Abstract
Remnant proviral sequences in the genome resulting from the ancient germline infection of exogenous retroviruses are called endogenous retroviruses (ERVs). The transcriptional activation of human ERVs (HERVs) in the brain of patients with some neurologic diseases suggests that ERVs may participate in certain disease processes in the central nervous system. In this study, we identified putative murine ERVs (MuERVs) which are transcriptionally active in the brain and characterized their biological properties to better understand the ERVs’ roles in the brain pathophysiology. The brain and selective non-nervous tissues (heart, muscle, adrenal gland, and salivary gland) of female C57BL/6J mice were subjected to RT-PCR analyses of MuERV expression by amplifying the 3′-end U3 regions and full-length/subgenomic transcripts. The expression patterns of the U3 regions and subgenomic transcripts in the brain were unique compared to the other tissues as well as the genomic MuERV profile. Two putative MuERVs (8,027 and 5,668 bp) were mapped on the mouse genome (chromosome 10, and chromosomes 4 and 8, respectively) using the MuERV U3 sequences, which were evidently expressed in the brain, as probes. Biological properties of these putative MuERVs, such as transcription potential, primer binding site, coding potential, integration age, recombination, and flanking host genes, were characterized. In particular, one of the two putative MuERV isolates had coding potentials for intact group specific antigen (gag), and truncated polymerase (pol) and envelope (env) polypeptides, while the other was defective for all three polypeptides. The findings from this study suggest that a specific group of MuERVs are constitutively expressed in the brain and they may participate in normal and pathogenic events pertaining to the brain through their replication gene products (e.g., gag and env polypeptides) as well as interactions with flanking host genes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
An ancient infection of exogenous retroviruses into the germline leads to a permanent fix of the proviral form of the retroviruses in the host genome. These retroviral elements in the germline genome are called endogenous retroviruses (ERVs) and are inherited to the offspring in a Mendelian manner [1]. The retroviral sequences in the genome are amplified during evolution through repetitive events of reverse transcription-mediated reintegration and infection into the germline. All vertebrates have ERVs in their genome, and the human genome is presumed to contain ∼300,000 to ∼500,000 copies of retroviral elements, constituting approximately 8% of the entire genome [2]. About 10% of the mouse genome is occupied by sequences of retroviral origin [3]. A substantial fraction of ERVs retain the coding potentials for all three retroviral polypeptides essential for virus assembly and completion of the infection cycle; however, the rest are presumed to have defective coding potentials.
Recent studies demonstrated that ERVs are involved in and/or correlated with a range of disease processes in humans and experimental animal models, such as multiple sclerosis, schizophrenia, bipolar disorder, insulin-dependent diabetes mellitus, experimental systemic lupus erythematosus, and burn injury [4–10]. The role of ERVs in inflammatory diseases is exemplified by the recent finding that syncytin, an envelope glycoprotein encoded by a human endogenous retrovirus (HERV)-W, is directly involved in the pathogenic processes of multiple sclerosis through its proinflammatory properties [11]. Syncytin-mediated release of redox oxidants from astrocytes is reported to be directly linked to the demyelination and death of oligodendrocytes. Increased transcriptional activities of the HERV-W family members and ERVs within the murine leukemia virus genus are observed in the brain of schizophrenia patients [7, 8]. In another study, a superantigen encoded by HERV-K, which is related to mouse mammary tumor virus, was characterized as a candidate autoimmune gene responsible for the systemic activation of autoreactive T cells targeting pancreatic β-cells [6].
The transcription regulatory elements located on the long terminal repeats (LTRs) of ERVs control expression of their own genes as well as neighboring host genes through various mechanisms, such as intergenic splicing, primary polyadenylation signal, and suppression of host gene translation [12–14]. The majority of ERVs are known to be transcriptionally inactive, probably due to epigenetic modifications, such as the methylation status of the promoter [15, 16]. It is likely that the transcription regulatory regions of actively expressed ERVs contain different profiles of transcription factor binding elements and epigenetic modifications, including histone modification (e.g., acetylation) near the MuERV loci, compared to silent ones [17]. However, transcriptionally silent ERVs can be activated by various stimuli from the environment, such as cytokines and viral infection [18, 19].
In this study, to better understand the ERVs’ role in normal physiology as well as in pathologic states of the brain, we identified and mapped the putative MuERVs which were actively expressed and characterized their biological properties.
Materials and methods
Animals
Female C57BL/6J mice from the Jackson Laboratory (Bar Harbor, ME) were housed according to guidelines of the National Institutes of Health. The Animal Use and Care Administrative Advisory Committee of the University of California, Davis, approved the experimental protocol. Three mice were sacrificed by cervical dislocation for tissue collection (brain, heart, muscle, adrenal gland, and salivary gland) without any pretreatments and the samples were snap-frozen.
RT-PCR analysis
Total RNA isolation and cDNA synthesis were performed based on protocols described previously [20]. Total RNA was extracted using an RNeasy kit (Qiagen, Valencia, CA). Subsequently, 100 ng of total RNA from each tissue sample was subjected to reverse transcription using Sensiscript reverse transcriptase (Qiagen). Primers ERV-U1 (5′-CGG GCG ACT CAG TCT ATC GG-3′) and ERV-U2 (5′-CAG TAT CAC CAA CTC AAA TC-3′) were used to amplify the 3′ U3 regions of non-ecotropic MuERVs [21]. The U3 regions were selected for expression analysis because of their polymorphic sequences within the MuERV population compared to the rest of the viral genome. The primers used for the amplification of full-length (∼8 kb) as well as subgenomic transcripts (variable in size), were designed based on a murine acquired immuno deficiency syndrome (MAIDS) virus-related provirus (Genbank accession No. S80082): MV1K (5′-CAT TTG GAG GTC CCA CCG AGA-3′) and MV2D (5′-CTC AGT CTG TCG GAG GAC TG-3′). The comparability between samples was determined by the electrophoresis of an equal amount (500 ng) of total RNA from each sample.
Genomic DNA PCR
Genomic DNA was isolated from the liver tissue using a DNeasy tissue kit (Qiagen). To amplify the U3 regions of MuERVs on the genome, 100 ng of the liver genomic DNA was subjected to PCR analysis using ERV-U1 and ERV-U2 primers.
Cloning of MuERV U3 regions and full-length/subgenomic transcripts
RT-PCR products resolved as distinctive bands representing the U3 regions from each tissue were gel purified using a QIAquick Gel Extraction kit (Qiagen) and cloned into the pGEM®-T Easy vector (Promega, Madison, WI). Plasmid DNAs for sequencing analysis were prepared using a miniplasmid kit from Qiagen. Sequencing was performed at Davis Sequencing Inc. (Davis, CA) and Molecular Cloning Laboratory (South San Francisco, CA).
In silico cloning of putative MuERVs, open reading frame (ORF) analysis, and identification of flanking genes
Putative MuERV sequences were identified by probing the NCBI (National Center for Biotechnology Information) mouse genome database using U3 clones as probes (greater than 98% homology between U3 probe and database) with the “BLAST the mouse genome” program. The parameters for identifying viral sequences of interest included a 5–9 kb size flanked by LTRs containing the U3 probe sequences. Subsequently, the ORFs of each putative MuERV were analyzed using the Vector NTI program (Invitrogen, Carlsbad, CA). The translation products were then compared to the following MuLV references retrieved from NCBI (M17327, AY219567.2, and AF033811) using the Vector NTI program (Invitrogen). Host genes located within 200 kb of the putative MuERV loci were surveyed as flanking genes in this study.
Multiple alignment and phylogenetic tree construction
The Lasergene (DNASTAR, Madison, WI) and Vector NTI (Invitrogen) programs were used to perform multiple alignment and phylogenetic analyses of the MuERV U3 clones. A phylogenetic tree was obtained using the neighbor-joining protocol within the MEGA4 program [22, 23].
Tropism analysis
The putative tropism of each unique U3 clone was determined by comparison to the reference sequence features (direct repeat and unique region) first reported by Tomonaga et al. [21, 24]. A total of four direct repeats (1/1*, 4/4*, 5/5*, and 6/6*) and a single unique region [2] were utilized for the tropism analysis.
Profiling transcription regulatory elements in U3 clones
The MatInspector program (Genomatix, Munich, Germany) was used to determine the presence of transcription regulatory elements within each unique U3 clone with a core similarity of greater than 90%, and the matrix similarity was optimized to minimize false positives.
Integration age, recombination event, and primer binding site (PBS)
The integration age was calculated based on a formula of “0.13% mutation rate between two flanking LTRs per one MYr” [25]. To examine the genomic rearrangement between MuERVs, a short stretch of sequences (∼4 bp) flanking each MuERV was surveyed for a direct repeat, which is formed during the initial proviral integration. A segment of 18 bp, located immediately downstream of the 5′ U5 region, was examined to determine PBSs for the putative MuERVs. The conserved PBS sequences for tRNAProline(P) and tRNAGlutamine(Q) were used as references [26, 27].
Results and discussion
Unique MuERV expression profile in the brain compared to the other tissues
The MuERV expression profile of the brain was examined by RT-PCR amplification of the 3′ U3 regions and it was compared to several non-nervous tissues (heart, muscle, adrenal gland, and salivary gland) (Fig. 1). It revealed that two major U3 regions (labeled as “a” and “b”) were amplified in the brain and the intensity of the b-U3 region was evidently stronger than the a-U3 regions. In addition, the MuERV expression profile of the brain was substantially different from the other tissues examined as well as the genomic MuERV profile. A total of 48 MuERV U3 sequences were initially cloned from all five tissues and subsequent multiple alignment analyses identified 16 unique U3 clones spanning seven different sizes (Fig. 2A). There is at least a one nucleotide mismatch between the unique sequences. Two unique U3 clones were isolated from the brain, BR-a-3 (542 bp) and BR-b-1 (346 bp), corresponding to a-U3 and b-U3 regions, respectively. It needs to be noted that the U3 regions in the PCR products (Fig. 1) include flanking sequences ∼121 bp upstream and downstream of the exact U3 sequence. The flanking sequences were trimmed off before the alignment analysis. The U3 clones derived from the a-U3 regions were highly homologous to each other except for a deletion in the middle of four U3 clones, while the U3 clones from the b-U3 and c-U3 (b/c-U3) regions were relatively divergent. In addition, the TATA box was well conserved in all U3 clones examined. The results from the tropism analysis revealed eight xenotropic and eight polytropic traits (Table 1). Interestingly, all U3 clones isolated from a-U3 regions except for the one from the brain (BR-a-3 clone [542 bp]) had a direct repeat “1/1*.” Furthermore, phylogenetic analysis revealed that both U3 clones isolated from the brain (BR-a-3 and BR-b-1) were placed on distinct branches (Fig. 2B).
RT-PCR analysis of the MuERV expression profile in the brain. Brain and four non-nervous tissues (heart, muscle, adrenal gland, and salivary gland) from C57BL/6J mice were analyzed for MuERV expression by RT-PCR using a primer set (ERV-U1 and ERV-U2) capable of amplifying the 3′ U3 regions. The brain had a unique MuERV expression profile compared to the others. The three different U3 regions analyzed, are indicated (a, b, and c). The MuERV U3 profile of the C57BL/6J genomic DNA serves as a reference. A schematic diagram depicting locations of ERV-U1 and ERV-U2 primers is presented
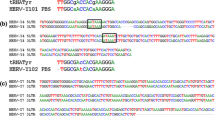
(A) Comparison analysis of MuERV U3 clones. A multiple alignment analysis was performed using 16 unique U3 clones isolated from the brain and four other tissues. Gray shades indicate different levels of sequence homology and dashes represent the absence of sequences. The direct repeats (1/1*, 4/4*, 5/5*, and 6/6*), unique region (2), and TATA box are indicated in solid boxes. An insertion of ∼190 bp is in a dotted box. (B) Phylogenetic analysis of MuERV U3 clones. A phylogenetic tree of 16 unique MuERV U3 clones was established. The values at the branch nodes represent the percentage confidence of a specific branching. The U3 clones derived from the brain (BR-a-3 and BR-b-1) are highlighted in gray
These findings suggest that although there are numerous copies of MuERVs on the genome, only a certain group of MuERVs are constitutively expressed in the brain. It will be of interest to identify specific cell types in the brain responsible for this unique MuERV expression profile. Furthermore, it is likely that certain stress signals, such as proinflammatory cytokines and glucocorticoids, may alter the MuERV expression profile in the brain in a cell type-specific manner. In fact, recent reports demonstrated that the HERV-W encoded envelope protein, called syncytin, participates in the autoimmune processes of multiple sclerosis in the brain through its proinflammatory properties [11].
Identification and characterization of two putative MuERVs presumed to be constitutively expressed in the brain
In this study, putative MuERVs whose sequences match to the two U3 clones (BR-a-3 and BR-b-1) derived from the brain were mapped on the C57BL/6J genome and their biological properties were characterized. Using the U3 clones (BR-a-3 and BR-b-1) as probes, the two putative MuERVs presumed to be constitutively expressed in the brain, were mapped to chromosome 10 (BR-a-3) and chromosomes 4 and 8 (BR-b-1) (Fig. 3). The putative MuERV derived from the BR-a-3 U3 clone was 8,027 bp in size and had four bp direct repeats (GTAC) at the integration junction suggesting no sign of genome rearrangement. Based on the mutation rate (0%) between its two flanking LTRs, the integration age of this putative MuERV is estimated to be <1 Myr. In addition, it had an intact gag ORF as well as truncated forms of pol and env ORFs due to a premature stop codon and the absence of 51 amino acids on the N-terminus, respectively. Furthermore, a survey of host genes flanking within 200 kb upstream and downstream of the genomic locus of this putative MuERV identified three genes (Scml4 [sex comb on midleg-like 4], Sobp [sine oculis-binding homolog], and Pdss2 [prenyl diphosphate synthase subunit-2]) (Fig. 3A). Interestingly, it has been reported that these genes are somewhat highly expressed in the brain suggesting that the transcriptional activities of the BR-a-3-derived MuERV and its neighboring genes might be closely linked [28–30]. The second putative MuERV was isolated using the BR-b-1 U3 clone and was 5,668 bp long with four bp direct repeats (ATAT) at the integration site. There was no mutation between its two flanking LTRs suggesting an estimated integration age of <1 Myr. Due to substantial deletions throughout the genome (primarily in the pol and env genes), this putative MuERV was not capable of encoding any of the three essential polypeptides (gag, pol, and env) (Fig. 3B). There was one putative gene (LOC100040466) within the survey region (200 kb upstream and downstream) of the proviral locus on chromosome 8 and it is presumed to encode a protein similar to cyclic nucleotide gated channel β1. No flanking gene was identified from the chromosome 4 proviral locus. The two putative MuERVs derived from BR-a-3 and BR-b-1 shared the same tRNAGlutamine(Q) PBS sequence.
Mapping of putative MuERVs associated with brain and their biological properties. Two putative MuERVs isolated using the U3 clones (BR-a-3 (A) and BR-b-1 (B)) from the brain as probes were mapped to relevant chromosomes. Numbers on both LTRs indicate chromosomal location and four bp direct repeats are marked at each proviral integration site. Coding potentials of the putative MuERV derived from the BR-a-3 clone are indicated with the total number of amino acids within a parenthesis. Host genes flanking (200 kb upstream and downstream) each putative MuERV locus were mapped. PBS (primer binding site), Q (tRNAglutamine)
The findings that both putative MuERVs isolated using the brain-derived U3 clones did not retain intact coding potentials for the polypeptides necessary for virus assembly suggest that they may need a helper virus for their replication, if any. The putative MuERV derived from the BR-a-3 U3 clone had an intact gag ORF and an env ORF retaining a partial SU (surface) domain and complete TM (transmembrane) domain. The gag protein of the MAIDS virus, particularly p12, has been known to participate in immune disorders within infected mice [31, 32]. Alignment of the gag sequence of the putative MuERV derived from the BR-a-3 U3 probe with the MAIDS virus (GenBank accession number: AY140895) demonstrated 92.6% sequence homology within the entire gag polypeptide and 94% within the p12 protein (data not shown). In addition, the truncated form of the env protein retains an intact TM domain that may participate in a network of signaling pathways associated with pathophysiologic processes in the brain and probably in other tissues. Furthermore, it has been well documented that certain transcription control elements, such as enhancers and negative regulatory elements on retroviral LTRs are often capable of modulating the expression of neighboring host genes. Thus, it will be interesting to investigate whether the LTRs of these putative MuERV isolates influence the expression of neighboring genes in the brain.
Constitutive expression of subgenomic MuERV transcripts in the brain
In the next experiment, we examined whether the full-length as well as subgenomic MuERV transcripts, such as a spliced env transcript, are detectable in the brain. RT-PCR using a set of primers capable of amplifying full-length and subgenomic transcripts revealed a distinct band of ∼5 kb, a faint doublet of ∼2.7 and ∼2.9 kb, and a ∼1 kb band. The expression pattern of the brain was markedly different from the adrenal gland and salivary gland (Fig. 4). The ∼5 kb transcript may be transcribed from the BR-b-1 U3-derived putative MuERV (5,668 bp); however, there was no detectable transcript of ∼7.5 kb, which may represent the BR-a-3 U3-derived MuERV (8,027 bp). To confirm the presence of the full-length transcript of the BR-a-3 U3-derived MuERV, we examined the expression of a pol region spanning a unique genomic deletion of 926 bp in this provirus using another set of primers. An RT-PCR product of the expected size was amplified from the brain and sequencing analysis confirmed a match to the BR-a-3 U3-derived MuERV sequence (data not shown). Primarily based on the size of the doublets (∼2.7 and ∼2.9 kb) and primer locations, it is likely that they were amplified from spliced env transcripts present in the RNA pool of the brain [5]. In addition, the proviral structure and locations of the envelope splicing signals within the BR-b-1 U3-derived MuERV suggest that its amplified envelope transcript is expected to be 926 bp. It is highly likely that the distinct ∼1 kb PCR product is the envelope transcript of the BR-b-1 U3-derived MuERV. Sequencing and alignment analyses of the ∼5 kb transcript band revealed that this transcript (5,064 bp) had a high level of sequence homology (greater than 99% identity) with a previously reported transcript (MuLV(LI-12), AY140896) whose expression was induced in the liver after burn injury [20]. Moreover, these findings suggested that the 5,064 bp transcript was presumed to be a non-spliced transcript of the putative MuERV of 5,668 bp.
Identification of subgenomic MuERV transcripts in the brain. (A) A schematic drawing depicts locations of primers (MV1K and MV2D) used to amplify full-length/subgenomic transcripts of MuERVs. (B) The brain and two non-nervous tissues (adrenal gland and salivary gland) were examined for the expression of MuERV full-length/subgenomic transcripts. A ∼5 kb band and several others, including a doublet of ∼2.7 and ∼2.9 kb sizes, and ∼1 kb band, were present in the brain. (C) The genomic structure of the ∼5 kb transcript was identified and is presented in comparison to a full-length reference MuLV (JO2255)
It was of interest to note that the same transcript, which was induced in the liver after burn injury, was constitutively expressed at a substantial level in the brain implicating potential pathophysiologic roles of this BR-b-1-derived putative MuERV.
Transcription potentials of U3 promoters of putative MuERVs constitutively expressed in the brain
To examine the transcription potentials of the putative MuERVs isolated using the BR-a-3 and BR-b-1 U3 clones as probes, profiles of transcription regulatory elements on these U3 promoters were determined and compared to the 14 other unique U3 clones isolated from four different tissues (Table 2). Nine elements (CCAAT/enhancer binding protein, TATA box, E2F, member of RSRF protein family, nuclear matrix protein 4, PAX6 paired domain, v-Myb, Yin and Yang 1 repressor site, and zinc finger transcription factor RU49) were present in all 16 U3 promoter sequences. None of the a-U3 derived U3 promoters had a transcription regulatory element profile that matched any of the profiles from the b/c-U3 derived U3 promoters. In addition, there were no transcription regulatory elements which are present only in the brain-derived BR-a-3 and BR-b-1 U3 promoters.
Further investigation into the roles of these elements in MuERV expression will aid in understanding how the transcription environment within the brain controls the constitutive and differential expression of MuERVs. In addition to these regulatory elements, there are two main factors contributing to the distinct MuERV expression profile of the brain. First, a pool of transcription factors and associated machinery (e.g., splicing) in the brain generally control the expression of individual MuERVs. Second, epigenetic genomic configuration (e.g., DNA methylation, histone acetylation) within the brain may limit or enhance the expression of MuERVs at certain chromosomal loci [33, 34].
References
D.N. Cooper, Nature encyclopedia of the human genome (Nature Publishing Group New York, 2003)
E.S. Lander, L.M. Linton, B. Birren, C. Nusbaum, M.C. Zody, J. Baldwin, K. Devon, K. Dewar, M. Doyle, W. FitzHugh, R. Funke, D. Gage, K. Harris, A. Heaford, J. Howland, L. Kann, J. Lehoczky, R. LeVine, P. McEwan, K. McKernan, J. Meldrim, J.P. Mesirov, C. Miranda, W. Morris, J. Naylor, C. Raymond, M. Rosetti, R. Santos, A. Sheridan, C. Sougnez, N. Stange-Thomann, N. Stojanovic, A. Subramanian, D. Wyman, J. Rogers, J. Sulston, R. Ainscough, S. Beck, D. Bentley, J. Burton, C. Clee, N. Carter, A. Coulson, R. Deadman, P. Deloukas, A. Dunham, I. Dunham, R. Durbin, L. French, D. Grafham, S. Gregory, T. Hubbard, S. Humphray, A. Hunt, M. Jones, C. Lloyd, A. McMurray, L. Matthews, S. Mercer, S. Milne, J.C. Mullikin, A. Mungall, R. Plumb, M. Ross, R. Shownkeen, S. Sims, R.H. Waterston, R.K. Wilson, L.W. Hillier, J.D. McPherson, M.A. Marra, E.R. Mardis, L.A. Fulton, A.T. Chinwalla, K.H. Pepin, W.R. Gish, S.L. Chissoe, M.C. Wendl, K.D. Delehaunty, T.L. Miner, A. Delehaunty, J.B. Kramer, L.L. Cook, R.S. Fulton, D.L. Johnson, P.J. Minx, S.W. Clifton, T. Hawkins, E. Branscomb, P. Predki, P. Richardson, S. Wenning, T. Slezak, N. Doggett, J.F. Cheng, A. Olsen, S. Lucas, C. Elkin, E. Uberbacher, M. Frazier, R.A. Gibbs, D.M. Muzny, S.E. Scherer, J.B. Bouck, E.J. Sodergren, K.C. Worley, C.M. Rives, J.H. Gorrell, M.L. Metzker, S.L. Naylor, R.S. Kucherlapati, D.L. Nelson, G.M. Weinstock, Y. Sakaki, A. Fujiyama, M. Hattori, T. Yada, A. Toyoda, T. Itoh, C. Kawagoe, H. Watanabe, Y. Totoki, T. Taylor, J. Weissenbach, R. Heilig, W. Saurin, F. Artiguenave, P. Brottier, T. Bruls, E. Pelletier, C. Robert, P. Wincker, D.R. Smith, L. Doucette-Stamm, M. Rubenfield, K. Weinstock, H.M. Lee, J. Dubois, A. Rosenthal, M. Platzer, G. Nyakatura, S. Taudien, A. Rump, H. Yang, J. Yu, J. Wang, G. Huang, J. Gu, L. Hood, L. Rowen, A. Madan, S. Qin, R.W. Davis, N.A. Federspiel, A.P. Abola, M.J. Proctor, R.M. Myers, J. Schmutz, M. Dickson, J. Grimwood, D.R. Cox, M.V. Olson, R. Kaul, C. Raymond, N. Shimizu, K. Kawasaki, S. Minoshima, G.A. Evans, M. Athanasiou, R. Schultz, B.A. Roe, F. Chen, H. Pan, J. Ramser, H. Lehrach, R. Reinhardt, W.R. McCombie, de la M. Bastide, N. Dedhia, H. Blocker, K. Hornischer, G. Nordsiek, R. Agarwala, L. Aravind, J.A. Bailey, A. Bateman, S. Batzoglou, E. Birney, P. Bork, D.G. Brown, C.B. Burge, L. Cerutti, H.C. Chen, D. Church, M. Clamp, R.R. Copley, T. Doerks, S.R. Eddy, E.E. Eichler, T.S. Furey, J. Galagan, J.G. Gilbert, C. Harmon, Y. Hayashizaki, D. Haussler, H. Hermjakob, K. Hokamp, W. Jang, L.S. Johnson, T.A. Jones, S. Kasif, A. Kaspryzk, S. Kennedy, W.J. Kent, P. Kitts, E.V. Koonin, I. Korf, D. Kulp, D. Lancet, T.M. Lowe, A. McLysaght, T. Mikkelsen, J.V. Moran, N. Mulder, V.J. Pollara, C.P. Ponting, G. Schuler, J. Schultz, G. Slater, A.F. Smit, E. Stupka, J. Szustakowski, D. Thierry-Mieg, J. Thierry-Mieg, L. Wagner, J. Wallis, R. Wheeler, A. Williams, Y.I. Wolf, K.H. Wolfe, S.P. Yang, R.F. Yeh, F. Collins, M.S. Guyer, J. Peterson, A. Felsenfeld, K.A. Wetterstrand, A. Patrinos, M.J. Morgan, de P. Jong, J.J. Catanese, K. Osoegawa, H. Shizuya, S. Choi, Y.J. Chen, Nature 409, 860–921 (2001)
R.H. Waterston, K. Lindblad-Toh, E. Birney, J. Rogers, J.F. Abril, P. Agarwal, R. Agarwala, R. Ainscough, M. Alexandersson, P. An, S.E. Antonarakis, J. Attwood, R. Baertsch, J. Bailey, K. Barlow, S. Beck, E. Berry, B. Birren, T. Bloom, P. Bork, M. Botcherby, N. Bray, M.R. Brent, D.G. Brown, S.D. Brown, C. Bult, J. Burton, J. Butler, R.D. Campbell, P. Carninci, S. Cawley, F. Chiaromonte, A.T. Chinwalla, D.M. Church, M. Clamp, C. Clee, F.S. Collins, L.L. Cook, R.R. Copley, A. Coulson, O. Couronne, J. Cuff, V. Curwen, T. Cutts, M. Daly, R. David, J. Davies, K.D. Delehaunty, J. Deri, E.T. Dermitzakis, C. Dewey, N.J. Dickens, M. Diekhans, S. Dodge, I. Dubchak, D.M. Dunn, S.R. Eddy, L. Elnitski, R.D. Emes, P. Eswara, E. Eyras, A. Felsenfeld, G.A. Fewell, P. Flicek, K. Foley, W.N. Frankel, L.A. Fulton, R.S. Fulton, T.S. Furey, D. Gage, R.A. Gibbs, G. Glusman, S. Gnerre, N. Goldman, L. Goodstadt, D. Grafham, T.A. Graves, E.D. Green, S. Gregory, R. Guigo, M. Guyer, R.C. Hardison, D. Haussler, Y. Hayashizaki, L.W. Hillier, A. Hinrichs, W. Hlavina, T. Holzer, F. Hsu, A. Hua, T. Hubbard, A. Hunt, I. Jackson, D.B. Jaffe, L.S. Johnson, M. Jones, T.A. Jones, A. Joy, M. Kamal, E.K. Karlsson, D. Karolchik, A. Kasprzyk, J. Kawai, E. Keibler, C. Kells, W.J. Kent, A. Kirby, D.L. Kolbe, I. Korf, R.S. Kucherlapati, E.J. Kulbokas, D. Kulp, T. Landers, J.P. Leger, S. Leonard, I. Letunic, R. Levine, J. Li, M. Li, C. Lloyd, S. Lucas, B. Ma, D.R. Maglott, E.R. Mardis, L. Matthews, E. Mauceli, J.H. Mayer, M. McCarthy, W.R. McCombie, S. McLaren, K. McLay, J.D. McPherson, J. Meldrim, B. Meredith, J.P. Mesirov, W. Miller, T.L. Miner, E. Mongin, K.T. Montgomery, M. Morgan, R. Mott, J.C. Mullikin, D.M. Muzny, W.E. Nash, J.O. Nelson, M.N. Nhan, R. Nicol, Z. Ning, C. Nusbaum, O(Connor M.J., Y. Okazaki, K. Oliver, E. Overton-Larty, L. Pachter, G. Parra, K.H. Pepin, J. Peterson, P. Pevzner, R. Plumb, C.S. Pohl, A. Poliakov, T.C. Ponce, C.P. Ponting, S. Potter, M. Quail, A. Reymond, B.A. Roe, K.M. Roskin, E.M. Rubin, A.G. Rust, R. Santos, V. Sapojnikov, B. Schultz, J. Schultz, M.S. Schwartz, S. Schwartz, C. Scott, S. Seaman, S. Searle, T. Sharpe, A. Sheridan, R. Shownkeen, S. Sims, J.B. Singer, G. Slater, A. Smit, D.R. Smith, B. Spencer, A. Stabenau, N. Stange-Thomann, C. Sugnet, M. Suyama, G. Tesler, J. Thompson, D. Torrents, E. Trevaskis, J. Tromp, C. Ucla, A. Ureta-Vidal, J.P. Vinson, A. Von, C. Niederhausern, C.M. Wade, M. Wall, R.J. Weber, R.B. Weiss, M.C. Wendl, A.P. West, K. Wetterstrand, R. Wheeler, S. Whelan, J. Wierzbowski, D. Willey, S. Williams, R.K. Wilson, E. Winter, K.C. Worley, D. Wyman, S. Yang, S.P. Yang, E.M. Zdobnov, M.C. Zody, E.S. Lander, Nature 420, 520–562 (2002)
K. Cho, T. Pham, L. Adamson, D. Greenhalgh, J Surg Res 115, 318–324 (2003)
K. Cho, L.K. Adamson, D.G. Greenhalgh, J Surg Res 104, 53–62 (2002)
B. Conrad, R.N. Weissmahr, J. Boni, R. Arcari, J. Schupbach, B. Mach, Cell 90, 303–313 (1997)
O. Frank, M. Giehl, C. Zheng, R. Hehlmann, C. Leib-Mosch, W. Seifarth, J Virol 79, 10890–10901 (2005)
H. Karlsson, S. Bachmann, J. Schroder, J. McArthur, E.F. Torrey, R.H. Yolken, Proc Natl Acad Sci USA 98, 4634–4639 (2001)
N. Talal, R.F. Garry, P.H. Schur, S. Alexander, M.J. Dauphinee, I.H. Livas, A. Ballester, M. Takei, H. Dang, J Clin Invest 85, 1866–1871 (1990)
H.B. Urnovitz, W.H. Murphy, Clin Microbiol Rev 9, 72–99 (1996)
J.M. Antony, van G. Marle, W. Opii, D.A. Butterfield, F. Mallet, V.W. Yong, J.L. Wallace, R.M. Deacon, K. Warren, C. Power, Nat Neurosci 7, 1088–1095 (2004)
P.E. Kowalski, J.D. Freeman, D.L. Mager, Genomics 57, 371–379 (1999)
P.E. Kowalski, D.L. Mager, J Virol 72, 6164–6168 (1998)
D.L. Mager, D.G. Hunter, M. Schertzer, J.D. Freeman, Genomics 59, 255–263 (1999)
C.P. Walsh, J.R. Chaillet, T.H. Bestor, Nat Genet 20, 116–117 (1998)
J.A. Yoder, C.P. Walsh, T.H. Bestor, Trends Genet 13, 335–340 (1997)
A. Nakamura, Y. Okazaki, J. Sugimoto, T. Oda, Y. Jinno, J Hum Genet 48, 575–581 (2003)
Y. Stauffer, S. Marguerat, F. Meylan, C. Ucla, N. Sutkowski, B. Huber, T. Pelet, B. Conrad, Immunity 15, 591–601 (2001)
N. Sutkowski, B. Conrad, D.A. Thorley-Lawson, B.T. Huber, Immunity 15, 579–589 (2001)
K. Cho, D. Greenhalgh, Virus Res 93, 189–198 (2003)
K. Tomonaga, J.M. Coffin, J Virol 73, 4327–4340 (1999)
S. Kumar, K. Tamura, M. Nei, Brief Bioinform 5, 150–163 (2004)
N. Saitou, M. Nei, Mol Biol Evol 4, 406–425 (1987)
K. Tomonaga, J.M. Coffin, J Virol 72, 8289–8300 (1998)
E.D. Sverdlov, Bioessays 22, 161–171 (2000)
F. Harada, G.G. Peters, J.E. Dahlberg, J Biol Chem 254, 10979–10985 (1979)
K.N. Nikbakht, C.Y. Ou, L.R. Boone, P.L. Glover, W.K. Yang, J Virol 54, 889–893 (1985)
B. Weasner, C. Salzer, J.P. Kumar, Dev Biol 303, 756–771 (2007)
D. Bornemann, E. Miller, J. Simon, Development 122, 1621–1630 (1996)
R. Saiki, A. Nagata, T. Kainou, H. Matsuda, M. Kawamukai, FEBS J 272, 5606–5622 (2005)
M. Huang, P. Jolicoeur, J Virol 64, 5764–5772 (1990)
S. Gayama, B.A. Vaupel, O. Kanagawa, Int Immunol 7, 861–868 (1995)
M.A. Ramirez, E. Pericuesta, R. Fernandez-Gonzalez, P. Moreira, B. Pintado, A. Gutierrez-Adan, Reprod Biol Endocrinol 4, 55 (2006)
J. Pothlichet, M. Mangeney, T. Heidmann, Int J Cancer 119, 1869–1877 (2006)
Acknowledgments
This study was supported by grants from Shriners of North America (No. 8680) and National Institutes of Health (R01 GM071360).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kwon, DN., Nguyen, S., Chew, A. et al. Identification of putative endogenous retroviruses actively transcribed in the brain. Virus Genes 36, 439–447 (2008). https://doi.org/10.1007/s11262-008-0216-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11262-008-0216-2