Abstract
Background
The microbial community structure in forest soils is expected to change in response to global environmental change, such as climate warming and nitrogen deposition. Community responses to these environmental changes may further interact with the site’s land-use history and understory light availability. Uncovering the relative importance of these global change drivers is crucial to understand and predict soil microbial communities’ changes.
Methods
A full-factorial in situ mesocosm experiment was conducted and the soil microbiota were analyzed by phospholipid fatty acid and neutral lipid fatty acid. The soils in the mesocosms were sampled from forests with different land-use history, and mesocosms contained typical forest understory plants. The mesocosms were exposed to experimental treatments of warming, nitrogen addition and subcanopy illumination.
Results
Among the treatments, past land-use had the strongest effect shaping the microbial community structure. We found a significantly higher abundance of arbuscular mycorrhizal fungi and Actinobacteria in ancient forests. The soil microbial and plant communities were co-structured in ancient forests, but not in past-agricultural forests. Warming and nitrogen fertilization did not affect the soil microbial community composition, yet illumination resulted in slight changes in soil microbial composition.
Conclusions
Our results underpin the role of land-use legacies in shaping soil microbial communities. The stronger plant-microbe linkages in ancient forest soils compared to post-agricultural secondary forest soils may contribute to a higher resilience against environmental changes. Our results advocate for more multifactor global change experiments that investigate the mechanisms underlying the potential effects of land-use legacies on plant-microbe relationships in forest.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Global change drivers are reshaping the structure and processes of ecosystems, which influence the biodiversity and functioning of the ecosystems (Crowther et al., 2015; Rillig et al., 2019). In the face of rapid environmental change, soil microbiota are of importance to regulate ecosystem functioning at local or wider spatial scales (Crowther et al., 2015; Zhang et al., 2021). These microbial communities can respond to global change drives, such as atmospheric N deposition and climate change (Smith et al., 2009; Grimm et al., 2013). Besides, abrupt disturbances from natural dynamics or anthropogenic activities in the canopy may affect availability in resources, such as light (van Loon et al., 2014; Rui et al., 2016; Williams et al., 2020; Bochet et al., 2021). Soil microbiota community responses to environmental changes may further interact with legacies from previous changes, such as past land-use (Vellend et al., 2007; Perring et al., 2016). Elucidating the contribution of these global change drivers and land-use legacies to the soil microbial community is necessary to improve predictions of soil microbial community development, and to understand how ecosystem functions (e.g., carbon storage and nutrient cycling) may shift in response to multiple interacting global changes (Burke et al., 2011; Eisenhauer et al., 2012; Rillig et al., 2019; van der Plas, 2019).
Soil microbiota, dominated by bacteria and fungi, can rapidly respond to environmental changes, and the alteration of microbes is initially reflected at the individual performance (De Long et al., 2016; Nguyen et al., 2021; Wieczynski et al., 2021). When global changes occur, those microorganisms that benefit from the changes could proliferate rapidly at the cost of other microbes. Along with the individual population changes, the community would re-assemble and a shift of local microbial composition would be expected (Smith et al., 2009; Yang et al., 2019). However, the sensitivity of soil microbiota to multiple global change drivers varies to a large degree, and different environmental drivers may mutually interact to obfuscate any single effect (Rinnan et al., 2007; Sun et al., 2019). Although a growing body of evidence suggests that ecosystem processes and functions respond to multiple global changes, multi-factorial experiments that study microbial composition are largely lacking to date (Rillig et al., 2019).
Changes in soil microbial community responding to global environmental drivers can further depend on previous environmental changes, such as past land-use (Perring et al., 2016). Land-use change, like forest clearing for agriculture, can affect the soil microbial community structure and assembly through shifts in plant communities and soil management practices (Veldkamp et al., 2020; Zhou et al., 2022). To date, effects of land-use history are well-documented in temperate forest ecosystems, with dominance on community dynamics focusing on the aboveground part in particular (Perring et al., 2016). The altered plant community caused by land-use changes can directly influence the soil microbial community through release of root exudates, or indirectly influence the belowground components via alterations in quality and quantity of litter fall (Wardle et al., 2004; Feng et al., 2022). Interactions between specific soil microorganisms with their host species (i.e., mutualistic or parasitic) would also be expected to affect the belowground microbial composition (Miki, 2012; Fabianska et al., 2019). Among forest plants, herbaceous understory layer was particularly well-studied as the life-history traits of these species were closely correlated with land-use history (Verheyen et al., 2003). To better understand the mechanisms behind the soil microbial changes under different land-use legacies, it is of significance to not only focus on the responses of the understory plants or the responses of the soil microbes separately, but also quantify co-structure between them (von Rein et al., 2016).
Here we assess the microbial communities, in terms of relative abundance of identified functional groups and overall community composition, in response to four environmental change drivers, i.e., warming, N enrichment and enhancing light availability, with a distinct focus on the land-use legacies. We used phospholipid fatty acid (PLFA) and neutral lipid fatty acid (NLFA)-based analyses to quantify the total living biomass and especially focus on the profiles of the active part of the microbial community (Frostegard et al., 2011; Orwin et al. 2018), which could expand our knowledge on the multiple interactive effects of global change on active belowground communities. We conducted a full-factorial mesocosm experiment including all these treatments on soils from ancient temperate deciduous forests. We hypothesized that: (1) land-use history will affect the composition of the soil microbial communities, wherein less arbuscular mycorrhizal fungi (AMF) were expected in post-agricultural forests due to the loss of forest specialists; (2) light availability and warming may induce higher soil microbial biomass, whereas N addition may not in this case as numerous European temperate forests are N saturated (De Schrijver et al., 2008); (3) the soil microbial and plant community will co-vary, as both communities are expected to respond concurrently to the applied global environmental change drivers or land-use legacies.
Materials and methods
Mesocosm setup
Study area
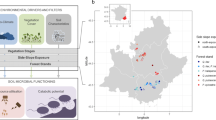
The mesocosms were set up in the Aelmoeseneie forest (50°58’30″ N, 3°48’16″ E, 20 m a.s.l), which is an ancient temperate deciduous forest in northern Belgium. The mean annual temperature (MAT) is 10.6 °C and the mean annual precipitation (MAP) is 768 mm (DEIMS-SDR Database) over the period November 2015-February 2016 and measured in the weather station of long-term ecological research (LTER) site. The dominant tree species are Fagus sylvatica L., Quercus robur L, Acer pseudoplatanus L., and Fraxinus excelsior L. The total throughfall subcanopy atmospheric nitrogen deposition in 2016 was 15.5 kg ha-1 (DEIMS-SDR Database, 2019).
Experimental design
We performed a full-factorial experiment with 64 mesocosms (2 regions× 2 pairs of forest patches × 8 treatments) to reveal the effects of global environmental changes, i.e., warming, illumination, N addition and past land-use, on the forest soil microbial community (Fig. S1).
Land-use legacies: ancient forests vs post-agricultural forests
We collected soil from two regions with different soil characteristics, and contrasting past agricultural intensity within the local forests (further details in Blondeel et al., 2019). The selected forests were located in southern Sweden (SW) and northern Belgium (VL). Within each selected region, two adjacent pairs of ancient or post-agricultural broadleaf forest with a similar canopy composition were included to compare the land-use history. The ancient forest patches exist since at least 1850; while the post-agricultural forests were identified as reforestations on abandoned fields after 1950. The detailed site location and major attributes of these forests can be found in Table S1. The surface soil (15 cm depth) in each forest patch was collected, and all the collected soil samples were transferred to Aelmoeseneie forest and used to build mesocosms after sieving (5-mm mesh).
Plant community assembly: slow, intermediate and fast colonizers
The plant species in the mesocosms were selected according to the recorded observations of understory communities in ancient forest or post-agricultural forests across Western Europe (Verheyen et al., 2003). The species were classified as three functional groups based on the capacity to successfully colonize into post-agricultural forests: slow colonizers and strictly forest specialists (group A), intermediate colonizers and species regularly found in post-agricultural forests but not strictly forest specialists (group B), and fast colonizing nitrophilous species (group C). The classifications and combinations of species can be found in Table S2 and S3 (more details see Blondeel et al., 2020). We used 64 mesocosms, resulting in 1280 plant individuals (64 mesocosms × 5 species × 4 individuals). We performed three rounds of weeding to remove spontaneous species during the first growing season, i.e., May, June, and September 2016.
Manipulating the environment: warming, illumination, and N addition
The first treatment is experimental warming the air and soil by open-top chambers (OTCs, 75 cm wide). During the first half of sampling period (March 2017 to May 2017), the daily mean temperature of the air was significantly increased by 1.04 ± 0.47 °C (p < 0.05), while soil temperature only increased 0.46 °C at the surface and 0.13 °C at 5 cm depth compared with the control treatments (p > 0.05). During the later sampling period (June 2017 to August 2017), no significant warming effects were observed between the warming and control treatments for both air and soil (Blondeel et al., 2020). The second treatment is enhancing light availability by two 18 W fluorescent tubes, which were suspended at 75 cm above the soil surface in each plot. In illuminated plots, photosynthetic active radiation (PAR) was significantly increased to 23.98 ± 4.40 μmol m-2 s-1 compared with the unilluminated plots (7.79 ± 0.68 μmol m-2 s-1 in 2017). The unilluminated plots only received ambient light but were equipped with dummy lamps to avoid other side effects caused by lamp installation (e.g., precipitation). The third treatment is N addition by adding 250 mL of a 2.01 g L-1 solution of NH4NO3 (50 kg N ha yr-1 eq.) to the N addition-treatment mesocosm, and an additional amount of 250 mL demineralized water was used to rinse the leaves to avoid the adherence of N to the plant tissues. The control plots received 500 mL of demineralized water. All the treatments (for both N addition and control) were performed four times per year at the beginning of each season. More details about the mesocosm setup can be found in Blondeel et al., 2020.
Data collection
The experiment started in April 2016 and data for the microbial analysis was collected in April 2020. From each mesocosm, five subsamples of surface soil (5 cm depth) were collected and then thoroughly mixed. To avoid contamination, the sampling spoon was disinfected with 75% ethanol between every mesocosm. The mixed soil was sieved through a 1 mm mesh and divided into two bags, as one subsample was used for microbial analysis and another for soil biogeochemical characterization.
Soil microbial community
The phospholipid fatty acid (PLFA) and neutral lipid fatty acid (NLFA) were extracted to evaluate the soil microbial biomass and the composition of the microbial community. The subsamples for microbial analysis were stored in the freezer at -20 °C, and the PLFA and NLFA extraction were performed according to Quideau et al. (2016). PLFA/NLFA analysis mainly included three steps: extraction, lipid fractionation and lipid methylation. Methyl nonadecanoate (MeC 19:0) was used as internal standard and the concentration of each biomarker was expressed in μg g-1. More details regarding the microbial experiment are provided in Yang et al., 2022a.
A total of 25 PLFA biomarkers and 5 NLFA biomarkers were detected, wherein 6 PLFA (C18:0, C18: 1ω11c, C18: 2ω9c, C18: 3ω3c, C18: 3ω6c, C20:0) and 4 NLFA (C14:0, C15:0, C16:0, C16: 1ω7c) were discarded in the following analysis because they could not easily be assigned to a particular taxonomic group or present low frequency in our dataset. The PLFA and NLFA biomarkers were classified to six functional groups. We used the sum of PLFA iC 14:0, iC 15:0,aC 15:0, iC 16:0, iC 17:0 and aC 17:0 to indicate Gram-positive bacteria (Gm+) (Farrell et al., 2013; Kaiser et al., 2015); the sum of PLFA 16:1ω7c, cy 17:0 and cy 19:0 to indicate Gram-negative bacteria (Gm-) (Mitchell et al., 2015); the sum of PLFA 10MeC16:0, 10MeC17:0 and 10MeC18:0 to indicate Actinobacteria (Act) (Xu et al., 2019); the sum of PLFA C14:0, C15:0, C16:0 and C17:0 to indicate non-specific bacteria (NSB) (Steinbeiss et al., 2009; Willers et al., 2015); and finally, the sum of PLFA 18:2ω6c and 18:1ω9c to indicate saprotrophic fungi (SF) (Zhang et al., 2014). To minimize the influence of the background amounts (from bacteria) of PLFA 16:1ω5c on the estimation of arbuscular mycorrhizal fungi (AMF) biomass in soil (Olsson, 1999; Ngosong et al., 2012), the ratio of NLFA and PLFA 16:1ω5c was used to indicate arbuscular mycorrhizal fungi. Since the NLFA/PLFA ratio was high (> 1) for most of samples in this study, we consider the PLFA 16:1ω5c to be a good proxy for AMF biomass herein (Zhang et al., 2020; Lekberg et al., 2022). The total microbial biomass was calculated by the sum of all representative biomarkers of functional groups. The relative abundances of microbial groups (i.e., Gm+, Gm-, Act, NSB, SF and AMF) were calculated as the percentage of the representative biomarkers divided by the total microbial biomass.
Edaphic properties
Soil moisture was gravimetrically determined by air-drying the soil samples at 50 °C for 48 hours in a drying oven (Reynolds, 1970). Additional subsamples used for chemical analyses were first dried at 40 °C before the further measurements. In brief, total carbon and nitrogen were quantified by combustion at 1200 °C using CNS elemental analyzer (vario Macro Cube, Elementar, Germany) (Kowalenko, 2001). Total P was measured after a complete soil digestion by using HClO4 (65%), HNO3 (70%) and H2SO4 (98%) in Teflon bombs for 4 h at 150 °C (Lajtha et al., 1999). Other mobile soil cations including potassium, magnesium and aluminum were extracted by lactic acid (88%), acetic acid (99%) and ammonium acetate (25%) at pH 3.74 and then measured by atomic absorption spectrophotometry (Blondeel et al., 2019). To calculate the proportion of exchangeable base cations (BC), we convert their concentration from mg/kg to meq/kg in which the charge of the cations would be considered (Reganold and Harsh, 1985). Soil pH was measured with a glass electrode (Orion, model 920A) after shaking a soil/H2O mixture (1:5) at 300 rpm for 5 min.
Plant community
We measured total vegetation cover (%) as the one-sided projection of vegetation in each mesocosm, which was measured by taking digital RGB photographs perpendicular to the ground surface in April 2020. The Canopy Area tool was used to assess the green pixels of vegetation recalculated as percentage cover (Easlon and Bloom, 2014).
Statistical analysis
All statistical analyses were carried out in R (ver.4.0.3) (Team, R.C, 2008). First, we calculated the relative abundance (%) of each microbial group by using the sum of representative PLFA biomarkers divided by the total microbial biomass. To assess how environmental changes and land-use history affected the microbial abundance as well as total microbial biomass, we used linear mixed-effects models (LMMs) to test factor variables (warming, illumination, nitrogen addition, and land-use legacy) effects on microbial communities. A total of eight response variables were tested with LMMs, i.e., relative abundance of six functional groups, ratio of Gm+ and GM- (GM+: GM-), and total microbial biomass (which was log-transformed to meet the normality assumption of the statistical tests). Then we tested the above four treatments (warming, illumination, nitrogen addition and past land-use) as fixed effects separately in our LMMs, with the “region” factor variable as a random effect. To check the effect of environmental changes and land-use history on edaphic properties, LMMs with the same fixed and random effects were built, where response variables soil K concentration and total P were log-transformed to fit normal distribution before the LMMs construction. To explain the relationship between edaphic properties and microbial abundance/ total biomass, a model including predictor variables soil total C, total P, pH, moisture, BC, C/N, and plant coverage was assessed for the response variables, i.e., relative abundance of each microbial group and total microbial biomass. To limit multicollinearity issues in the models, we performed Spearman correlation (Fig. S2) among the predictors, wherein soil total N was highly correlated with other variable (total C, r = 0.95) so as to discard in the following LMMs (Fig. S2). To further simplify the models, a single best model was selected based on the Akaike’s Information Criterion (AIC) using the dredge function of the MuMIn package (Bartoń, 2018).
To examine the variability of the microbial community composition among environmental changes, land-use history as well as sampling regions, non-metric multidimensional scaling (NMDS) based on the Bray Curtis distance matrix were performed using metaMDS function in the vegan package (Oksanen et al., 2017). As the ordinations were strongly disturbed by the fungal biomarkers (with very low abundances), we only focused the bacterial community in this case. The goodness of fit for data calculation (NMDS) could be found in Shepard diagram (Fig. S3). To test the significance of the variation of the microbial composition among different treatments, land-use history and regions, we used analysis of similarities (ANOSIM) with function anosim (package vegan). Besides, to investigate the relationship between microbial composition and soil abiotic properties, and further screen out the most important drivers affecting the microbial composition in the soils, we performed canonical correspondence analysis (CCA). The best model was selected based on AIC by step function, and the collinearity among the constraining variables was checked by Variance Inflation Factor by vif.cca function (only variables with VIF<10 were preserved).
To assess the co-structure between the soil microbial communities and plant communities, we performed co-inertia analysis (COIA) which allows measuring the concordance between two multivariate datasets within the same object (mesocosms in this case). We first performed Hellinger transformation to both of the datasets and then applied COIA on a different level: i) microbial functional groups v.s. plant species, ii) PLFA biomarkers v.s. plant species (function coinertia in ade4 package) (Dray and Dufour, 2007). Besides, to better discern the co-structure between plant and different microbial communities, PLFA biomarkers were further split into bacterial biomarkers and fungal biomarkers, which were then used to conduct the COIA with the plant community, respectively (plant vs bacterial communities and plant vs fungal communities). We evaluated the strength of the coupling pairs of datasets with the RV coefficient, and this global similarity measurement could give an overview of the correlation of the dataset pairs. This coefficient value varies between 0 and 1; the closer the RV value approaches to 1, the stronger the correlation between the two datasets. We finally used Monte-Carlo test (999 random permutations) to test the significance of the co-structure between the datasets.
Results
Soil microbial abundance and associated environmental properties
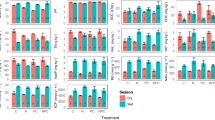
In general, the total microbial biomass in soils ranged from 2.94 to 37.50 μg g-1. Except for illumination (marginally significant), no significant differences were found among treatments. Most (~80%) of the obtained biomarkers (excluding undetermined biomarkers) were indicative for bacteria, with Gram-positive bacteria having the highest relative abundance (33.71% on average). Several fungal biomarkers were also detected in the soil samples, but the relative abundance (6.73% on average) was comparably lower than those of bacteria (21.64% on average). When we compared the relative abundances of these functional groups among four different treatments, we found that past land-use exerted a significant impact on the soil microbial abundance, in which AMF and Actinobacteria showed a higher relative abundance in ancient forests (+4.33 ± 1.79 and 1.09± 0.46, respectively), while non-specific bacteria showed a higher relative abundance in post-agricultural forests (+3.63 ± 0.91, Fig. 1 and Table 1). The illumination had a marginally significant effect and led to a higher microbial biomass (+3.33 ± 1.53, p = 0.06). Saprotrophic fungi had a higher relative abundance (+3.51 ± 1.74, p = 0.08), while Gram-positive bacteria had lower abundance (-3.89 ± 1.93, p = 0.08) in response to light addition. In contrast, the soil microbial abundance was not significantly affected by the warming nor nitrogen addition (p> 0.1).
Relative abundances (%) of microbial groups, ratio of Gram-positive and Gram-negative bacteria, and total microbial biomass (μg g-1) of the experimental treatments. Gm+, Gram-positive bacteria; Gm-, Gram-negative bacteria; Act, Actinobacteria; NSB, non-specific bacteria; SF, saprotrophic fungi; AMF, arbuscular mycorrhizal fungi; SW, southern Sweden; VL, northern Belgium
Past land use showed also significant effects on the soil abiotic properties and plant coverage. As shown in Table S4, post-agricultural forests had higher soil pH (+0.37 ± 0.07), total P (+277.92 ± 34.60) as well as plant coverage (+18.59 ± 5.13) than ancient forests (p < 0.001), while ancient forests had higher soil moisture (+9.98 % ± 1.87, snapshot data during our sampling), total C (+0.57 % ± 0.24) and C/N ratio (+1.14 ± 0.34) instead (p < 0.01). Then we used the simplified LMMs to reveal the correlations of these properties to the relative abundance of microbial groups and total microbial biomass and found significant correlations of microbial abundance and soil C and C/N ratio (Table 2). The relative abundance of Gram-positive bacteria was positively correlated with soil C and negatively correlated with C/N ratio, while Actinobacteria were only positively correlated with soil C.
Soil microbial composition and associated environmental properties
NMDS ordination and ANOSIM revealed significantly different community composition in soil bacteria between both past land-use classes, but no significant effects were found for other experimental treatments (warming, illumination and nitrogen addition) (ANOSIM, p >0.05; Fig. 2). CCA was performed to investigate which environmental properties contributed to explain the variation in soil microbial community composition. As shown in Fig. 3, the first two axes explained 14.47% of the variance in bacterial community composition, and the variation was significantly related to soil pH and C/N ratio (p = 0.04 and p = 0.01, respectively & Table S5).
Non-metric multidimensional scaling (NMDS) results of the bacterial composition based on PLFA biomarkers. Variations of the bacterial composition between different experimental treatments were calculated based on the Bray-Curtis distance matrix. Dissimilarity values based on analysis of similarities (ANOSIM) are also shown in plots. SW, southern Sweden; VL, northern Belgium
The relationship between bacterial community composition and abiotic properties (soil characteristics) based on canonical correspondence analysis (CCA). The first axis accounted for 10.58% of the variability and the second axis accounted for 3.89%. The significance levels of CCA1 and CCA2 were p = 0.02 and p = 0.08, respectively (permutation test). (a) The distribution of samples collected in different regions. Different colors in this panel indicated the soil sources. SW, southern Sweden; VL, northern Belgium. (b) The distribution of single PLFA biomarkers. Different symbols in this panel indicated the classified microbial groups. Gm+, Gram-positive bacteria; Gm-, Gram-negative bacteria; Act, Actinobacteria; NSB, non-specific bacteria
Co-structure between the soil microbial and the plant community
There was no significant co-structure between the soil microbial groups or PLFA biomarkers (for both bacterial and fungal biomarkers) and plant communities across all plots (Table 3). However, when we split the dataset into different past land-use classes, we found a significant co-structure in the ancient, but not in the post-agricultural soils (Table 3). In order to visualize the relative abundance of plant community that affected microbial community, projections of the first two axes of COIA were presented for all 6 microbial groups and 19 plant species (Fig. 4a). The first two axes explained 97% of the total co-variation between the soil microbial and plant communities. A shorter arrow in ancient forests (1.22 ± 0.05, Mean ± S.E.) than post-agricultural forests (1.34 ± 0.07) indicated a stronger relationship between plant and microbial communities in the former. Besides, the positions of each indicator in panels (b) and (c) represent the importance and the direction of their contribution to the distribution of sites in the COIA shown in panel (a). In that case, microbial functional groups (in panel b) and plant species (in panel c) projecting in the same direction from the origin have a strong association. Based on our results, CIOA showed that bacterial groups (Gram-negative bacteria, Actinobacteria, Gram-positive bacteria and non-specific bacteria) were strongly related to plant species like Quercus robur, Geranium robertianum, Rubus fruticosus, and Fraxinus excelsior L. In contrast, saprotrophic fungi were associated with Polygonatum multiflorum L. and Carex sylvatica Huds., and AMF were associated with Hyacinthoides non-scripta and Poa trivialis (Fig. 4b & c).
The co-inertia analysis of the soil microbial and plant community composition across all plots. (a) The positions of each plot on the first two axes of the COIA is conditional on the plant (arrow tails) or microbial (arrow heads) communities. The colours of the arrows show the site groups according to the past land-use. The percentages of variance explained by axes are shown on each axis. (b) Projections of the microbial groups and (c) projection of the plant species. The abbreviations of the bacterial phyla in (b): Gm+, Gram-positive bacteria; Gm-, Gram-negative bacteria; Act, Actinobacteria; NSB, non-specific bacteria; SF, saprotrophic fungi; AMF, arbuscular mycorrhizal fungi. The abbreviations of the plant species in (c): Acp, Acer pseudoplatanus L.; Ajr, Ajuga reptans L.; Ann, Anemone nemorosa L.; Cas, Carex sylvatica Huds.; Dry, Dryopteris; Fre, Fraxinus excelsior L.; Gao, Galium odoratum L.; Ger, Geranium robertianum L.; Heh, Hedera helix L.; Hyn, Hyacinthoides non-scripta L.; Oxa, Oxalis acetosella L.; Pom, Polygonatum multiflorum L.; Pon, Poa nemoralis L.; Pot, Poa trivialis L.; Qur, Quercus robur L.; Raf, Ranunculus ficaria; Ruf, Rubus fruticosus agg.; Urd, Urtica dioica L.; Vim, Vinca minor L. We refer to the text for further info on the interpretation of the figure
Discussion
Land-use history: keystone driver in shaping microbial community structure
We found that land-use history was the prominent driver for shaping microbial community structure, rather than the in situ applied environmental changes. The land-use legacy significantly changed the microbial abundance and community composition (Fig. 1 & 2d). We showed a higher abundance of arbuscular mycorrhizal fungi (AMF) and Actinobacteria in ancient forests, which confirms our first hypothesis. In ancient forests, functional traits of understory plants show adaptation to low-light conditions (Valladares & Niinemets, 2008). These understory species often emerge in early spring and benefit from nutrients stored in belowground tissues (Verheyen et al. 2003). When the photosynthesis of plants decreased due to the low light intensity or shading, AMF communities could aid plants with nutrient and water acquisition with the nutrient uptake via plant root system, hence highlighting their dominant ecological niche in ancient forests (Xu et al., 2018; Kaur and Suseela, 2020; Wen et al., 2022). Ancient forests soils are also expected to have higher carbon accumulation in soils compared to those in post-agricultural forests (Fanin and Bertrand, 2016). Actinobacteria are one of the most representative copiotrophic bacterial members, i.e., preferring to colonize soils rich in organic matter and characterized with a relatively rapid growth rate (Selene Gomez-Acata et al., 2016; Yao et al., 2017). Therefore, the abundant C accumulation in ancient forest soils facilitates the enrichment of Actinobacteria herein (see Table S4). Furthermore, previous literature has shown that α-diversity of AMF and several bacterial species will decrease as the land use changes (Berkelmann et al., 2020; Ji et al., 2022), which could partially support our findings (increase of AMF and Actinobacteria abundance in ancient forest soils). The colonization of soil microbial communities is mainly dependent on the organic layers and nutrient availability in forests (Wilson and Puri, 2001; Zhou et al., 2006), and this would reasonably result in the microbial species in ancient forests generally having a longer time for colonization (Boeraeve et al., 2018).
The beta-diversity analysis (NMDS ordination), which displayed the overall community composition of soil bacteria, showed clear changes between both land-use legacies (Fig. 2d). The differences between land-use types may be in part be explained by differences in soil chemical properties, as soil pH was one of the most important drivers affecting the microbial composition in the soils (Fig. 3 and Table S5). Due to past fertilizing, liming and manuring practices, post-agricultural deciduous forests with an intensive agricultural history generally present a higher pH compared to ancient forests (Verheyen et al., 1999; von Oheimb et al., 2008; Orczewska, 2009; Blondeel et al., 2019). On the one hand, pH can affect the microbial structure directly, where the microbial community changes from acidophilic microbes to neutrophilic microbes during the agricultural period (Chen et al., 2021; Sridhar et al., 2022). On the other hand, these edaphic properties might have an effect on aboveground plant species composition first and then exert a legacy effect on the underground community. For example, the changes of litter composition and humus type caused by altered plant community composition can result in indirect effects on the microbial community structure in different land-use legacies (Santonja et al., 2017; Bayranvand et al., 2021). However, the response strength of the understory traits to the edaphic property variations can be further dependent on the background flora (local plant species pool) and detailed land-use practice (Falkengren-Grerup et al., 2006; Barlow et al., 2020). The soil microbial structuring mechanisms in forests with contrasting land-use legacies, and their linkages with understory species and edaphic properties, require further experimental tests and analysis.
Environmental change: additional illumination increases total microbial biomass and affect part of their abundance
Light availability emerged to increase the soil microbial biomass and also change the relative abundances of several microbial groups in our experiment. This result supports our second hypothesis that when illumination is applied, the aboveground plant productivity increases and more litter inputs and root exudates are expected to flow to the soil food web, which lead to a higher soil microbial biomass and activity. Indeed, additional illumination can also regulate the root exudates and rhizodeposition to enhance the microbial biomass or shift soil microbial composition (Hristozkova et al., 2017; Xi et al., 2019). For example, Balasooriya et al. (2014) used 13C pulse-labeling to assess the role of soil microbes in the cycling of rhizodeposit-C. They found that fungi enable to form a rapid channel of rhizodeposit-C into the soil microbes, while Actinomycetes and gram-positive bacteria presented a delayed utilization. Above conclusions support the results in this study, i.e., saprotrophic fungi showed a higher relative abundance under the illuminating treatment while Gram-positive bacteria showed a lower relative abundance instead (Fig. 1). These results may indicate the increasing root exudates caused by illumination can accelerate the enrichment of saprotrophic fungi rather than gram-positive bacteria, which may prefer other C sources instead of rhizodeposition (Bird et al., 2011; Balasooriya et al., 2014). However, no significant enrichment of AMF in soils was observed when additional illumination was applied. This is to say, even with the additional illumination, the plant investment in an already established mycorrhizal network could impose a lower carbon cost (Teste et al., 2009; Unger et al., 2021), which may inhibit the response of AMF to the additional light availability. Furthermore, the light effects on AMF were equivocal in previous literature, and more detailed data on the soil nutrient availability, AMF taxa and ecosystem types should be explicitly considered in the future (Liu et al., 2015; Ballhorn et al., 2016; Ficano et al., 2021; Yang et al., 2022b).
Plant communities: co-varied with soil microbial community, but only in ancient forest
The significant relationship between the soil microbial and plant communities in ancient forest soils supports the idea that land-use history affects the soil microorganisms through a legacy effect on the understory plants. First, we found that the fast-colonizing, generalist plant species including Poa trivialis were positively correlated with AMF, whereas slow-colonizing plant species such as Polygonatum multiflorum and Carex sylvatica were positively correlated with saprotrophic fungi based on the COIA analyses (Fig. 4b & c). These findings corroborate that in an acid nutrient-poor ancient forest soil, communities with slow-colonizing species are expected to be dominant, whereas in less acid and nutrient-rich post-agricultural forest, the plant community is dominated by fast-colonizing species (Verheyen et al., 2003; Wardle et al., 2004). Second, the significant linkages of plant and soil microbial communities in ancient forest soils could be the result of similar compositional responses of aboveground and belowground communities responding to the same soil characteristics. Additionally, the indirect effects of land-use history on belowground community compositions due to changes that occur in plant functional traits can also contribute, as we also found a significant difference of total vegetation cover and cover-weighed community weighted means of SLA (specific leaf area) between ancient and post-agricultural forest in the second growing season of the experiment (March to September in 2017, see Blondeel et al., 2019 and Marri, 2020).
However, we found no significant co-structure between plant and soil microbial community in post-agricultural forest soils. It is known that plant-microbe community dynamics are complex and characterized by instability, therefore different land-use legacies could evoke a series of plant-microbe community shift (Bardgett et al., 2005; Kardol et al., 2006). For example, due to past fertilizing and manuring practice, post-agricultural forest soils represent chronic nutrient input, as such it would facilitate the growth of those plant/microbes beneficial from high nutrient availability (Holt, 2008; Veresoglou et al., 2011) and eventually distort the plant/microbial linkages compared to those in ancient forests. Besides, the different co-structure between plant and microbial community in this study would also be linked to the divergence of plant communities between the two past land-use types. For ancient forests, we see the dominance of forest specialists and other species have been filtered out. By contrast, a wide array of planted species (mixture of fast- and slow colonizing plants) were present on post-agricultural soils. Furthermore, whether the strong co-structured relationship between plant and soil microbes in ancient forest soils may act as a buffer, such as weakening understory control over other environmental changes (i.e., against extreme droughts), should be tested in future experiments. In this respect, it is essential to focus on plant-microbial relationship when studying the response of soil microbial community under different land-use legacies, certainly when interested in mechanisms of community response to environmental changes.
Conclusion
In our full-factorial mesocosm experiments with three global change drivers and past land-use included, we found that land-use legacy emerged as a critical driver for soil microbial community. In ancient forests, there was a higher abundance of AMF and Actinobacteria, a differential composition of soil bacterial communities, and the composition of the soil microbial and plant communities was also co-structured. Warming and additional N did not affect the soil microbes, while illumination did. This treatment resulted in an increased soil microbial biomass, a higher abundance of SF and a lower abundance of Gram-positive bacteria. Our results shed light on the importance of past land-use for belowground biota and the linkages with aboveground vegetation. A logical next step will be to test whether the strong linkages with plant communities in ancient forest soils may contribute to the high ecosystem resilience.
Data availability
Data related to this manuscript are available on Figshare: https://doi.org/10.6084/m9.figshare.21788738.v1.
References
Balasooriya WK, Denef K, Huygens D, Boeckx P (2014) Translocation and turnover of rhizodeposit carbon within soil microbial communities of an extensive grassland ecosystem. Plant Soil 376:61–73
Ballhorn DJ, Schaedler M, Elias JD, Millar JA, Kautz S (2016) Friend or Foe-Light Availability Determines the Relationship between Mycorrhizal Fungi, Rhizobia and Lima Bean (Phaseolus lunatus L.). PloS One 11
Bardgett RD, Bowman WD, Kaufmann R, Schmidt SK (2005) A temporal approach to linking aboveground and belowground ecology. Trends Ecol Evol 20:634–641
Barlow KM, Mortensen DA, Drohan PJ (2020) Soil pH influences patterns of plant community composition after restoration with native-based seed mixes. Restor Ecol 28
Bartoń, K. (2018). MuMIn: Multi-Model Inference.
Bayranvand M, Akbarinia M, Jouzani GS, Gharechahi J, Kooch Y, Baldrian P (2021) Composition of soil bacterial and fungal communities in relation to vegetation composition and soil characteristics along an altitudinal gradient. FEMS Microbiol Ecol 97
Berkelmann D, Schneider D, Meryandini A, Daniel R (2020) Unravelling the effects of tropical land use conversion on the soil microbiome. Environmen Microbiome 15
Bird JA, Herman DJ, Firestone MK (2011) Rhizosphere priming of soil organic matter by bacterial groups in a grassland soil. Soil Biol Biochem 43:718–725
Blondeel H, Perring MP, Berges L, Brunet J, Decocq G, Depauw L, Diekmann M, Landuyt D, Liira J, Maes SL et al (2019) Context-Dependency of Agricultural Legacies in Temperate Forest Soils. Ecosystems 22:781–795
Blondeel H, Perring MP, De Lombaerde E, Depauw L, Landuyt D, Govaert S, Maes SL, Vangansbeke P, De Frenne P, Verheyen K (2020) Individualistic responses of forest herb traits to environmental change. Plant Biology 22:601–614
Bochet E, Jose Molina M, Monleon V, Espigares T, Manuel Nicolau J, de Las M, Heras M, Garcia-Fayos P (2021) Interactions of past human disturbance and aridity trigger abrupt shifts in the functional state of Mediterranean holm oak woodlands. Catena 206
Boeraeve M, Honnay O, Mullens N, Vandekerkhove K, De Keersmaeker L, Thomaes A, Jacquemyn H (2018) The impact of spatial isolation and local habitat conditions on colonization of recent forest stands by ectomycorrhizal fungi. Forest Ecol Manag 429:84–92
Burke DJ, Weintraub MN, Hewins CR, Kalisz S (2011) Relationship between soil enzyme activities, nutrient cycling and soil fungal communities in a northern hardwood forest. Soil Biol Biochem 43:795–803
Chen, Z., Maltz, M.R., Zhang, Y., O'Brien, B.J., Neff, M., Wang, Y., and Cao, J. (2021). Plantations of Cinnamomum camphora (Linn) Presl with Distinct Soil Bacterial Communities Mitigate Soil Acidity within Polluted Locations in Southwest China. Forests, 12
Crowther TW, Thomas SM, Maynard DS, Baldrian P, Covey K, Frey SD, van Diepen LTA, Bradford MA (2015) Biotic interactions mediate soil microbial feedbacks to climate change. P Natl Acad Sci USA 112:7033–7038
De Long JR, Dorrepaal E, Kardol P, Nilsson M-C, Teuber LM, Wardle DA (2016) Contrasting Responses of Soil Microbial and Nematode Communities to Warming and Plant Functional Group Removal Across a Post-fire Boreal Forest Successional Gradient. Ecosystems 19:339–355
De Schrijver, A., Verheyen, K., Mertens, J., Staelens, J., Wuyts, K., and Muys, B. (2008). Nitrogen saturation and net ecosystem production. Nature, 451, 0-0.
Dray S, Dufour A-B (2007) The ade4 Package: Implementing the Duality Diagram for Ecologists. J Stat Softw 22
Easlon HM, Bloom AJ (2014) Easy Leaf Area: Automated digital image analysis for rapid and accurate measurement of leaf area. Appl. Plant Sci 2
Eisenhauer N, Cesarz S, Koller R, Worm K, Reich PB (2012) Global change belowground: impacts of elevated CO2, nitrogen, and summer drought on soil food webs and biodiversity. Global Change Biol 18:435–447
Fabianska I, Sosa-Lopez E, Bucher M (2019) The role of nutrient balance in shaping plant root-fungal interactions: facts and speculation. Curr Opin Microbiol 49:90–96
Falkengren-Grerup U, ten Brink DJ, Brunet J (2006) Land use effects on soil N, P, C and pH persist over 40-80 years of forest growth on agricultural soils. Forest Ecol Manag 225:74–81
Fanin N, Bertrand I (2016) Aboveground litter quality is a better predictor than belowground microbial communities when estimating carbon mineralization along a land-use gradient. Soil Biol Biochem 94:48–60
Farrell M, Kuhn TK, Macdonald LM, Maddern TM, Murphy DV, Hall PA, Singh BP, Baumann K, Krull ES, Baldock JA (2013) Microbial utilisation of biochar-derived carbon. Sci Total Environ 465:288–297
Feng J, He K, Zhang Q, Han M, Zhu B (2022) Changes in plant inputs alter soil carbon and microbial communities in forest ecosystems. Global Change Biol 28:3426–3440
Ficano N, Porder S, McCulloch LA (2021) Tripartite legume-rhizobia-mycorrhizae relationship is influenced by light and soil nitrogen in Neotropical canopy gaps. Ecology 102
Frostegard A, Tunlid A, Baath E (2011) Use and misuse of PLFA measurements in soils. Soil Biol Biochem 43:1621–1625
Grimm NB, Chapin FS III, Bierwagen B, Gonzalez P, Groffman PM, Luo Y, Melton F, Nadelhoffer K, Pairis A, Raymond PA et al (2013) The impacts of climate change on ecosystem structure and function. Front Ecol Environ 11:474–482
Holt RD (2008) Theoretical perspectives on resource pulses. Ecology 89:671–681
Hristozkova M, Geneva M, Stancheva I, Velikova V (2017) Led spectral composition effects on mycorrhizal symbiosis formation with tomato plants. Appl Soil Ecol 120:189–196
Ji L, Yang X, Zhu C, Ma L, Chen Y, Ling N, Zhou Z, Ni K, Guo S, Helgason T et al (2022) Land-use changes alter the arbuscular mycorrhizal fungal community composition and assembly in the ancient tea forest reserve. Agr Ecosyst Environ 339
Kaiser C, Kilburn MR, Clode PL, Fuchslueger L, Koranda M, Cliff JB, Solaiman ZM, Murphy DV (2015) Exploring the transfer of recent plant photosynthates to soil microbes: mycorrhizal pathway vs direct root exudation. New Phytol 205:1537–1551
Kardol P, Bezemer TM, van der Putten WH (2006) Temporal variation in plant-soil feedback controls succession. Ecol Lett 9:1080–1088
Kaur S, Suseela V (2020) Unraveling Arbuscular Mycorrhiza-Induced Changes in Plant Primary and Secondary Metabolome. Metabolites 10
Kowalenko CG (2001) Assessment of Leco CNS-2000 analyzer for simultaneously measuring total carbon, nitrogen, and sulphur in soil. Commun Soil Sci Plan 32:2065–2078
Lajtha K, Driscoll CT, Jarrel WM, Elliott ET (1999) Soil phosphorus - Characterization and total element analysis. Standard Soil Methods for Long-Term Ecological Research, Oxford University Press 2:115–142
Lekberg Y, Baath E, Frostegard A, Hammer E, Hedlund K, Jansa J, Kaiser C, Ramsey PW, Rezanka T, Rousk J et al (2022) Fatty acid 16:1ω5 as a proxy for arbuscular mycorrhizal fungal biomass: current challenges and ways forward. Biol Fert Soils 58:835–842
Liu Y, Mao L, Li J, Shi G, Jiang S, Ma X, An L, Du G, Feng H (2015) Resource availability differentially drives community assemblages of plants and their root-associated arbuscular mycorrhizal fungi. Plant Soil 386:341–355
Marri G (2020) Diverging trajectories of forest herb community development in a multi-factor global change experiment. Ghent University, Master thesis
Miki T (2012) Microbe-mediated plant-soil feedback and its roles in a changing world. Ecol Res 27:509–520
Mitchell PJ, Simpson AJ, Soong R, Simpson MJ (2015) Shifts in microbial community and water-extractable organic matter composition with biochar amendment in a temperate forest soil. Soil Biol Biochem 81:244–254
Ngosong C, Gabriel E, Ruess L (2012) Use of the signature fatty acid 16: 1ω5 as a tool to determine the distribution of arbuscular mycorrhizal fungi in soil. J Lipids 2012
Nguyen J, Lara-Gutierrez J, Stocker R (2021) Environmental fluctuations and their effects on microbial communities, populations and individuals. FEMS Microbiol Rev 45
Oksanen, J., Blanchet, F.G., Friendly, M., Kindt, R., and others (2017). vegan: Community Ecology Package. R Package Version 2.4-3. 2017.
Olsson PA (1999) Signature fatty acids provide tools for determination of the distribution and interactions of mycorrhizal fungi in soil. FEMS Microbiol Ecol 29:303–310
Orczewska A (2009) The impact of former agriculture on habitat conditions and distribution patterns of ancient woodland plant species in recent black alder (Alnus glutinosa (L.) Gaertn.) woods in south-western Poland. Forest Ecol Manag 258:794–803
Orwin KH, Dickie IA, Holdaway R, Wood JR (2018) A comparison of the ability of PLFA and 16S rRNA gene metabarcoding to resolve soil community change and predict ecosystem functions. Soil Biol Biochem 117: 27–35
Perring MP, De Frenne P, Baeten L, Maes SL, Depauw L, Blondeel H, Caron MM, Verheyen K (2016) Global environmental change effects on ecosystems: the importance of land-use legacies. Global Change Biol 22:1361–1371
Quideau SA, McIntosh ACS, Norris CE, Lloret E, Swallow MJB, Hannam K (2016) Extraction and Analysis of Microbial Phospholipid Fatty Acids in Soils. J Vis Exp 114:e54360
Reganold JP, Harsh JB (1985) Expressing cation exchange capacity in milliequivalents per 100 grams and in SI units. J Agron Educ 14:84–90
Reynolds SG (1970) The gravimetric method of soil moisture determination Part I A study of equipment, and methodological problems. J Hydrol 11:258–273
Rillig MC, Ryo M, Lehmann A, Aguilar-Trigueros CA, Buchert S, Wulf A, Iwasaki A, Roy J, Yang G (2019) The role of multiple global change factors in driving soil functions and microbial biodiversity. Science 366:886-+
Rinnan R, Michelsen A, Baath E, Jonasson S (2007) Fifteen years of climate change manipulations alter soil microbial communities in a subarctic heath ecosystem. Global Change Biol 13:28–39
Rui Y, Murphy DV, Wang X, Hoyle FC (2016) Microbial respiration, but not biomass, responded linearly to increasing light fraction organic matter input: Consequences for carbon sequestration. Sci Rep 6
Santonja M, Rancon A, Fromin N, Baldy V, Hattenschwiler S, Fernandez C, Montes N, Mirleau P (2017) Plant litter diversity increases microbial abundance, fungal diversity, and carbon and nitrogen cycling in a Mediterranean shrubland. Soil Biol Biochem 111:124–134
Selene Gomez-Acata E, Valencia-Becerril I, Valenzuela-Encinas C, Socorro Velasquez-Rodriguez A, Ebenezer Navarro-Noya Y, Montoya-Ciriaco N, Cecilia Suarez-Arriaga M, Rojas-Valdez A, Gabriel Reyes-Reyes B, Luna-Guido M et al (2016) Deforestation and cultivation with maize (zea mays l.) has a profound effect on the bacterial community structure in soil. Land Degrad Dev 27:1122–1130
Smith MD, Knapp AK, Collins SL (2009) A framework for assessing ecosystem dynamics in response to chronic resource alterations induced by global change. Ecology 90:3279–3289
Sridhar B, Lawrence GB, Debenport SJ, Fahey TJ, Buckley DH, Wilhelm RC, Goodale CL (2022) Watershed-scale liming reveals the short- and long-term effects of pH on the forest soil microbiome and carbon cycling. Environ Microbiol 24:6184–6199
Steinbeiss S, Gleixner G, Antonietti M (2009) Effect of biochar amendment on soil carbon balance and soil microbial activity. Soil Biol Biochem 41:1301–1310
Sun S, Wu Y, Zhang J, Wang G, DeLuca TH, Zhu W, Li A, Duan M, He L (2019) Soil warming and nitrogen deposition alter soil respiration, microbial community structure and organic carbon composition in a coniferous forest on eastern Tibetan Plateau. Geoderma 353:283–292
Team, R.C (2008) R: A language and environment for stastiscal computing. Vienna, Austria, R Foundation for Statistical computing
Teste FP, Simard SW, Durall DM, Guy RD, Jones MD, Schoonmaker AL (2009) Access to mycorrhizal networks and roots of trees: importance for seedling survival and resource transfer. Ecology 90:2808–2822
Unger S, Habermann FM, Schenke K, Jongen M (2021) Arbuscular Mycorrhizal Fungi and Nutrition Determine the Outcome of Competition Between Lolium multiflorum and Trifolium subterraneum. Front Plant Sci 12:778861
Valladares F, Niinemets U (2008) Shade Tolerance, a Key Plant Feature of Complex Nature and Consequences. Annu Rev Ecol Evol S 39:237–257
van der Plas F (2019) Biodiversity and ecosystem functioning in naturally assembled communities. Biol Rev 94:1220–1245
van Loon MP, Schieving F, Rietkerk M, Dekker SC, Sterck F, Anten NPR (2014) How light competition between plants affects their response to climate change. New Phytol 203:1253–1265
Veldkamp E, Schmidt M, Powers JS, Corre MD (2020) Deforestation and reforestation impacts on soils in the tropics. Nat Rev Earth Env 1:590–605
Vellend M, Verheyen K, Flinn KM, Jacquemyn H, Kolb A, Van Calster H, Peterken G, Graae BJ, Bellemare J, Honnay O et al (2007) Homogenization of forest plant communities and weakening of species-environment relationships via agricultural land use. J Ecol 95:565–573
Veresoglou SD, Mamolos AP, Thornton B, Voulgari OK, Sen R, Veresoglou DS (2011) Medium-term fertilization of grassland plant communities masks plant species-linked effects on soil microbial community structure. Plant Soil 344:187–196
Verheyen K, Bossuyt B, Hermy M, Tack G (1999) The land use history (1278-1990) of a mixed hardwood forest in western Belgium and its relationship with chemical soil characteristics. J Biogeogr 26:1115–1128
Verheyen K, Honnay O, Motzkin G, Hermy M, Foster DR (2003) Response of forest plant species to land-use change: a life-history trait-based approach. J Ecol 91:563–577
von Oheimb G, Haerdtle W, Naumann PS, Westphal C, Assmann T, Meyer H (2008) Long-term effects of historical heathland farming on soil properties of forest ecosystems. Forest Ecol Manag 255:1984–1993
von Rein I, Gessler A, Premke K, Keitel C, Ulrich A, Kayler ZE (2016) Forest understory plant and soil microbial response to an experimentally induced drought and heat-pulse event: the importance of maintaining the continuum. Global Change Biol 22:2861–2874
Wardle DA, Bardgett RD, Klironomos JN, Setala H, van der Putten WH, Wall DH (2004) Ecological linkages between aboveground and belowground biota. Science 304:1629–1633
Wen Z, White PJ, Shen J, Lambers H (2022) Linking root exudation to belowground economic traits for resource acquisition. New Phytol 233:1620–1635
Wieczynski DJ, Singla P, Doan A, Singleton A, Han Z-Y, Votzke S, Yammine A, Gibert JP (2021) Linking species traits and demography to explain complex temperature responses across levels of organization. P Natl Acad Sci USA 118
Willers C, Jansen van Rensburg P, Claassens S (2015) Phospholipid fatty acid profiling of microbial communities–a review of interpretations and recent applications. J Appl Microbiol 119:1207–1218
Williams LJ, Cavender-Bares J, Paquette A, Messier C, Reich PB (2020) Light mediates the relationship between community diversity and trait plasticity in functionally and phylogenetically diverse tree mixtures. J Ecol 108:1617–1634
Wilson B, Puri G (2001) A comparison of pinewood and moorland soils in the Abernethy Forest Reserve, Scotland. Global Ecol Biogeogr 10:291–303
Xi N, Bloor JMG, Wang Y, Chu C (2019) Contribution of conspecific soil microorganisms to tree seedling light responses: Insights from two tropical species with contrasting shade tolerance. Environ Exp Bot 166
Xu J, Liu S, Song S, Guo H, Tang J, Yong JWH, Ma Y, Chen X (2018) Arbuscular mycorrhizal fungi influence decomposition and the associated soil microbial community under different soil phosphorus availability. Soil Biol Biochem 120:181–190
Xu Y, Seshadri B, Bolan N, Sarkar B, Ok YS, Zhang W, Rumpel C, Sparks D, Farrell M, Hall T (2019) Microbial functional diversity and carbon use feedback in soils as affected by heavy metals. Environ Int 125:478–488
Yang J, Blondeel H, Meeussen C, Govaert S, Vangansbeke P, Boeckx P, Lenoir J, Orczewska A, Ponette Q, Hedwall PO et al (2022a) Forest density and edge effects on soil microbial communities in deciduous forests across Europe. Appl Soil Ecol 179
Yang J, Wang Y, Cui X, Xue K, Zhang Y, Yu Z (2019) Habitat filtering shapes the differential structure of microbial communities in the Xilingol grassland. Sci Rep 9
Yang M, Fu Y, Zhao R, Koorem K, Li B, Siemann E, Yang Q (2022b) The effects of light availability on plant-soil interactions and salinity tolerance of invasive tree species, Triadica sebifera. Forest Ecol Manag 506
Yao F, Yang S, Wang Z, Wang X, Ye J, Wang X, DeBruyn JM, Feng X, Jiang Y, Li H (2017) Microbial Taxa Distribution Is Associated with Ecological Trophic Cascades along an Elevation Gradient. Front Microbiol 8
Zhang B, Li Y, Ren T, Tian Z, Wang G, He X, Tian C (2014) Short-term effect of tillage and crop rotation on microbial community structure and enzyme activities of a clay loam soil. Biol Fert Soils 50:1077–1085
Zhang J, Ekblad A, Sigurdsson BD, Wallander H (2020) The influence of soil warming on organic carbon sequestration of arbuscular mycorrhizal fungi in a sub-arctic grassland. Soil Biol Biochem 147:107826
Zhang N, Nunan N, Hirsch PR, Sun B, Zhou J, Liang Y (2021) Theory of microbial coexistence in promoting soil-plant ecosystem health. Biol Fert Soils 57:897–911
Zhou G, Liu S, Li Z, Zhang D, Tang X, Zhou C, Yan J, Mo J (2006) Old-Growth Forests Can Accumulate Carbon in Soils. Science 314:1417–1417
Zhou Z, Krashevska V, Widyastuti R, Scheu S, Potapov A (2022) Tropical land use alters functional diversity of soil food webs and leads to monopolization of the detrital energy channel. Elife 11
Acknowledgement
This work was supported by the European Research Council via ERC Consolidator Grant PASTFORWARD (Number 614839), ERC Starting Grant FORMICA (Number 757833), the FWO Scientific research network FLEUR, China Postdoctoral Science Foundation (Number 2023M733462), and the scholarship provided by the University of Chinese Academy of Sciences (UCAS). We thank Luc Willems and Greet De bruyn during the chemical analyses, as well as Samuel Bodé and Katja Van Nieuland for providing help during the PLFA experiment.
Author information
Authors and Affiliations
Contributions
J.Y., H.B., P.D.F., and K.V. conceived the ideas and designed methodology; all authors collected data; J.Y. performed statistical analyses; J.Y., with contributions from H.B., P.D.F., and K.V. wrote the paper; all authors discussed the results and commented on the manuscript drafts.
Corresponding author
Ethics declarations
Competing Interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Additional information
Responsible Editor: Paul Bodelier.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
ESM 1
(DOCX 1831 kb)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yang, J., Blondeel, H., Boeckx, P. et al. Responses of the soil microbial community structure to multiple interacting global change drivers in temperate forests. Plant Soil 496, 641–656 (2024). https://doi.org/10.1007/s11104-023-06388-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-023-06388-7