Abstract
Dengue is an endemic arboviral disease with continuous transmission in Indonesia for more than five decades. A recent outbreak in Jember, East Java province, demonstrated the predominance of DENV-4, a serotype known for its low global spread and limited transmission. While epidemiological factors such as new serotype introduction and lacking herd immunity may explain its predominance, viral factors may also contribute. Using next-generation sequencing, we generated 13 representative complete genomes of DENV-4 responsible for the outbreak. Phylogenetic and evolutionary analyses on complete genomes were performed to understand the spatial and temporal dynamics of the viruses. Further analyses were done to study amino acid variations in DENV genes, as well as the potential events of recombination and selection pressure within the genomes. We revealed the DENV-4 genetic factors that may lead to its predominance in the 2019 Jember dengue outbreak. A combination of selection pressure and mutational genetic changes may contribute to the DENV-4 predominance in East Java, Indonesia. The possible intra-serotype recombination events involving the non-structural protein 5 (NS5) gene were also observed. Altogether, these genetic factors may act as additional factors behind the complex dengue outbreak mechanism.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Dengue is an arboviral disease with approximately 40% of the global population at risk of developing the disease, especially in the tropical and sub-tropical regions of the world [1]. Dengue virus (DENV) has four genetically distinct serotypes (DENV-1, -2, -3, and -4). Although DENV infection can be asymptomatic in the majority of cases, dengue may manifest as mild dengue fever (DF, with or without warning signs) or severe dengue hemorrhagic fever (DHF) and dengue shock syndrome (DSS) [2, 3].
The genome of DENV consists of single-stranded positive-sense RNA encoding three structural proteins: C, prM/M, and E, and seven non-structural (NS) proteins: NS1, NS2A, NS2B, NS3, NS4A, NS4B, and NS5 [4]. Genomic information of DENV is important in reconstructing the spatial and temporal history of dengue outbreaks/epidemics and to supplement the epidemiological data [5, 6]. Virus evolution has been identified as a significant force that shapes the genetic structure of virus populations and drives epidemiologic changes [7]. Analysis of the complete sequences of viral genomes is an important tool for virus evolution studies, as well as for surveillance and dengue epidemic preparedness [6,7,8].
Indonesia has been affected by all four DENV serotypes for more than five decades [9]. The predominance of DENV serotypes is not constant, with documented reports of predominance switch between serotypes and evidence of genotype switching within each serotype [10,11,12]. A recent 2019 dengue outbreak in Jember, a regency in East Java, Indonesia, recorded the predominance of DENV-4, which is rarely seen during dengue outbreaks in Indonesia [13].
The emergence of DENV-4 dates back to 1958, when it was first reported in the Philippines and Thailand, and continued to spread in Sri Lanka and the Indochina region, including Indonesia, and to the Americas in early 1980s [14]. Despite being the first DENV serotype to diverge in flavivirus group phylogenetic analysis, DENV-4 has always been considered as serotype with the least global spread, including in Indonesia, also marked by the smallest number of viral genome sequences available in public databases [15]. Moreover, the transmission and circulation of DENV-4 seem to be relatively limited, ultimately due to faltering competition with other DENV serotypes [16].
Although having the lowest prevalence, DENV-4 still has the potential to cause severe dengue manifestation (DHF), as reported in a previous study [17]. The dengue outbreak in Jember manifested as milder (DF) disease, with only 26.7% of patients showing DHF manifestation [13]. Nevertheless, the predominance of DENV-4 during the outbreak raised awareness and scientific questions on whether there are virus factors that contributed to this phenomenon.
This study aimed to understand the spatial and temporal evolutionary patterns of DENV-4 using complete genome sequences generated from the Jember 2019 dengue outbreak. We performed analyses of genetic factors that may contribute to DENV-4 serotype predominance. The information gained from establishing the DENV evolution is important to increase our understanding of the basic mechanisms of evolutionary change, and may also assist in designing control, treatment, and eradication strategies, as well as potentially predicting any future emergence [7]. The information gained from this study will be beneficial in preparing for future outbreaks/epidemics in the region.
Materials and methods
Sample collection and preparation
Serum samples were collected during a DENV surveillance study in Jember, East Java, from May 2019 to March 2020, which revealed the predominance of DENV-4 [13]. The use of human serum samples in the study was granted ethical approval from the Research Ethics Committee of the Faculty of Medicine Universitas Airlangga, Surabaya, Indonesia (Ethical Approval No. 20/EC/KEPK/FKUA), and written informed consent was obtained from all patients and/or children’s legal guardians [13]. Virus RNA was extracted from serum sample using QIAamp Viral RNA Mini kit (Qiagen, Hilden, Germany) and proceeded to simultaneous DENV detection and serotyping using abTES DEN/CHIKU 5 qRT-PCR (AIT Biotech, Singapore) qRT-PCR [13]. A total of 13 DENV-4 RNAs were then used in next-generation sequencing (NGS) library preparation.
NGS library preparation
Viral RNAs were prepared into an NGS library using a multiplex PCR method and Illumina sequencing by synthesis approach [18, 19]. Briefly, RNAs were reverse-transcribed into single-stranded cDNA using random hexamer (Thermo Fisher Scientific, Carlsbad, CA) and Superscript III Reverse Transcriptase (Thermo Fisher Scientific), according to the manufacturers’ instructions. The cDNA was subsequently processed to second-strand synthesis step using Large Klenow Fragment (New England Biolabs, Ipswich, MA) and then used in two multiplex PCR reactions using primer set for DENV-4, according to a protocol described previously [19]. The multiplex PCR amplification generated amplicons sized approximately 350 bp. Following amplification, the two reaction pools were combined, and amplicons were purified using an equal volume of AMPure beads (Beckman Coulter, Indianapolis, IN), with 20 μL of final elution using RNase-free water. The eluted DNA concentration was measured using Qubit dsDNA HS assay kit (Thermo Fisher Scientific).
An amount of 100 ng of DNA was used in the NGS library preparation using TruSeq DNA Nano (Illumina, San Diego, CA), according to the protocol described by the manufacturer. The library preparation excluded the DNA fragmentation step and started with the sequential processing of DNA end-repair, 3′-end adenylation, adapter ligation, and DNA fragment enrichment, prior to library normalization and pooling steps. The normalized pooled library proceeded to sequencing with MiSeq Reagent Kit v2 Nano (2 × 250 cycles) on the MiSeq System (Illumina).
NGS reads analysis, sequence assembly, and alignment
Reads as fastq files from Illumina machine were paired, trimmed for adapters, and used in mapping to DENV-4 Refseq (NC_002640) using Geneious mapper and medium sensitivity/fast fine-tuning setting, available in Geneious v.11 software. Assembled sequences were aligned and checked for possible sequence gaps. Gap-filling was performed using the long amplicon (1–2.5 kb) approach for DENV-4 [11], which was generated using cDNA and Pfu DNA Polymerase (Strategene-Agilent, Thermo Fisher Scientific). The amplicon was sequenced using BigDye Dideoxy terminator kit v.3.1 (Applied Biosystems, Waltham, MA) in the Sanger capillary sequencer ABI 3130xl (Applied Biosystems). The sequence generated was used to fill in the sequence gaps. Complete genome sequences of Jember DENV-4 were aligned using MUSCLE [20] in MEGA-X software package [21] and used in the following phylogenetic and evolutionary analyses.
DENV-4 phylogenetic and evolutionary analyses
Complete genome sequences or sequences with complete coding sequences (CDS) were retrieved from GenBank as of May 2021. A total of 285 sequences were selected and used as reference strains in phylogenetic and evolutionary analyses. The reference sequences and 13 Jember DENV-4 complete genome sequences were aligned together using MAFFT software [22] and used as a dataset in a phylogenetic reconstruction approach using the maximum likelihood method with GTR + G model as the selected best-fit model inferred by jModelTest2 [23]. The analysis was run for 500 bootstraps. The resulting tree was annotated in FigTree v.1.4.4 (available from http://tree.bio.ed.ac.uk/software/figtree/).
The phylogenetic and evolutionary analyses were then performed on a subset of data, where Jember sequences were grouped in a clade of closely related strains as inferred by the maximum likelihood analysis. A total of 55 genome coding sequences (CDS) were used in a dataset generation using BEAUti v.2.6 by excluding the possible recombination events. Each node was calibrated using the year of isolation and run in Bayesian Markov Chain Monte Carlo (MCMC) analysis using GTR + G, Bayesian skyline prior, and relaxed lognormal molecular clock with an initial rate of 7.6 × 10–4 substitutions/site/year [5]. Computation was done using BEAST v.2.6 software [24]. The generated tree was then annotated using TreeAnnotator v.2.6 with 10% burn-in and edited in FigTree showing posterior probability values in each node and the time to the most recent common ancestors (TMRCA) as median year with 95% Highest Posterior Density (HPD).
Amino acid variations in gene, recombination, and selection pressure analyses
Analyses were performed using a dataset containing 55 taxa of CDS genome sequences, including Jember strains, showing genetic close-relatedness. Amino acid alignment of Jember DENV-4 CDS was performed using MUSCLE, showing only variable sites in each DENV gene. Amino acid variations analysis was performed in comparison to a strain from China isolated in 1978 (Genbank accession no. FJ196849), which is located in the basal position of the phylogenetic tree. The recombination analysis was done using RDP4 software [25] by performing a full exploratory recombination scan using RDP, GENECONV, BootScan, MaxChi, Chimaera, SiScan, 3Seq, LARD, and Phylpro. The selection pressure analysis was performed using the Datamonkey selective and evolutionary bioinformatics online server [26], incorporating methods to detect selection using BUSTED, MEME, FEL, SLAC, and FUBAR.
Results
RNA template, NGS library preparation, and sequence analysis
The DENV-4 samples were collected from our previous surveillance study in Jember, East Java in 2019, showing the predominance of DENV-4 during a dengue outbreak [13]. Among 43 samples detected as DENV-4, a total of 13 RNA templates were sufficient in amount and quality for sequencing library preparation. These represented samples from DF and DHF clinical manifestations, as well as samples with primary and secondary infection status (Table 1). The sequencing run of Jember samples generated sequences with variable sequencing depth and the calculated mean of average sequencing depth of 50.5 × . This mean of depth was sufficient to generate complete genome sequences mapped to the DENV-4 reference sequence. In terms of genome coverage, Jember sequences have a mean coverage to the DENV-4 reference sequence of 98.58% (95% CI 96.98–100.18) (Table 2). Following the quality control and gap filling processes, all 13 sequences were curated and deposited to GenBank with accession numbers of OL314735 –OL314747 as listed in Table 2.
Phylogenetic analysis
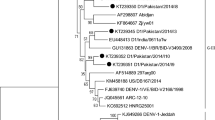
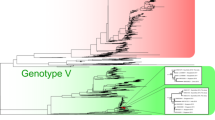
Phylogenetic analysis of Jember DENV-4 complete genomes with global reference strains available in public database revealed the classification of Jember viruses into Genotype IIa of DENV-4 (Fig. 1A). Phylogenetic analysis of 55 genome sequences including Jember sequences resulted in a maximum clade credibility tree having an age of 54 (95% HPD of 42–79) years. The overall mean evolutionary rate was recorded at 10.3 × 10–4 subs/site/year (95% HPD of 8.0–12.7 × 10–4 subs/site/year) and coefficient of variation of 0.682 (95% HPD of 0.521–0.879). All statistical indicators were supported by sufficient effective sampling size (> 150). Phylogenetic and evolutionary analyses showed further lineages grouping with 12 Jember DENV-4 strains clustered into Lineage 1 and closely related to strains from Malaysia and Bali, Indonesia (Fig. 1B). The Jember strains was estimated to diverge and group into a single sub-lineage at circa 2008. A single Jember DENV-4 strain was clustered into Lineage 2 and is closely related to strains from China and Jambi, Indonesia. The estimated TMRCA of this lineage was dated back at circa 1988. Evolutionary analysis of each Jember DENV-4 strain ranged from 7.30 to 13.80 × 10–4 subs/site/year (Table 2) with the mean evolutionary rate of 10.6 × 10–4 subs/site/year.
Maximum likelihood tree of Jember complete genome sequences together with global DENV-4 reference strains available in public database (A). Jember DENV-4 is grouped into Clade IIa of DENV-4 Genotype II based on global Maximum Likelihood analysis of all DENV-4 complete genome sequences found in GenBank repository. Closely related sequences in Genotype IIa (within red square) were selected to generate a Maximum Clade Credibility (MCC) tree of Jember DENV-4 strains (red font) together with most closely related DENV-4 reference strains from Indonesia (blue font) and other countries (B). The number in each node represents the posterior probability of the particular node. The time to the most common recent ancestor (TMRCA) is shown as year [95% Highest Posterior Density (HPD)]. Arabic numbers show classification into Lineages
Amino acid variations in gene, recombination, and selection pressure
Amino acid (AA) analysis of DENV-4 CDS sequences from Jember showed AA variation in genes, with 308 out of 3387 (9.1%) AA changes, compared to a reference strain at basal position from China in 1978. AA changes were found in NS5 gene (33 AA changes or 3.7% of total AA encoding the gene) (Fig. 2). Other genes showing AA changes were NS1, NS4B, and NS3 with 22 (6.25%), 21 (8.6%), and 20 (3.2%) AA changes, respectively.
Recombination analysis detected 12 potential recombination events in the analyzed dataset. Among them, two events showed significant detection by two or more methods with significant p values. Both events detected strain JBB-007 as the recombinant strain involving parents from other Jember strains. The first event predicted strain JBB-051 as the major parent and JBB-049 as the minor parent detected using RDP, BootScan, MaxChi, and Chimaera methods. The beginning of breakpoint was calculated at nucleotide (nt) position 9711 (99% confidence interval/CI 9486–9885) up to the end of sequence nt 10,660, which covers NS5 gene and 3′-UTR regions. The second event also involved strain JBB-051 as the major parent and JBB-008 as the minor parent. The event was detected by RDP, GENECONV, BootScan, MaxChi, Chimaera, and SiScan methods, with the predicted breakpoint beginning at nt position 2,538 (99% CI 1939–2717) and breakpoint end at nt 3554 (99% CI 3395–4793), which covers the NS1-NS2A genes.
Within the genomes, there was majority of negative purifying selection pressure with some detectable positive selection pressure by one method. There was only one site detected as under positive/diversifying selection by two methods. The AA site number 2772 (NS5 region) was detected as positively selected by FUBAR (p = 0.971) and MEME (p = 0.04). The AA change from glutamine (Q) to arginine (R) was observed in most of Jember DENV-4 genomes (Fig. 2). In addition, BUSTED analysis revealed the evidence of gene-wide episodic diversifying selection (p < 0.001), where at least one site on at least one test branch has experienced diversifying selection.
Discussion
The region of Jember in East Java province, Indonesia, was alerted by a dengue outbreak in 2019 in which a predominance of DENV-4 cases was reported [13]. This was a new finding since there have been no reports of outbreaks dominated by DENV-4 in Indonesia, especially in small cities far from the capital city of Jakarta. The outbreak affected mostly children under 10 years old (27.2% of the total cases), with a higher proportion of primary infections, concomitant with milder forms of dengue (DF) [13]. We proposed earlier that the introduction of a new DENV serotype, in this case DENV-4, in a population with a low herd immunity would lead to dengue outbreak. While this seems to be the case for Jember outbreak, virus factors may also contribute to the outbreak and explain how DENV-4 became predominant.
Using NGS approach, we successfully sequenced complete genomes of 13 DENV-4 strains from RNAs that were directly extracted from the patients’ serum samples in order to limit/exclude the adaptive mutations that can occur when DENV is passaged into cell culture [27]. The genome sequences have been deposited in public genome repository and these provide important addition to the limited number of DENV-4 genomes available, especially for strains originated from Indonesia. To our knowledge, currently, only seven complete genomes of Indonesia DENV-4 available in GenBank database, therefore, addition of 13 new genomes will be beneficial for future DENV genome study.
Jember DENV-4 strains were found to have evolved into two main lineages (Fig. 1B), based on phylogenetic tree and evolutionary analyses. They were grouped together in Genotype IIa of DENV-4 (Fig. 1A). The first lineage (Lineage 2, Fig. 1B) is represented by a strain with older evolutionary age, estimated to be circulating since around 1988. This lineage may be the parental lineage of DENV-4 in Jember, with a close relationship with an imported case from Malaysia to China and a strain from Jambi, Indonesia [28, 29]. This may suggest that DENV-4 viruses have been endemic in Indonesia and the region for more than three decades. The evolution process may then have led to the emergence of the second lineage (Lineage 1, Fig. 1B), comprising the majority of Jember 2019 strains, estimated to have emerged around the year of 2000 and Jember strains started to diverge and generating a new sub-lineage in circa 2008. This younger lineage may reflect the substantial evolutionary changes in Jember DENV-4 that enabled the virus mutants to infect a naïve population.
We identified changes in DENV genomes that may be correlated to their transmission. During the evolution, DENV-4 in Jember seems to gain many mutations as part of its evolution process (Fig. 2). We observed that the number of amino acid changes was higher in the NS5 gene, accounting for 3.7% of the total amino acid within the gene (Fig. 2). Moreover, the selection pressure analysis revealed that the viruses have primarily undergone purifying selection and only one site was detected to be positively selected, located in the NS5 gene. The highest percentage of AA changes was detected in NS4B gene (8.6%), encoding for a transmembrane protein that anchors the viral replication complex to the endoplasmic reticulum membrane [30], although none was positively selected. Positive selection may cause an increase in the rate of sequence variation in the virus genome and enhance the overall fitness of a virus sub-population, which may occur when there are changes in environmental factors, such as host immunity or the effects of antiviral drugs, as reported in HCV and HIV [31,32,33]. NS5 is one of DENV’s key catalytic enzymes, involved in viral genome replication and its capping [34]. The N‐terminal region of NS5 has a methyltransferase domain, while the C‐terminal region contains a conserved domain for RNA‐dependent RNA polymerase for de novo synthesis of viral RNA [35]. NS5 presence in the nucleus has roles in host splicing machinery and DENV pathogenesis [36]. All of these properties of NS5 have made it a potential target of directly acting antiviral drugs and vaccine development [37, 38]. The positive selection pressure in the Jember DENV-4 NS5 gene may be the possible cause of viral genomic changes that lead to the enhanced virus fitness, allowing it to gain predominance during the dengue outbreak. Regarding the AA substitutions related to disease severity, we did not observe any mutational changes that specifically correlate with patients’ clinical manifestations (DF vs DHF).
In addition, our in-silico analysis detected a recombination event involving breakout at the NS5 gene of Jember DENV-4 viruses. Recombination is considered a common phenomenon in RNA viruses and plays an important role in the biology of the virus and its evolution, which may contribute to an epidemic [39]. Evidence of intra-serotype recombination among DENV-4 virus strains has been reported previously and the event may contribute to the emergence of a new distinct genotype [40, 41]. Although the recombination event in Jember DENV-4 did not generate a new distinct genotype, the event may still be the cause of the emergence of the more recent phylogenetic clade Lineage 1 consisting of the majority of DENV-4 viruses isolated during the outbreak (Fig. 1B).
Regarding the possible recombination, we are aware that bioinformatic analysis alone may not be able to conclusively confirm the recombination in DENV-4. It is important that the recombination breakpoints be sequenced from a single cDNA molecule (e.g., by cDNA cloning) to ensure it is originated from a single virus. Additionally, the recombination should be shown in clonal populations of a live virus (e.g., by plaque purification or limited dilution) and during post-recombination evolution, sufficient level of sequence conservation should be maintained by the recombinant viruses. Therefore, further in vitro investigation is needed to confirm the recombination events occurred in our DENV-4 strains.
In conclusion, genetic factors may have contributed to DENV-4 viral fitness, causing its serotype predominance in the Jember 2019 dengue outbreak. A combination of selection pressures and mutational genetic changes may be the key factors of DENV-4 dynamics during the Jember dengue outbreak. Additionally, intra-serotype recombination events, although it need to be confirmed in vitro, may contribute to the fitness of the DENV-4. Altogether, these findings may act as additional factors besides the epidemiological aspects within the complex dengue outbreak mechanism. Continuous surveillance and study of DENV genomes and their evolutionary aspects are needed to monitor the dynamics of DENV in Indonesia and to improve preparedness for future outbreaks or epidemics.
References
Bhatt S, Gething PW, Brady OJ, Messina JP, Farlow AW, Moyes CL et al (2013) The global distribution and burden of dengue. Nature 496:504–507. https://doi.org/10.1038/nature12060
Halstead SB (2015) Pathogenesis of dengue: dawn of a new era. F1000Research. https://doi.org/10.12688/f1000research.7024.1
TDR/WHO (2009) Dengue: guidelines for diagnosis, treatment, prevention and control. World Health Organization, Geneva
Guzman MG, Halstead SB, Artsob H, Buchy P, Farrar J, Gubler DJ et al (2010) Dengue: a continuing global threat. Nat Rev Microbiol 8:S7-16. https://doi.org/10.1038/nrmicro2460
Costa RL, Voloch CM, Schrago CG (2012) Comparative evolutionary epidemiology of dengue virus serotypes. Infect Genet Evol 12:309–314. https://doi.org/10.1016/j.meegid.2011.12.011
Yohan B, Wardhani P, Trimarsanto H, Aryati A, Sasmono RT (2018) Genomic analysis of dengue virus serotype 1 (DENV-1) genotypes from Surabaya, Indonesia. Virus Genes 54:461–465. https://doi.org/10.1007/s11262-018-1558-z
Moya A, Holmes EC, González-Candelas F (2004) The population genetics and evolutionary epidemiology of RNA viruses. Nat Rev Microbiol 2:279–288. https://doi.org/10.1038/nrmicro863
Ong SH, Yip JT, Chen YL, Liu W, Harun S, Lystiyaningsih E et al (2008) Periodic re-emergence of endemic strains with strong epidemic potential-a proposed explanation for the 2004 Indonesian dengue epidemic. Infect Genet Evol 8:191–204
Harapan H, Michie A, Mudatsir M, Sasmono RT, Imrie A (2019) Epidemiology of dengue hemorrhagic fever in Indonesia: analysis of five decades data from the national disease surveillance. BMC Res Notes 12:350. https://doi.org/10.1186/s13104-019-4379-9
Harapan H, Michie A, Yohan B, Shu P-Y, Mudatsir M, Sasmono RT et al (2019) Dengue viruses circulating in Indonesia: a systematic review and phylogenetic analysis of data from five decades. Rev Med Virol 29:e2037. https://doi.org/10.1002/rmv.2037
Sasmono RT, Wahid I, Trimarsanto H, Yohan B, Wahyuni S, Hertanto M et al (2015) Genomic analysis and growth characteristic of dengue viruses from Makassar, Indonesia. Infect Genet Evol 32:165–177. https://doi.org/10.1016/j.meegid.2015.03.006
Yamanaka A, Mulyatno KC, Susilowati H, Hendrianto E, Ginting AP, Sary DD et al (2011) Displacement of the predominant dengue virus from type 2 to type 1 with a subsequent genotype shift from IV to I in Surabaya, Indonesia 2008–2010. PLoS ONE 6:e27322. https://doi.org/10.1371/journal.pone.0027322
Aryati A, Wrahatnala BJ, Yohan B, Fanny M, Hakim FKN, Sunari EP et al (2020) Dengue virus serotype 4 is responsible for the outbreak of dengue in East Java city of Jember, Indonesia. Viruses. https://doi.org/10.3390/v12090913
Messina JP, Brady OJ, Scott TW, Zou C, Pigott DM, Duda KA et al (2014) Global spread of dengue virus types: mapping the 70 year history. Trends Microbiol 22:138–146. https://doi.org/10.1016/j.tim.2013.12.011
Sang S, Liu-Helmersson J, Quam MBM, Zhou H, Guo X, Wu H et al (2019) The evolutionary dynamics of DENV 4 genotype I over a 60-year period. PLoS Negl Trop Dis. https://doi.org/10.1371/journal.pntd.0007592
Acosta POA, Maito RM, Granja F, da Silva Cordeiro J, Siqueira T, Cardoso MN et al (2011) Dengue virus serotype 4, Roraima State, Brazil. Emerg Infect Dis 17:1979–1980. https://doi.org/10.3201/eid1710.110776
Nisalak A, Endy TP, Nimmannitya S, Kalayanarooj S, Thisayakorn U, Scott RM et al (2003) Serotype-specific dengue virus circulation and dengue disease in Bangkok, Thailand from 1973 to 1999. Am J Trop Med Hyg 68:191–202
Quick J, Grubaugh ND, Pullan ST, Claro IM, Smith AD, Gangavarapu K et al (2017) Multiplex PCR method for MinION and Illumina sequencing of Zika and other virus genomes directly from clinical samples. Nat Protoc 12:1261–1276. https://doi.org/10.1038/nprot.2017.066
Stubbs S, Blacklaws B, Yohan B, Yudhaputri FA, Schwem B, Salvana EM, et al. (2018) A nanopore-based method for generating complete coding region sequences of dengue virus in resource-limited settings. bioRxiv. https://doi.org/10.1101/499111
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797. https://doi.org/10.1093/nar/gkh340
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547–1549. https://doi.org/10.1093/molbev/msy096
Katoh K, Asimenos G, Toh H (2009) Multiple alignment of DNA sequences with MAFFT. Methods Mol Biol 537:39–64. https://doi.org/10.1007/978-1-59745-251-9_3
Darriba D, Taboada GL, Doallo R, Posada D (2012) jModelTest 2: more models, new heuristics and parallel computing. Nat Methods 9:772. https://doi.org/10.1038/nmeth.2109
Bouckaert R, Vaughan TG, Barido-Sottani J, Duchêne S, Fourment M, Gavryushkina A et al (2019) BEAST 2.5: an advanced software platform for Bayesian evolutionary analysis. PLoS Comput Biol 15:e1006650. https://doi.org/10.1371/journal.pcbi.1006650
Martin DP, Murrell B, Golden M, Khoosal A, Muhire B (2015) RDP4: detection and analysis of recombination patterns in virus genomes. Virus Evol 1:vev003. https://doi.org/10.1093/ve/vev003
Weaver S, Shank SD, Spielman SJ, Li M, Muse SV, Kosakovsky Pond SL (2018) Datamonkey 2.0: a modern web application for characterizing selective and other evolutionary processes. Mol Biol Evol 35:773–777. https://doi.org/10.1093/molbev/msx335
Dolan PT, Taguwa S, Rangel MA, Acevedo A, Hagai T, Andino R et al (2021) Principles of dengue virus evolvability derived from genotype-fitness maps in human and mosquito cells. Wittkopp PJ, Sanjuan R, Illingworth CJ. eLife 10:61921. https://doi.org/10.7554/eLife.61921
Haryanto S, Hayati RF, Yohan B, Sijabat L, Sihite IF, Fahri S et al (2016) The molecular and clinical features of dengue during outbreak in Jambi, Indonesia in 2015. Pathog Glob Health 110:119–129. https://doi.org/10.1080/20477724.2016.1184864
Yohan B, Dhenni R, Hayati RF, Yudhaputri FA, Denis D, Pamai YWB et al (2018) Whole genome sequencing of Indonesian dengue virus isolates using next-generation sequencing. Indones J Biotechnol 23:74–83
Xie X, Wang Q-Y, Xu HY, Qing M, Kramer L, Yuan Z et al (2011) Inhibition of dengue virus by targeting viral NS4B protein. J Virol 85:11183–11195. https://doi.org/10.1128/JVI.05468-11
Kuntzen T, Timm J, Berical A, Lewis-Ximenez LL, Jones A, Nolan B et al (2007) Viral sequence evolution in acute hepatitis C virus infection. J Virol 81:11658–11668. https://doi.org/10.1128/JVI.00995-07
Schmidt DJ, Pickett BE, Camacho D, Comach G, Xhaja K, Lennon NJ et al (2011) A phylogenetic analysis using full-length viral genomes of South American dengue serotype 3 in consecutive Venezuelan outbreaks reveals a novel NS5 mutation. Infect Genet Evol J Mol Epidemiol Evol Genet Infect Dis 11:2011–2019. https://doi.org/10.1016/j.meegid.2011.09.010
von Hahn T, Yoon JC, Alter H, Rice CM, Rehermann B, Balfe P et al (2007) Hepatitis C virus continuously escapes from neutralizing antibody and T-cell responses during chronic infection in vivo. Gastroenterology 132:667–678. https://doi.org/10.1053/j.gastro.2006.12.008
Nasar S, Rashid N, Iftikhar S (2020) Dengue proteins with their role in pathogenesis, and strategies for developing an effective anti-dengue treatment: a review. J Med Virol 92:941–955. https://doi.org/10.1002/jmv.25646
Issur M, Geiss BJ, Bougie I, Picard-Jean F, Despins S, Mayette J et al (2009) The flavivirus NS5 protein is a true RNA guanylyltransferase that catalyzes a two-step reaction to form the RNA cap structure. RNA N Y N 15:2340–2350. https://doi.org/10.1261/rna.1609709
De Maio FA, Risso G, Iglesias NG, Shah P, Pozzi B, Gebhard LG et al (2016) The dengue virus NS5 protein intrudes in the cellular spliceosome and modulates splicing. PLoS Pathog 12:e1005841. https://doi.org/10.1371/journal.ppat.1005841
Bhatnagar P, Sreekanth GP, Murali-Krishna K, Chandele A, Sitaraman R (2021) Dengue virus non-structural protein 5 as a versatile, multi-functional effector in host-pathogen interactions. Front Cell Infect Microbiol 11:574067. https://doi.org/10.3389/fcimb.2021.574067
Maddipati VC, Mittal L, Mantipally M, Asthana S, Bhattacharyya S, Gundla R (2020) A review on the progress and prospects of dengue drug discovery targeting NS5 RNA- dependent RNA polymerase. Curr Pharm Des 26:4386–4409. https://doi.org/10.2174/1381612826666200523174753
Lai MM (1992) Genetic recombination in RNA viruses. Curr Top Microbiol Immunol 176:21–32. https://doi.org/10.1007/978-3-642-77011-1_2
Weaver SC, Vasilakis N (2009) Molecular evolution of dengue viruses: contributions of phylogenetics to understanding the history and epidemiology of the preeminent arboviral disease. Infect Genet Evol J Mol Epidemiol Evol Genet Infect Dis 9:523–540. https://doi.org/10.1016/j.meegid.2009.02.003
Worobey M, Rambaut A, Holmes EC (1999) Widespread intra-serotype recombination in natural populations of dengue virus. Proc Natl Acad Sci U S A 96:7352–7357
Acknowledgements
We would like to thank the National Genome Center (Pusat Genom Nasional) of Eijkman Institute and PT. Pandu Biosains for their help and assistance in NGS preparation.
Funding
This study was funded by a research grant from the Universitas Airlangga (Grant No: 965/UN3.14/LT/2019) to AA.
Author information
Authors and Affiliations
Contributions
PW, BY, AA, and RTS conceived and designed the study. BY, RFH, AR, MT, EPS, BJW, DH, JTOH, and FKNH performed sample collection and laboratory experiments. PW, BY, and AR did data analysis. BY, PW, and RTS wrote the paper. JTOS, RFH, MSS, and RTS contributed to the final draft of the manuscript. AA and RTS performed study supervision.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Edited by Joachim J. Bugert.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wardhani, P., Yohan, B., Tanzilia, M. et al. Genetic characterization of dengue virus 4 complete genomes from East Java, Indonesia. Virus Genes 59, 36–44 (2023). https://doi.org/10.1007/s11262-022-01942-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11262-022-01942-4