Abstract
It is important to study the molecular properties of vertically transmitted viruses in early infancy to understand disease progression. P24 having an important role in virus assembly and maturation was selected to explore the genotypic characteristics. Blood samples, obtained from 82 HIV-1 positive infants, were categorized into acute (≤ 6 months) and early (> 6–18 months) age groups. Of the 82 samples, 79 gave amplification results for p24, which were then sequenced and analysed. Amino acid heterogeneity analysis showed that substitutions were more frequent. Several substitution mutations were present in some of the sequences of both the age groups in the functional motifs of the gene namely Beta hairpin, CyPA binding loop, residues L136 and L190, linker region and major homology region. In the acute age group, an insertion of Asparagine residue (N5NL6) was observed in the β hairpin region in one of the sequences. This insertion was accompanied with analogous substitutions of N5Q, Q7L and G8R. In the early age group, a deletion of two residues; VK181−182, was observed at the C-terminal end in one of the sequences. These mutations may impair the structure of the protein leading to defective virus assembly. Protein variation effect analyzer software showed that deleterious mutations were more in the acute than the early age group. Variability analysis revealed that the amino acid heterogeneity was comparatively higher in the acute than the early age group. Variability in the virus was decreasing with the increasing age of the infants indicating that the virus is gradually evolving under positive selection pressure. HLA class 1 binding peptide analysis showed that the epitopes TPQDLNTML and RMYSPVSIL may be helpful in designing epitope based vaccine.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
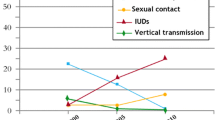
Transmission of Human Immunodeficiency Virus type-1 (HIV-1) (Species: Human Immunodeficiency Virus-1, genus: Lentivirus, Family: Retroviridae) infection from mother to her child is the most important mode of transmission in the children [1, 2]. Disease progression in vertically infected infants/neonates occurs rapidly as compared to the adults [3]. Although ART in HIV-1 infected pregnant women has significantly reduced the rate of mother to child transmission (MTCT), still it continues to remain a major concern in the children worldwide [4].
The P24 capsid protein of HIV-1 plays an important role in virus assembly, maturation and in early post-entry steps. It is released from the central part of HIV-1 group specific antigen (Gag) polyprotein, forming the conical shell of the virus and enclosing the viral genome [5,6,7,8]. The protein is made up of approximately 240 amino acids and is divided into two different domains, an N-terminal domain (NTD) and a Carboxyl-terminal domain (CTD). Both the domains are joined via a four residue inter-domain flexible linker region [6, 8, 9]. NTD is rich in proline residues and is essential in formation of functional mature virion core while CTD is significant in particle assembly and oligomerisation of Gag polyprotein [5, 10].
Identifying the genetic variability in vertically transmitted viruses in early infancy is important to understand the disease progression. Data on how heterogeneity in viral genes develops in the early and later ages of the infant are very limited. Previously, we have characterized HIV-1 nef gene from HIV-1 infected infants and reported a decreasing trend of amino acid variability with the increasing age of the infants [11]. In the present study molecular characterization of p24 gene in HIV-1 infected infants born to HIV-1 infected mothers from northern states of India was carried out. Though p24 gene being conserved and under low evolutionary pressure, it has a dominant role in gag specific CTL epitopes, and thus play a significant role for vaccine formulation [12]. Heterogeneity within subtype C strains in different population and their HLA profile influences the variety of epitopes recognized [13]. Therefore, identification of different epitopes from different antigenic regions in different population is important for designing vaccine construct.
In the current study, patients were categorized in “acute age group” (≤ 6 months) and an “early age group” (> 6–18 months) to understand the evolutionary changes in the gene at very early age and at later age group. Functional motifs of the gene essential for p24 activity were analysed and evolutionary relationships were studied. Similarity analysis of the gene with the vaccine candidate sequences and HLA-binding motifs was also predicted that may provide useful data for designing an efficient multi-epitope vaccine.
Materials and methods
Study population
A total of 82 whole blood samples (WBS) of infants aged 6 weeks to 18 months, born to HIV-1 positive mothers, were included in the present study. These samples were received from various ART centres of northern states of India to National Centre for Disease Control (NCDC), Delhi for early diagnosis of HIV-1. The study was carried out at NCDC with proper ethical clearance.
Sample preparation
WBSs were processed to obtains the pellet of PBMCs using the BLD Wash solution provided in the AMPLICOR HIV-1 DNA test kit, version 1.5 (Roche Molecular Systems) according to the manufacture instructions.
Genomic DNA extraction
Genomic DNA was extracted from the stored PBMCs using Qiagen DNeasy kit (Qiagen GmbH, Germany) according to the manufacturer’s protocol. Presence of the genomic DNA was confirmed by gel electrophoresis in 1% agarose using 0.5× Tris–acetate EDTA (TAE) buffer. DNA was stored at − 20 °C till used.
PCR amplification
All the samples which were included in the study (n = 82) were diagnosed by PCR technique using the primers SK145 (5′-AGTGGGGGGACATCAAGCAGC CATGCAAAT-3′) and SKCC1B (5′-TACTAGTAGTTCCTGCTATGTCACTTCC-3′). These primers amplified a sequence of 155 nucleotides within the highly conserved region of gag gene. All the 82 samples gave amplification results for 155 nucleotides in the gag gene. In these positive samples, the p24 gene with amplicon size 717 bp was amplified through nested PCR as described earlier [14]. Sequences of the forward and reverse primers used in the outer PCR were G00 (5′-GACTAGCGGAGGCTAGAAG-3′ nt. position 764–782) and G01 (5′-AGGGGTCGTTGCCAAAGA-3′ nt. position 2264–2281) respectively. For the inner PCR the primer pair used was G60 (5′-CAGCCAAAATTACCCTATAGTGCAG-3′ nt. position 1173–1197) and G25 (5′-ATTGCTTCAGCCAACTCTTG-3′ nt. position 1867–1889). Amplification was carried out using the Green Master Mix PCR kit (Promega, Madison, USA) according to the manufacturer’s instructions. The thermal profile for outer PCR was initial denaturation at 95 °C for 2 min followed by amplification 30 cycles of 92 °C for 10 s; 55 °C for 30 s; 72 °C for 1 min; final extension at 72 °C for 7 min. For nested PCR, it was initial denaturation at 95 °C for 2 min followed by amplification 30 cycles of 92 °C for 10 s; 56 °C for 20 s; 72 °C for 1 min; final extension at 72 °C for 7 min. Quality of the PCR products was checked on 1% (w/v) agarose gel electrophoresis in 1× Tris Acetate EDTA (TAE) buffer (40 mM Tris, 20 mM acetic acid and 1 mM EDTA), pH 8.3 (Merck). Agarose gels were stained with Ethidium Bromide and were visualized under UV transilluminator (Gel Documentation System, AlphaImager EC, USA).
Sequencing
Amplified PCR products were subjected to automated nucleotide sequencing separately with both forward and reverse primers of inner PCR. Sequencing was carried out using the Big Dye Terminator Cycle Sequencing Kit v 3.1 (Applied Biosystems, CA, USA) containing four dye labelled dideoxynucleotide terminators according to the manufacturer’s instructions. Cycle sequencing parameters included 25 cycles at 96 °C for 10 s, 50 °C for 5 s and 60 °C for 4 min. Purification of the reaction mixture was carried out manually using 3M sodium acetate (pH 4.6) and ethanol precipitation method. Purified DNA was lyophilized and re-suspended in 12 µl Hi-Di formamide (Applied Biosystems) followed by heat (95 °C for 2–3 min) and immediate chilling on ice (+ 4 °C) for 5–10 min. DNA was finally loaded onto 3130xl Genetic Analyzer (Applied Biosystems) for nucleotide sequencing and data collection.
Genetic analysis
All the sequences collected from Genetic Analyzer were resolved with the help of bioinformatics software MEGA v 6.06 [15] and BioEdit software v. 7.0.9.0 [16]. All the sequences were analysed for the peaks in the electropherogram with the help of sequencing analysis software v 5.3. Only dominant peaks having high Normalized fluorescence intensity (peak values) were used for analysis. Resolved sequences were submitted to the GeneBank and accession numbers were obtained (KY926622–KY926694, KY930920–KY930924).
Virus subtyping
Subtypes of all the isolates were determined through NCBI Blast and RIP tool available at http://www.hiv.lanl.gov/content/sequence/RIP/RIP.html.
Phylogenetic analysis and inter-patient nucleotide distances
Phylogenetic tree was constructed using the sequences of the present study and previously reported sequences. The tree was constructed by maximum likelihood method using MEGA v 6.06 and the reliability of the branching orders was determined using bootstrapping. The inter-patient nucleotide distance and genetic diversity were calculated separately for both the age groups using MEGA, v 6.06 [15].
Synonymous/nonsynonymous substitution ratio (dS/dN)
This ratio helps in analysing regions of proteins evolving under positive selection. A value of dS/dN > 1 indicates amino acid conservation due to structural and functional constraints, whereas dS/dN < 1 suggests the diversity in amino acids [15]. The dS/dN substitution ratio was calculated separately for both the age groups using the SNAP programme available at http://www.hiv.lanl.gov/content/sequence/SNAP/SNAP.html [17].
Multiple alignments
Multiple sequence alignment was carried out using BioEdit software v. 7.0.9.0 [16]. Alignment was done separately for both the age groups. All the study sequences were aligned with the consensus subtype C sequence consisting of 231 amino acid residues (source: HIV sequence database) in both the age groups. Position numbering of amino acids in the sequences is according to their positions in HIV-1 subtype C consensus sequence.
Protein variation effect analyzer software (PROVEAN)
It is a software tool which is available at http://provean.jcvi.org/index.php. The tool predicts whether an amino acid mutation affects the protein function [18]. It can predict any kind of substitutions, insertions and deletions. When an amino acid sequence is submitted, the software tool searches it’s homologs against the NCBI database using BLAST that are clustered by software CD-HIT. On the basis of the selected homologs, the PROVEAN scores are computed for each of the mutation. The scores are then averaged within and across clusters to generate the final PROVEAN score. If the PROVEAN score is equal to or below a predefined threshold (e.g. − 2.5), the protein variant is predicted to have a “deleterious” effect. If the PROVEAN score is above the threshold, the variant is predicted to have a “neutral” effect. The default score threshold is − 2.5 for binary classification i.e. deleterious vs neutral [19, 20]. The PROVEAN score in this study was calculated for each amino acid mutation separately in both the age groups.
Measure of variability
Entropy measures the variability of amino acids at a given position. It takes into account possible amino acids and their frequency at a given position. Sequences from both the age groups were examined for variability in amino acids of the functional motif of the protein. Variability was calculated as entropy using the tool, Entropy-Two available at http://www.hiv.lanl.gov/content/sequence/ENTROPY/entropy.html. The tool uses Claude Shannon Entropy to measure the variation in protein sequence alignments. A Shannon entropy score was calculated for each amino acid in the alignment. Entropy of two age groups was compared to find the difference in variability between the alignments of two groups. Entropy plots of both the age groups and a plot showing the difference in amino acid residue with statistically significant difference in variability were constructed. Statistical significance was calculated using Monte Carlo randomization (5 out of 100) with replacement. The sign of entropy difference in the dataset indicates whether the observed variations/conservation was in acute age group or in early age group.
Prediction of host HLA class-1 binding peptide motif
In Indian population, there is high occurrence of HLA-A1, A2, A-0201, A-0205, B-7, B-8, B-2705, Cw401 and Cw602 alleles [21,22,23,24]. These HLA types were studied to identify the binding epitopes in HIV-1 p24 sequences of the study. The software programme ProPred-I available at http://www.imtech.res.in/raghava/propred1/ was used for analysis of HLA-1 binding sites/epitopes in the p24 sequences [25]. Percentage of isolates showing conservation of each epitope was calculated and the epitopes which were most likely to be expressed in circulating HLA strains were identified. These epitopes were then compared with the potential vaccine candidate sequences AB023804 (93IN101), AY043175 (DU422), AF286227 (97ZA012), U52953 (92BR025) and AF286224 (96ZM651) to assess which vaccine candidate(s) sequence encodes the largest number of relevant epitope sequences [26].
Results
Of the 82 HIV-1 positive samples amplified, 79 were found to be PCR positive for the p24 gene, which were then sequenced and analysed. NCBI Blast and RIP tool categorized 74 sequences (93.67%) as subtype C; 3 (3.79%) as subtype A1 (IND_UP215, IND_PB67, IND_UP107), while 2 (2.53%) were categorized as recombinants (A1C and 01_AE). Further analyses were conducted using only subtype C sequences (n = 74). Out of the 74 subtype C sequences, 24 belonged to the acute and 50 to the early age groups. In the Phylogenetic tree (Fig. 1), it was observed that subtype C sequences of the present study clustered together and were interdigitated with those of previously reported subtype C p24 sequences from India and other countries. Inter-patient nucleotide distances were 0.045 and 0.051 in acute and early age groups respectively. The average of dS/dN substitution ratio was 7.4081 and 8.1957 in acute and early age groups respectively.
Amino acid heterogeneity
Multiple alignment of the sequence data indicated that the p24 in the subtype C sequences ranged from 231 to 232 residues in the acute age group (≤ 6 months) while in the early age group, it ranged from 229 to 231 residues (Fig. 2A, B). All the sequences had an intact ORF indicating the presence of a functional protein. Substitutions were more frequent than insertion and deletions. Several substitution mutations were present in some of the sequences of both the age groups in the functional motifs of the gene namely Beta hairpin, CyPA binding loop, residues L136 and L190, Linker region and MHR.
A Multiple sequence alignment of p24 subtype C sequences of infants with the consensus sequence in the acute age group. The sequence names are shown on the left side. Dots indicate identity with the consensus sequence. A ruler with the block of 10 amino acid residues is shown. Motifs with mutations are highlighted in a box. B i Multiple sequence alignment of p24 subtype C sequences of infants with the consensus sequence in the early age group. Motifs with mutations are highlighted in a box. ii Multiple sequence alignment of p24 subtype C sequences of infants with the consensus sequence in the early age group. Motifs with mutations are highlighted in a box
In the acute age group an insertion of Asparagine (N) residue between N5 and L6 was observed in the β hairpin structure formed by the residues PIVQNLQGQMVHQA1−14 in the sequence IND_DL80. In the same sequence residues N5, Q7 and G8 were substituted by homologue Glutamine, analogues Leucine and Arginine respectively. In another sequence of the same age group, residue H12 was substituted by Tyrosine (IND_UP106) while in another two sequences; A14 was substituted by Proline (IND_RJ187 and IND_UP208). In the early age group, residue L6 of the β hairpin was substituted in 3 of the sequences (IND_DL39, IND_UP101 and IND_RJ216). Residues Q9 and H12 were substituted by Leucine in IND_UP79 and IND_UP48 respectively, while the residue Q13 was substituted by Serine in the sequence IND_UP79. Residue A14 was substituted in 12 sequences (Leucine in 2, Proline in 9 and Serine in 1 sequence).
In the NTD, a loop is formed by the residues PVHAGPIAPG85−94 which catalyzes the binding of CyPA (cellular peptidyl-prolyl cis–trans isomerase cyclophilin A) into the virion. In the acute age group, residue V86 in this loop was substituted in 5 of the sequences (Isoleucine in 4 and Glutamine in 1 sequence); H87 was substituted in 2 of the sequences (Glutamine in 1 and Proline in 1 sequence); I91 was substituted in 7 of the sequences (Valine in 4, Asparagine in 2 and Leucine in 1 sequence) and A92 was substituted by Proline in 1 of the sequences. In the early age group, residue V86 was substituted in 13 of the sequences (Isoleucine in 6, Glutamine in 3, Alanine in 3 sequences and Threonine in 1 sequence); H87 was substituted in 15 of the sequences (Glutamine in 12 and Proline in 3 sequences); I91 was substituted in 24 of the sequences (Valine in 16, proline in 1, Asparagine in 2, Leucine in 2 and Alanine in 3 sequences) and A92 was substituted by Proline in 3 of the sequences.
Glutamine at position 112, important in viral particle production, was conserved in all the sequences of acute age group while in the early age group this residue was substituted by Alanine in 1 of the sequences (IND_UP202). Leucine residue at positions 136 and 190 are important for intracellular assembly of Gag precursor. In the acute age group, L136 was substituted by Isoleucine in 1 of the sequences (IND_UTH66) while in the early age group; this residue was substituted by Methionine in 1 of the sequences (IND_RJ216). In the acute age group, L190 was conserved in all the sequences while in the early age group this residue was substituted by Methionine in 1 of the sequences (IND_UP130).
In the flexible linker region between the NTD and CTD, YSPVSIL145−151, the residue V148 was substituted in 8 of the sequences of acute age group (Threonine in 6, Serine in 1 and Alanine in 1 sequence) and 22 of the sequences in early age group (Threonine in 17, Alanine in 2 and Serine in 3 sequences). Residue S149 was substituted by Glycine in 1 of the sequences of early age group (IND_RJ216).
A conserved epitope with the sequence DRFYKTLRAE166−175 present at the C-terminal end of the Major Homology Region (MHR) was found to be changed at Y position (DRFFKTLRAE) in the current study. In the acute age group T171 in this epitope was substituted by Cysteine in the sequences IND_CHD46 while in the early age group this residue was substituted in 4 of the sequences (by Cysteine in 1 and Valine in 3 sequences). In the early age group residue K170R was also substituted in the sequence IND_RJ105. At C-terminal to this MHR, a deletion of 2 residues, VK181−182, was also observed in 1 of the sequences of early age group (IND_RJ105). Summary of isolates showing mutations in various functional motifs of the gene is shown in Tables 1 and 2. Mutations observed in each motif of the gene were compared in both the age groups and were tested for statistical significance. Statistical analysis of sequences in each age group displaying variability in each motif of p24 gene is shown in Table 3.
Protein variation effect analyzer software (PROVEAN)
The PROVEAN score was calculated for each amino acid mutation separately in both the age groups (Tables 4, 5). In the acute age group there were a total of 18 types of amino acid mutations. Out of the 18 mutations, 11 were deleterious (61.1%) and 7 were neutral (38.9%). While in the early age group there were a total of 30 types of amino acid mutations. Out of the 30 mutations, 14 were deleterious (46.7%) and 16 were neutral (53.3%).
Measure of variability
The entropy score of amino acids in the sequence alignment were plotted for the two age groups (Fig. 3). It was found that the number of residues in the functional motifs of p24 having high entropy score were more in early age group (14 residues) as compared to the acute age group (6 residues) as shown in Table 6. The entropy difference between acute and early age groups was tested for statistical significance using Monte Carlo randomization. The amino acid residues with significant p values were determined (Table 7). It was observed that 8 amino acids namely A15, S45, M69, E76, H88 (part of the CyPA Binding loop), E129, R133 and L212 had statistically significant p values. Out of these 8 significant residues, 5 residues had high entropy score and belonged to the acute age group while only 3 residues having high score belonged to the early age group. A plot showing difference in entropy at each amino acid between the two age groups was also plotted (Fig. 4).
a Entropy plot of p24 amino acid sequences in acute age group. The bars indicate the entropy score of each amino acid in the sequence alignment. The entropy score of amino acids, which are part of functional protein motifs are shown by horizontal brackets. The motifs are numbered from 1 beta hairpin, 2 D51, 3 CyPA binding loop, 4 Q112, 5 L136, 6 linker region, 7 MHR, 8 WM, 9 L190. b Entropy plot of p24 amino acid sequences from early age group
Predicted HLA-binding peptide motif
Study sequences of p24 gene were compared with the circulating HLA types to identify the binding epitopes. A total of 11 epitopes namely YVDRFFKTL, IILGLNKIV, WIILGLNKI, KRWIILGLN, DRFFKTLRA, YKRWIILGL, DLNTMLNTV FRDYVDRFF, TPQDLNTML, SPRTLNAWV and PFRDYVDRF were present in > 90% of the sequences. Out of these 11 epitopes, 4 (FRDYVDRFF, TPQDLNTML, SPRTLNAWV and PFRDYVDRF) were present in all the 74 sequences. Epitope RMYSPVSIL was present in only 56.75% sequences, but it was present in maximum number of circulating HLA types (HLA-A2, HLA-A*0201, HLA-A*0205, HLA-B*2702, HLA-B*2705). Searching all the epitopes in potential vaccine candidate sequences, it was observed that the epitopes TPQDLNTML, DLNTMLNTV, KIVRMYSPV, RMYSPVSIL, FRDYVDRFF, YVDRFFKTL, DRFFKTLRA and EMMTACQGV were present in all the 5 vaccine candidate sequences as shown in Table 8.
Discussion
The p24 gene is significant in virus assembly and maturation and plays important role in early post entry steps. Molecular characterization of this gene will help in dissecting the epidemiology of the virus strains and genomic heterogeneity in young infants. As observed in earlier reports from Indian subcontinent, in this study too, the virus belonged to the subtype C (93.67%) followed by subtype A1 (3.79%) [27, 28]. In the phylogenetic tree, interdigitated sequences of the present study with those of previously reported varied subtype C sequences suggested that the source of infection was not common. The inter-patient nucleotide distance was low in both acute and early age groups suggesting that virus population in both the groups was genetically related. Also the difference in nucleotide distances between both the age groups was narrow indicating that the viruses in two age groups were not genetically diverse too [29]. The average dS/dN substitution ratio, an indicator of sequence conservation was > 1 in both the age groups, confirming that synonymous substitutions were more frequent signifying a purifying selection towards robust viral population [30].
Analyses of heterogeneity of amino acid sequences in infant groups revealed that most of the functional motifs of the gene were conserved. An insertion of Asparagine (N) residue between N5 and L6 was observed in 1 of the sequences of acute age group. This insertion was accompanied with the substitution of N5Q, Q7L and G8R residues. This was the unique finding in the study. These mutations were present in the β hairpin structure, which is formed by the residues PIVQNLQGQMVHQA1−14 present at the NTD of the protein. It is reported that the mutations present in the NTD can interfere in the formation of capsid in early phases of the virus [31, 32]. Therefore, we may predict that the insertion of N and substitutions of N5Q, Q7L, G8R, H12Y and A14P may distort the loop structure and functioning of the protein.
In the Proline-rich loop (PVHAGPIAP85−93), residues V86, H87, I91, A92 were substituted in 4 and 10 sequences of acute and early age groups respectively. This loop catalyzes the incorporation of CyPA into the virion. The CyPA regulates the protein folding and prevents the aggregation of protein folding intermediates [33,34,35]. Reports have suggested that the mutations near the CyPA binding site reduce the interaction between the virion and CyPA [36]. Therefore, the substitutions are suggestive of altered binding of the virion with and CyPA that may disturb the protein folding.
A Glutamine residue at position 112, important in viral particle production, was substituted with analogue Alanine (Q112A) in 1 of the sequences of early age group. Q112A mutation reduces the viral particle production, abolishes capsid formation and blocks the viral infectivity [34]. Therefore, it may be predicted that the substitution of this residue in this study may also contribute in blocking the viral infectivity. Another substitution was observed in Leucine residue at positions 136 and 190. The L136 was substituted in 1 sequence each in both the age groups while L190 was substituted in 1 of the sequences of early age group. Both the residues are reported to be highly conserved in HIV-1 database. Substitutions of these residues may affect the virus assembly as these residues are important for intracellular assembly of Gag precursor [37].
In the flexible linker region (YSPVSIL145−151), residue V148 was substituted in 8 and 22 of the sequences of acute and early age group respectively while residue S149 was substituted in 1 of the sequences of early age group. This linker region is important for correct core assembly, viral replication and infectivity [32]. Mutations in this region are reported to abrogate the correct core assembly of capsid [9], therefore, the substitutions of V148 and S149 observed in the current study may also alter the assembly of the virus particle.
In the highly conserved, immunodominant CTL epitope (DRFFKTLRAE166−175) present at the C-terminal end of the MHR, residue T171 was substituted in 1 and 4 of the sequences of acute and early age groups respectively and residue K170 was substituted in 1 of the sequences of early age group. Reports suggest that variability in these two residues results in impaired viral replication [38]. The substitution of K170 and T171 observed in this study may also diminish the CTL recognition leading to immune escape and disease progression. Another unique finding of the study was the deletion of 2 residues, VK181−182, at C-terminal to the MHR in one of the sequences of early age group. It is reported that the sequences within the C-terminal of HIV-1 capsid protein are important for particle formation and the deletions in this region causes defects in viral replication, reducing the ability to form viral particles [39].
PROVEAN score analysis revealed that percentage of deleterious mutations were more in the acute age group (61.1%) compared to the early age group (46.7%). Similar kind of results was also observed in one of our previous study on nef gene [11]. This suggested that amino acid mutations which may give nonfunctional protein were more in the acute age group and less in the early age group indicating that the virus is gradually evolving towards positive selection pressure. Entropy analysis revealed that out of the 8 statistically significant residues which were having high entropy score, 5 belonged to the acute and only 3 belonged to the early age groups. This suggested that heterogeneity of the residues was higher in the acute than the early age group. This indicated that the infants have received heterogeneous virus from their mothers and the virus is gradually evolving towards positive selection. Similar kind of result has also been found in our previous study on nef gene [11]. Analysis of HLA-binding peptide motif suggested that the epitopes TPQDLNTML and RMYSPVSIL may be helpful in designing an epitope based vaccine suitable in Indian subcontinent as the same were present in maximum number of circulating HLA types and vaccine candidate sequences.
In conclusion the study revealed that the amino acid variability was comparatively higher in the acute age group and the variability in the virus gene was decreasing with the increasing age of the infant signifying the positive selection. Deleterious mutations in the gene were more in the acute than the early age group. The unique finding of this study were, insertion of Asparagine in 1 of the sequences of acute age group accompanied with substitution of N5Q, Q7L and G8R residues and deletion of 2 residues (VK181−182) in 1 of the sequences of early age group. Epitopes TPQDLNTML and RMYSPVSIL may be a good candidate for vaccine design suitable for subtype C strains in Indian subcontinent. More studies, however, with inclusion of more HIV-1 genes and with the larger cohort of infants may provide better understanding of genetic characteristics of the virus in early age groups.
References
Read JS (2005) Prevention of mother-to-child transmission of HIV. In: Zeichner SL, Read JS (eds) Textbook of pediatric HIV care. Cambridge University Press, Cambridge
Read JS (2007) Diagnosis of HIV-1 infection in children younger than 18 months in the United States. Pediatric 120(6):e1547–e1562
Little K, Thorne C, Luo C, Bunders M, Ngongo N, McDermott P, Newell ML (2007) Disease progression in children with vertically-acquired HIV infection in sub-Saharan Africa: reviewing the need for HIV treatment. Curr HIV Res 5(2):139–153
Ahmad N (2005) The vertical transmission of human immunodeficiency virus type 1: molecular and biological properties of the virus. Crit Rev Clin Lab Sci 42(1):1–34
Freed EO (1998) HIV-1 Gag proteins: diverse functions in the virus life cycle. Virology 251:1–15
Iroegbu J, Birk M, Lazdina U, So’Nnerborg A, Sa’Llberg M (2000) Variability and immunogenicity of human immunodeficiency virus type 1 p24 gene quasispecies. Clin Diagn Lab Immunol 7(3):377–383
Gottlinger HG (2001) HIV-1 Gag: a molecular machine driving viral particle assembly and release. HIV Seq Compend 2001:2–28
Bayro MJ, Chen B, Yau WM, Tycko R (2014) Site-specific structural variations accompanying tubular assembly of the HIV-1 capsid protein. J Mol Biol 426(5):1109–1127
Jianga J, Ablanb S, Derebaila S, Hercika K, Soheilianc F et al (2011) The interdomain linker region of HIV-1 capsid protein is a critical determinant of proper core assembly and stability. Virology 421(2):253–265
Fitzon T, Leschonsky B, Bieler K, Paulus C, Schroeder J, Wolf H, Wagner R (2000) Proline residues in the HIV-1 NH2-terminal capsid domain: structure determinants for proper core assembly and subsequent steps of early replication. Virology 268:294–307
Sharma U, Gupta P, Singhal M, Singh S, Gupta S, Venkatesh S, Rai A, Husain M (2017) Comparative genetic variability in HIV-1 subtype C nef gene in early age groups of infants. J Med Virol 89(9):1606–1619
Gupta RM, Prasad VVSP, Rai A, Seth P (2005) Analysis of HIV type 1 subtype C full-length gag gene sequences from India: novel observations and plausible implications. AIDS Res Hum Retroviruses 21(12):1066–1072
Thakar MR, Bhonge LS, Lakhashe SL, Shankarkumar U, Sane SS et al (2005) Cytolytic T lymphocytes (CTLs) from HIV-1 subtype C–infected Indian patients recognize CTL epitopes from a conserved immunodominant region of HIV-1 Gag and Nef. J Infect Dis 192:749–759
Sanders-Buell E, Salminen MO, McCutchan FE (1995) Sequencing primers for HIV-1. In: Myers G (ed) Human retroviruses and AIDS, part III. Los Alamos National Laboratory, Los Alamos, NM, pp 15–21
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA 5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucl Acids Symp 41:95–98
Korber B (2000) HIV signature and sequence variation analysis. In: Rodrigo AG, Learn GH (eds) Computational analysis of HIV molecular sequences, Chap. 4. Kluwer Academic Publishers, Dordrecht, pp 55–72
Choi Y, Chan AP (2015) PROVEAN web server: a tool to predict the functional effect of amino acid substitutions and indels. Bioinformatics 31(16):2745–2747
Choi Y, Sims GE, Murphy S, Miller JR, Chan AP (2012) Predicting the functional effect of amino acid substitutions and indels. PLoS ONE 7(10):e46688
Choi Y (2012) A fast computation of pairwise sequence alignment scores between a protein and a set of single-locus variants of another protein. In: Proceedings of the ACM conference on bioinformatics, computational biology and biomedicine (BCB ‘12). ACM, New York, NY, pp 414–417
Rani R, Vina MAF, Stastny P (1998) Association between HLA class II alleles in a North Indian population. Tissue Antigens 52:37–43
Mehra NK, Jaini R, Rajalingam R, Balamurugan A, Kaur G (2001) Molecular diversity of HLA-A*02 in Asian Indians: predominance of A*0211. Tissue Antigens 57:502–507
Shankarkumar U, Thaka M, Mehendale S, Praranjape RS, Mohanty D (2003) Association of HLA B*3520, B*1801, and Cw*1507 with HIV-1 Infection Maharashtra, India. JAIDS 34:113–114
Kumar M, Jain SK, Pasha ST, Chattopadhayay D, La S, and Rai A (2006) Genomic diversity in the regulatory nef gene sequences in Indian sequences of HIV type 1: emergence of a distinct subclade and predicted implications. AIDS AIDS Res Hum Retroviruses 22:1206–1219
Singh H, Raghava GPS (2003) ProPred1: prediction of promiscuous MHC class-I binding sites. Bioinformatics 19(8)1009–1014
Gaschen B, Taylor J, Yusim K, Foley B, Gao F, Lang D, Novitsky V, Haynes B, Hahn BH, Bhattacharya T, Korber B (2002) Diversity considerations in HIV-1 vaccine selection. Science 296(5577):2354–2360
Siddappa NB, Dash PK, Mahadevan A, Jayasuryan N, Hu F, Dice B, Keefe R, Satish KS, Satish B (2004) Identification of subtype C human immunodeficiency virus type 1 by subtype-specific PCR and its use in the characterization of viruses circulating in the southern parts of India. J Clin Microbiol 42(6):2742–2751
Pandey SS, Cherian S, Thakar M, Paranjape RS (2016) Short communication: phylogenetic and molecular characterization of six full-length HIV-1 genomes from India reveals a monophyletic lineage of indian sub-subtype A1. AIDS Res Hum Retroviruses 32(5):489–502
Geffin R, Wolf D, Muller R, Hill MD, Stellwag E, Freitag M, Sass G, Scott GB, Baur AS (2000) Functional and structural defects in HIV type 1 nef genes derived from pediatric long-term survivors. AIDS Res Hum Retroviruses 16:1855–1868
Gojobori T, Yamaguchi Y, Ikeo K, Mizokami M (1994) Evolution of pathogenic viruses with special reference to the rates of synonymous and nonsynonymous substitutions. Jpn J Gene 69:481–488
Abdurahman S, Youssefi M, Hoglund S, Vahlne A (2007) Characterization of the invariable residue 51 mutations of human immunodeficiency virus type 1 capsid protein on in vitro CA assembly and infectivity. Retrovirology 4:69
Bell NM, Lever AML (2013) HIV Gag polyprotein: processing and early viral particle assembly. Trends Microbiol 21(3):136–144
Braaten D, Aberham C, Franke EK, Yin L, Phares W, Luban J (1996) Cyclosporine A-resistant human immunodeficiency virus type 1 mutants demonstrate that Gag encodes the functional target of cyclophilin A. J Virol 70(8):5170–5176
von Schwedler UK, Stray KM, Garrus JE, Sundquist WI (2003) Functional surfaces of the human immunodeficiency virus type 1 capsid protein. J Virol 77(9):5439–5450
Franke EK, Yuan HE, Luban J (1994) Specific incorporation of cyclophilin A into HIV-1 virions. Nature 372(6504):359–362
Dorfman T, Weimann A, Borsetti A, Walsh CT, Gottlinger HG (1997) Active-site residues of cyclophilin A are crucial for its incorporation into human immunodeficiency virus type 1 virions. J Virol 71(9):7110–7113
Hong SS, Boulanger P (1993) Assembly-defective point mutants of the human immunodeficiency virus type 1 Gag precursor phenotypically expressed in recombinant baculovirus-infected cells. J Virol 67(5):2787–2798
Wagner R, Leschonsky B, Harrer E, Paulus C, Weber C et al (1999) Molecular and functional analysis of a conserved CTL epitope in HIV-1 p24 recognized from a long-term nonprogressor: constraints on immune escape associated with targeting a sequence essential for viral replication. J Immunol 162:3727–3734
Dorfman T, Bukovsky A, Ohagen A, Hoglund S, Gottlinger HG (1994) Functional domains of the capsid protein of human immunodeficiency virus type 1. J Virol 68(12):8180–8187
Acknowledgements
The authors are deeply indebted to all the study subjects to carry out this study.
Author information
Authors and Affiliations
Contributions
All the authors have contributed sufficiently to carry out this study.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Informed consent
For this type of study formal consent was not required.
Research involving human participants and/or animals
All procedures performed in this study involving human participants were in accordance with the ethical standards of the NCDC ethical research committee. This was a retrospective study so formal consent was not required.
Additional information
Edited by Wolfram Gerlich.
Rights and permissions
About this article
Cite this article
Sharma, U., Gupta, S., Venkatesh, S. et al. Comparative genetic variability in HIV-1 subtype C p24 Gene in early age groups of infants. Virus Genes 54, 647–661 (2018). https://doi.org/10.1007/s11262-018-1588-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11262-018-1588-6