Abstract
Herpesviruses maintain a dynamic balance between latency and productive infection. This is a complex process regulated by viral and cellular factors. We have developed a Murine gammaherpesvirus 68 (MHV-68) model system in which to study mechanisms underlying balance between latency and lytic infection. We have generated an epithelial cell line that carries MHV-68 in a tightly latent form by using a bacterial artificial chromosome clone of the virus genome with a mutation in the MHV-68 major lytic R transactivator gene. Complementation of this defect in trans by transfection with a plasmid encoding R transactivator initiated and restored the productive cycle. This cell line model was used to investigate transcription factor occupancy (CCCTC binding factor [CTCF] and Sp1) of the two internal repeat elements in the viral genome during latency and reactivation using chromatin immunoprecipitation. Our results show that CTCF can bind to the 40-bp and the 100-bp repeat sequences during latency, whereas binding is reduced upon reactivation. In contrast, Sp1 only bound to the 100-bp repeat after reactivation. Our results indicate that the large internal repeat sequences in MHV-68 have different functions. We hypothesise that the 40-bp repeat may be involved in regulation of gene expression during the maintenance of latency, while the 100-bp repeat domain may be involved in regulation of the lytic cycle.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The human γ-herpesviruses, Epstein-Barr virus (EBV) and Kaposi’s sarcoma-associated herpesvirus (KSHV; alternatively human herpesvirus 8 [HHV-8]), cause significant human disease, most of which are associated with persistence of these viruses in the host. However, strict host preferences of EBV and KSHV limit assessment of the mechanisms that contribute to their persistence and pathogenesis. Consequently, there has been considerable effort to develop an amenable small animal model for human γ-herpesviruses. Murine γ-herpesvirus 68 (MHV-68 or γHV68; officially Murid herpesvirus 4 [MuHV-4]) is an endogenous pathogen of free-living rodents of the Apodemus genus, e.g. wood mice [1–6]. Experimental infection of laboratory mice with MHV-68 has therefore been developed and utilised to good effect as a model of γ-herpesvirus infection [7–25]. Following intranasal inoculation of mice with MHV-68, a productive infection occurs in the lung [26]. This is cleared around day 10–14 post-infection (p.i.) by CD8+ T cells [27], though the virus persists in a latent or non-productive form in epithelial cells at this site [28]. MHV-68 spreads to the spleen, where it also becomes latent, predominantly not only within B lymphocytes, but also in macrophages and dendritic cells [29–32].
Latency and reactivation from latency are central to the pathogenesis of γ-herpesviruses [33]. Infection of mice with MHV-68 has enabled many aspects of γ-herpesvirus biology to be elucidated. Likewise, MHV-68 readily infects and undergoes productive replication in a range of cell lines in vitro, enabling the study of the productive cycle. However, there is no ideal system with which to study latency and the reactivation from latency in vitro. The S11 B cell tumour cell line [34] which is predominantly latently infected is relevant and has been used to define aspects of MHV-68 latency [35]. However, it has a measurable rate of spontaneous reactivation making the discrimination of latency and reactivation hard. Likewise, latent infection of A20 mouse B cells with a selectable MHV-68 has value but still suffers from spontaneous reactivation [36]. There is therefore a need for a cell culture system that supports MHV-68 in a tightly latent form with which to study latency and reactivation.
The R transactivator homologue (RTA) encoded by MHV-68 open reading frame 50 (ORF50) is an immediate-early gene product that is conserved among all characterised γ-herpesviruses and is a critical regulator of lytic replication and reactivation from latency [6, 37–44]. MHV-68 RTA is responsible for transactivating its own promoter in addition to other virus promoters to reactivate latent virus [38, 45]. It has previously been shown that an RTA-null mutant is incapable of viral protein synthesis, viral DNA replication or virion production. This phenotype can be rescued by expressing RTA in trans [46].
The MHV-68 genome contains two internal repetitive sequences known as the 40-bp and 100-bp repeats (Fig. 1) [39]. The 40-bp repeat is located between nt 26778-28191 in the genome, and the 100-bp repeat is between nt 98981-101170 (Genbank AF105037). The 40-bp repeat has been identified as an enhancer for latency and is important for the expression of mK3 (ORF12) and ORF72 in the Ag8 cell line that is derived from B cells [47]. Moreover, a region of both the 40- and 100-bp repeat sequence is essential for the lytic origins of replication to function [48–50]. We surmised, therefore, regulation of these loci may be important for maintenance of latency and reactivation.
MHV-68 internal repeats contain a high density of tandem CTCF and Sp1 clusters. A diagrammatic representation of the MHV-68 genome is shown at the top. Unique regions are indicated by shaded boxes and repetitive elements by open boxes. The genome is bounded by multiple copies of a terminal repeat element (TR). The positions of the regions amplified by PCR in the ChIP assay are shown by solid bars. Beneath this, the sequences of the 40- and 100-bp repeats (forward strand) are expanded showing the location of the clustered CTCF and Sp1 motifs as indicated. The sequence of the repeats is shown in upper case and the surrounding sequence in lower case. Motifs for CTCF and Sp1 are highlighted in either red (forward strand) or blue (reverse strand) where motifs overlap are highlighted in green. The sequence of the PCR primers used are shown in purple (Color figure online)
CCCTC-binding factor (CTCF) is a zinc finger DNA-binding protein that is highly conserved in vertebrates. It is expressed in most cell types and has transcriptional activator activity. Described originally as binding to the sequence CCCTC, it has now been shown that CTCF has a much larger 20-bp consensus binding sequence [51]. The presence of a single CTCF DNA-binding motif is sufficient for binding. However, clustering of these consensus sequences leads to a higher CTCF-binding affinity. CTCF is associated with several distinct activities, including transcriptional activation/repression, the formation of chromatin insulators, imprinting and X chromosome inactivation [52].
Sp1 is a zinc finger-containing DNA binding protein that is ubiquitously expressed and can either activate or repress transcription. It binds to GC-rich motifs (such as 5′-G/T-GGGCGG-G/A-G/A-C/T-3′ or 5′-G/T-G/A-GGCG-G/T-G/A-G/A-C/T-3′). Sp1 has also been linked to chromatin remodelling via interactions with chromatin-modifying factors such as p300 and histone deacetylases (HDACs) [53]. Sp1 and CTCF have been demonstrated to bind to a GC-rich triplet domain in the c-myc gene and proposed to alter transcription start site usage of c-myc [54, 55].
Both CTCF and Sp1 have been shown to play a role in herpesvirus gene regulation. CTCF binds during latency to clusters of consensus binding motifs that are present in the repetitive regions of the HSV-1 genome. CTCF could therefore be important in the maintenance of herpes virus latency [56]. A number of promoters for HSV-1 immediate-early genes (e.g. ICP4) have consensus binding sites that have been shown to interact with Sp1. This factor therefore has an important role during reactivation and the lytic cycle [57–59].
The aim of this study was to analyse the potential role of CTCF and Sp1 in MHV-68 gene expression during the switch from latency to reactivation. We describe the construction of an epithelial cell line model for MHV-68 that was capable of analysing latency and reactivation. We describe the clustering of CTCF and Sp1 DNA-binding motifs in the two internal repetitive sequences and go on to use the cell line model to analyse transcription factor occupancy and the epigenetics of MHV-68 during latency and after reactivation.
Materials and methods
Generation of the C127ΔRTA cell line and reactivation from latency
The C127 epithelial cell line (ATCC CRL 1616; mouse mammary epithelial cells) was grown in Dulbecco’s Modified Eagles Medium (DMEM) supplemented with 10 % fetal bovine serum, 2 mM l-glutamine, 70 μg/ml penicillin and 10 μg/ml streptomycin. The BAC clone containing the MHV-68 genome with a transposon-insertion mutation rendering ORF50 non-functional (BACMHV68ΔORF50) has been described previously [60]. This BAC also contains a functional puromycin cassette enabling selection in mammalian cells.
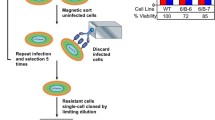
BACMHV68ΔORF50 DNA was transfected into the C127 cell line by the calcium phosphate method as described previously [61, 62]. Three days post-transfection, cells were transferred into culture medium containing puromycin (3 μg/ml). Surviving cell colonies containing MHV-68 genomes were isolated by ring cloning [63]. This was repeated three times to obtain pure cell clones. Colonies were screened by PCR analysis for the presence of MHV-68 genomes using primers specific for gp150 as described previously [28]. One clone that grew and stably maintained the MHV-68 genome was selected (C127ΔRTA) for further analysis.
To reactivate virus from latency, C127ΔRTA cells were transfected with a mammalian expression vector (pFLAG-CMV1, Sigma) containing ORF50 (pFLAG-CMV1-ORF50) using 1.5 μg plasmid per cm2 cells with Lipofectamine 2000 (Invitrogen) according to the manufacturer’s protocol.
Immunostaining
The percentage of cells reactivating MHV-68 was assessed by immunofluorescence using a polyclonal rabbit antiserum to MHV-68 structural antigens [6] as described previously [64, 65]. C127ΔRTA cells were plated onto glass cover slips and transfected as above with varying concentrations of pFLAG-CMV1-ORF50 or GFP (pEGFP-N1, Clontech). The secondary antibody was anti-rabbit IgG conjugated to Texas Red (Jackson Immunoresearch Laboratories). The cells were visualised on a Zeiss Axioscop 40 microscope, and images were captured using Axiovision software.
qRT-PCR for viral gp150 transcripts
Total RNA was extracted, treated with DNase and checked for quality, followed by reverse transcription using a Reverse Transcription System (Promega). The qRT-PCR was carried out using the iQ SYBR Green supermix (BioRad) with 0.18 μM primers and 30 ng of each cDNA sample. Primer sequences were as follows: RPL8 (forward, 5′-CAGTGAATATCGGCAATGTTTTG-3′; reverse, 5′-TTCACTCGAGTCTTCTTGGTCTC-3′) and gp150 (forward, 5′-GAACCTCCCACCTCCAATGC -3′; reverse, 5′-TTGTGGGGGTGTCTCATGGTTCG-3′). PCR was performed in an iQ5 cycler (BioRad) under the following conditions: 95 °C for 10 min, and then 45 cycles at 94 °C for 30 s and 60–61 °C for 40 s. Melting curve analysis was carried out to confirm the specificity of the products between 65 and 95 °C at 0.2 °C increments. Data were analysed using Bio-Rad iQ5 optical system software.
ChIP, PCR and densitometry
ChIP was performed using the ChIP-IT Express kit (Active Motif) as per the manufacturer’s instructions. C127ΔRTA cells were fixed with 1 % formaldehyde for 10 min and then washed and lysed, and the DNA sheared by sonication to produce DNA fragments in the range of 500–1500 bp. The chromatin/DNA complexes were incubated with antibodies to Sp1 (Santa Cruz Biotechnology), CTCF (BD Transduction Laboratories), or non-specific IgG (Active Motif) for 16 h at 4 °C. The immune complexes were then precipitated, washed, eluted, reverse cross-linked and treated with proteinase K. The resulting DNA fragments were amplified with primers that adjoined the repeats as described in Table 1. (Genbank NC001826).
Each PCR reaction contained 35 ng DNA template, the appropriate concentration of MgCl2 (2.5-4 mM), 0.2 mM of each dNTP (except dGTP where 0.1 mM of deoxy-7-deazaguanosine triphosphate and 0.1 mM of dGTP was used), 0.2 pM of each primer, 1× Diamond buffer, 1.5 U Diamond DNA polymerase (Bioline) and 0.5 M betaine. The PCR conditions were 96 °C for 3 min, followed by 36 cycles of 94 °C (1 min), 58 °C (30 s) and 72 °C (20 s). PCR products were analysed by agarose gel electrophoresis in the presence of ethidium bromide, followed by visualisation using a UV transilluminator. Signal intensities from PCR reactions obtained from ChIP assays or from whole cell lysates (control for PCR) were quantified from the TIFF images with ImageJ software (National Institutes of Health; http://rsbweb.nih.gov/ij/). Area, fraction and mean gray values were taken. Mean gray values were used as a measure of relative number of pixels. Fold enrichment = specific antibody/IgG was calculated for each independent experiment.
Results and discussion
The MHV-68 internal repeat elements contain clusters of binding motifs for the cellular proteins CTCF and Sp1
To determine the location of consensus CTCF and Sp1 binding sites within the MHV-68 genome, the DNA sequence was scanned using open-source prediction algorithms. CTCF motifs were predicted using the CTCF binding site prediction tool [66] (http://insulatordb.uthsc.edu/storm.php) and Sp1 motifs were predicted using Consite [67](http://asp.ii.uib.no:8090/cgi-bin/CONSITE/consite/) which utilises the JASPAR database [68]. Initial scans revealed that the 40-bp and 100-bp tandem direct repeat sequences contained a high density of consensus binding sequences for both CTCF and Sp1 (Fig. 1). These motifs were only rarely found throughout the unique regions of the genome (not shown). The 40-bp internal repeat contains 26 high-fidelity CTCF and 23 Sp1 consensus binding sequences, whereas the 100-bp repeat contains 28 CTCF and 50 Sp1 consensus binding sequences. The number and clustering of these sequences were highly striking and suggested a significant role for these repeat regions in controlling virus gene expression.
Generation and validation of a cell line containing latent MHV-68
Analysis of factor binding during viral latency and reactivation required a cell line that was latently infected with MHV-68, had a minimal level of spontaneous reactivation and that was capable of being reactivated into the lytic cycle. All of the cell lines available (e.g. S11) had a relatively high level of spontaneous reactivation. Thus, to generate a cell line that contained MHV-68 in a tightly latent form, a bacterial artificial chromosome (BAC) containing the MHV-68 genome with an insertional (transposon) mutation of ORF50 (that encodes RTA) was used to stably transfect C127 epithelial cells. Epithelial cells are a relevant cell type to study since MHV-68 both replicates and becomes latent in epithelial cells in vivo [28]; furthermore, C127 is a mouse epithelial cell line that supports MHV-68 productive replication [69]. ORF50/RTA is critical for the initiation of the MHV-68 lytic cycle, and productive gene expression and replication should not occur in its absence. One cell clone (C127ΔRTA) that contained MHV-68 DNA as determined by PCR analysis was selected for further analysis.
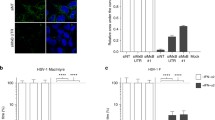
The C127ΔRTA cell line was examined over several passages in culture and did not develop viral plaques and thus, appeared to be latently infected. To confirm that the lytic cycle was not occurring, and that RTA provided in trans could induce reactivation of virus, C127ΔRTA cells were analysed for lytic cycle antigens and transcripts before and after transient transfection with an expression plasmid that encodes RTA (pFLAG-CMV1-ORF50). The expression of MHV-68 structural antigens was analysed in cells by immunofluorescence staining using a polyclonal antibody to MHV-68 [6]. Transfection with a control plasmid encoding GFP (pEGFP-N1) revealed highly efficient transfection levels with >90 % of C127ΔRTA cells expressing GFP (not shown). There was no evidence of the expression of structural antigens in untreated cells or after transfection with the empty expression vector pFLAG-CMV1 (Fig. 2a, b). However, after transient transfection of C127ΔRTA with pFLAG-CMV1-ORF50 (supplying RTA), structural antigens were detected (Fig. 2c–h). The dose- and time-dependent effect of RTA on reactivation was assessed using concentrations from 2 to 4 μg of pFLAG-CMV1-ORF50, assayed at both 24 and 48 h post-transfection. Reactivation occurred at all concentrations; however, it occurred in the highest proportion of cells when 4 μg of pFLAG-CMV1-ORF50 was used and fluorescence analysed at 48 h post-transfection (Fig. 2h), suggesting a dose dependant effect.
Reactivation of MHV-68 occurs after transfection of C127ΔRTA cells with RTA. C127ΔRTA cells were cultured on glass cover slips and reactivation was assessed at 24 and 48 h post-transfection with varying concentrations of the RTA expression plasmid pFLAG-CMV1-ORF50, as indicated. Reactivation was assessed by indirect immunofluorescence for expression of MHV-68 structural antigens using a rabbit polyclonal anti-MHV68 and Texas red anti-rabbit secondary. Nuclei were counter-stained (blue) with DAPI. The cells were visualised on a Zeiss Axioscop 40 microscope; images were captured using Axiovision software. Scale bars represent 20 μm. a Texas red conjugated secondary antibody only; b Transfection with pFLAG-CMV1 backbone alone; c 2 μg of pFLAG-CMV1-ORF50, 24 h; d 3 μg of pFLAG-CMV1-ORF50, 24 h; e 4 μg of pFLAG-CMV1-ORF50, 24 h; f 2 μg of pFLAG-CMV1-ORF50, 48 h; g 3 μg of pFLAG-CMV1-ORF50, 48 h; h 4 μg of pFLAG-CMV1-ORF50, 48 h (Color figure online)
To quantify the time-course of reactivation, we analysed the expression of transcripts specific for the major MHV-68 glycoprotein gp150 that is known to be expressed with late kinetics [64]. Expression was quantified by qRT-PCR. The results (Fig. 3) showed that expression of gp150 mRNA was not detectable after transfection with pFLAG-CMV1 and was not significantly increased 3 h after transfection with pFLAG-CMV1-ORF50. However, expression thereafter increased rapidly to plateau at around 12 h post-transfection. Levels were then maintained through 36 h post-transfection, and declined slightly by 48 h. The kinetics of gp150 expression observed was similar to that seen after infection of cells with MHV-68 [64], implying a rapid and efficient induction of the productive cycle in C127ΔRTA cells that mirrors natural infection.
qRT-PCR analysis of gp150 mRNA after transfection of C127∆RTA cells with pFLAG-CMV1-ORF50. RNA was extracted from C127∆RTA cells at the indicated times after either transfection with pFLAG-CMV1 (filled triangle) or the RTA-expressing vector pFLAG-CMV1-ORF50 (filled circle). The fold change in gp150 mRNA was normalised to the levels of the cellular ribosomal protein L8 mRNA and expressed as relative to the 0 h control as described elsewhere [74]. The quantitative cycle (Cq) values of test samples spanning the time-course were plotted. Error bars depict the standard error of the means for 6 replicates. Statistical analysis was performed by two-way ANOVA with Bonferroni post-tests. Points where a significant (P < 0.001) difference in gp150 mRNA levels were seen are indicated by asterisks
Thus, the C127ΔRTA cell line contains a tightly latent MHV-68 genome that can be made to reactivate by supplying RTA in trans by transfection. While this manuscript was in preparation, a similar approach to generating a cell line containing latent MHV-68 was described [70]. This utilised a doxycycline-inducible promoter to control RTA expression. While this system is roughly equivalent to the one described here, the use of fibroblasts may limit its usefulness, as MHV-68 does not become latent in this cell type in vivo.
CTCF and SP1 associate with the 40 bp and 100 bp repeat sequences
Since our bioinformatic analyses showed that the MHV-68 internal tandem repeat sequences contained clustered CTCF and Sp1 motifs (Fig. 1), we determined whether these transcription factors bound to the internal repeats during latency and reactivation. Chromatin immunoprecipitation (ChIP) analysis was performed on chromatin extracted from the C127ΔRTA cell line. In initial pilot studies, we found that primers directed to the internal repeat sequences generated a large number of non-specific products, which is likely due to their extremely high GC content and repetitive nature. We therefore used primers in the ChIP assay that amplified flanking sequences directly adjoining the repeats. Random DNA fragments generated by sonication in this protocol were of such a size (500–1500 bp) that the primers specific for the directly adjoining flanking regions would still amplify precipitated DNA containing internal repeat sequences (Fig. 1). Analysing for enrichment of a sequence adjacent to a repeat has been validated previously as an indicator of factor association with the repeat sequence [56]. Furthermore, we analysed sequences flanking both sides of the repeat to more accurately assess factor binding.
We first addressed by ChIP whether the 40-bp tandem repeat sequence could associate with CTCF using an anti-CTCF antibody. The analysis was performed on C127ΔRTA cells both before and after reactivation of MHV-68 from latency by transfection with pFLAG-CMV1-ORF50. We demonstrated CTCF-binding to the 40-bp repeat flanking regions during latency as illustrated by an enriched number of PCR amplicons both 5′ and 3′ to the 40 bp tandem repeat region, whereas after reactivation there was a reduction in CTCF-binding to both flanking regions (Fig. 4a). Likewise, analysis of the flanking regions of the 100-bp repeat demonstrated CTCF-binding during latency, but after reactivation this was also reduced (Fig. 4a).
Chromatin immunoprecipitation (ChIP) analysis of CTCF and Sp1 at the MHV-68 internal tandem repeat sequences. Latently infected C127ΔRTA cells were assayed either without treatment or after reactivation from latency by delivering RTA with pFLAG-CMV1-ORF50. Proteins were cross-linked in vivo at 36 h after transfection and subjected to ChIP using anti-CTCF, anti-Sp1 or non-specific IgG as a control. PCR products were analysed by agarose gel electrophoresis and densitometric image analysis of agarose gels. The results are expressed as the mean fold enrichment (n = 2) of transcription factor relative to the level seen for IgG alone from C127ΔRTA cells either untreated (solid bars) or after reactivation (open bars). a anti-CTCF; b anti-Sp1
Thus, it appears CTCF can bind to the 40-bp and the 100-bp repeat sequences during latency, whereas binding is reduced upon reactivation. CTCF is important for regulation at the KSHV major latency control region, at which it can associate with three repeated consensus sequences; at these sites CTCF may be important for the creation of intragenomic domains important for transcription during latency [71, 72]. Thus, this might point to a conserved mechanism between herpesviruses that uses CTCF to segregate active from inactive genes during latency. Further, our observations fit with reverse genetic analysis showing that the MHV-68 40-bp repeat is important for latency amplification [47].
We next addressed whether the repeats could associate with Sp1, repeating ChIP with an anti-Sp1 antibody. Significant Sp1 binding was not detected at either the 40- or 100-bp repeat domains prior to reactivation (Fig. 4b). However, after reactivation, while Sp1 still was not significantly enriched at the 40-bp repeat (Fig. 4b), there was enrichment at both sides of the 100-bp repeat. Specifically, we observed a more than threefold enrichment at the 5′ end and a fivefold enrichment at the 3′ end of the 100-bp repeat (Fig. 4b).
During the lytic cycle, DNA is replicated and transcription of the ordered cascade of lytic cycle-associated viral genes occurs. Interestingly, the 3′ end of the 100-bp repeat is essential for the function of the lytic origin of replication [48]. Therefore, binding of Sp1 at this region may be involved in the regulation of concomitant transcription and DNA replication.
In summary, we have developed a tightly latent but reactivatable MHV-68-containing cell line which provides a model to enable analysis of factors regulating the MHV-68 genome, both during latency and during and after reactivation. Our investigation of both the 40-bp and 100-bp internal repeat domains revealed differences in factor binding before and after reactivation. This indicates that the large internal repeat sequences in MHV-68 have different functions. We hypothesise that the 40-bp repeat may be involved in regulation of gene expression during the maintenance of latency, while the 100-bp repeat domain may be involved in regulation of the lytic cycle. Furthermore, this model will be of use in further dissecting the regulation of MHV-68 by transcription factors.
The role of CTCF and Sp1 to regulate an important switch in herpesvirus latency is also consistent with our data on the regulation of the HSV-1 repeat RE1 adjacent to the LAT region. This domain demonstrates tissue specific and inducible transcriptional properties which are in part regulated by these transcription factors. This may point to the conserved role of this signalling pathway in herpesvirus evolution [73].
References
B. Ehlers, J. Kuchler, N. Yasmum, G. Dural, S. Voigt, J. Schmidt-Chanasit, T. Jakel, F.R. Matuschka, D. Richter, S. Essbauer, D.J. Hughes, C. Summers, M. Bennett, J.P. Stewart, R.G. Ulrich, J. Virol. 81, 8091–8100 (2007)
D.J. Hughes, A. Kipar, J.T. Sample, J.P. Stewart, J. Virol. 84, 3949–3961 (2010)
K. Blasdell, C. McCracken, A. Morris, A.A. Nash, M. Begon, M. Bennett, J.P. Stewart, J. Gen. Virol. 84, 111–113 (2003)
S.D. Becker, M. Bennett, J.P. Stewart, J.L. Hurst, Lab. Anim. 41, 229–238 (2007)
D.J. Hughes, A. Kipar, G.H. Leeming, E. Bennett, D. Howarth, J.A. Cummerson, R. Papoula-Pereira, B.F. Flanagan, J.T. Sample, J.P. Stewart, PLoS Pathog. 7, e1001321 (2011)
D.J. Hughes, A. Kipar, S.G. Milligan, C. Cunningham, M. Sanders, M.A. Quail, M.A. Rajandream, S. Efstathiou, R.J. Bowden, C. Chastel, M. Bennett, J.T. Sample, B. Barrell, A.J. Davison, J.P. Stewart, J. Gen. Virol. 91, 867–879 (2010)
A.A. Nash, B.M. Dutia, J.P. Stewart, A.J. Davison, Philos. Trans. R. Soc. Lond. B Biol. Sci. 356, 569–579 (2001)
P.G. Stevenson, S. Efstathiou, Viral Immunol. 18, 445–456 (2005)
M.A. Blackman, E. Flano, E. Usherwood, D.L. Woodland, Mol. Med. Today 6, 488–490 (2000)
S.H. Speck, H.W. Virgin, Curr. Opin. Microbiol. 2, 403–409 (1999)
E. Flano, I.J. Kim, D.L. Woodland, M.A. Blackman, J. Exp. Med. 196, 1363–1372 (2002)
P.C. Doherty, J.P. Christensen, G.T. Belz, P.G. Stevenson, M.Y. Sangster, Philos. Trans. R. Soc. Lond. B Biol. Sci. 356, 581–593 (2001)
J.P. Simas, S. Efstathiou, Trends Microbiol. 6, 276–282 (1998)
A.A. Nash, E.J. Usherwood, J.P. Stewart, Semin. Virol. 7, 125–130 (1996)
S.S. Lok, Y. Haider, D. Howell, J.P. Stewart, P.S. Hasleton, J.J. Egan, Eur. Respir. J. Off. J. Eur. Soc. Clin. Respir. Physiol. 20, 1228–1232 (2002)
A.I. Macrae, B.M. Dutia, S. Milligan, D.G. Brownstein, D.J. Allen, J. Mistrikova, A.J. Davison, A.A. Nash, J.P. Stewart, J. Virol. 75, 5315–5327 (2001)
A.I. Macrae, E.J. Usherwood, S.M. Husain, E. Flano, I.J. Kim, D.L. Woodland, A.A. Nash, M.A. Blackman, J.T. Sample, J.P. Stewart, J. Virol. 77, 9700–9709 (2003)
J.J. Obar, D.C. Donovan, S.G. Crist, O. Silvia, J.P. Stewart, E.J. Usherwood, J. Virol. 78, 10829–10832 (2004)
D. Verzijl, C.P. Fitzsimons, M. Van Dijk, J.P. Stewart, H. Timmerman, M.J. Smit, R. Leurs, J. Virol. 78, 3343–3351 (2004)
J.P. Stewart, A. Kipar, H. Cox, C. Payne, S. Vasiliou, J.P. Quinn, PLoS ONE 3, e1673 (2008)
B.M. Dutia, J.P. Stewart, R.A. Clayton, H. Dyson, A.A. Nash, J. Gen. Virol. 80, 2729–2736 (1999)
L.A. Terry, J.P. Stewart, A.A. Nash, J.K. Fazakerley, J. Gen. Virol. 81, 2635–2643 (2000)
B.M. Dutia, D.J. Roy, B. Ebrahimi, B. Gangadharan, S. Efstathiou, J.P. Stewart, A.A. Nash, J. Gen. Virol. 85, 1393–1400 (2004)
J.P. Stewart, O.J. Silvia, I.M.D. Atkin, D.J. Hughes, B. Ebrahimi, H. Adler, J. Virol. 78, 10449–10459 (2004)
J.P. Quinn, A. Kipar, D.J. Hughes, E. Bennett, H. Cox, L. McLaughlin, A. Zimmer, S.P. Hunt, J.P. Stewart, Neuropeptides 45, 49–53 (2011)
N.P. Sunil-Chandra, S. Efstathiou, J. Arno, A.A. Nash, J. Gen. Virol. 73, 2347–2356 (1992)
S. Ehtisham, N.P. Sunil-Chandra, A.A. Nash, J. Virol. 67, 5247–5252 (1993)
J.P. Stewart, E.J. Usherwood, A. Ross, H. Dyson, T. Nash, J. Exp. Med. 187, 1941–1951 (1998)
N.P. Sunil-Chandra, S. Efstathiou, A.A. Nash, J. Gen. Virol. 73, 3275–3279 (1992)
E.J. Usherwood, J.P. Stewart, K. Robertson, D.J. Allen, A.A. Nash, J. Gen. Virol. 77, 2819–2825 (1996)
K.E. Weck, S.S. Kim, H.W. Virgin, S.H. Speck, J. Virol. 73, 3273–3283 (1999)
E. Flano, S.M. Husain, J.T. Sample, D.L. Woodland, M.A. Blackman, J. Immunol. 165, 1074–1081 (2000)
M. Ottinger, D. Pliquet, T. Christalla, R. Frank, J.P. Stewart, T.F. Schulz, J. Virol. 83, 4423–4434 (2009)
E.J. Usherwood, J.P. Stewart, A.A. Nash, J. Virol. 70, 6516–6518 (1996)
S.M. Husain, E.J. Usherwood, H. Dyson, C. Coleclough, M.A. Coppola, D.L. Woodland, M.A. Blackman, J.P. Stewart, J.T. Sample, Proc. Natl. Acad. Sci. USA 96, 7508–7513 (1999)
J.C. Forrest, S.H. Speck, J. Virol. 82, 7688–7699 (2008)
R. Sun, S.F. Lin, L. Gradoville, Y. Yuan, F. Zhu, G. Miller, Proc. Natl. Acad. Sci. U.S.A. 95, 10866–10871 (1998)
T.T. Wu, E.J. Usherwood, J.P. Stewart, A.A. Nash, R. Sun, J. Virol. 74, 3659–3667 (2000)
H.W. Virgin, P. Latreille, P. Wamsley, K. Hallsworth, K.E. Weck, A.J. Dal Canto, S.H. Speck, J. Virol. 71, 5894–5904 (1997)
V.L. van Santen, J. Virol. 67, 773–784 (1993)
A. Whitehouse, I.M. Carr, J.C. Griffiths, D.M. Meredith, J. Virol. 71, 2550–2554 (1997)
G. Miller, A. El-Guindy, J. Countryman, J. Ye, L. Gradoville, Adv. Cancer Res. 97, 81–109 (2007)
T–.T. Wu, L. Tong, T. Rickabaugh, S. Speck, R. Sun, J. Virol. 75, 9262–9273 (2001)
J. Hart, M. Ackermann, G. Jayawardane, G. Russell, D.M. Haig, H. Reid, J.P. Stewart, J. Gen. Virol. 88, 28–39 (2007)
K.S. Gray, R.D. Allen 3rd, M.L. Farrell, J.C. Forrest, S.H. Speck, J. Virol. 83, 314–328 (2009)
I.V. Pavlova, H.W.t. Virgin, S.H. Speck, J. Virol. 77, 5731–5739 (2003)
N.N. Thakur, S. El-Gogo, B. Steer, K. Freimuller, A. Waha, H. Adler, PLoS ONE 2, e733 (2007)
H. Adler, B. Steer, K. Freimuller, J. Haas, J. Virol. 81, 7300–7305 (2007)
H. Deng, J.T. Chu, N.-H. Park, R. Sun, J. Virol. 78, 9123–9131 (2004)
D. Gong, J. Qi, V. Arumugaswami, R. Sun, H. Deng, Virology 387, 285–295 (2009)
T.H. Kim, Z.K. Abdullaev, A.D. Smith, K.A. Ching, D.I. Loukinov, R.D. Green, M.Q. Zhang, V.V. Lobanenkov, B. Ren, Cell 128, 1231–1245 (2007)
J.E. Phillips, V.G. Corces, Cell 137, 1194–1211 (2009)
N.Y. Tan, L.M. Khachigian, Mol. Cell. Biol. 29, 2483–2488 (2009)
N. Hay, J.M. Bishop, D. Levens, Genes Dev. 1, 659–671 (1987)
V.V. Lobanenkov, R.H. Nicolas, V.V. Adler, H. Paterson, E.M. Klenova, A.V. Polotskaja, G.H. Goodwin, Oncogene 5, 1743–1753 (1990)
A.L. Amelio, P.K. McAnany, D.C. Bloom, J. Virol. 80, 2358–2368 (2006)
I.K. Chung, S.M. Soisson, M.T. Muller, J. Biochem. 117, 19–22 (1995)
A.N. Imbalzano, D.M. Coen, N.A. DeLuca, J. Virol. 65, 565–574 (1991)
K.A. Jones, R. Tjian, Nature 317, 179–182 (1985)
M.J. Song, S. Hwang, W.H. Wong, T.T. Wu, S. Lee, H.I. Liao, R. Sun, Proc. Natl. Acad. Sci. U.S.A. 102, 3805–3810 (2005)
G. Jayawardane, G.C. Russell, J. Thomson, D. Deane, H. Cox, D. Gatherer, M. Ackermann, D.M. Haig, J.P. Stewart, J. Gen. Virol. 89, 2447–2455 (2008)
B. Steer, B. Adler, S. Jonjic, J.P. Stewart, H. Adler, PLoS ONE 5, e11672 (2010)
D.J. Roy, B.C. Ebrahimi, B.M. Dutia, A.A. Nash, J.P. Stewart, Arch. Virol. 145, 2411–2420 (2000)
J.P. Stewart, N.J. Janjua, S.D. Pepper, G. Bennion, M. Mackett, T. Allen, A.A. Nash, J.R. Arrand, J. Virol. 70, 3528–3535 (1996)
J.P. Stewart, J.R. Arrand, Hum. Pathol. 24, 239–242 (1993)
L. Bao, M. Zhou, Y. Cui, Nucleic Acids Res. 36, D83–D87 (2008)
A. Sandelin, W.W. Wasserman, B. Lenhard, Nucleic Acids Res. 32(suppl 2), W249–W252 (2004)
A. Sandelin, W. Alkema, P. Engstrom, W.W. Wasserman, B. Lenhard, Nucleic Acids Res. 32, D91–D94 (2004)
J.P. Stewart, N.J. Janjua, N.P. Sunil-Chandra, A.A. Nash, J.R. Arrand, J. Virol. 68, 6496–6504 (1994)
J.S. May, N.J. Bennett, P.G. Stevenson, PLoS ONE 5, e11080 (2010)
H. Kang, P.M. Lieberman, J. Virol. 83, 6199–6210 (2009)
W. Stedman, H. Kang, S. Lin, J.L. Kissil, M.S. Bartolomei, P.M. Lieberman, EMBO J. 27, 654–666 (2008)
H.C. Stevens, C. Fiskerstrand, V.J. Bubb, R. Dalziel, J.P. Quinn, FEBS Lett. 583, 3335–3338 (2009)
M.W. Pfaffl, Nucleic Acids Res. 29, e45 (2001)
Acknowledgments
HCS was funded by the Wellcome Trust Prize Studentship. JPQ and VJB were funded by the BBSRC (award BB/D016754/9). This work was funded in part by U.S. Public Health Service grant CA090208 from the National Cancer Institute and the Penn State Hershey Cancer Institute. JPS was funded by a Royal Society (London) University Research Fellowship. The authors would like to thank Dr Bahram Ebrahimi for the kind gift of pFLAG-CMV1-ORF50.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Stevens, H.C., Cham, K.SW., Hughes, D.J. et al. CTCF and Sp1 interact with the Murine gammaherpesvirus 68 internal repeat elements. Virus Genes 45, 265–273 (2012). https://doi.org/10.1007/s11262-012-0769-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11262-012-0769-y