factor (F1) of the fungi species accounts for 20.0% of the variance and has high to moderate positive loadings with P. ochrochloron, T. viride, and A. niger at 2.205. F1 signifies that fungi species controls the degradation of total petroleum hydrocarbon variability and hierarchical cluster analysis (HCA) as a dendrogram with a total of five observations and three variables were used for the analysis and the half-life showed that A. niger was the shortest with t1/2 = 21.280 day corresponding to the highest percent degradation of 71.19%. Therefore, the degradation of total petroleum hydrocarbon by fungi shows a potential and study is still open for further investigation as the biochemical and metabolic make-up on the studied fungi that were isolated from the olive oil mill effluent site that made them capable of degrading total petroleum hydrocarbon is still not known.
References
Abdulkadir, E., Abdulaziz, Y., AlKindi, S. A.-B., & Charles Bakheit, S. N. (2007). Biodegradation of crude oil and n-alkanes by fungi isolated from Oman. Marine Pollution Bulletin, 54, 1692–1696.
Acevedoa, F., Pizzul, L., Castillo, M. P., Cuevas, R., & Diez, M. C. (2011). Degradation of polycyclic aromatic hydrocarbons by the Chilean white-rot fungus Anthracophyllum discolor. Journal of Hazardous Materials, 185, 212–219.
Adnan, B., Jialong, Z., Shue, L., Jiashu, L., Hussein, B., Xiaoyu, Z., & Fuying, M. (2018). Biodegradation of n-hexadecane by Aspergillus sp. RFC-1 and its mechanism. Ecotoxicology and Environmental Safety, 164, 398–408.
Agarry, S. E., & Oghenejoboh, K. M. (2015). Enhanced aerobic biodegradation of naphthalene in soil: kinetic modelling and half-life study. International Journal of Environmental Bioremediation & Biodegradation, 3(2), 48–53.
Agarry, S. E., Aremu, M. O., & Aworanti, O. A. (2013). Kinetic modelling and half-life study on enhanced soil bioremediation of bonny light crude oil amended with crop and animal-derived organic wastes. Journal of Petroleum & Environmental Biotechnology, 4, 137. https://doi.org/10.4172/2157-7463.1000137.
Agarry, S. E., Oghenejoboh, K. M., & Solomon, B. O. (2015). Kinetic modelling and half-life study of adsorptive bioremediation of soil artificially contaminated with bonny light crude oil. Journal of Ecological Engineering, 16(3), 1–13.
Al-Nasrawi, H. (2012). Biodegradation of crude oil by fungi isolated from Gulf of Mexico. Journal of Bioremediation & Biodegradation, 3, 147. https://doi.org/10.4172/2155-6199.1000147.
Amechi, S. N., & Chukwudi, O. O. (2017). Bioremediation of gasoline contaminated agricultural soil by bioaugmentation. Environmental Technology & Innovation., 7, 1–11.
Anna, K., Ewa, B., Jarosław, L., Agnieszka, K., & Piotr, W. (2015). Influence of oil contamination on physical and biological properties of forest soil after chainsaw use. Water, Air, and Soil Pollution, 226, 389. https://doi.org/10.1007/s11270-015-2649-2.
Barathi, S., & Vasudevan, N. (2003). Bioremediation of crude oil contaminated soil by bioaugmentation of Pseudomonas fluorescens NS1. Journal of Environmental Science and Health, Part A: Toxic/Hazardous Substances and Environmental Engineering, 38(9), 1857–1866.
Behnood, M., Nasernejad, B., & Nikazar, M. A. (2014). Biodegradation of crude oil from saline waste water using white rot fungus Phanerochaete chrysosporium. Journal of Industrial and Engineering Chemistry, 20, 1879–1885.
Bovio, E., Gnavi, G., Prigione, V., Spina, F., Denaro, R., Yakimov, M., Calogero, R., Crisafi, F., & Varese, G. C. (2017). The culturable mycobiota of a Mediterranean marine site after an oil spill: isolation, identification and potential application in bioremediation. Science of the Total Environment, 576, 310–318. https://doi.org/10.1016/j.scitotenv.2016.10.064.
Caliz, J., Genoveva, M., Esther, M., Jordi, S., Robert, C., Garau, M. A., Xavier, T. M., & Xavier, V. (2012). The exposition of a calcareous Mediterranean soil to toxic concentrations of Cr, Cd and Pb produces changes in the microbiota mainly related to differential metal bioavailability. Chemosphere, 89, 494–504.
Calvo, C., Manzanera, M., Silva-Castro, G., Uad, I., & González-López, J. (2009). Application of bioemulsifiers in soil oil bioremediation processes. Future prospects. Sci Total Environ., 407(12), 3634–3640.
Chaillan, F., Fleche, A. L., & Bury, E. (2004). Identification and biodegradation potential of tropical aerobic hydrocarbon degrading microorganisms. Research in Microbiology, 155, 587–595.
Chikere, C. B., Okpokwasili, G. C., & Chikere, B. O. (2011). Monitoring of microbial hydrocarbon remediation in the soil. Review article. 3 Biotech, 1, 117–138.
Chima, C. W., Ndifreke, E. W., Edidiong, A. E., Emmanuel, U. E., & Akhayere, E. (2016). Modelling changes in soil chemical parameters of Obio-Akpor, Rivers state, Nigeria using geographic information system. Journal of Scientific and Engineering Research, 3(6), 207–221.
Chouhan, S., Garg, R. K., Sairkar, P., atav, N., Silawat, N., Sharma, R., Singh, R. K., & Mehrotra, N. N. (2014). Characterization of genetic variance within and among five populations of Sperata seenghala (Skyes, 1839) revealed by random amplified polymorphic DNA markers. Journal of Genetic Engineering and Biotechnology, 12(1), 7–14.
David, T. T., Otilio, A. S., Blanca, R. R. P., & Martha, G. C. (2017). Phylogeny and polycyclic aromatic hydrocarbons degradation potential of bacteria isolated from crude oil-contaminated site. Journal of Environmental Science and Health, Part A, 0(0), 1–8.
de Amorim, R. C., Makarenkov, V., & Mirkin, B. (2016). A-Wardpb effective hierarchical clustering using the Minkowski metric and a fast k-means initialization. Information Sciences, 370–371, 343–354.
Dobler, R., Matthias, S., & Reinhard, B. (2000). Population changes of soil microbial communities induced by hydrocarbon and heavy metal contamination. Bioremediation Journal, 4, 41–56.
Eman, K., Esmaeil, S., Arturo, A., Mohamed, T., Nagalakshmi, H., Tanvi, H., Paul, D., & Andrew, S. (2016). Bioremediation potential of diesel-contaminated Libyan soil. Ecotoxicology and Environmental Safety, 133, 297–305.
Eman, K., Esmaeil, S., Arturo, A.-M., Mohamed, T., Nagalakshmi, H., Tanvi, H. M., Morrison, P. D., & Ball, A. S. (2017). Bioremediation potential of diesel-contaminated Libyan soil. Ecotoxicology and Environmental Safety, 133, 297–305.
Environmental Protection Agency, U. S. (1986). Test method for evaluating solid waste, SW-846, third ed (p. 1A). Washington: U.S. EPA.
Fritsche, W., & Hofrichter, M. (2008). Aerobic degradation by microorganisms. In J. Klein (Ed.), Environmental processes soil decontamination (pp. 146–155). Weinheim: Wiley- VCH.
Hadibarata, T., Tachibana, S., & Itoh, K. (2009). Biodegradation of chrysene, an aromatic hydrocarbon by Polyporus sp. S133 in liquid medium. Journal of Hazardous Materials, 164, 911–917.
Hasan, I. (2014). Biodegradation of kerosene by Aspergillus niger and Rhizopus stolonifer. Applied and Environmental Microbiology, 2, 31–36.
Hasan, S. W., Ghannam, M. T., & Esmail, N. (2010). Heavy crude oil viscosity reduction and rheology for pipe line transportation. Fuel, 89(5), 1095–1100.
Hassan, G., Hamid, M., & Seyed, M. D. (2018). Evaluation of heavy petroleum degradation using bacterial-fungal mixed cultures. Ecotoxicology and Environmental Safety, 164, 434–439.
Hung, S., Pius, M., Makoto, S., & Chae, G. (2008). Bioremediation of oil-contaminated soil using Candida catenulate and food waste. Environmental Pollution., 156, 891–896.
Hwanhwi, L., Yeongseon, J., Young, M., Hanbyul, L., Gyu-Hyeok, K., & Jae-Jin, K. (2015). Enhanced removal of PAHs by Peniophora incarnata and ascertainment of its novel ligninolytic enzyme genes. Journal of Environmental Management, 164, 10–18.
Kanaly, R. A., & Harayama, S. (2010). Advances in the field of high-molecular-weight polycyclic aromatic hydrocarbon biodegradation by bacteria. Microbial Biotechnology, 3, 136–164.
Kauppi, S., Sinkkonen, A., & Romantschuk, M. (2011). Enhancing bioremediation of diesel-fuel-contaminated soil in a boreal climate: comparison of biostimulation and bioaugmentation. International Biodeterioration and Biodegradation, 65(2), 359–368.
Kiran, G. S., Hema, T. A., Gandhimathi, R., Selvin, J., Thomas, T. A., Ravji, T. R., & Natarajaseenivasan, K. (2009). Optimization and production of a biosurfactant from the sponge-associated marine fungus Aspergillus ustus MSF3. Colloids and Surfaces B: Biointerfaces, 73(2), 250–256.
Kriti, S., & Subhash, C. (2014). Treatment of petroleum hydrocarbon polluted environment through bioremediation: a review. Pakistan Journal of Biological Sciences, 17, 1–8. https://doi.org/10.3923/pjbs.2014.1.8.
Lee, H., Yeong, Y., Seo, J., Seokyoon, K., Gyu-Hyeok, K., & Jae, J. (2015). Bioremediation of polycyclic aromatic hydrocarbons in creosote-contaminated soil by Peniophora incarnata KUC8836. Bioremediation Journal, 19, 1–8.
Li, Y., Liu, H., Tian, Z., Zhu, L., Wu, Y., & Tang, H. (2008). Diesel pollution biodegradation: synergetic effect of mycobacterium and filamentous fungi. Biomedical and Environmental Sciences, 21, 181–187.
Maletic, S., Dalmacija, B., Roncevic, S., 2013. Petroleum hydrocarbon biodegradability in soil—implications for bioremediation. INTECH, https://doi.org/10.5772/50108.
Manli, W., Warren, A. D., Wei, L., Xiaochang, W., Qian, Y., Tingting, W., Limei, M., & Liming, C. (2016). Bioaugmentation and biostimulation of hydrocarbon degradation and the microbial community in a petroleum-contaminated soil. International Biodeterioration & Biodegradation., 107, 158–164.
Marquez-Rocha, F. J., Olmos-Soto, J., & Rosano-Hernandez, M. C. (2005). Determination of the hydrocarbon-degrading metabolic capabilities of tropical bacterial isolates. International Biodeterioration and Biodegradation, 55, 17–23.
McGenity, T. J., Folwell, B. D., McKew, B. A., & Sanni, G. O. (2012). Marine crude-oil biodegradation: a central role for interspecies interactions. Aquatic Biosystems, 8(10), 10–1186.
Montagnolli, R. N., Lopes, P. R., & Bidoia, E. D. (2015). Screening the toxicity and biodegradability of petroleum hydrocarbons by a rapid colorimetric method. Archives of Environmental Contamination and Toxicology, 68, 342–353.
Naga R. M., Ricardo, B., Venkateswarlu, K., Andrea, R. C., Manjunatha, B (2016). Removal of petroleum hydrocarbons from crude oil in solid and slurry phase by mixed soil microorganisms isolated from Ecuadorian oil fields. International Biodeterioration & Biodegradation 108, 85–90. https://doi.org/10.1016/j.ibiod.2015.12.015.
Naga, R. M., Laura, S., & Kadiyala, V. (2017). Microbial degradation of total petroleum hydrocarbons in crude oil: a field-scale study at the low-land rainforest of Ecuador. Environmental Technology, 38(20), 2543–2550. https://doi.org/10.1080/09593330.2016.1270356.
Nilanjana, D., & Preethy, C. (2011). Microbial degradation of petroleum hydrocarbon contaminants: an overview. Biotechnology Research International, 941810, 1–13. https://doi.org/10.4061/2011/941810.
Olga, M., Ewa, K., Dorota, W., & Tadeusz, A. (2015). Biodegradation of diesel oil hydrocarbons enhanced with Mucor circinelloides enzyme preparation. International Biodeterioration & Biodegradation, 104, 142–148.
Onojake, M. C., Omokheyeke, O., & Osakwe, J. O. (2014). Petroleum hydrocarbon contamination of the environment: a case study. Bulletin of Earth Sciences of Thailand, 6(1), 67–79.
Oriomah, C., Olufemi, A., Olawale, A., & Abimbola, O. (2014). Bacteria from spent engine-oil-contaminated soils possess dual tolerance to hydrocarbon and heavy metals, and degrade spent oil in the presence of copper, lead, zinc and combinations thereof. Annales de Microbiologie, 65, 207–215.
Palanisamy, N., Ramya, J., Kumar, S., Vasanthi, N., Chandran, P., & Khan, S. (2014). Diesel biodegradation capacities of indigenous bacterial species isolated from diesel contaminated soil. Journal of Environmental Health Science and Engineering, 12, 142.
Passarini, M. R., Sette, L. D., & Rodrigues, M. V. (2011). Improved extraction method to evaluate the degradation of selected PAHs by marine fungi grown in fermentative medium. Journal of the Brazilian Chemical Society, 22(3), 564–570.
Rhodes, C. J. (2010). Seaweeds and their role in globally changing environments, cellular origin, life in extreme. Habitats and Astrobiology, 15, 229–248.
Saowakon, S., Thanaphorn, R., Tewan, Y., Poonsuk, P., & Kanokphorn, S. (2017). The production of biodiesel using residual oil from palm oil mill effluent and crude lipase from oil palm fruit as an alternative substrate and catalyst. Fuel, 195, 82–87.
Saravanan, R., & Sivakumar, T. (2013). Biodiversity and biodegradation potentials of fungi isolated from marine systems of East Coast of Tamil Nadu, India. International Journal of Current Microbiology and Applied Sciences, 2, 192–201.
Sayara, T., Sarra, M., & Sanchez, A. (2010). Effects of compost stability and contaminant concentration on the bioremediation of PAHs-contaminated soil through composting. Journal of Hazardous Materials, 179, 999–1006.
Sayara, T., Borras, E., Caminal, G., Sarra, M., & Sanchez, A. (2011). Bioremediation of PAHs-contaminated soil through composting: influence of bioaugmentation and biostimulation on contaminant biodegradation. International Biodeterioration and Biodegradation, 65, 859–865.
Shaieb, F. M., Elghazawani, A. H., & Issa, A. (2015). Studies on crude oil degrading bacteria isolated from Libyan desert. International Journal of Current Microbiology and Applied Sciences, 4(2), 920–927.
Sheppard, P. J., Adetutu, E. M., Makadia, T. H., & Ball, A. S. (2011). Microbial community and ecotoxicity analysis of bioremediated, weathered hydrocarbon contaminated soil. Soil Research, 49(3), 261–269.
Tahhan, R. A., Ammari, T. G., Goussous, S. J., & Al-Shdaifat, H. I. (2011). Enhancing the biodegradation of total petroleum hydrocarbons in oily sludge by a modified bioaugmentation strategy. International Biodeterioration and Biodegradation, 65, 130–134.
Toshio, M., Masashi, W., Hisato, T., Tasuku, K., & Ichiro, K. (2015). Degradation of chlorinated dioxins and polycyclic aromatic hydrocarbons (PAHs) and remediation of PAH-contaminated soil by the entomopathogenic fungus, Cordyceps militaris. Journal of Environmental Chemical Engineering, 3, 2317–2322.
Varjani, S. J. (2017). Microbial degradation of petroleum hydrocarbons. Bioresource Technology, 223, 277–286.
Weiwei, C., Junde, L., Xiangnan, S., Jun, M., & Xiaoke, H. (2017). High efficiency degradation of alkanes and crude oil by a salt-tolerant bacterium Dietzia species CN-3. International Biodeterioration & Biodegradation, 118, 110–118.
Yao, L., Teng, Y., Luo, Y., Christie, P., Ma, W., Liu, F., Wu, Y., Luo, Y., & Li, Z. (2015). Biodegradation of polycyclic aromatic hydrocarbons (PAHs) by Trichoderma reesei FS10-C and effect of bioaugmentation on an aged PAH-contaminated soil. Bioremediation Journal, 19, 9–17.
Zhao, D., Liu, C., Liu, L., Zhang, Y., Liu, Q., & Wu, W. M. (2011). Selection of functional consortium for crude oil-contaminated soil remediation. International Biodeterioration and Biodegradation, 65(8), 1244–1248.
Zuzanna, S., Paweł, C., Wojciech, J., Jakub, C., Justyna, S., & Agnieszka, P. (2015). Antibacterial effect of the Trichoderma viride fungi on soil microbiome during PAH’s biodegradation. International Biodeterioration & Biodegradation, 104, 170–177.
Acknowledgements
The authors sincerely acknowledge Environmental Research Centre, Cyprus International University’s (North Cyprus, Mersin 10, Turkey) facilities for the research.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Essabri, A.M.A., Aydinlik, N.P. & Williams, N.E. Bioaugmentation and Biostimulation of Total Petroleum Hydrocarbon Degradation in a Petroleum-contaminated Soil with Fungi Isolated from Olive Oil Effluent. Water Air Soil Pollut 230, 76 (2019). https://doi.org/10.1007/s11270-019-4127-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11270-019-4127-8