Abstract
Bamboo shoots grow quickly through the rapid elongation of internodes, but the precise molecular mechanisms underlying this process remain unknown. We used a combination of suppressive subtractive hybridization (SSH), dot blotting, sequencing and bioinformatics to identify Phyllostachys pubescens genes that are differentially expressed in rapidly elongating vs. static internodes (SIs). We isolated 1020 expressed sequence tags (ESTs) by SSH, 173 of which were shown to be differentially expressed by dot blotting. We then sequenced the 20 ESTs showing the greatest difference in expression, 13 of which were preferentially expressed in elongating internodes and seven in SIs. Functional characterization of the ESTs showed that rapid internode elongation requires meristem initiation and proliferation, high-level protein synthesis, cellular respiration, and cell wall synthesis, as well as the regulation of the activated methyl cycle, gibberellin and brassinosteroid biosynthesis, and their signal transduction pathways.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The bamboo subfamily (Bambusoideae) is a division of the grasses (Poaceae), which is further divided into nine subtribes comprising 77 genera and about 1030 species worldwide (Soderstrom and Ellis 1987; Dransfield and Widjaja 1995). In China, there are 48 genera and nearly 500 species, among which Phyllostachys pubescens accounts for over two thirds of commercially planted bamboo because it is valued as a source of building material, fodder, and food (Fu 2001). Bamboo generally grows faster than other woody plants, with culm elongation rates of 24.5 cm/day for Phyllostachys bambusoides, 9.7 cm/day for Bambusa oldhamii (Lin 1958), and >100 cm/day for P. pubescens (Ueda 1960; Li et al. 1997).
Internode elongation involves a number of steps including cell division within the intercalary meristem, cell elongation, and lignification (Martin 1988; Morrison et al. 1994). Several genes required for internode elongation have been indentified, such as ZmREV (RLD1), which is expressed in the adaxial portion of the leaf following the emergence of primordia (Juarez et al. 2004), and OsGLU1, which is involved in the synthesis and deposition of fibers in the extending cell walls (Zhou et al. 2006). Hormones such as gibberellin, ethylene, and abscisic acid are also thought to play an important role (Suge 1985; Azuma et al. 1990; Sauter et al. 1995). The precise molecular mechanism of rapid internode elongation in bamboo remains unknown, but the regulation of carbohydrate metabolism is likely to play a pivotal role because the bamboo shoot has no leaves, and the sheath-covered internodes are etiolated, so energy for growth must be derived from stored carbon sources imported from connected photosynthetic tissues (Liese and Weiner 1995; Okahisa et al. 2006). The respiration rate of P. pubescens shoots is also much higher than that of mature culms (Isagi et al. 1997), suggesting there is active metabolism and efficient energy production during elongation. Additionally, several different sucrose synthases play roles during bamboo growth, both in terms of polysaccharide biosynthesis and energy production (Chiu et al. 2006).
Unlike model plant species such as Arabidopsis thaliana and rice (Oryza sativa), there is relatively little genomic information available for bamboo species (Tang 2009). It is therefore necessary to develop tools and resources specific for bamboo in order to gain insight into its unique characteristics, such as rapid internodal growth. Suppressive subtractive hybridization (SSH) allows the construction of a cDNA library enriched for differentially expressed genes, which helps to identify genes that are developmentally regulated or induced/repressed in response to intracellular or external signals. In this study, we used cDNA derived from rapidly elongating internodes (REIs) and static internodes (SIs) of P. pubescens to develop an SSH library enriched for genes involved in the elongation of bamboo shoots.
Materials and Methods
Plant Materials and RNA Isolation
We selected a P. pubescens culm 1.82 m in length containing 43 visible internodes, which were numbered from the base to the canopy (Fig. 1). Internode 13 was the longest (16.2 cm) and was already lignified, whereas internodes 31, 32, and 33 were shorter (8.1, 7.3, and 6.9 cm, respectively) and were still very soft. This implied that internodes 31, 32, and 33 had the potential for rapid elongation, whereas internode 13 was likely to be static (Li et al. 1997). Wall tissue from internode 13 (static) and pooled wall tissue from REIs 31, 32, and 33 were frozen in liquid nitrogen and stored at −80°C. For RNA extraction, wall tissue was ground in liquid nitrogen and total RNA was isolated using Trizol reagent (Invitrogen, USA) according to the manufacturer's instructions. For each 500 μg of total RNA, 5 μg of poly(A)+ RNA was isolated using the Oligotex mRNA Mini Kit (QIAGEN, USA).
Internodes used for RNA extraction. Internode 13 was the longest and was static, whereas internodes 31, 32, and 33 were shorter and retained the capacity for rapid elongation. RNA from internode 13 and pooled RNA from internodes 31, 32, and 33 were used as templates in the two reciprocal SSH reactions
Suppressive Subtractive Hybridization
SSH was carried out using the PCR-Select cDNA Subtraction Kit (Clontech, California, USA) according to the manufacturer's instructions. After digesting the cDNA samples with RsaI, two SSH reactions were performed, one with cDNA derived from REIs as the driver and cDNA from the SIs as the tester, and another with the reverse arrangement. The subtractive products were amplified in two rounds of polymerase chain reaction (PCR) using oligonucleotide primers complementary to the adapters. The first round comprised a 5-min incubation at 95°C followed by 25 cycles at 94°C for 30 s, 66°C for 30 s, and 72°C for 90 s. The second nested PCR comprised 15 cycles at 94°C for 30 s, 66°C for 30 s, and 72°C for 90 s. The final PCR products in the two SSH experiments represented genes expressed predominantly in SIs and REIs, respectively.
Dot Blotting
The subtracted cDNAs identified above were purified using the QIAquick PCR Purification Kit (QIAGEN) and inserted into vector pMD18-T (TaKaRa Biotechnology Co., China). After transformation and selection in Escherichia coli JM109, 1020 independent colonies were used as PCR templates. T7prom/SP6 universal primers derived from pUC18T were used as forward and reverse primers to amplify the inserted cDNAs. PCR products were separated by 1% agarose gel electrophoresis, and 0.2 μg of each recovered product was dissolved in 0.3 M NaOH and spotted onto a positively charged nylon membrane (Roche, USA). Two membranes with the same lattice pattern were produced and baked in vacuo at 80°C for 2 h. The membranes were prehybridized for 30 min and then hybridized with denatured DIG-labeled probes for 12 h using the DIG Easy Hyb kit (Roche). Two sets of labeled probes were prepared, one from the forward-subtracted products and one from the reverse-subtracted products described in the preceding section. The probes were labeled using the DIG High Prime DNA Labeling and Detection Starter Kit II (Roche). Posthybridization washes with 0.5× standard saline citrate (SSC), 0.1% sodium dodecyl sulfate (SDS) were followed by incubations in blocking solution, antibody solution, and washing solutions according to the manufacturers' protocols before immunological detection with ready-to-use chloro-5-substituted adamantyl-1,2-dioxetane phosphate and exposure to X-ray film at 20°C for 15 to 20 min. The membranes were scanned using PowerLook 3000, and grayscale images were assessed visually and signal intensities were quantified using TINA 2.09e software (Raytest, Straubenhardt, Germany). Clones corresponding to dots whose signal intensity differed significantly between the REI and SI preparations were selected for further analysis.
Sequencing and Sequence Analysis
The selected cDNAs were sequenced using the BigDye Terminator V3.1 Cycle Sequencing Kit and an ABI 3100-Avant DNA Analyzer (Applied Biosystems, USA). The predicted unigene sequences were used to query the nonredundant nucleotide database at GenBank (http://www.ncbi.nlm.nih.gov/) using BLASTX.
Semiquantitative PCR
First-strand cDNAs were synthesized using mRNA isolated from REIs and SIs, and those showing significantly different expression levels were validated by semiquantitative reverse transcriptase (RT)-PCR, with actin mRNA used as a control (Matsui et al. 2004). The primers specific for each gene are listed in Table 1. The PCR conditions were as described above for the colony amplification procedures.
Results
Construction of the Subtracted Library
RNA derived from REIs and SIs was fractionated by agarose gel electrophoresis revealing bands >1 kb in length (Fig. 2a). Other quality indicators such as the 28 S:18 S intensity ratio (∼2:1), A260/A280 (>1.9), and A260/A230 (>1) confirmed that the RNA was suitable for further experiments (Schultz et al. 1994). A 2-μg aliquot of mRNA produced about 1.2 μg of double-stranded cDNA with a size range of 0.5 to 3 kb (Fig. 2b), which was reduced to 0.1 to 2 kb following digestion with RsaI (Fig. 2c). The efficiency of subtraction was confirmed by testing for the abundance of actin cDNA before and after subtraction. An actin product appeared after 15 PCR cycles for the unsubtracted cDNA and after 35 cycles for the subtracted cDNA (Fig. 2d).
Agarose gel electrophoresis of RNA and DNA from REIs and SIs. a RNA. Lane 1: REIs. Lane 2: SIs. Lane M: 1 kb plus DNA ladder (Invitrogen). b cDNA. Lane 1: REIs. Lane 2: SIs. Lane M: 1 kb plus DNA ladder. c Comparison of intact cDNA and cDNA digested with RsaI. 1: REIs. 2: SIs. Lane M: 1 kb plus DNA ladder. d Reduction of actin abundance by PCR-Select subtraction. PCR was performed with the actin forward and reverse primers (Table 1) using the subtracted cDNA (lanes 1–3) and the unsubtracted cDNA (lanes 5–7) as the template. Lanes 1–3: 15, 25, 35 cycles. Lanes 5–7: 15, 25, 35 cycles. Lane M: 100-bp DNA ladder (Biobasic Inc, USA)
Confirmation of Expressed Sequence Tags with Differential Expression
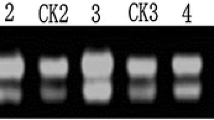
Differentially expressed cDNA products were cloned in vector pMD-18T and introduced into bacteria. In total, 1020 colonies were isolated. Colony PCR analysis revealed insert fragments >0.25 kb in length (Fig. 3). Dot blots confirmed that 173 genes were differentially expressed, 126 predominantly expressed in REIs, and 47 in SIs. This included 20 genes with a tenfold difference in signal intensity between the samples (a REI/SI signal intensity ratio of ≥10 or ≤0.1 based on dot blotting results), 13 of which were expressed predominantly in REIs and seven of which were expressed predominantly in SIs (Table 2). The expression profiles of ten of the genes were validated by semiquantitative RT-PCR, generating results that were approximately consistent with the dot blot experiments (Table 2 and Fig. 4).
Identification of Differentially Expressed Sequence Tags
The 20 candidate expressed sequence tags (ESTs) were sequenced (GenBank: GT917652–GT917660 and GT917662–GT917672), and BLASTX analysis revealed that 13 of the sequences matched genes with known functions (83–94% identity), three matched genes with unknown functions (REun-12, REun-13, and Stun-4; 86–89% identity), and the remaining four were orphan genes, lacking homology to any other sequence in GenBank (REun-3, STun-5, STun-6, and Stun-7). The 13 ESTs matching sequences with known functions included ten that were predominantly expressed in REIs and three predominantly expressed in SIs. The genes represented a number of different physiological and biochemical functions including meristem initiation and proliferation (REfa-1), cell growth and development (REce-2), protein metabolism (REpro-4, REpro-5), cellular respiration (REen-6), sugar metabolism (REsu-7), methylation (REmet-8, REmet-9), cell wall structure (REwa-10, REwa-11), hormone metabolism (STga-2), and signaling (STmo-1).
Discussion
The rapid growth of bamboo shoots is one of the most distinctive characteristics of bamboo development in comparison to other woody plants, but little is known about the molecular mechanisms underpinning this process. We used SSH to generate a library enriched for genes that are differentially expressed in rapidly elongating vs. SIs. Among 173 clones identified by dot blotting, we sequenced 20 ESTs whose expression level differed significantly between the two tissues, 13 of which were homologous to genes with a known function. The other seven were either orphans or homologous to genes whose functions have not been determined.
Previous studies have implicated several physiological processes in internode elongation. One of the earliest events is the initiation and proliferation of meristem tissue, which is regulated by REVOLUTA (REV)-like class III homeodomain leucine zipper (HD-Zip) proteins (Otsuga et al. 2001), such as ZmREV (RLD1) in maize (Juarez et al. 2004). REV proteins are also associated with fiber deposition and wood formation (Ohashi-Ito et al. 2005), e.g., LFL1 in A. thaliana (Zhong and Ye 1999) and PtaHB1 in aspen (Ko et al. 2005). We found that REfa-1, a homolog of the rice NIF3 gene encoding a REV protein, is up-regulated in REIs, suggesting that high-level expression of this REV gene in bamboo may be required for meristem initiation and fiber deposition. In Pleione praecox, the REV homolog PpHB1 is also expressed within the developing procambium, suggesting it may underlie the extensive lignification of bamboo shoots (Peng et al. 2007). After initiation, rapid but tightly regulated cell growth and division within the meristem precedes cell elongation and lignification. The SNARE superfamily (soluble N-ethylmaleimide-sensitive fusion protein attachment protein receptors; Sutter et al. 2006; Lipka et al. 2007) plays an important role in this process, particularly SNARE 11, a component of the cell plate membrane fusion machinery involved in membrane trafficking and fusion during cytokinesis (Zheng et al. 2002). The identification of REce-2 as a SNARE 11 homolog indicates that rapid but correctly regulated cell division is critical in the early stages of bamboo shoot growth.
The rapid growth in bamboo shoots also requires very efficient protein synthesis and recycling, so it is unsurprising that two of the differentially expressed ESTs (REpro-4 and REpro-5) correspond to genes involved in protein metabolism. REpro-4 corresponds to the 25 S ribosomal RNA gene. Its high-level expression in growing bamboo shoots indicates that the number of ribosomes may become a limiting factor during culm elongation. REpro-5 is homologous to a rice S10 serine carboxypeptidase gene. Increasing the availability of the protease may increase the plant's capacity for C-terminal processing, protein turnover, and recycling (Remington and Breddam 1994; Mortensen et al. 1999). In addition to the raw materials necessary for shoot elongation, a large amount of energy is also required. We identified REen-5 as a homolog of the aspen mitochondrial lipoamide dehydrogenase (LPD1) gene, which encodes a pivotal component of several supraenzymatic complexes involved in carbohydrate and energy metabolism. LPD1 is often associated with the Gly decarboxylase complex, which ultimately provides ATP (Williams 1992; Vettakkorumakankav and Patel 1996). The high-level expression of REen-5 in REIs shows that active cellular respiration is required for internode elongation (Lutziger and Oliver 2001), consistent with the much higher respiration rate observed in P. pubescens shoots compared to mature culms (Isagi et al. 1997).
Cell elongation is dependent on the laying down of fibers in the nascent cell walls. Wang et al. (2010) detected the high-level expression of starch-branching enzyme I in bamboo buds, and likewise, we found that REsu-7, which is expressed preferentially in REIs, is homologous to starch-branching enzyme genes. We found that REwa-10 was homologous to a maize cell wall hydroxyproline-rich glycoprotein gene (HRGP), whereas REwa-11 was homologous to the rice endo-β-1, 4-d-glucanase gene (OsGLU1), both of which were involved in the synthesis of cell wall components (Stiefel et al. 1990; Zhou et al. 2006). HRGP is expressed at sites of early vascular differentiation in embryos, coleoptiles, leaves, hypocotyls and both primary and lateral roots, as well as at much lower levels throughout developing maize plants (Stiefel et al. 1990). Loss of function mutations in the OsGLU1 gene result in rice plants with shorter and narrower cells than wild-type plants (Zhou et al. 2006). The slightly elevated expression of REwa-10 and REwa-11 in REIs may facilitate the synthesis and deposition of fibers in the extending cell walls.
Two further ESTs up-regulated in REIs (REmet-8 and REmet-9) were found to be homologous to a putative rice methyltransferase gene (DUF248) and a wheat S-adenosyl-l-homocysteine hydrolase (SAHH) gene. Methyltransferases are predominantly part of the activated methyl cycle, providing key precursors in amino acid and nucleotide biosynthesis (Shu et al. 2006). SAHH is a key enzyme involved in DNA methylation (Tanaka et al. 1997), which is essential for normal plant development (Chan et al. 2005). Both genes would need to be expressed more strongly in REIs to facilitate the increase in cell number and biomass that occurs during rapid internode elongation.
Genes that affect plant height are often found to encode enzymes involved in hormone metabolism or components of the corresponding signaling pathways (Peng et al. 1997, 1999). We identified two ESTs (STmo-1 and STga-2) predominantly expressed in SIs. STmo-1 was homologous to a rice shaggy-related protein kinase (ASK-eta), which is a negative regulator of brassinosteroid signaling (Clouse and Sasse 1998; Li and Nam 2002), whereas STga-2 was homologous to rice GA 2-oxidase, whose role is to inactivate gibberellins by 2β-hydroxylation (Pimenta and Lange 2006). The preferential expression of these ESTs in SIs suggested that rapid stem elongation in bamboo requires the derepression of a negative regulatory system.
In conclusion, we identified 20 promising candidates from a panel of >173 ESTs that were differentially expressed in rapidly elongating vs. SIs in bamboo, using a combination of SSH, dot blotting, and semiquantitative RT-PCR for verification. Thirteen of the sequences could be assigned putative functions on the basis of homology searching, revealing the involvement of proteins involved in meristem initiation and proliferation, protein biosynthesis, cellular respiration, the regulation of methylation as part of the activated methyl cycle, starch accumulation, cell wall synthesis, hormone synthesis, and signal transaction. These data provide insight into the molecular mechanisms underlying the rapid internode elongation that characterizes the growth of bamboo shoots.
Abbreviations
- SSH:
-
suppressive subtractive hybridization
- ESTs:
-
expressed sequence tags
- HD-Zip:
-
homeodomain leucine zipper
- SNAREs:
-
soluble N-ethylmaleimide-sensitive fusion protein attachment protein receptors
- ASK:
-
shaggy-related protein kinase
- SAHH:
-
S-adenosyl-l-homocysteine hydrolase
- REIs:
-
rapidly elongating internodes
- SIs:
-
static internodes
References
Azuma T, Mihara F, Uchida N, Yasuda T, Yamaguchi T (1990) Plant hormonal regulation of internodal elongation of floating rice stem sections. Jpn J Trop Agr 34:271–275
Chan SWL, Henderson IR, Jacobsen SE (2005) Gardening the genome: DNA methylation in Arabidopsis thaliana. Nat Rev Genet 6:351–360
Chiu WB, Lin CH, Chang CJ, Hsieh MH, Wang AY (2006) Molecular characterization and expression of four cDNAs encoding sucrose synthase from green bamboo Bambusa oldhamii. New Phytol 170:53–63
Clouse SD, Sasse JM (1998) Brassinosteroids: essential regulators of plant growth and development. Ann Rev Plant Physiol Plant Mol Biol 49:427–451
Dransfield S, Widjaja EA (1995) Plant resources of south-east Asia No. 7 Bamboos. Backhuys Publishers, Leiden
Fu J (2001) Chinese moso bamboo: its importance. Bamboo 22:5–7
Isagi Y, Kawahara T, Kamo K, Ito H (1997) Net production and carbon cycling in a bamboo Phyllostachys pubescens stand. Plant Ecol 130:41–52
Juarez MT, Kui JS, Thomas J, Heller BA, Timmermans MCP (2004) MicroRNA-mediated repression of rolled leaf1 specifies maize leaf polarity. Nature 428:84–88
Ko JH, Prassinos C, Han K-H (2005) Developmental and seasonal expression of PtaHB1, a Populus gene encoding a class III HD-Zip protein, is closely associated with secondary growth and inversely correlated with the level of microRNA (miR166). New Phytol 169:469–478
Li J, Nam KH (2002) Regulation of brassinosteroid signaling by a GSK3/SHAGGY-like kinase. Science 295:1299–1301
Li R, Zhong ZC, Werger MJA (1997) Studies on the dynamics of the bamboo shoots in Phyllostachys pubescens. Acta Phytoecol Sin 21:53–59
Liese W, Weiner G (1995) Ageing of bamboo culms. A review. Wood Sci Technol 30:77–89
Lin WC (1958) Studies on the growth of bamboo species in Taiwan. Taiwan For Exp Inst Rep No. 248, Taibei
Lipka V, Kwon C, Panstruga R (2007) SNARE-Ware: the role of SNARE-domain proteins in plant biology. Annu Rev Cell Dev Biol 23:147–174
Lutziger I, Oliver DJ (2001) Characterization of two cDNAs encoding mitochondrial lipoamide dehydrogenase from Arabidopsis. Plant Physiol 127:615–623
Martin GG (1988) Cell growth in the maize stem. Botany 45:35–39
Matsui T, Bhowmik PK, Yokozeki K (2004) A cDNA sequence encoding actin gene in moso bamboo shoot and its phylogenetic analysis. Asian J Plant Sci 3:128–131
Morrison TA, Kessler JR, Buxton DR (1994) Maize internode elongation patterns. Crop Sci 34:1055–1060
Mortensen UH, Olesen K, Breddam K (1999) Carboxypeptidase C including carboxypeptidase. In: Barrett AJ, Rawlings ND, Woessner JF (eds) Handbook of proteolytic enzymes, vol 132. Academic Press, London, pp 389–393
Ohashi-Ito K, Kubo M, Demura T, Fukuda H (2005) Class III homeodomain leucine-zipper proteins regulate xylem cell differentiation. Plant Cell Physiol 46:1646–1656
Okahisa Y, Yoshimur T, Imamura Y (2006) Seasonal and height-dependent fluctuation of starch and free glucose contents in moso bamboo (Phyllostachys pubescens) and its relation to attack by termites and decay fungi. J Wood Sci 52:445–451
Otsuga D, DeGuzman B, Prigge MJ, Drews GN, Clark SE (2001) REVOLUTA regulates meristem initiation at lateral positions. Plant J 25:223–236
Peng J, Carol P, Richards DE, King KE, Cowling RJ, Murphy GP, Harberd NP (1997) The Arabidopsis GAI gene defines a signaling pathway that negatively regulates gibberellin responses. Genes Dev 11:3194–3205
Peng J, Richards DE, Hartley NM, Murphy GP, Devos KM, Flintham JE, Beales J, Fish LJ, Worland AJ, Pelica F, Sudhakar D, Christou P, Snape JW, Gale MD, Harberd NP (1999) “Green Revolution” genes encode mutant gibberellin response modulators. Nature 400:256–261
Peng HZ, Lin EP, Sang QI, Yao S, Jin YQ, Hua XQ, Zhu MY (2007) Molecular cloning, expression analyses and primary evolution studies of REV- and TB1-like genes in bamboo. Tree Physiol 27:1273–1281
Pimenta LMJ, Lange T (2006) Gibberellin biosynthesis and the regulation of plant development. Plant Biol 8:281–290
Remington SJ, Breddam K (1994) Carboxypeptidases C and D. Methods Enzymol 244:231–248
Sauter M, Mekhedov SL, Kende H (1995) Gibberellin promotes histone H1 kinase activity and the expression of cdc2 and cyclin genes during the induction of rapid growth in deepwater rice internodes. Plant J 7:623–632
Schultz DJ, Craig R, Cox-Foster DL, Mumma RO, Medford JI (1994) RNA isolation from recalcitrant plant tissue. Plant Mol Biol Report 12:310–316
Shu S, Mahadeo DC, Liu X, Liu WL, Parent CA, Korn ED (2006) S-Adenosylhomocysteine hydrolase is localized at the front of chemotaxing cells, suggesting a role for transmethylation during migration. Proc Natl Acad Sci USA 103:19788–19793
Soderstrom TR, Ellis RP (1987) The position of bamboo genera and allies in a system of grass classification. In: Soderstrom TR, Hilu KW, Campbell CS, Barkworth ME (eds) Grass systematics and evolution. Smithsonian Institution Press, Washington DC, pp 225–238
Stiefel V, Ruiz-Avila L, Raz R, Vallés MP, Gómez J, Pagés M, Martínez-Izquierdo JA, Ludevid MD, Langdale JA, Nelson T, Puigdomènech P (1990) Expression of a maize cell wall hydroxyproline-rich glycoprotein gene in early leaf and root vascular differentiation. Plant Cell 2:785–793
Suge H (1985) Ethylene and gibberellin: regulation of internodal elongation and nodal root development in floating rice. Plant Cell Physiol 26:607–614
Sutter JU, Campanoni PA, Blatt MR, Paneque M (2006) Setting SNAREs in a different wood. Traffic 7:627–638
Tanaka H, Masuta C, Uehara K, Kataoka J, Koiwai A, Noma M (1997) Morphological changes and hypomethylation of DNA in transgenic tobacco expressing antisense RNA of the S-adenosyl-l-homocysteine hydrolase gene. Plant Mol Biol 35:981–986
Tang D-Q (2009) Genomic sequencing and its application for biological and evolutional research in bamboo. Bamboo J 26:1–10
Ueda K (1960) Studies on the physiology of bamboo, with reference to practical application. Bulletin Kyoto Univ Forests 30:1–169
Vettakkorumakankav N, Patel MS (1996) Dihydrolipoamide dehydrogenase: structural and mechanistic aspects. Indian J Biochem Biophys 33:168–176
Wang K, Peng H, Lin E, Jin Q, Hua X, Yao S, Bian H, Han N, Pan J, Wang J, Deng M, Zhu M (2010) Identification of genes related to the development of bamboo rhizome bud. J Exp Bot 61:551–561
Williams CH Jr (1992) Lipoamide dehydrogenase, glutathione reductase, thioredoxin reductase, and mercuric ion reductase: a family of flavoenzyme transhydrogenases. In: Mueller F (ed) Chemistry and biochemistry of flavoenzymes, vol 3. CRC Press, Boca Raton, FL, pp 121–211
Zheng HY, Bednarek YS, Sanderfoot AA, Alonso J, Ecker JR, Raikhel NV (2002) NPSN11 is a cell plate-associated SNARE protein that interacts with the syntax in KNOLLE. Plant Physiol 129:530–539
Zhong R, Ye ZH (1999) IFL1, a gene regulating interfascicular fiber differentiation in Arabidopsis, encodes a homeodomain–leucine zipper protein. Plant Cell 11:2139–2152
Zhou HL, He SJ, Cao YR, Chen T, Du BX, Chu CC, Zhang JS, Chen SY (2006) OsGLU1, a putative membrane-bound endo-1, 4-β-d-glucanase from rice, affects plant internode elongation. Plant Mol Biol 60:137–151
Acknowledgements
This work was supported by a special grant from the National Natural Science Foundation of China (no. 30371181) and the Natural Science Foundation of Zhejiang Province (nos. Y3080002 and R303420).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhou, Mb., Yang, P., Gao, Pj. et al. Identification of Differentially Expressed Sequence Tags in Rapidly Elongating Phyllostachys pubescens Internodes by Suppressive Subtractive Hybridization. Plant Mol Biol Rep 29, 224–231 (2011). https://doi.org/10.1007/s11105-010-0222-0
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-010-0222-0