Abstract
Neuregulin (NRG), an epidermal growth factor is known to promote the growth of various cell types, including human melanoma cells through ErbB family of tyrosine kinases receptors. Tanapoxvirus (TPV)-encoded protein TPV-15L, a functional mimic of NRG, also acts through ErbB receptors. Here, we show that the TPV-15L protein promotes melanoma proliferation. TPV recombinant generated by deleting the 15L gene (TPVΔ15L) showed replication ability similar to that of wild-type TPV (wtTPV) in owl monkey kidney cells, human lung fibroblast (WI-38) cells, and human melanoma (SK-MEL-3) cells. However, a TPV recombinant with both 15L and the thymidine kinase (TK) gene 66R ablated (TPVΔ15LΔ66R) replicated less efficiently compared to TPVΔ15L and the parental virus. TPVΔ15L exhibited more robust tumor regression in the melanoma-bearing nude mice compared to other TPV recombinants. Our results indicate that deletion of TPV-15L gene product which facilitates the growth of human melanoma cells can be an effective strategy to enhance the oncolytic potential of TPV for the treatment of melanoma.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Melanoma is a common skin cancer with significant morbidity. It accounts for approximately 4 % of all skin cancers, but for nearly 80 % of all skin cancer-related deaths [1–3]. If not detected at the early stage, metastatic melanoma is highly aggressive with the five-year survival rate of approximately 15 % [4]. Significant progress has been made in the development of treatment options for melanoma, where current therapeutic options include surgery, chemotherapy, radiotherapy, immunotherapy, cryotherapy, and targeted drugs. Two such targeted drugs are FDA approved vemurafenib and dabrafenib (BRAF inhibitors) which inhibit a member of Raf kinase family, referred to as B-Raf. Most melanomas and nevi harbor the mutation in BRAF gene, which results in over-activation of B-Raf kinase involved in cell division. These advances have not significantly changed the survival of melanoma patients with advanced disease [5]. In the continuous quest for development of new therapeutic agents for melanoma, apoptosis-inducing agents and oncolytic viruses (OVs) are relatively new additions.
The OVs, which either selectively or preferentially infect and destroy tumor cells by lysis, and possibly by inducing an anti-tumor immune response, appear to be one of the most promising approaches to treat melanoma [6–8]. Some of the common broad strategies used during the development of OVs include increasing tumor selectivity and induction of a strong anti-cancer immune response. Infecting and replicating in tumor cells in a selective manner is an inherent feature of certain viruses. For viruses without significant native tumor specificity, oncoselectivity can usually be achieved by targeted genetic engineering to increase the viral tropism and/or the efficiency of replication in the cancerous cells [9, 10]. For example, thymidine kinase (TK) and ribonucleotide reductase (RR) are important for DNA synthesis and their levels of expression are elevated in cancerous cells as compared to normal cells [11]. Therefore, deletion of genes encoding TK and RR in OVs is a commonly used strategy to increase tumor selectivity of DNA viruses [9]. Further, OVs can be genetically modified and equipped to express immune stimulatory proteins, such as granulocyte-monocyte colony-stimulating factor (GM-CSF), macrophage chemotactic protein-1, flagellin, and interleukin 2 (IL-2), to activate the host innate and adaptive immune systems to trigger tumor regression [9, 11, 12]. Several DNA and RNA viruses, including vesicular stomatitis virus (VSV), adenovirus, and Newcastle disease virus (NDV), have been genetically modified and/or “armed” with immune-modulatory genes for use in advanced clinical trials [6, 13–16]. A genetically engineered adenovirus (H101) was approved by China’s State Food and Drug Administration (CFDA) in 2005 for treating head and neck cancer [17, 18]. Talimogene laherparepvec (T-VEC), a herpes simplex virus-1 (HSV) strain modified to express human GM-CSF, was approved by the U.S. FDA for melanoma treatment in October 2015 [19].
Tanapoxvirus (TPV), which belongs to the family Poxviridae (genus Yatapoxvirus), is a large virus containing double-stranded (ds) DNA genome (approximately 144 kbp) [20]. TPV is antigenically distinct and does not immunologically cross react with other poxviruses, including vaccinia virus (VV). It causes a mild self-limiting disease in humans, and most of the global population (with the exception of equatorial Africa) is immunologically naïve. Additionally, no man-to-man transmission has been reported [21–23]. With a large genome for genetic modification and highly attenuated virulence, TPV can potentially serve as an ideal candidate for oncolytic virotherapy. Previous studies have revealed that poxviruses employ various strategies to modulate the host anti-viral immune responses, including the secretion of cytokine and cytokine- and/or chemokine-binding receptor homologs [24–27]. It has been shown that a 45-kDa protein is encoded by the TPV-2L gene, which functions as a high-affinity inhibitor of human tumor necrosis factor (TNF) [28, 29]. Our previous studies have also shown that TPV-15L-encoded protein biologically mimics neuregulin (NRG). As one of the epidermal growth factors (EGF), NRG is the ligand for ErbB3 and ErbB4 which are associated with therapeutic resistance in many cancers, such as melanoma, breast cancer, and prostate cancer [30–33]. We have also demonstrated that TPV-15L protein is capable of binding and phosphorylating the NRG receptor heterodimer ErbB2/3 in an established NRG bioassay [34].
The ErbB family of tyrosine kinases receptors includes ErbB1 (epidermal growth factor receptor [EGFR]), ErbB2, ErbB3, and ErbB4 [35, 36]. The binding of ligands to the ErbB receptors leads to the formation of homo- or heterodimers and the activation of the intrinsic kinase domain. These activities, in turn, initiate a signal transduction cascade that ultimately leads to DNA synthesis and cell proliferation [37–39]. NRG specifically binds to ErbB3 and ErbB4 receptors. Binding of NRG to ErbB3/ErbB4 changes the conformation of receptors and induces heterodimerization with ErbB2, which results in the activation of downstream signaling [40, 41]. ErbB3 expression has been found to be enhanced in malignant melanoma cells and is associated with poor prognosis and reduced patient survival rate [42–44]. NRG-treated melanocytes show increased proliferation and invasion, altered morphology, and increased expression of progression and metastasis genes [40]. Further, it has been shown that the knock-down of NRG results in significant inhibition of melanoma growth, and those melanoma tumors treated with anti-ErbB3 antibody showed slower proliferation [45]. Recent research advances have shown success with melanoma therapies targeting activated BRAF. It has been demonstrated that NRG is highly expressed in dermal fibroblast and cancer-associated fibroblasts, and enhances the growth of RAF-inhibited BRAF V600E/D melanoma cells. Further, the safeguarding effects of fibroblast-derived NRG on melanoma cells treated with RAF inhibitors can be impeded effectively by ErbB2- and ErbB3-neutralizing antibodies. Upregulation of ErbB3 and enhanced responsiveness to NRG were observed as a form of adaptive resistance to RAF/MEK inhibitors in mutant BRAF melanoma [46–50]. These studies provide a solid theoretical basis for targeting ErbB signaling in combination with RAF inhibitors in mutant BRAF melanoma [51].
In this study, we show that the NRG mimicking protein encoded by the TPV-15L gene exhibits a similar growth-promoting effect in vitro on human mutant BRAF melanoma cells SK-MEL-3, which harbor BRAF V599E mutation and the elevated kinase activity of B-Raf. In light of this, we genetically engineered TPV with a 15L gene deletion and without/with the deletion of 66R gene that encodes TK, thus generating TPVΔ15L and TPVΔ15LΔ66R, respectively. TPVΔ66R, with 66R/TK gene deletion was generated earlier in our laboratory [11]. Deletion of the 15L gene had no significant effect on TPV replication in vitro. However, TPVΔ15LΔ66R replicated less efficiently compared to wtTPV, TPVΔ66R, and TPVΔ15L. Our in vivo study showed that TPVΔ15L regressed human melanoma tumors in nude mice more significantly compared to that of wtTPV, TPVΔ66R, or TPVΔ15LΔ66R. Taken together, our results suggest that TPVΔ15L can be a promising candidate for oncolytic virotherapy of melanoma.
Materials and methods
Cell lines, virus, and reagents
Owl monkey kidney (OMK) cells, human lung fibroblasts (WI-38), and human melanoma cell line SK-MEL-3 were purchased from American Type Culture Collection (ATCC product numbers CRL-1556, CCL-75, HTB-69 respectively). OMK and WI-38 were cultured and propagated in complete growth medium containing earle’s minimum essential medium supplemented with 10 % (v/v) fetal bovine serum (FBS) (Atlanta Biologicals), 2 mM l-glutamine (Sigma-Aldrich), and 50 μg/ml gentamicin sulfate (AMRESCO). SK-MEL-3 was cultured in growth medium consisting of McCoy’s 5A medium (Sigma-Aldrich) with 15 % FBS. Macoy’s 5A medium with no serum was used as starving medium for melanoma growth. All cell monolayers infected with virus were maintained in maintenance medium with all components being the same as the growth medium except for FBS concentration being 2 % v/v. All cell lines were incubated at 37 °C in a 5 % CO2 atmosphere. All cell counting and viability assays were conducted using an Improved Neubauer hemacytometer and 0.2 % (w/v) trypan blue in a normal saline solution. The wtTPV (Kenya strain) was originally a gift from Dr. Joseph Esposito (Centers for Disease Control, Atlanta, GA, USA). NRG1 was purchased from R&D Systems, MN.
Construction of the recombinant TPVs
A plasmid derived from a commercially available cloning vector pBluescript II KS (+) was used to include the genomic sequences flanking the right and left sides of TPV-15L open reading frame (ORF). The left flanking was generated by using polymerase chain reaction (PCR) with forward primer 5′-TAGGTACTCGAGAAAAACACCAATA-3′ (XhoI) and reverse primer 5′-GTTTAAATCGATGGACCTG-3′ (ClaI). Two primers were designed for amplifying the right flanking as forward primer 5′-CATATTTTGCGGCCGCGGTAAACAATT-3′ (NotI) and reverse primer 5′-GTTAAAAATGGAAAAGAGCTCTAATTTTAACAACAG-3′. A synthetic poxvirus early/late (E/L) promoter and a green fluorescence protein (GFP) gene were in between left and right flanking sequences. The plasmid was named as p15LGFP plasmid for generating a TPV-15L knockout virus (TPVΔ15L). This plasmid was transfected (Superfect transfection reagent; Qiagen) into 35-mm tissue culture dishes plated with OMK cell monolayer infected with wtTPV at 1 multiplicity of infection (MOI). The cells with transfection and infection were incubated at 37 °C incubator with 5 % CO2 until visible fluorescence caused by GFP expression appeared. The virus was harvested, and recombinant virus TPVΔ15L was plaque purified at least three times until no visible non-fluorescence plaques and no 15L gene were detected by confirmation PCR. Plaque assays were carried out as described earlier [52].
A second plasmid based on pBluescript II KS (+) was generated to produce a TPV-66R gene knockout virus. The left flanking and right flanking of 66R ORF were inserted in the plasmid, with an E/L synthetic promoter driving the expression of mCherry in between two flanking sequences. The left flanking was amplified using PCR with SacI forward 5′-AATGGATCACATAAAGGAGCTCTTAACG-3′ and NotI reverse 5′-CAGAAAACATGCGGCCGCATATAATCT-3′. The right flanking was generated using EcoRI forward 5′-GGAGATGAACAAGAAATAGAATTCATAGG-3′ and HindIII reverse 5′-TGTTCTTTATCACAAGCTTCTATCGGGTG-3′. The generated plasmid was named as p66RmCherry plasmid. By using TPV genomic flanking sequences for either 15L or 66R, the engineered plasmid could be guided to a specific point where a targeted ablation occurs (ablation of 15L or 66R gene) with the simultaneous replacement of a fluorescent reporter (GFP/mCherry). All plasmids were confirmed by DNA sequencing.
This p66RmCherry plasmid was transfected into OMK cell monolayer infected with wtTPV at 1 MOI. By the process of transfection and infection, the recombinant virus TPVΔ66R was generated with visible fluorescence caused by mCherry expression. The virus was harvested, and TPVΔ66R was plaque purified at least three times by using a plaque purification assay, with the selection enabled by the expression of mCherry as described earlier [52]. Also, the p66RmCherry plasmid was transfected into TPVΔ15L-infected OMK cells, and recombinant TPV with both TPV-15L and TPV-66R ablated (TPVΔ15LΔ66R) was generated. TPVΔ15LΔ66R was plaque purified with the selection by both mCherry and GFP expression. The purity of the generated recombinant viruses was determined by confirmation PCR [11]. The TPV-15L deletion was confirmed by amplifying using internal forward primer 5′-CACACCTTTTTCCGTTAAATTGCC-3′ and internal reverse primer 5′-GTTTTTTACTTTATCATGTGTCATTTTAGC-3′. TPV-66R deletion was confirmed by amplifying internal forward primer 5′-CGGTATCAAATTGCTAGGTATACTTGC-3′ and internal reverse primer 5′-TCCAATTCGTTTAGAAAACGATGCG-3′. Internal primers for TPV-136R gene are forward 5′-GTATTTATGTACTGTTTCAACTAACAAAAGC-3′ and reverse 5′-CTTTAGGTGTTAGGATATATCAATTATACAG-3′. The primers for amplifying ampicillin gene in the plasmid are forward primer 5′-CTCACGTTAAGGGATTTTGGTCATGAG-3′ and reverse primer 5′-CCGCTCATGAGACAATAACCCTG-3′.
Expression and purification of TPV-15L protein
TPV-15L gene was amplified using forward primer 5′-GCGGGATCCATGAAAAACAAATTTATG-3′ (BamHI) and reverse primer 5′-CGCTCGAGATTTACTATTTTATTTTCAC-3′ (XhoI). The 15L amplicon was cloned into pcDNA3.1/myc/His ver C (Invitrogen), producing a fusion-tagged construct which was subsequently cloned into the pFastBac-Dual-eGFP cassette. Recombinant baculovirus was generated using Bac-to-Bac system (Invitrogen). The recombinant baculovirus was used to infect sf21 insect cells and the cell supernatant was collected, and TPV-15L protein was purified using a hexa-His Co2+ chelate resin affinity column (BD Sciences) routinely employed in our laboratory [34].
Growth of human melanoma cells in the presence of TPV-15L protein
Human melanoma cells (SK-MEL-3) were plated in a 96 well plate in McCoy’s 5A medium with 15 % FBS and incubated overnight at 37 °C with 5 % CO2 in humidified chambers. The next day, cells were switched to the serum-free McCoy’s 5A medium and incubated for 24 h before the addition of NRG or TPV-15L protein. Cells were treated with either TPV-15L purified protein (500 ng/ml) or NRG (500 ng/ml). Untreated cells were used as mock. Total number of viable cells was determined in each group at day 1, 3, and 5 post-treatment, by counting cells on a hemocytometer chamber using trypan blue stain to exclude non-viable cells. Each experiment was repeated three times independently and standard deviations were calculated.
Replication of TPV recombinants
Virus replication was determined in OMK, WI-38, and SK-MEL-3 cells. Cell monolayers planted in 24-well plates were infected with wtTPV or one of the recombinants of TPV (TPV∆66R, TPV∆15L, and TPV∆15L∆66R) at 0.1 and 5 MOI. Infected cells were harvested at 49, 96, and 240 h post infection (hpi), and total virus titer was determined by viral plaque assay on OMK monolayers in 6-well plates. Each experiment was repeated three times.
Animals
Male athymic nude mice (Crl:NU(NCr)-Foxn1nu) were purchased from Charles River company (Wilmington, Massachusetts, USA) at 6–8 weeks of age. Mice were housed individually with food and water available in pathogen-free animal facility at Western Michigan University. Animals were allowed to acclimatize for 1 week before experimentation. All protocols for treatment and manipulations were approved by the Institutional Animal Care and Use Committee of Western Michigan University (IACUC protocol number 13-07-01).
Virotherapy of human melanoma tumor xenografts
Melanoma tumors were induced in nude mice by injecting 5 × 106 viable SK-MEL-3 cells suspended in 100 µl of phosphate saline buffer (PBS), subcutaneously on the dorsal surface in the inter-scapular region. Cell viability assays were conducted using trypan blue following each injection to test the cell viability. Tumor volume was calculated by using the digital calipers to measure three-dimensionally according to the formula [4/3 × 3.1415 × (length/2) × (width/2) × (height/2)] in mm3. Mice were randomly segregated into control and experimental groups, when the tumor size reached 45 ± 4.5 mm3, with each group containing five mice (n = 5). The number of animals per group was determined by power analysis, and n of 5 is standard procedure for in vivo experiments including quantitative pathology. Each mouse in experimental group received a single intratumoral injection of 5 × 106 pfu virus particles in 100 µl medium. Animals in mock group received 100 µl medium only. The day of treatment administration was considered as day 0, and mouse weights and tumor volumes were measured and recorded every day. All the mice were sacrificed at 35 days post-treatment, and the tumors were harvested. The blood sample of each mouse was collected and centrifuged for the collection of the serum. Tumors were weighed and stored for histological analyses.
Statistical analysis
All in vitro experiments were done in triplicates and the measurements were presented as mean ± SD. The two-sample unequal variance Student’s t test analysis was applied for testing the differences. The significance level used was P < 0.05. Student’s t-test was used for assessing the treatment effect of each experimental virotherapy group compared to the mock group in vivo. The tumor volumes of each experimental group were compared with those in the mock group, and significance was considered if P < 0.05.
Results
TPV-15L protein and NRG increase melanoma cell proliferation
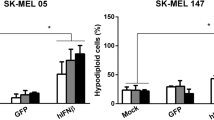
We have shown that TPV-15L is a secreted early protein that phosphorylates NRG receptors and binds to heparin [34]. Since NRG has also been shown to promote melanoma progression and metastasis [40], we sought to investigate if TPV-15L protein could have a similar effect on melanoma cells. SK-MEL-3 cells in the serum-free medium were treated with NRG or TPV-15L protein (500 ng/ml) and incubated for 1, 3, and 5 days. Cells in the control wells received no treatment. Compared to control, cell numbers of those treated with NRG and TPV-15L were significantly higher as early as on day 1 (P < 0.05). At day 3, SK-MEL-3 cells treated with NRG and TPV-15L increased by 110 and 86 %, respectively, demonstrating continued proliferation, while those with no treatment achieved only 2.5 % growth. At day 5, NRG- and TPV-15L-treated cells achieved 125 and 120 % growth, respectively, while those with no treatment gained only 69 % growth (Fig. 1). These results provide compelling evidence that TPV-15L protein, like NRG, promoted melanoma growth. The proliferative efficacy of TPV-15L was indistinguishable from NRG (P < 0.05).
Effect of TPV-15L protein and NRG on the proliferation of human melanoma SK-MEL-3 cells. Each well in 96-well plate was planted with 2 × 104 SK-MEL-3 cells. Growth medium was replaced with serum-free (starving) medium containing purified TPV-15L protein (500 ng/ml) or NRG (500 ng/ml). Viable cells were counted on day 1, 3, and 5 following treatment. Untreated cells served as mock. Each experiment was repeated three times independently and standard deviations are shown
Generation of TPVΔ15L and TPVΔ15LΔ66R
In the light of functional similarities between TPV-15L protein and NRG in enhancing melanoma proliferation, we sought to delete 15L gene from TPV (TPVΔ15L) and to test as an OV for melanoma virotherapy. The recombinant virus TPVΔ15L was plaque purified at least three times until no visible wtTPV plaques appeared. Deletion of 15L ORF was then confirmed by PCR. Similarly, ORF 66R which encodes TK was deleted from TPVΔ15L. TK is present in abundant quantities in cancerous cells and its deletion from virus has widely been used to enhance tumor selectivity of DNA viruses [9]. For this purpose, TPVΔ15L-infected cells were transfected with p66R-mCherry plasmid-containing mCherry reporter gene under E/L synthetic promoter. The resulting TPVΔ15LΔ66R was plaque purified at least three times as previously described. The exhibition of both mCherry and GFP was confirmed by fluorescence microscopy. The deletions of both 15L and 66R genes were further confirmed by PCR, using primers described in Materials and Methods. Figure 2 shows that wtTPV contained both 15L and 66R ORFs, while ORF 15L was deleted from TPVΔ15L. Similarly, ORF 15L and ORF 66R were absent in TPVΔ15LΔ66R. ORF 136R was used as a positive control in these experiments and was present in all three viruses, namely wtTPV, TPVΔ15L, and TPVΔ15LΔ66R. Ampicillin gene contained in plasmid was amplified in all three viruses to exclude single-crossover recombinants. These experiments provide compelling evidence that 15L and/or 66R gene were deleted, and that these viruses were generated as a result of double-crossover recombination events during transfection and infection.
Verification of gene ablations in TPV recombinants. PCRs were used for verifying the ablations of 15L and 66R genes. Genomic DNA from each virus was isolated and probed for sequences internal in 15L, 66R, 136R, and ampicillin genes. Genes ablated showed no bands in the gel. TPV-15L internal fragment is 197 bp; 66R region is 379 bp; and 136R internal fragment used as a control has a predicted amplicon length of 531 bp. Ampicillin gene was probed for determining the insertion of ampicillin or other genes from plasmid into viral DNA during the transfection–infection process when homologous recombination occurs. Lanes: wtTPV (control); TPV∆15L (15L is ablated); TPV∆15L∆66R (both 15L and 66R are ablated). Moreover, no insertion of ampicillin gene is shown in the viral DNA of TPV∆15L and TPV∆15L∆66R
Replication of TPV recombinants in human and monkey cells
Replication curves of wtTPV, TPVΔ66R, TPVΔ15L, and TPVΔ15LΔ66R were generated using 0.1 and 5 MOI in OMK cells and human fetal lung fibroblasts (WI-38) and are shown in Fig. 3. The infected cells were harvested, and virus was titrated on OMK monolayers at 48, 96, and 240 hpi. The first time point for harvesting (48 hpi) was decided based on the eclipse phase period of TPV which is around 48 hpi, as TPV unlike VV is a slow replicating virus [52]. Results in Fig. 3 demonstrate that wtTPV and recombinant TPVs achieved the maximum titer at around 96 hpi as expected [34, 52, 53]. At 0.1 and 5 MOI in OMK cells, TPVΔ15L and wtTPV exhibited similar replication kinetics. At 0.1 MOI, the yield of TPVΔ15LΔ66R was time dependent, with the titer being lower compared to wtTPV and TPVΔ66R at 48 hpi. By 96 hpi, however, the titer had attained levels similar to those of wtTPV and TPVΔ66R. At 5 MOI on OMK cells, the replication of TPVΔ15LΔ66R was not significantly lower compared to wtTPV and TPVΔ66R. At 0.1 MOI in WI-38 cells, TPVΔ15L showed similar replication kinetics as wtTPV and more effective replication compared to TPVΔ66R. TPVΔ15LΔ66R exhibited less efficient replication compared to TPVΔ15L, with the maximum yield at 96 hpi being 7-8 fold lower compared to that of TPVΔ15L. When infecting WI-38 cells at 5 MOI, TPVΔ15L, TPVΔ66R, and TPVΔ15LΔ66R showed similar titers which were lower compared to those of wtTPV. The above data supports the earlier results that (1) TPV-15L is non-essential for the replication of TPV in cell culture, and that (2) deletion of TPV-66R adversely affects the viral replication.
TPV∆15L and TPV∆15L∆66R replication compared to wtTPV and TPV∆66R. OMK and WI-38 cell monolayers in 24-well plates were infected with wtTPV, TPV∆66R, TPV∆15L, and TPV∆15L∆66R at 0.1 and 5 MOI. The virus was harvested at 48, 96, and 240 hpi and titrated on OMK monolayers. Each experiment was repeated three times independently. a Viral infection at 0.1 MOI in OMK cells b Viral infection at 0.1 MOI in WI-38 cells c Viral infection at 5 MOI in OMK cells d Viral infection at 5 MOI in WI-38 cells
Replication of TPVΔ15L and TPVΔ15LΔ66R in SK-MEL-3 cells
The replication characteristics of TPVΔ15L and TPVΔ15LΔ66R were also examined in BRAF mutant melanoma SK-MEL-3, and compared with wtTPV and TPVΔ66R. SK-MEL-3 cells were infected at 0.1 and 5 MOI and the total virus titer was determined at 48, 96, and 240 hpi. As shown in Fig. 4, at 0.1 MOI, the replication of TPVΔ15L displayed a slight delay in the onset of the replication. However, the virus titer of TPVΔ15L reached a similar level as wtTPV at 96 hpi. In contrast, TPVΔ66R and TPVΔ15LΔ66R replicated less efficiently compared to wtTPV and TPVΔ15L. At 96 hpi, the titer of TPVΔ15LΔ66R was fivefold to sixfold lower compared to TPVΔ15L and the titer of TPVΔ66R was threefold to fourfold lower compared to that of TPVΔ15L. At 5 MOI, TPVΔ15L exhibited similar replication kinetics with wtTPV. At 96 hpi, virus titers reached the maximum, with the titer of TPVΔ15L being twofold to threefold higher compared to that of TPVΔ66R and threefold to fourfold higher compared to that of TPVΔ15LΔ66R. The results showed that deletion of 66R gene but not 15L gene resulted in a significant delay of TPV replication in SK-MEL-3 melanoma cells in vitro.
Replication of TPV∆15L and TPV∆15L∆66R in melanoma cells. SK-MEL-3 cell monolayers in 24-well plates were infected with wtTPV, TPV∆66R, TPV∆15L, and TPV∆15L∆66R at a 0.1 MOI and b 5 MOI. The virus was harvested at 48, 96, and 240 hpi and titrated on OMK monolayers. Each experiment was repeated three times independently
Oncolytic virotherapy of human melanoma tumors xenografted in nude mice
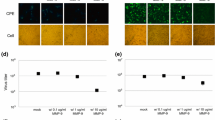
To evaluate the oncolytic effectiveness of wtTPV and TPV recombinants, human melanoma tumors were induced by injecting BRAF mutant SK-MEL-3 cells subcutaneously on the dorsal surface in the inter-scapular region of athymic nude mice. Treatments consisted of intratumoral mock or virus injections, when the tumor volumes reached 45 ± 4.5 mm3. While mock-treated tumor xenografts continued to grow exponentially, xenografts treated with viruses exhibited continued but slow tumor growth. As shown in Fig. 5, there were significant differences in the tumor growth between virus-treated groups and mock group (P < 0.05). Compared to mock-treated tumors achieving nearly 552 % growth, tumors treated with wtTPV and TPVΔ66R displayed 250 and 244 % growth, respectively, after 35 days of treatment. TPVΔ15L produced significantly greater tumor growth delay compared to wtTPV and TPVΔ66R (P < 0.05). At 35 days post-treatment, TPVΔ15L-treated tumors exhibited only 68 % growth compared to day 0. Interestingly, TPVΔ15LΔ66R showed markedly less tumor growth inhibition efficacy (286 % tumor growth) compared to TPVΔ15L, perhaps due to the poor replication of the virus when ORF 66R was deleted (Figs. 3, 4). These results showed that TPVΔ15L was able to significantly reduce the tumor progression at more time points in comparison to the mock than other TPVs. We believe that the enhanced oncolytic potential of TPVΔ15L was due to deletion of TPV-15L gene which encodes a viral protein that contributes to melanoma proliferation, as demonstrated by our in vitro studies. As TPV replicates only in humans and monkeys, nude mice with xenografted melanoma tumors were not suitable for studies related to the bio-distribution and safety of the virus.
Effectiveness of mutant viruses in tumor suppression in melanoma tumor models in vivo. Melanoma tumors were induced in 6–8 week old athymic nude mice by subcutaneously injecting 5 × 106 SK-MEL-3 cells onto the dorsal surface. Mice were randomly segregated into the control or experimental groups with each group containing five mice (n = 5) when the tumor size reached 45 ± 4.5 mm3. Mice in the experimental groups were treated with intratumoral injections of 5 × 106 pfu of virus, while the mice in the mock group were intratumorally injected with medium only. Tumor volumes were measured using the digital calipers every day. In each graph shown here, the y-axis represents the average percentage of tumor growth (%) and the x-axis is time (days post virotherapeutic treatment). All experimental groups are compared to the mock group in which the mice were treated with only medium injection. Bars show the standard error of the mean (± 1 SEM). Points indicated with an asterisk (*) refer to tumor growth that was significantly reduced from the control (P ≤ 0.05). Asterisks were marked every 3 days if there was significance. a wtTPV b TPV∆66R c TPV∆15L d TPV∆15L∆66R
Discussion
In the search for novel anti-cancer strategies, OVs have emerged as one of the most promising therapeutic options to eliminate cancer cells. They also overcome tumor-induced immune evasion in host and elicit an effective anti-tumor immune response. Viruses which are under evaluation for therapeutic purposes can be divided into two major groups: (a) viruses that possess a degree of innate oncoselectivity, such as reovirus, VSV, and NDV [10, 54–56], and (b) viruses which require genetic modifications to enhance their tumor selectivity and immune stimulatory capability, such as VV with ablation of TK gene [57, 58]. Although OVs have shown encouraging results in experimental and clinical trials, the efficacy of an OV is still potentially limited by several factors. First, the anti-viral immune response developed by the host immune system, such as the induction of neutralizing anti-viral cellular and humoral immune responses, inactivates the virus and impedes the effectiveness of virus administered systemically. The “barriers,” such as neutralization and sequestration of the virus, make it uncertain for OVs to be “one shot” therapeutic agents, and also make that virus less effective during successive treatment cycles. Strategies have been devised to prime the immune response with one viral vector and boost with another vector, in order to focus the immune response on the tumor antigens and avoid anti-viral immune responses [59, 60]. Although an effective virus-specific immune tolerance strategy will ultimately solve this issue, presently it may be necessary to have a set of antigenically distinct OVs for serial injections to avoid this problem. Second, the safety profile of a virus determines its feasibility as the virotherapy, and transmission from human to human in combination of continued virus evolution pose a serious concern for virus application. Only a small number of viruses, such as HSV and adenovirus, have been demonstrated as safe for application in clinical trials [54]. In light of these concerns, TPV appears to be an ideal OV candidate due to its characteristics. These include: (1) most of the global population is immunologically naïve to TPV, (2) TPV causes a mild, self-limiting febrile disease in humans making it relatively safer to use, (3) TPV is not transmitted from human to human, and (4) the large genome size of TPV (144 kbp) allows incorporation of several immune stimulatory transgenes.
At present, advanced malignant melanoma is one of the diseases with very few effective treatments. Although significant advances have been made in monitoring and treatment of melanoma, the mortality associated with the disease remains largely un-altered [5]. Due to the limited efficiency and significant side effects of chemotherapies, a variety of immunological approaches have been devised, among which interferon-α (IFN-α) and IL-2 have shown significant efficacy and have been approved by U.S. FDA for melanoma treatment [61–63]. The OVs appear to be an appealing addition to melanoma therapies, due to the viral oncoselectivity and lysis of tumor cells [13]. Combination of virotherapy and other therapies, such as chemotherapy, have also shown synergistic effects [64].
Our previous studies of TPV genomic analyses revealed that TPV-66R gene encodes TK, and that TPV-15L gene encodes EGF-like growth factor often referred to as NRG [34, 65]. Further, we have demonstrated that TPV-15L protein functions as a mimetic of NRG, capable of binding and phosphorylating the most potent NRG heterodimer receptor ErbB2/3 [34]. Elevated NRG levels have been shown to contribute to melanoma proliferation by binding to ErbB3, while melanoma treated with anti-ErbB3 has shown reduced cell proliferation [30, 45]. Thus, it would be interesting and worthwhile to further explore the application of neutralizing agents against ErbB receptors or NRG for melanoma therapy.
A variety of virokines carrying EGF-like sequence, such as vaccinia virus growth factor and myxoma virus growth factor, have been demonstrated to act on ErbB receptors and promote cell proliferation [66, 67]. In this study, we showed that TPV-15L protein and NRG both significantly increased the proliferation of the human melanoma cells (SK-MEL-3) in vitro. This further supports our earlier claim about the functional similarity between TPV-15L and NRG. In the light of these findings, we deleted TPV-15L gene both with and without the deletion of TPV-66R (TK) gene, generating two TPV recombinants TPVΔ15L and TPVΔ15LΔ66R. The replication kinetics of TPVΔ15L and TPVΔ15LΔ66R was studied in OMK, WI-38, and SK-MEL-3 cells. We demonstrated that TPVΔ15L had similar replication efficacy as wtTPV and that TPVΔ15LΔ66R replicated less efficiently compared to TPVΔ15L and wtTPV. The data supports our earlier results showing that TPV-15L is a non-essential gene for TPV replication and that TPV-66R exerts more significant effect on viral replication [34].
We further studied whether deletion of TPV-15L gene (encoding NRG mimicking protein) will abolish the growth-promoting effect of TPV-15L protein on human melanoma cells (SK-MEL-3), and enhance oncolytic efficacy of TPV in melanoma-bearing nude mice. Tumor xenografts were established using SK-MEL-3 cells, which were treated with intratumoral injection of wtTPV, or one of the TPV recombinants (TPVΔ66R or TPVΔ15L or TPVΔ15LΔ66R). Our results demonstrated that TPVΔ15L exhibited a more robust tumor reduction compared to wtTPV, TPVΔ66R, and TPVΔ15LΔ66R. TPVΔ15LΔ66R was less effective in regressing tumor, possibly due to its slower replication, which has been demonstrated in vitro. The greater tumor reduction efficacy of TPVΔ15L compared with wtTPV and other TPV recombinants suggests that (1) the reduced melanoma proliferation possibly relates to the absence of tumor-enhancing properties caused by TPV-15L protein, and that (2) deletion of viral genes encoding NRG-like proteins is potentially an effective strategy to be used in genetic engineering of OVs for melanoma. Considering the pathological power of abnormal ErbB signaling and its contribution to oncogenesis in cancer biology, deletion of EGF-like growth factor genes from certain OVs and/or blocking the ErbB receptors would likely offer additional tumor suppression effects for oncolytic virotherapies. Similarly, Saydam et al. [68] have shown that HSV expressing small interference RNA (siRNA) directed against EGFR inhibits human glioblastoma cell growth more significantly compared to the control virus both in vitro and in vivo.
The results of this study warrant further investigation of TPVΔ15L as an effective vector for oncolytic virotherapy, while more strategies could be incorporated to further enhance its oncolytic efficacy in the therapy of melanoma and other cancers. TPV-2L gene encodes a 38 kDa secretory protein which binds and neutralizes TNFα [29] with high affinity [28]. It has been shown that TNF downregulates the expression of ErbB2 in pancreatic tumor cells, which demonstrates the inverse relation between TNF and ErbB2 [69]. In addition, ErbB2 overexpression induces resistance of breast cancer cells to the cytotoxic effect of TNFα [70]. Therefore, it would be appropriate to consider the construction of an oncolytic TPV recombinant with both 15L and 2L genes ablated, which will synergistically decrease the ErbB-related tumor proliferation and increase the viral oncolysis. Also, it is conceivable to delete TPV-136R gene encoding type I IFN binding receptor from TPVΔ15L backbone, as host IFN will potentially be “saved” from being neutralized and exert antiproliferative efficacy.
Coupling viral oncolytic activity with immune cells recruitment proves to be another attractive strategy for attaining the optimal clinical efficacy. Oncolytic VV JX-594 and more recently Amgen’s oncolytic HSV, T-vec, are both armed with GM-CSF, suggesting the clinical benefit of enhancing host anti-tumor immunity [8]. In addition to virotherapies expressing GM-CSF, many other viruses expressing cytokines, such as IL-2, IL-12, and IL-24, have shown encouraging tumor regression efficacy [9, 71, 72]. Other strategies, such as promoting virus spread and increasing cytotoxicity, have also been applied in OV modification. Engineered virus expressing relaxin and decorin, known as extracellular matrix-modulating proteins, have shown enhanced penetration, persistence, and spread compared to the control virus in melanoma, glioma, and lung cancer [73, 74]. Introduction of suicide genes, such as FCU1-encoding bifunctional fusion protein which converts non-toxic 5-FC to toxic 5-fluorouracil and 5-fluorouridine monophosphate, has also been proven to be a feasible strategy for enhancing OV’s effectiveness in different cancers [75]. Based on these findings and the oncolytic efficacy of TPVΔ15L shown in this study, we speculate that the modifications described above to TPV or TPVΔ15L may further improve the outcome of TPV as an oncolytic virotherapy.
References
H. Tsao, M.B. Atkins, A.J. Sober, New Engl. J. Med. 351(10), 998–1012 (2004)
T.M. Johnson, A.M. Yahanda, A.E. Chang, D.J. Fader, V.K. Sondak, J. Am. Acad. Dermatol. 38(5), 731–741 (1998)
B.S. Kim, H. Seo, H.J. Kim, S.M. Bae, H.N. Son, Y.J. Lee, S. Ryu, R.W. Park, J.O. Nam, J. Med. Food 18(10), 1121–1127 (2015)
A. Sandru, S. Voinea, E. Panaitescu, A. Blidaru, J. Med. Life 7(4), 572–576 (2014)
E. Maverakis, L.A. Cornelius, G.M. Bowen, T. Phan, F.B. Patel, S. Fitzmaurice, Y. He, B. Burrall, C. Duong, A.M. Kloxin, H. Sultani, Acta Derm. Venereol. 95(5), 516–527 (2015)
R.M. Eager, J. Nemunaitis, Cancer Gene Ther. 18(5), 305–317 (2011)
E.A. Chiocca, Nat. Rev. Cancer 2(12), 938–950 (2002)
S.J. Russell, K.W. Peng, J.C. Bell, Nat. Biotechnol. 30(7), 658–670 (2012)
H. Zhao, M. Janke, P. Fournier, V. Schirrmacher, Virus Res. 136(1), 75–80 (2008)
A.L. Angelova, K. Geletneky, J.P. Nüesch, J. Rommelaere, Front. Bioeng. Biotechnol. 3, 55 (2015)
S.J. Conrad, M. El-Aswad, E. Kurban, D. Jeng, B.C. Tripp, C. Nutting, R. Eversole, C. Mackenzie, K. Essani, J. Exp. Clin. Cancer Res. 34(1), 1–15 (2015)
K. Ikeda, T. Ichikawa, H. Wakimoto, J.S. Silver, T.S. Deisboeck, D. Finkelstein, G.R. Harsh, D.N. Louis, R.T. Bartus, F.H. Hochberg, E.A. Chiocca, Nat. Med. 5(8), 881–887 (1999)
A.U. Blackham, S.A. Northrup, M. Willingham, R.B. D’Agostino, D.S. Lyles, J.H. Stewart, Surgery 153(3), 333–343 (2013)
A. Sorensen, R.J. Mairs, L. Braidwood, C. Joyce, J. Conner, S. Pimlott, M. Brown, M. Boyd, J. Nucl. Med. 53(4), 647–654 (2012)
F.L. Bai, Y.H. Yu, H. Tian, G.P. Ren, H. Wang, B. Zhou, X.H. Han, Q.Z. Yu, D.S. Li, Cancer Biol. Ther. 15(9), 1226–1238 (2014)
S. Tayeb, Z. Zakay-Rones, A. Panet, Oncolytic Virother 4, 49–62 (2015)
R. Alemany, Mol. Asp. Med. 28(1), 42–58 (2007)
C. Larson, B. Oronsky, J. Scicinski, G.R. Fanger, M. Stirn, A. Oronsky, T.R. Reid, Oncotarget 6(24), 19976 (2015)
E.S. Appleton, S. Turnbull, C. Ralph, E. West, K. Scott, K. Harrington, H. Pandha, A. Melcher, Expert Opin. Biol. Ther. 15(10), 1517–1530 (2015)
C.R. Brunetti, H. Amano, Y. Ueda, J. Qin, T. Miyamura, T. Suzuki, X. Li, J.W. Barrett, G. McFadden, J. Virol. 77(24), 13335–13347 (2003)
A.W. Downie, C.H. Taylor-Robinson, A.E. Caunt, G.S. Nelson, P.E.C. Manson-Bahr, T.C. Matthews, Br. Med. J. 1(5745), 363–368 (1971)
S.H. Nazarian, J.W. Barrett, M.M. Stanford, J.B. Johnston, K. Essani, G. McFadden, Virology 368(1), 32–40 (2007)
Z. Jezek, I. Arita, M. Szczeniowski, K.M. Paluku, K. Ruti, J.H. Nakano, Bull. World Health Organ. 63(6), 1027 (1985)
B.T. Seet, J.B. Johnston, C.R. Brunetti, J.W. Barrett, H. Everett, H.C. Cameron, J. Sypula, S.H. Nazarian, A. Lucas, G. McFadden, Annu. Rev. Immunol. 21(1), 377–423 (2003)
A. Alcami, U.H. Koszinowski, Immunol. Today 21(9), 447–455 (2000)
D. Tortorella, B.E. Gewurz, M.H. Furman, D.J. Schust, H.L. Ploegh, Annu. Rev. Immunol. 18(1), 861–926 (2000)
B. Moss, J.L. Shisler, Semin. Immunol. 13(1), 59–66 (2001)
C.R. Brunetti, M. Paulose-Murphy, R. Singh, J. Qin, J.W. Barrett, A. Tardivel, P. Schneider, K. Essani, G. McFadden, Proc. Natl. Acad. Sci. USA 100(8), 4831–4836 (2003)
M. Paulose-Murphy, B.L. Bennett, A.M. Manning, K. Essani, Microb. Pathog. 25, 33–41 (1998)
J.A. Engelman, K. Zejnullahu, T. Mitsudomi, Y. Song, C. Hyland, J.O. Park, N. Lindeman, C.M. Gale, X. Zhao, J. Christensen, T. Kosaka, Science 316(5827), 1039–1043 (2007)
Y. Zhang, D. Linn, Z. Liu, J. Melamed, F. Tavora, C.Y. Young, A.M. Burger, A.W. Hamburger, Mol. Cancer Ther. 7(10), 3176–3186 (2008)
C. Desbois-Mouthon, A. Baron, M.J. Blivet-Van Eggelpoël, L. Fartoux, C. Venot, F. Bladt, C. Housset, O. Rosmorduc, Clin. Cancer Res. 15(17), 5445–5456 (2009)
C.H. Kugel, E.J. Hartsough, M.A. Davies, Y.Y. Setiady, A.E. Aplin, Cancer Res. 74(15), 4122–4132 (2014)
D. Jeng, Z. Ma, J.W. Barrett, G. McFadden, J.A. Loeb, K. Essani, J. Virol. 87(6), 3018–3026 (2013)
M. Sibilia, R. Kroismayr, B.M. Lichtenberger, A. Natarajan, M. Hecking, M. Holcmann, Differentiation 75(9), 770–787 (2007)
R. Pinkas-Kramarski, L. Soussan, H. Waterman, G. Levkowitz, I. Alroy, L. Klapper, S. Lavi, R. Seger, B.J. Ratzkin, M. Sela, Y. Yarden, EMBO J. 15(10), 2452 (1996)
J.H. Fallon, K.B. Seroogy, S.E. Loughlin, R.S. Morrison, R.A. Bradshaw, D.J. Knaver, D.D. Cunningham, Science 224(4653), 1107–1109 (1984)
Y. Yarden, M.X. Sliwkowski, Nat. Rev. Mol. Cell Biol. 2(2), 127–137 (2001)
J. Schlessinger, Cell 110(6), 669–672 (2002)
K. Buac, M. Xu, J. Cronin, A.T. Weeraratna, S.M. Hewitt, W.J. Pavan, Pigment Cell Melanoma Res. 22(6), 773–784 (2009)
D.N. Amin, K. Hida, D.R. Bielenberg, M. Klagsbrun, Cancer Res. 66(4), 2173–2180 (2006)
M. Reschke, D. Mihic-Probst, E.H. Van Der Horst, P. Knyazev, P.J. Wild, M. Hutterer, S. Meyer, R. Dummer, H. Moch, A. Ullrich, Clin. Cancer Res. 14(16), 5188–5197 (2008)
L. Xu, S.S. Shen, Y. Hoshida, A. Subramanian, K. Ross, J.P. Brunet, S.N. Wagner, S. Ramaswamy, J.P. Mesirov, R.O. Hynes, Mol. Cancer Res. 6(5), 760–769 (2008)
S. Tiwary, M. Preziosi, P.G. Rothberg, N. Zeitouni, N. Corson, L. Xu, Oncogenesis 3(7), e110 (2014)
K. Zhang, P. Wong, L. Zhang, B. Jacobs, E.C. Borden, J.C. Aster, B. Bedogni, Oncogene 31(43), 4609–4618 (2012)
E.V. Abel, K.J. Basile, C.H. Kugel, A.K. Witkiewicz, K. Le, R.K. Amaravadi, G.C. Karakousis, X. Xu, W. Xu, L.M. Schuchter, J.B. Lee, J. Clin. Invest. 123(5), 2155 (2013)
C. Capparelli, S. Rosenbaum, A.C. Berger, A.E. Aplin, J. Biol. Chem. 290(40), 24267–24277 (2015)
H. Davies, G.R. Bignell, C. Cox, P. Stephens, S. Edkins, S. Clegg, J. Teague, H. Woffendin, M.J. Garnett, W. Bottomley, N. Davis, Nature 417(6892), 949–954 (2002)
E.V. Abel, A.E. Aplin, Cancer Res. 70(7), 2891–2900 (2010)
K.J. Basile, E.V. Abel, A.E. Aplin, Oncogene 31(19), 2471–2479 (2012)
K. Tworkoski, G. Singhal, S. Szpakowski, C.I. Zito, A. Bacchiocchi, V. Muthusamy, M. Bosenberg, M. Krauthammer, R. Halaban, D.F. Stern, Mol. Cancer Res. 9(6), 801–812 (2011)
S. Mediratta, K. Essani, Can. J. Microbiol. 45(1), 92–96 (1999)
H.L. Lee, K. Essani, Open Virol. J. 4, 1 (2010)
B. Everts, H.G. van der Poel, Cancer Gene Ther. 12(2), 141–161 (2005)
T.C. Liu, D. Kirn, Gene Ther. 15(12), 877–884 (2008)
T.C. Liu, D. Kirn, Cancer Res. 67(2), 429–432 (2007)
H.J. Zeh, D.L. Bartlett, Cancer Gene Ther. 9(12), 1001–1012 (2002)
K.A. Parato, C.J. Breitbach, F. Le Boeuf, J. Wang, C. Storbeck, C. Ilkow, J.S. Diallo, T. Falls, J. Burns, V. Garcia, F. Kanji, Mol. Ther. 20(4), 749–758 (2012)
J.L. Marshall, R.J. Hoyer, M.A. Toomey, K. Faraguna, P. Chang, E. Richmond, J.E. Pedicano, E. Gahen, R.A. Peck, P. Arlen, K.Y. Tsang, J. Schlom, J. Clin. Oncol. 18(23), 3964–3973 (2000)
K.R. Irvine, R.S. Chamberlain, E.P. Shulman, D.R. Surman, S.A. Rosenberg, N.P. Restifo, J. Natl. Cancer Inst. 89(21), 1595–1601 (1997)
W. Sun, L.M. Schuchter, Curr. Treat. Options Oncol. 2(3), 193–202 (2001)
J.M. Kirkwood, M.H. Strawderman, M.S. Ernstoff, T.J. Smith, E.C. Borden, R.H. Blum, J. Clin. Oncol. 14(1), 7–17 (1996)
J.A. Sparano, R.I. Fisher, M. Sunderland, K. Margolin, M.L. Ernest, M. Sznol, M.B. Atkins, J.P. Dutcher, K.C. Micetich, G.R. Weiss, J. Clin. Oncol. 11(10), 1969–1977 (1993)
M. Aghi, R.L. Martuza, Oncogene 24(52), 7802–7816 (2005)
S.H. Nazarian, J.W. Barrett, J.W.A.M. Frace, M. Olsen-Rasmussen, M. Khristova, M. Shaban, S. Neering, Y. Li, I.K. Damon, J.J. Esposito, K. Essani, Virus Res. 129(1), 11–25 (2007)
A. Opgenorth, N. Nation, K. Graham, G. McFadden, Virology 192, 701–709 (1993)
D.R. Twardzik, J.P. Brown, J.E. Ranchalis, G.J. Todaro, B. Moss, Proc. Natl. Acad. Sci. USA 82, 5300–5304 (1985)
O. Saydam, D.L. Glauser, I. Heid, G. Turkeri, M. Hilbe, A.H. Jacobs, M. Ackermann, C. Fraefel, Mol. Ther. 12(5), 803–812 (2005)
H. Kalthoff, C. Roeder, J. Gieseking, I. Humburg, W. Schmiegel, Proc. Natl. Acad. Sci. USA 90(19), 8972–8976 (1993)
R.M. Hudziak, G.D. Lewis, M.R. Shalaby, T.E. Eessalu, B.B. Aggarwal, A. Ullrich, H.M. Shepard, Proc. Natl. Acad. Sci. USA 85(14), 5102–5106 (1988)
S. Varghese, S.D. Rabkin, R. Liu, P.G. Nielsen, T. Ipe, R.L. Martuza, Cancer Gene Ther. 13(3), 253–265 (2006)
L. Zhao, J. Gu, A. Dong, Y. Zhang, L. Zhong, L. He, Y. Wang, J. Zhang, Z. Zhang, J. Huiwang, Q. Qian, Hum. Gene Ther. 16(7), 845–858 (2005)
I.K. Choi, Y.S. Lee, J.Y. Yoo, A.R. Yoon, H. Kim, D.S. Kim, D.G. Seidler, J.H. Kim, C.O. Yun, Gene Ther. 17(2), 190–201 (2010)
J.W. Choi, J.S. Lee, S.W. Kim, C.O. Yun, Adv. Drug Deliv. Rev. 64(8), 720–729 (2012)
J.D. Dias, I. Liikanen, K. Guse, J. Foloppe, M. Sloniecka, I. Diaconu, V. Rantanen, M. Eriksson, T. Hakkarainen, M. Lusky, P. Erbs, Clin. Cancer Res. 16(9), 2540–2549 (2010)
Acknowledgments
This study was partially supported by a National Institutes of Health (NIH) grant 1R15CA156262-01 to KE. The authors are grateful to Drs. Chris Fisher and John Spitsbergen for their suggestions and editorial comments.
Author contributions
Conception or design of the work—Tiantian Zhang and Karim Essani. Data collection—Tiantian Zhang, Yogesh R. Suryawanshi, Dennis H. Kordish and Helene M. Woyczesczyk. Data analysis and interpretation—Tiantian Zhang, Yogesh R. Suryawanshi, David Jeng and Karim Essani. Drafting the article—Tiantian Zhang, Yogesh R. Suryawanshi, David Jeng and Karim Essani. Critical revision of the article—Tiantian Zhang, Yogesh R. Suryawanshi, David Jeng and Karim Essani. Final approval of the version to be published—Tiantian Zhang, Yogesh R. Suryawanshi, Dennis H. Kordish, Helene M. Woyczesczyk, David Jeng and Karim Essani.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Ethical approval
Prior to initiation, all study protocols for treatment and manipulations of mice were approved by the Institutional Animal Care and Use Committee of Western Michigan University (IACUC protocol number 13-07-01).
Informed consent
Informed consent was obtained from all individual participants included in the study.
Additional information
Edited by Keizo Tomonaga.
Rights and permissions
About this article
Cite this article
Zhang, T., Suryawanshi, Y.R., Kordish, D.H. et al. Tanapoxvirus lacking a neuregulin-like gene regresses human melanoma tumors in nude mice. Virus Genes 53, 52–62 (2017). https://doi.org/10.1007/s11262-016-1402-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11262-016-1402-2