Abstract
The plant used in the characterization of Fragaria chiloensis latent virus (FClLV) contained double-stranded RNA (dsRNA) bands that did not correspond in size to the genomic or subgenomic RNAs of known strawberry viruses. Using shotgun cloning, sequences of a cryptic virus, named Fragaria chiloensis cryptic virus (FCCV), were obtained. This communication presents the complete genome sequence of FCCV. The genome of FCCV consists of three monocistronic RNAs, an unusual feature for cryptic viruses that are normally bipartite. The largest molecule encodes an RNA-dependent RNA polymerase, while the other two encode closely related proteins predicted to be the coat proteins of the virus. Phylogenetic analysis showed that FCCV is related most closely to members of the fungi-infecting Partitivirus genus rather than members of the plant-infecting Alphacryptovirus, providing a possible insight into the evolution of cryptic viruses. More than 300 strawberry plants from North and South America were tested for FCCV, and the virus was only detected in plants from South America.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
A rapid decline of strawberry has emerged in the western North America beginning in 2000 in British Columbia, Canada. In 2002 and 2003 there were significant losses in California. Plants decline rapidly with leaves turning red and becoming necrotic. Many of the diseased plants die and yield is reduced dramatically [1]. The decline is of viral etiology and manifests itself when plants are infected with three or more viruses [2]. Virus species vary depending on the geographic region, with the whitefly-borne viruses being predominant but in complex with one or more aphid-borne viruses in California and the aphid-borne viruses causing the disease in the Pacific Northwest [3]. Since symptoms are expressed after the accumulation of several but not any one specific strawberry virus complex, development of sensitive detection protocols for the poorly characterized viruses that could be involved in the disease and remained undetected was essential. One of the poorly characterized viruses at the molecular level was Fragaria chiloensis latent virus (FClLV), a virus described in Fragaria chiloensis plants (Chilean strawberry) from Chile [4]. Double-stranded RNA (dsRNA) extracted from the plant used in FClLV characterization contained additional bands that did not correspond to the genomic or subgenomic RNAs of the virus and were absent from virus-free strawberry plants. Several clones obtained from shotgun cloning from this plant corresponded to a novel cryptic virus [5] designated as Fragaria chiloensis cryptic virus (FCCV). We decided to characterize FCCV and test for it in major strawberry producing areas of North America and Chile to estimate the geographic distribution of the virus.
Materials and methods
Plant material
The sequence of FCCV was obtained from the National Clonal Germplasm Repository (Corvallis, OR) asymptomatic F. chiloensis accession CFRA 9089. This accession is a clone of a seedling grown from seed collected near Lonquimay, Chile [6]. In addition to CFRA 9089, dsRNA was extracted from accessions CFRA 9087, 748.001, 757.001, 758.001, 891.003, 891.006, and 892.008 (Fig. 1). All accessions are clones of seedlings, with accession 9087 and 9089 obtained from Lonquimay and all others from Lake Conguillo, Chile. A total of 322 strawberry samples from Florida, North Carolina, Maryland, New York, Ohio, California, Oregon, Washington, British Columbia, Guelph, and Ontario in North America and Chile in South America were tested for FCCV (Table 1).
Double-stranded RNA (dsRNA) extracted from National Clonal Germplasm Repository accessions CFRA 891.003, 891.006, and 892.008 infected with Fragaria chiloensis cryptic virus. Lane 1: Molecular marker (NEB); lanes 2–4: dsRNA from NCGR 891.003, 891.006, and 892.008, respectively; lane 5: dsRNA extracted from healthy strawberry plant
Virus characterization and phylogenetic analysis
Virus purification was done by following the protocol of MacDonald et al. [7] and virus preparations were visualized after negative staining with 2% molybdenum acetate in a Philips CM-12 electron microscope. DsRNA extraction, cloning, and sequencing of FCCV were done as described for FClLV [8] with the exception of the 5′ termini of the RNAs, where 5′ RACE was used (Invitrogen). In order to determine the 5′ terminal nucleotide of the RNAs two tailing reactions with different nucleotides were performed [9]. The sequence of FCCV was assembled using CAP3 with clones and reverse transcription-polymerase chain reaction (RT-PCR) products that gave an at least 3 × genome coverage [10] and deposited in Genbank under accession numbers DQ093961, DQ355440, and DQ355439 for RNA-1, -2, and -3, respectively. Phylogenetic analysis sequence comparisons were performed with ClustalW [11] using bootstrap analysis of 1000 pseudoreplicates. The secondary structure of FCCV and other cryptic virus proteins were predicted using PSIPRED v2.5 [12].
Detection
For detection, total RNA was extracted and reverse transcribed as described [13]. For PCR amplification either Taq polymerase (GenScript) or Platinum Taq (Invitrogen) were used. The integrity of the RNA was tested using internal control primers ICPF/ICPR [14]. Primers FCCV1F/ FCCV1R were used for the detection of FCCV [5] (Fig. 2). The dsRNA from the eight accessions mentioned above was additionally used for RT-PCR amplification of regions of each RNA using primers FCCV1F/FCCV1R for RNA 1, FCCV2F (5′-TGGCGACTGACAATAGCGAC-3′), FCCV2R (5′-TTACTGG TGTAGGCGTGCGA-3′) and FCCV3F (5′-CGAATGATCGAGCCTGCTGC-3′), FCCV3R (5′-GATGTATTGAGCGCAGTTTA-3′) that amplify a 300 base region of RNA 2 and a 400-base region RNA 3, respectively, to verify the presence of the three RNAs in all isolates.
Detection of Fragaria chiloensis cryptic virus in Chilean Fragaria chiloensis plants. Lane 9: 100 base pairs molecular marker (NEB); lanes 1–8 and 10–13: F. chiloensis plants grown from seed collected from Lonquimay, Chile; lanes 14–17 F. chiloensis plants grown from seed collected from near Lake Conguillio, Chile
Cryptic viruses are closely related to fungal viruses. To eliminate the possibility that FCCV infects an endophytic fungus, DNA was extracted using the DNasy Plant Minikit (Qiagen) from the accessions used for the dsRNA extractions and a Botrytis cinerea colony and amplified using primers ITS 1/4 [15]. PCR was performed in a Stratagene Robocycler and consisted of denaturation for 3 min at 94°C followed by 40 cycles of 30 sec at 94°C, 30 sec at 57°C for primers FCCV1F/ FCCV1R, FCCV2F/2R, and FCCV3F/3R and 66°C for ITS 1/4 and 60 sec at 72°C. The program terminated after 5-min incubation at 72°C.
Results and discussion
Unlike other viruses that could be transmitted to herbaceous hosts mechanically or by vectors, cryptic viruses are transmitted solely with pollen and seed [16, 17]. For this reason we had to use strawberry, a recalcitrant host, for FCCV purification. This fact, in addition to the low titer of cryptic viruses in plants [16, 18, 19], made purification a challenging undertaking. We tried to purify the virus from CFRA 9089 using FClLV as an internal control for our ability to purify spherical viruses from the host. This protocol has been used successfully with herbaceous hosts for the purification of other ilarviruses and Strawberry latent ringspot virus, but we were unable to visualize virus particles. Attempts to visualize particles from the plant reproductive organs, fruit, and achenes were also unsuccessful.
The sizes of FCCV RNAs are 1734, 1479, and 1465 nt for RNAs −1, −2, and −3, respectively; and the first ten and the last three nucleotides of each of the three RNAs are identical. RNA 1 encodes a single protein that starts at nt 183 and terminates at nt 1622. Database searches indicated that the 56 kDa protein has core RNA-dependent RNA polymerases (RdRp) motifs between residues 164 and 363 [20] and is most closely related to the RdRps of RNA 3 from Raphanus sativus-root (RasR 3) [21] with 60% amino acid (aa) identity (76% aa similarity) and Beet cryptic virus-3 (BCV-3) [22] with 34% aa identity (56% aa similarity) [23] (Fig. 3).
Alignment of the polymerase conserved motifs [12] of Fragaria chiloensis cryptic virus (FCCV) and closely related cryptic viruses. Abbreviations and GenBank accession numbers: FCCV, AAZ06131; Raphanus sativus-root RNA 3, RasR 3, ABB04855; Pinus sylvestris virus, PinSV, AAY51483; Beet cryptic virus-3, BCV 3, AAB27624; Pyrus pyrifolia cryptic virus, PPCV, AB012616. Globally conserved residues are in black background, identical in gray, similar in light gray. The consensus line indicates identical residues in capital and most common residues in lower case letters
RNAs 2 and 3 are also monocistronic with open reading frames (ORF) spanning between nt 191–1237 of RNA 2 and nt 172–1212 of RNA 3. The putative 39 kDa proteins are most closely related to each other (23% aa identity, 43% aa similarity) and the putative proteins encoded by RNAs 4 and 5 (RasR 4 and RasR 5, respectively) from Raphanus sativus-root [21], than any other proteins found in the databases. Structural analysis of the putative proteins (data not shown) [12] show near identical folding of the proteins indicating that they are probably involved in the same function. Since RasR 3, 4, and 5 were purified from virus-like particles [21], we predict that the proteins encoded by FCCV RNA 2 and 3 and RasR 4 and 5 are the coat proteins of the viruses. While Chen et al. [21] speculated that one of the molecules may be a satellite RNA, we provide evidence to the contrary. We have tested plants grown from seed that originated from different locations and all had three genomic molecules (Fig. 4). In addition, a new cryptic virus in rose shares the same genomic characteristics as FCCV with a tripartite genome and RNAs 2 and 3 encoding closely related proteins (Sabanadzovic, personal communication). The idea that three different viruses from divergent hosts share satellite RNAs encoding highly similar proteins is unprecedented. Furthermore, in the case of FCCV the three RNAs are present in all seed-transmitted isolates from different locations. More than 50 clones were sequenced from the original CFRA 9089 shotgun cloning and all corresponded to FClLV, FCCV, or host sequences. This minimizes the possibility that another RNA, encoding for an RNA-dependent RNA polymerase is present in CFRA 9089 and supports the hypothesis that RNAs 2 and 3 code for coat proteins of FCCV. Although there are other cryptic viruses composed of three genomic molecules [24 and references within], these reports did not provide any information on the proteins encoded by these viruses. The similarity of the proteins encoded by RNAs 2 and 3 of FCCV, RasR 4 and RasR 5 of Raphanus sativus-root and RNAs 2 and 3 of the novel rose cryptic virus indicates that there was a duplication event before the speciation event of the viruses and the proteins continue to function as the coat proteins of the viruses. Cryptic viruses are transmitted solely through seed and pollen and in most cases are presumed to be asymptomatic in plants. These characteristics indicate that cryptic viruses probably do not undergo many major ‘bottleneck’ events that constrain viruses transmitted by vectors and/or those that cause severe symptoms in their hosts, leading to rapid divergence in the evolving virus-vector and virus-host interactions.
Detection of the genomic molecules of Fragaria chiloensis cryptic virus. Lanes 10 and 20: 100 base pairs molecular marker (NEB); Lanes 1–9: amplicons from RNA 1; lanes 11–19: amplicons from RNA 2; lanes 21–29: amplicons from RNA 3. The amplicons were obtained from dsRNA extracted from National Clonal Germplasm Repository accessions CFRA 748.001, 757.001, 758.001, 891.003, 891.006, 892.008, 9087, 9089, and a healthy strawberry control, respectively
Fragaria and Raphanus belong to different plant families and in view of the mode of transmission of cryptic viruses, the common ancestor of FCCV and RasR cryptic virus would have to infect plants before the divergence of the two plant families as proposed for other cryptic viruses [25]. This is unlikely given the high similarity shared between the two viruses. A more plausible theory is that FCCV and RasR cryptic virus evolved from a virus of a plant pathogen, most likely a fungal virus. This hypothesis is supported by the phylogenetic analysis of the polymerases of members of the Partitiviridae (Fig. 5), where FCCV and RasR cryptic virus are more closely related to fungi-infecting members of the genus Partitivirus than members of the plant-infecting genus Alphacryptovirus with the exception of BCV-3. For this reason we investigated the possibility that FCCV infects an endophytic fungus. No amplicons were obtained when fungal specific ITS primers were used in an attempt to detect the presence of a fungus in the FCCV-infected plants, whereas the fungal control gave positive results suggesting that FCCV is a plant and not a fungal virus.
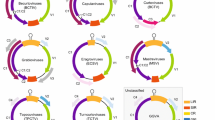
Unrooted phylogram of the polymerases of members of the Partitiviridae. Abbreviations and Genbank accessions: Beet cryptic virus-3, BCV 3, AAB27624; Fragaria chiloensis cryptic virus, FCCV, AAZ06131; Helicobasidium mompa virus, HMV, BAC23065; Helicobasidium mompa partitivirus V1–2, HMV V1–2, AB110980; Oyster mushroom isometric virus, OMIV, AAP74192; Penicillium stoloniferum virus F, PSV F, YP 271922; Pepper cryptic virus-1, PCV 1, ABC96789; Pinus sylvestris virus, PinSV, AAY51483; Pyrus pyrifolia cryptic virus, PPCV, AB012616; Raphanus sativus-root RNA 1, RasR 1, YP 656506; Raphanus sativus-root RNA 3, RasR 3, ABB04855; Rhizoctonia solani virus, RSV, NP 620659; Vicia cryptic virus, VCV, ABN71241; White clover cryptic virus-1, WCCV 1, YP 086754. Bootstrap values are shown on the clads. The bar represents 0.1 amino acid changes per site
In the light of these results, we speculate that fungal viruses escaped from fungal pathogens of plants and evolved to adapt to their new hosts. The remnants of this process would then be the two very divergent groups of plant cryptic viruses that are more closely related to their fungal virus ancestors than to each other.
Sequence comparisons and phylogenetic analyses (Figs. 3, 5) clearly indicate that BCV-3 does not belong to the genus Alphacryptovirus [26] but probably to the same genus as FCCV and RasR cryptic virus. There is no molecular information on members of the genus Betacryptovirus other than that betacryptoviruses have two dsRNA segments of over 2 kilobases [26]. This is in high contrast to FCCV and RasR cryptic virus that are tripartite and have genomic molecules smaller than 2 kilobases. We propose the development of a new genus of cryptic viruses that would include FCCV and RasR cryptic virus and closely related viruses including BCV-3, Pinus sylvestris virus, and Pyrus pyrifolia cryptic virus (Fig. 3).
Fragaria. chiloensis is one of the parents of modern strawberry, F. × ananassa, and is still used widely by breeders worldwide as a source of desirable traits in cultivar development. This practice could introduce FCCV into cultivars and the lack of symptoms could result in the virus being undetected. However, if there is an interaction between FCCV and strawberry that changes the physiology of the host as in the case of White clover cryptic virus-1 and Lotus japonicus [27] or FCCV and one of the other strawberry viruses it could have a significant impact on production. We tested more than 300 samples from important strawberry production areas from North and South America. All samples from North America were free of the virus while plants from both locations in Chile were infected with the virus, all of which were clones of seedlings verifying the seed transmission of the virus. These results also indicate that FCCV is an indigenous virus of South America that has probably not yet been introduced to North America.
References
R.R. Μartin, I.E. Tzanetakis, Plant Dis. 90, 384 (2006)
I.E. Tzanetakis, PhD dissertation, Oregon State University, USA, 2004, p.136
I.E. Tzanetakis, M. Bolda, R.R. Martin, Phytopathology. 94, S104 (2004)
S. Spiegel, R.R. Martin, F. Leggett, M. ter Borg, J. Postman, Phytopathology 83, 991 (1993)
I.E. Tzanetakis, R.R. Martin, Plant Dis. 89, 1241 (2005)
J.S. Cameron, T.M. Sjulin, J.R. Ballington, C.H. Shanks Jr, C.E. Munoz, A. Lavin, Acta Hort. 348, 65 (1993)
S.G. MacDonald, R.R. Martin, P.R. Bristow, Phytopathology 81, 210 (1991)
I.E. Tzanetakis, R.R. Martin, Virus Res. 112, 32–37 (2005)
I.E. Tzanetakis, R.R. Martin, Virus Genes 28, 239 (2004)
X. Huang, A. Madan, Genome Res. 9, 868 (1999)
J.D. Thompson, D.G. Higgins, T.J. Gibson, Nucleic Acids Res. 22, 4673 (1994)
L.J. McGuffin, K. Bryson, D.T. Jones, Bioinformatics 16, 404 (2000)
I.E. Tzanetakis, A.B. Halgren, N. Mosier, R.R. Martin, Virus Res. 127, 26 (2007)
I.E. Tzanetakis, J.D. Postman, R.R. Martin, Arch. Virol. 152, 2027 (2007)
T.J. White, T. Brun, S. Lee, J. Taylor, in PCR Protocols, a Guide to Methods and Applications, ed. by M.A. Innis, D.H. Gelfand, J.J. Sninsky, T.J. White (Academic Press, New York, 1990), p. 315
G. Boccardo, V. Lisa, E. Luisoni, R.G. Milne, Adv. Virus Res. 32, 171 (1987)
B. Kassanis, G.E. Russell, R.F. While, Phytopathol. Z. 91, 76 (1978)
K. Hammer, A. Stanarius, T. Kuhne, Euphytica 45, 23 (1990)
D. Veliceasa, N. Enunlu, P.B. Kos, S. Koster, E. Beuther, B. Morgun, S.D. Deshmukh, N. Lukacs, Virus Genes 32, 177 (2006)
A. Marchler-Bauer S.H. Bryant, Nucleic Acids Res. 32, W327 (2004)
L. Chen, J.S. Chen, H. Zhang, S.N. Chen, Arch. Virol. 151, 2077 (2006)
W.S. Xie, J.F. Antoniw, R.F. White, J. Gen. Virol. 74, 1467 (1993)
J.J. Campanell, L. Bitincka, J. Smalley, BMC Bioinformatics 4, 29 (2003)
M.A. Abou-Elnasr, A.T. Jones, M.A. Mayo, J. Gen. Virol. 66, 2453 (1985)
E. Luisoni, R.G. Milne, G.P. Accotto, G. Boccardo, Intervirology 28, 144 (1987)
C.M. Fauquet, M.A. Mayo, J. Maniloff, U. Desselberger, L.A. Ball, Virus Taxonomy: VIIIth Report of the International Committee on Taxonomy of Viruses. (Elsevier/Academic Press, London, 2005)
M. Nakatsukasa-Akune, K. Yamashita, Y. Shimoda, T. Uchiumi, M. Abe, T. Aoki, A. Kamizawa, S. Ayabe, S. Higashi, A. Suzuki, Mol. Plant Microbe Interact. 10, 1069 (2005)
Acknowledgments
We thank Sead Sabanadzovic (Mississippi State University) for sharing the data on the rose cryptic virus, Joseph Postman (NCGR, Corvallis, OR), Mark Bolda (UC Davis), Mike Ellis (Ohio State University), Courtney Weber (Cornell University), Adam Dale (University of Guelph), Craig Chandler (University of Florida), and Barclay Poling (North Carolina State University) that provided strawberry samples and Walter Mahaffee (USDA-ARS, Corvallis, OR) for providing the Botrytis cinerea colony.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Tzanetakis, I.E., Price, R. & Martin, R.R. Nucleotide sequence of the tripartite Fragaria chiloensis cryptic virus and presence of the virus in the Americas. Virus Genes 36, 267–272 (2008). https://doi.org/10.1007/s11262-007-0186-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11262-007-0186-9