Abstract
Viral enteritis is a significant cause of death among dogs younger than 6 months. In this study, the presence of canine chaphamaparvovirus (CaChPV), canine bufavirus (CBuV), and canine adenovirus (CAdV) was investigated in 62 diarrheal dogs previously tested for other viral pathogens (canine parvovirus type 2, canine coronavirus, and canine circovirus). CBuV was detected in two dogs (3.22%) and CaChPV in one dog (1.61%). One dog tested positive for three parvoviruses (CPV-2b, CBuV, and CaChPV). All dogs tested negative to CAdV-1/CAdV-2. A long genome fragment of one of the two identified CBuVs and of the CaChPV was obtained and analyzed. New Turkish CBuVs had high identity rates (96%–98% nt; 97%–98% aa) with some Italian CBuV strains (CaBuV/9AS/2005/ITA and CaBuV/35/2016/ITA). The phylogenetic analysis powerfully demonstrated that these viruses belonged to a novel genotype (genotype 2). A part of the genome ChPV-TR-2021–19 revealed high identity rates (> 98% nt and > 99% aa) with some Canadian CaChPV strains (NWT-W88 and NWT-W171) and the Italian CaChPV strain Te/37OVUD/2019/IT. This study is the first report on the detection of CBuV-2 and the concomitant presence of three canine parvoviruses in Turkey. The obtained data will contribute to the molecular epidemiology and the role in the etiology of enteric disease of new parvoviruses.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Parvoviruses belonging to the Parvoviridae family are small (25–30 nm in diameter), non-enveloped, and single-stranded (ss) DNA viruses of 3.9–6.3 kb in length [1,2,3]. Parvoviridae family is divided into three subfamilies: Parvovirinae, which includes viruses that infect vertebrate hosts; Densovirinae, which encompass viruses that infect arthropod hosts; and Hamaparvovirinae, which infect both vertebrate and arthropod hosts [3,4,5]. Their genome is predicted to encode for 3 or 4 proteins: nonstructural (NS) 1, nucleoprotein (NP) 1, and viral proteins (VP) 1 and 2 [1, 2, 6].
The most well-studied parvovirus of dogs is canine parvovirus type 2 (CPV-2; species Carnivore protoparvovirus 1, genus Protoparvovirus, and subfamily Parvovirinae), which emerged as a canine pathogen at the end of the 1970s. The antigenic variants of the virus (CPV-2a, CPV-2b, and CPV-2c) have been found to cause even severe enteritis and leukopenia and to be associated with high mortality in puppies [7, 8].
In 2012, a novel protoparvovirus, later named Bufavirus (BuV), was discovered in the viral metagenomic analysis of the fecal samples of diarrheic children in Burkina Faso [9]. Since then, protoparvoviruses related to BuVs have been reported as sporadic cases in human and non-human mammals (pigs, dogs, bats, rats, shrews, and non-human primates) [10,11,12,13,14]. In 2018, a virus with a close genetic relationship to the human bufavirus (HuBuV) was detected in dogs with either gastroenteric or respiratory disease in Italy and Hungary and it was named canine bufavirus [15]. Accordingly, the virus was prevalent in dogs with respiratory signs and also a common component of the canine fecal virome [15]. To date, canine bufavirus (CBuV; species Carnivore protoparvovirus 3, genus Protoparvovirus, and subfamily Parvovirinae) was reported in Italy, Hungary, China, Canada, and India [13, 15,16,17,18,19]. However, since the first discovery of CBuV in 2018, the virus has not yet been reported in Turkey.
Chaphamaparvovirus (ChPV) was originally discovered in the viral metagenomic analysis of the oropharyngeal swab of a fruit bat (Eidolon helvum) in Ghana, Africa. Since then, ChPV-like viruses have been described in several additional animal species [3], including dogs and cats [20,21,22,23]. More recently, canine chaphamaparvovirus (CaChPV) has been identified in Canadian wolves [24]. CaChPV (species Carnivore chaphamaparvovirus 1, genus Chaphamaparvovirus, and subfamily Hamaparvovirinae) was identified by metagenomic analysis in a fecal sample during an unexplained outbreak of diarrhea in dogs in Colorado, USA, in 2019 [20]. To date, CaChPV has been reported in dogs in Italy, China, and Turkey, but no association was found between the virus and enteric signs [21, 23, 25]. Nonetheless, reports about detecting ChPVs in Turkey are still limited [25, 26].
Canine adenovirus (CAdV; species Canine mastadenovirus A, genus Mastadenovirus, and family Adenoviridae) is a non-enveloped icosahedral double-stranded DNA virus and is an important and well-established pathogen of dogs worldwide [27, 28]. CAdV comprises canine adenovirus type 1 (CAdV-1) and type 2 (CAdV-2) serotypes. Canine adenovirus type 1 (CAdV-1) is the causative agent of a systemic, potentially fatal, and highly contagious viral disease of domestic and wild canids. Acute disease is variably characterized by the presence of fever, tonsillitis, conjunctivitis, lethargy, vomiting, hematemesis, diarrhea, cough, tachypnea, and icterus [27]. Diarrhea may contain frank blood or melena. CAdV-2 causes infectious tracheobronchitis characterized by respiratory signs [29].
Reports and information on new parvoviruses are limited. Whether they cause infection in the host or a disease with clinical signs has not yet been fully established. The aim of this study was to detect CBuV, CaChPV, CAdV-1, and CAdV-2 in fecal samples from 62 diarrheal dogs from Turkey.
Materials and methods
Samples and DNA extraction
To detect ChCPV, CBuV, CAdV-1, and CAdV-2, fecal samples of 62 dogs of different ages (from 1.5 to 60 month) and breeds that showed signs of gastroenteritis were included in the study.
Fecal samples were from symptomatic dogs admitted to veterinary clinics in Ankara (n = 7), Elazig (n = 24), Kayseri (n = 27) and Kocaeli (n = 4) provinces, from November 2019 to January 2022.
The CPV-2b positivity (48/62) and canine circovirus (CCV) and canine coronavirus (CCoV) negativity (0/62) of these samples were confirmed in the previous epidemiological survey [30]. Template DNA was extracted from the supernatants of the fecal samples using a commercial extraction kit (MinElute Virus Spin Kit, QIAGEN, Hilden, Germany), following the manufacturer's instructions. The extracted DNA was eluted in 100 μl of elution buffer, quantified using a NanoDrop2000 spectrophotometer (Thermo Fisher Scientific, USA), and stored at − 20 °C until use.
Detection of viral agents
Viral agents were detected by polymerase chain reaction (PCR) screening using the commercial Taq PCR master mix (Hibrigen, Ist, Turkey) with the primer sets listed in Table 1. Each PCR mixture consisted of 5 μl template DNA, 2 μl primers (20 μM each primer), 25 μl PCR master mix, and ddH2O to final volume of 50 μl. All PCR assays included positive and negative reaction controls. CBuV and CaChPV DNA detected in nasal samples from two dogs with respiratory tract disease in a shelter in Elazig province were used as positive controls. Negative controls consisted of only ultrapure water. For CBuV detection, PCR protocols and thermal conditions were applied as described in previous studies with minor modifications [21, 24, 31,32,33,34]. A primer pair named BuV-Screening (5ʹ-CACTATAAATGACTCTTACCATGCA-3ʹ and 5ʹAGTTCGATTGGTACACAGTTTTC-3ʹ) targeting part of the VP2 gene (480 bp) was preferred to detect BuVs [31]. To obtain the complete genome (4.2 kb), a random CBuV strain was selected and PCR-amplified overlapping partial fragments of this strain were generated using the primer sets (CBuV-1F and 2R, CBuV-2F and 2R, CBuV-5F and 5R, CBuV-6F and 6R) previously described [32]. An alternative primer pair (CBuV-3F and 3R) from different studies were included here, as some primer sets cannot amplify target regions [33]. However, because a new primer pair is needed for a single target region, a new primer set (BuV-4F and 4R) was designed from the conserved regions using the primer-BLAST tool [35].
The presence of CaChPV was assessed by using a nested PCR assay with diagnostic primer sets (CaChPV-Screening F: 5ʹ-GCARCGAGAGCTGCGAATG-3ʹ, R: 5ʹ -TGYTTCCCAYTCTATCTCGTC-3ʹ, and nested: 5ʹ -TGAYGCTAGATCCTGAGCTG-3ʹ) following the PCR protocol and thermal conditions previously described [24]. The PCR-positive sample was selected for amplifying the complete NS1 gene of CaChPV. Previously described primer sets (CaChPV-1F and 1R, CaChPV-2F and 2R, CaChPV-3F and 3R) and PCR conditions were used to amplify the 2.5-kb-long CaChPV containing the putative NS gene, NS1, and partial VP1 gene, with overlapping fragments [21].
Previously designed primers (CAdV-Screening F and R) targeting the E3 gene and flanking regions were used to detect and differentiate CAdV-1 and CAdV-2 in all clinical samples of DNA [34]. The DNA extracted from a CAdV-2-attenuated vaccine strain (Bioveta, Czech Republic) was used as the positive control.
Sequencing and phylogenetic analysis
All amplicons were separated by electrophoresing in 1.5% (w/v) agarose gel stained with ethidium bromide in 1X standard tris–acetate-EDTA (TAE) buffer. A 100-bp DNA ladder (NEB, USA) was used in each gel to check for the presence of amplicons of the expected size. Amplified products were purified by GeneJET Gel Extraction Kit (Thermo Fisher, MA, USA). Purified products were sequenced bidirectionally with the ABI Prism 3130 genetic analyzer (Applied Biosystems, USA) using the BigDye Terminator v3.1 cycle sequencing kit (Applied Biosystems, USA) and Sanger method by Macrogen company (The Netherlands). The viral genome was assembled by the sequential splicing of overlapping sequences. Sequence alignments were performed using the CLUSTALW algorithm in MEGA X software [36]. Aligned sequences were controlled in the BLASTn program and were submitted to the GenBank database by editing. Genetic distances by the p-distance method were calculated in Mega X software [36]. Phylogenies were performed by the maximum likelihood (ML) method using the MEGA X software [36] and constructed using the Tamura-Nei model, gamma-distributed variation among sites, and 1000 bootstrap replicates [37].
Results
Detection of viral agents
Of the 62 samples screened by PCR, two tested positive for CBuV (3.22%), and one resulted positive for CaChPV (1.61%). One of those positive samples tested positive for CaChPV, had a concomitant of CaChPV, CBuV, and CPV-2b. The other one CBuV positive sample tested negative for other viruses. The Kangal shepherd dog with three parvoviruses was 2 months old, female, and had never been vaccinated. It also showed signs of bloody diarrhea. The other dog with CBuV, a Golden retriever, was three months old, male, and vaccinated with two doses of a core vaccine consisting of canine parainfluenza virus, canine distemper virus, CPV-2, CAdV-2, and leptospira canicola. It also showed signs of mild yellowish diarrhea. Both dogs were collected in Kayseri province in Turkey, and clinical samples were from 2021. All PCR amplicons were confirmed by sequencing. No amplification was observed from the negative controls.
CAdV-1 and CAdV-2 positivity were not detected in any of the 62 clinical samples. Positive controls had expected size amplicons at the PCR screening.
Sequence and phylogenetic analysis of CBuV
After sequencing of the 480-bp PCR products, two CBuVs, TR/27/CBuV/2021 (OP879719) and TR/19/CBuV/2021 (OP879720), were identified, showing 85%–98% nt identity rates with the CBuV strains in GenBank. Two CBuVs, TR/27/CBuV/2021 (OP879719) and TR/19/CBuV/2021 (OP879720), were identified by sequencing the VP2 gene. One of the two CBuV DNAs was selected as a template in the PCR steps to obtain the complete genome. A 3735-nt long fragment of genome of the strain TR/27/CBuV/2021 was obtained (encompassing a 1430 nt-475 aa of the partial NS region, a 2136 nt-711 aa of the entire VP1 region, and 1707 nt-568 aa of the entire VP2 region) and used for further genetic analysis. TR/27/CBuV/2021 shared 55%–98% identity with the selected 34 BuVs from GenBank. TR/27/CBuV/2021 displayed the closest identity with some Italian strains (CaBuV/9AS/2005/ITA and CaBuV/35/2016/ITA): 97%–98% for the 3735-nt genome, 96%–97% (97%–98% aa) for partial NS, 97%–98% nt (98% aa) for VP1, and 98%–99% nt (98% aa) for VP2.
Similar to TR/27/CBuV/2021, the strain TR/19/CBuV/2021 was highly identical (96%–98% nt, 97% aa) with CaBuV/9AS/2005/ITA and CaBuV/35/2016/ITA.
On the basis of the phylogenetic tree including 34 BuVs (Fig. 1), CBuVs were included in the same branch in two separate clades (genotype 1 and 2) with a bootstrap value of 100% and sharing > 96 nt identities. TR/27/CBuV/2021, CaBuV/9AS/2005/ITA, and CaBuV/35/2016/ITA clustered in genotype 2, which is a different subgroup from the other 32 CBuVs (genotype 1) in Fig. 1. The CBuV branch was closely related to the sea otter BuV, followed by Eulipotyphla protoparvovirus 1 and porcine BuVs. It was distantly related to WUHARV parvovirus 1.
Phylogenetic tree based on the aligned 3735-nt long sequences of TR/27/CBuV/2021 (OP879719) and other 34 BuVs. The different host species are numbered one through seven in the figure. According to numerics, these were species of dog, sea otter, crocidura hirta, swine, bat, rhesus monkey, and human, respectively. Strain TR/27/CBuV/2021 obtained from this study was shown by a filled circle on the phylogenetic tree
The phylogenetic tree based on the VP2 gene (Fig. 2) of TR/19/CBuV/2021, TR/27/CBuV/2021, and other 83 selected viruses was consistent with branch and subgroups (%99 bootstrap value, > %98 nt identities) in Fig. 1. In addition to some Italian strains (CaBuV/9AS/2005/ITA and CaBuV/35/2016/ITA), the Canadian strain LSD41 with a shorter sequence (195 nt, partial VP2 gene) shared a high identity (> 98%) with Turkish BuVs and then clustered in the genotype 2 clade. Table S1 shows the identity ratios between the VP1 and VP2 genes of TR/27/CBuV/2021 and other CBuVs.
Phylogenetic tree based on the VP2 gene of TR/19/CBuV/2021, TR/27/CBuV/2021 and other selected 83 viruses. Strains TR/27/CBuV/2021 and TR/19/CBuV/2021 (OP879719-20) obtained from this study were shown by a filled circle on the phylogenetic tree. The black star marks Chinese (AH-001, AH-002, AH-003, D-9P-37, CBuV-88, Henan-38, Henan44, GXNN01-2018, and GXNN02-2018), Canadian (NWT-W25, NWT-W34, NWT-W116, NWT-W233, NWT-W236), and Indian (CaBuV/2020/PVNRTVU) strains
Sequence and phylogenetic analysis of CaChPV
The first- and second-round PCR products targeting for the NS1 gene of CaChPV (459 and 267 bp, respectively) were confirmed by sequencing. The partial NS1 gene sequence of the identified strain ChPV-TR-2021-19 showed high nt (> 99%) and aa (99%–100%, respectively) identity with Canadian strains NWT-W85 and NWT-W89. As a result of the PCR assay for CaChPV, a long genome fragment (2544-nt long) containing the entire NS1 gene (1992 nt; 663 aa) of the Turkish CaChPV strain (ChPV-TR-2021-19, accession number: OP879721) was obtained. Comparing the ChPV-TR-2021-19 strain and other 38 ChPV sequences from different host species origins, the NS1 gene shared 75%–100% identity. ChPV-TR-2021-19 was found closest to canine ChPVs with 94%–100% nt identity. Similarly, ChPV-TR-2021-19 had high identity ratios (> 98% nt and > 99% aa) with some Canadian (NWT-W88 and NWT-W171) and Italian canine ChPV strains sequences (Te/37OVUD/2019/ITA).
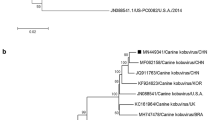
In the phylogenetic tree based on the complete NS1 nt sequence (Fig. 3), ChPV-TR-2021-19 was in the same branch as the canine ChPVs. ChPV-TR-2021-19 was in the same phylogroup as some Canadian (NWT-W88, NWT-W171, NWT-W85, and NWT-W89) and Italian ChPVs (Te/36OVUD/2019/ITA and Te/37OVUD/2019/ITA).
Amino acid sequence and substitutions
The ATP- or GTP-binding Walker loop motif GXXXXGK[T/S] (-GPASTGKS-) and a replication initiator motif (IVRYFLTKQP) were found in the partial NS1 of TR/27/CBuV/2021 [15]. VP1 gene contained several aa motifs that are highly conserved among the family Parvovirinae, including the Ca2 + binding loop (YLGPGN) and the catalytic center (HDLEY) of the phospholipase A2 (PLA2) motif. The N terminus of the TR/27/CBuV/2021 VP2 protein found glycine-rich sequence (-GGGGGGGSGVG-) [15]. An amino acid insertion (P; proline) was found between VP1 and VP2 proteins of the TR/27/CBuV/2021. This insertion was just after position 142 of VP1 and was just before VP2 starting codon (ATG; methionine). It was detected 56 aa subsitutions on VP2 of selected CBuVs in this study; 16 were common to CBuV − 2 s.
Substitutions on the VP2 proteins of TR/27/CBuV/2021, TR/27/CBuV/2021, CaBuV/35/2016/ITA, and CaBuV/9AS/2005/ITA are shown in Table 2. These were at amino acid positions 3, 6, 7, 9, 10, 14, 23, 34, 86, 113, 210, 220, 250, 267, 303, 366, and 519.
The NS protein aa sequence of ChPV-TR-2021-19 was compared with those of NWT-W88, NWT-W171, Te/37OVUD/2019/ITA, and Te/36OVUD/2019/ITA and had the original substitutions at positions 125 (K to R) and 599 (Y to C). Other substitutions were observed at amino acid positions 62 (R to P), 198 (H to R), 539 (I to V), and 581 (D to N).
Discussion
The advancement and significant development of sequencing technologies and metagenomics increase the possibility of identifying new infectious agents as the cause of disease. These technological advances reveal not only possible new causes of diseases but also provide essential benefits regarding the study of co-infections [38, 39].
In this study, the presence of CBuV, CaChPV, CAdV-1, and CAdV-2 in the fecal samples of dogs with parvoviral enteritis and dogs with similar signs but unknown etiology was investigated. We detected CBuV in two (3.22%) of 62 dogs. The low prevalence (1.74%–4.3%) of CBuV in the fecal samples of symptomatic dogs was consistent with previous reports in China and India [13, 17, 19, 33]. These rates also match with that of human bufavirus (HuBuV, 0.3%–4.1%) [40]. In China, the positivity rates of CBuV in Guangxi, Henan, and Anhui provinces ranged from 1.74% to 2.5% [13, 16, 17, 33]. In a study in India, the positivity rate of CBuV was 4.3% [19]. Contrary to our results, some reports in Italy, Hungary, and China had a relatively higher prevalence of CBuV, ranging from 7.7% to 42.5% [15, 16, 32].
Based on the phylogenetic tree of 34 BuVs (Fig. 1), CBuVs were included in the same branch. Within this phylogroup, two clear subgroups were defined (bootstrap value, 100%), supporting the evidence for two distinct CuBV genotypes. Until now, only one genotype of CBuVs was mainly reported. Genotype 1 included the strains of all CBuVs from Italy (except for a few), Hungary, China, Canada, and India. In contrast, genotype 2 includes the new Turkish strain TR/27/CBuV/2021 and the Italian strains 9AS/2005/ITA and 35/2016/ITA. These results were supported by the phylogenetic analysis based on the VP2 gene and that of the previous study [32]. In Humans, based on VP1 and VP2 sequences, three genotypes, HBuV-1, HBuV-2, and HBuV-3, have been determined; different genotypes of this virus show ıt had the highly diverse capsid gene [40]. VP2 of the carnivore protoparvoviruses is a significant capsid protein vital in determining antigenicity, host range, and tissue tropism [8, 30]. When the VP2 protein of the selected canine Bufaviruses in this study was examined, more than 50 substitutions were detected, and 16 were specific for CBuV-2. While parvoviruses are DNA viruses, the high mutation rate of the VP2 gene can increase the genetic and antigenic diversity of the virus [7]. Accordingly, additional researches are necessary to elucidate whether the VP2 aa changes may affect some biological properties of CBuVs. The CBuV-2 can have predictable differences in antigenic and other biological properties [32]. We found that TR/27/CBuV/2021 had a 3-nucleotide (-CCA-) insertion at nucleotide position 427–429 of VP1, unlike other CBuVs [13, 15,16,17,18,19]. Therefore, the VP1 protein of TR/27/CBuV/2021 contained 711 aa. The phenotypic effect of this change and whether it is predominant should be revealed with further research.
Reports of the prevalence of CaChPV are limited. Existing reports are from the United States, Italy, China, and Turkey [20, 21, 23, 25]. To date, the prevalence rate of CaChPV has ranged from 1.5% to 4.3% in diarrheic dogs and from 0% to 1.6% in healthy dogs. Palombieri et al. [23] detected CaChPV in 3 of 155 diarrheal dogs (1.9%) in Italy. In the same study, CaChPV DNA was found in co-infection with CPV-2 (3/3), CBuV (1/3), and CCoV (1/3). In China, Hu et al. [21] detected CaChPV in 5 rectal swabs out of 323 (1.55%) dogs with diarrhea, of which two had co-infection with CPV-2 and CDV or CCoV. Dinçer et al. [25] detected CaChPV DNA in 2.9% of dogs with and without diarrhea in Turkey. In these reports, no significant association was found with enteric disease. In our study, we found a 1.61% prevalence rate, and the low prevalence of CaChPV in diarrheic dogs was consistent with those in other reports [20, 21, 23, 25]. The complete NS1 gene of the strain ChPV-TR-2021–19 had high identity rates (> 98% nt and > 99% aa) with some Canadian strains (NWT-W88, NWT-W171) and the Italian strain Te/37OVUD/2019/ITA. The phylogenetic analyses based on the NS1 revealed that CaChPVs were grouped in the same phylogroup but in a different branch from the feline ChPVs. The NS1-based phylogenetic tree supported species discrimination.
According to our results, in one of CBuV and CaChPV positive dog, CPV-2b was also detected. While the striking finding in the other studies on CaChPV and CBuV prevalence was that the predominant virus in co-infections was CPV-2, here, we determined lower (1/48) [13, 19, 2332, 33]. Co-infections may cause severe gastroenteric signs in dogs [41]. In this study, enteric disease signs in the dog showing co-infections with three viral species were more severe than only BuV-positive dog, but this difference may originate from CPV-2 alone. Therefore, we have limited possibilities to assess whether CaChPV and CBuV in synergy with CPV-2b would cause more severe gastroenteric signs. Indeed, it is challenging to make this assessment without demonstrating the negativity of other enteric viruses or bacterial agents. Further pathogenic studies will help clarify the role of co-infection in the severity of gastroenteritis in dogs or the enteric part of these viruses.
In some countries, although modified live CAdV vaccines have significantly reduced the incidence of CAdV and rendered it negligible in veterinary practice, there have been recent remarkable reports on the epidemiology of these viruses in dogs. Balboni et al. [42] found an alarmingly high prevalence of CAdV-2 (27/51, 52.9%) in the feces of dogs with different clinical manifestations (gastrointestinal, dermatologic, musculoskeletal, etc.) in Italy, raising the question of whether this virus is an opportunistic agent or pathogen of the gastrointestinal tract. More recently, Balboni et al. [43] investigated CCV, CAdV-1, and CAdV-2 in fecal or intestine samples of dogs with parvoviral enteritis in Italy. They predominantly detected CAdV-2 in 19 of 28/95 CPV-2-infected dogs, while they detected CAdV-1 (2/28) and CCV (8/28) in the other dogs. Contrary to these reports, we did not find CAdV-1 and CAdV-2 in the dog feces examined. We can infer that this risk is low in the studied regions. However, more large-scale studies are needed to provide a complete picture.
In conclusion, knowledge about new parvoviruses is still limited. Herein, we report for the first time the detection and comprehensive characterization of CBuV in Turkey. All CBuV strains in this study belonged to a novel CBuV-2 genotype. Experimental attempts will reveal the role of new-type viruses, alone or with multiple pathogens in intestinal or extraintestinal organs. We think new genetic data are of great value for research on population dynamics and virus evolution. The epidemiological data must be increased, and the pathogenic roles of both CBuV and CaChPV in the intestinal or extraintestinal systems still require elucidation.
Data availability
All data generated or analysed during this study are included in this published article. The data that support the findings of this study are available from the corresponding author upon reasonable request.
Code availability
Not applicable.
References
Cotmore SF, Agbandje-McKenna M, Chiorini JA, Mukha DV, Pintel DJ, Qiu J, Soderlund-Venermo M, Tattersall P, Tijssen P, Gatherer D, Davison AJ (2014) The family Parvoviridae. Arch Virol 159:1239–1247. https://doi.org/10.1007/s00705-013-1914-1
Cotmore SF, Agbandje-McKenna M, Canuti M et al (2019) ICTV Virus taxonomy profile: Parvoviridae. J Gen Virol 100:367–368. https://doi.org/10.1099/jgv.0.001212
Pénzes JJ, de Souza WM, Agbandje-McKenna M, Gifford RJ (2019) An ancient lineage of highly divergent parvoviruses infects both vertebrate and invertebrate hosts. Viruses 11:525. https://doi.org/10.3390/v11060525
Souza WMD, Romeiro MF, Fumagalli MJ, Modha S, de Araujo J, Queiroz LH, Durigon EL, Figueiredo LTM, Murcia PR, Gifford RJ (2017) Chapparvoviruses occur in at least three vertebrate classes and have a broad biogeographic distribution. J Gen Virol 98:225–229. https://doi.org/10.1099/jgv.0.000671
Pénzes JJ, Söderlund-Venermo M, Canuti M, Eis-Hübinger AM, Hughes J, Cotmore SF, Harrach B (2020) Reorganizing the family Parvoviridae: a revised taxonomy independent of the canonical approach based on host association. Arch Virol 165:2133–2146. https://doi.org/10.1007/s00705-020-04632-4
Jager MC, Tomlinson JE, Lopez-Astacio RA, Parrish CR, Van de Walle GR (2021) Small but mighty: old and new parvoviruses of veterinary significance. Virol J 18:210. https://doi.org/10.1186/s12985-021-01677-y
Decaro N, Buonavoglia C (2012) Canine parvovirus—a review of epidemiological and diagnostic aspects, with emphasis on type 2c. Vet Microbiol 155:1–12. https://doi.org/10.1016/j.vetmic.2011.09.007
Miranda C, Thompson G (2016) Canine parvovirus: the worldwide occurrence of antigenic variants. J Gen Virol 97:2043–2057. https://doi.org/10.1099/jgv.0.000540
Phan TG, Vo NP, Bonkoungou IJ, Kapoor A, Barro N, O’Ryan M, Kapusinszky B, Wang C, Delwart E (2012) Acute diarrhea in West African children: diverse enteric viruses and a novel parvovirus genus. J Virol 86:11024–11030. https://doi.org/10.1128/JVI.01427-12
Kemenesi G, Dallos B, Gorfol T, Estok P, Boldogh S, Kurucz K, Oldal M, Marton S, Banyai K, Jakab F (2015) Genetic diversity and recombination within bufaviruses: detection of a novel strain in Hungarian bats. Infect Genet Evol 33:288–292
Sasaki M, Orba Y, Anindita PD, Ishii A, Ueno K, Hang’Ombe BM, Mweene Ito AS, Sawa H (2015) Distinct lineages of bufavirus in wild shrews and nonhuman primates. Emerg Infect Dis 21:1230–1233
Yang S, Liu D, Wang Y, Qu F, He Y, Sun Z, Shen Q, Li W, Fu X, Deng X, Zhang W, Delwart E (2016) Bufavirus Protoparvovirus in feces of wild rats in China. Virus Genes 52:130–133. https://doi.org/10.1007/s11262-015-1262-1
Sun W, Zhang S, Huang H, Wang W, Cao L, Zheng M, Yin Y, Zhang H, Lu H, Jin N (2019) First identification of a novel parvovirus distantly related to human bufavirus from diarrheal dogs in China. Virus Res 265:127–131. https://doi.org/10.1016/j.virusres.2019.03.020
Sun Y, Chen Y, Cai Y, Zhu D, Pan H, Wei Y, Han X, Ji C, Lu G, Wang H et al (2020) First report and genetic diversity of porcine bufavirus in China. Virol J. https://doi.org/10.1186/s12985-019-1278-6
Martella V, Lanave G, Mihalov-Kovács E, Marton S, Varga-Kugler R, Kaszab E, Di Martino B, Camero M, Decaro N, Buonavoglia C, Bányai K (2018) Novel parvovirus related to primate bufaviruses in dogs. Emerg Infect Dis 24:1061–1068. https://doi.org/10.3201/eid2406.171965
Li J, Cui L, Deng X, Yu X, Zhang Z, Yang Z, Delwart E, Zhang W, Hua X (2019) Canine bufavirus in faeces and plasma of dogs with diarrhoea. China Emerg Microbes Infect 8:245–247. https://doi.org/10.1080/22221751.2018.1563457
Shao R, Zheng F, Cai S, Ji J, Ren Z, Zhao J, Wu L, Ou J, Lu G, Li S (2020) Genomic sequencing and characterization of a novel group of canine bufaviruses from Henan province, China. Arch Virol 165:2699–2702. https://doi.org/10.1007/s00705-020-04785-2
Canuti M, Mira F, Sorensen RG, Rodrigues B, Bouchard É, Walzthoni N, Hopson M, Gilroy C, Whitney HG, Lang AS (2022) Distribution and diversity of dog parvoviruses in wild, free-roaming and domestic canids of Newfoundland and Labrador, Canada. Transbound Emerg Dis 69:e2694–e2705. https://doi.org/10.1111/tbed.14620
Ganji VK, Buddala B, Yella NR, Putty K (2022) First report of canine bufavirus in India. Arch Virol 167:1145–1149. https://doi.org/10.1007/s00705-022-05398-7
Fahsbender E, Altan E, Seguin MA, Young P, Estrada M, Leutenegger C, Delwart E (2019) Chapparvovirus DNA Found in 4% of dogs with diarrhea. Viruses 11:398. https://doi.org/10.3390/v11050398
Hu W, Liu Q, Chen Q, Ji J (2020) Molecular characterization of Cachavirus firstly detected in dogs in China. Infect Genet Evol 85:104529. https://doi.org/10.1016/j.meegid.2020.104529
Ji J, Hu W, Liu Q, Zuo K, Zhi G, Xu X, Kan Y, Yao L, Xie Q (2020) Genetic analysis of Cachavirus-related parvoviruses detected in pet cats: the first report From China. Front Vet Sci 7:580836. https://doi.org/10.3389/fvets.2020.580836
Palombieri A, Di Profio F, Lanave G, Capozza P, Marsilio F, Martella V, Di Martino B (2020) Molecular detection and characterization of Carnivore chaphamaparvovirus 1 in dogs. Vet Microbiol 251:108878. https://doi.org/10.1016/j.vetmic.2020.108878
Canuti M, Fry K, Cluff HD, Mira F, Fenton H, Lang AS (2022) Co-circulation of five species of dog parvoviruses and canine adenovirus type 1 among gray wolves (Canis lupus) in northern Canada. Transbound Emerg Dis 69:e1417–e1433. https://doi.org/10.1111/tbed.14474
Dinçer E, Timurkan MÖ, Dinçer PFP, Aydın H (2020) Co-circulation of canine chaphamaparvovirus and canine parvovirus 2 in dogs with diarrhea in Turkey. Thai j vet med 50:495–501
Abayli H, Can-Sahna K (2022) First detection of feline bocaparvovirus 2 and feline chaphamaparvovirus in healthy cats in Turkey. Vet Res Commun 46:127–136. https://doi.org/10.1007/s11259-021-09836-w
Decaro N, Martella V, Buonavoglia C (2008) Canine Adenoviruses and Herpesvirus. Vet Clin North Am Small Anim Pract 38:799–814. https://doi.org/10.1016/j.cvsm.2008.02.006
Adams MJ, Lefkowitz EJ, King AM et al (2016) Ratification vote on taxonomic proposals to the International Committee on Taxonomy of Viruses. Arch Virol 161:2921–2949. https://doi.org/10.1007/s00705-016-2977-6
Sykes JE (2013) Canine and feline infectious diseases. In: Sykes JE (ed) Infectious Canine Hepatitis, 1st edn. Saunders, Philadelphia, pp 182–186
Abayli H, Aslan O, Tumer KC, Can-Sahna K, Tonbak S (2022) Predominance and first complete genomic characterization of canine parvovirus 2b in Turkey. Arch Virol 167:1831–1840. https://doi.org/10.1007/s00705-022-05509-4
Shao R, Ye C, Zhang Y, Sun X, Cheng J, Zheng F, Cai S, Ji J, Ren Z, Zhong L, Lai Z, Ou J, Lu G, Li S (2021) Novel parvovirus in cats China. Virus Res 304:198529. https://doi.org/10.1016/j.virusres.2021.198529
Di Martino B, Sarchese V, Di Profio F, Palombieri A, Melegari I, Fruci P, Aste G, Bányai K, Fulvio M, Martella V (2021) Genetic heterogeneity of canine bufaviruses. Transbound Emerg Dis 68:802–812. https://doi.org/10.1111/tbed.13746
Wang Y, Guo X, Zhang D, Sun J, Li W, Fu Z, Liu G, Li Y, Jiang S (2020) Genetic and phylogenetic analysis of canine bufavirus from Anhui Province Eastern China. Infect Genet Evol 86:104600. https://doi.org/10.1016/j.meegid.2020.104600
Hu RL, Huang G, Qiu W, Zhong ZH, Xia XZ, Yin Z (2001) Detection and differentiation of CAV-1 and CAV-2 by polymerase chain reaction. Vet Res Commun 25:77–84. https://doi.org/10.1023/a:1006417203856
Ye J, Coulouris G, Zaretskaya I, Cutcutache I, Rozen S, Madden T (2012) Primer-BLAST: a tool to design target-specific primers for polymerase chain reaction. BMC Bioinform 13:134
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547–1549. https://doi.org/10.1093/molbev/msy096
Tamura K, Nei M (1993) Estimation of the number of nucleotide substitutions in the control region of mitochondrial DNA in humans and chimpanzees. Mol Biol Evol 10:512–526. https://doi.org/10.1093/oxfordjournals.molbev.a040023
Visser M, Bester R, Burger JT, Maree HJ (2016) Next-generation sequencing for virus detection: covering all the bases. Virol J 13:85. https://doi.org/10.1186/s12985-016-0539-x
Mohsin H, Asif A, Fatima M, Rehman Y (2021) Potential role of viral metagenomics as a surveillance tool for the early detection of emerging novel pathogens. Arch Microbiol 203:865–872. https://doi.org/10.1007/s00203-020-02105-5
Väisänen E, Fu Y, Hedman K, Söderlund-Venermo M (2017) Human Protoparvoviruses. Viruses 9:354. https://doi.org/10.3390/v9110354
Dowgier G, Lorusso E, Decaro N, Desario C, Mari V, Lucente MS, Lanave G, Buonavoglia C, Elia G (2017) A molecular survey for selected viral enteropathogens revealed a limited role of Canine circovirus in the development of canine acute gastroenteritis. Vet Microbiol 204:54–58. https://doi.org/10.1016/j.vetmic.2017.04.007
Balboni A, Mollace C, Giunti M, Dondi F, Prosperi S, Battilani M (2014) Investigation of the presence of canine adenovirus (CAdV) in owned dogs in Northern Italy. Res Vet Sci 97:631–636. https://doi.org/10.1016/j.rvsc.2014.10.010
Balboni A, Terrusi A, Urbani L, Troia R, Stefanelli SAM, Giunti M, Battilani M (2022) Canine circovirus and Canine adenovirus type 1 and 2 in dogs with parvoviral enteritis. Vet Res Commun 46:223–232. https://doi.org/10.1007/s11259-021-09850-y
Funding
This study has not been carried out with the funds of any institution or organization.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study’s conception and design. Material preparation and data collection were performed by HA, OA, and KÇT. Analysis was performed by HA, ST, and KC-S. The first draft of the manuscript was written by HA, and all authors commented on previous versions. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Ethical approval
No attempt was made to adversely affect animal health or disrupt the tissue integrity of the dogs selected in this study. All procedures performed in animal studies complied with the institution (Fırat University Animal Experiments Local Ethics Committee) and international ethical standards.
Informed consent
All the authors consent to publish. The human participants and their data are not included in this article. The consent of animal owners was obtained at the time of sampling.
Consent to participations
All the authors consented to participate in this study.
Consent for publications
All the authors consent to the publication of this article.
Additional information
Edited by Nicola Decaro.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Abayli, H., Aslan, O., Tumer, K.C. et al. Investigation of canine chaphamaparvovirus, canine bufavirus, and canine adenovirus in dogs with diarrhea: First report of novel canine bufavirus in Turkey. Virus Genes 59, 427–436 (2023). https://doi.org/10.1007/s11262-023-01982-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11262-023-01982-4