Abstract
Purpose
Understanding the impact of neighbor tree diversity on soil biodiversity at the individual tree scale and clarifying which facets of neighbor tree diversity have a decisive impact on soil biodiversity.
Methods
We collected and identified soil nematodes underneath 256 individual trees of 16 species at four species-richness levels (1, 2, 4, 8 species) in a large tree diversity experiment in southeast China. We analyzed how the taxonomic, phylogenetic, and functional diversity of the nematode community were influenced by neighbor tree diversity.
Results
Our analyses showed that nematode alpha diversity at the individual tree scale decreased with increasing neighbor tree richness while beta diversity increased at taxonomic, phylogenetic, and functional levels. Nematode alpha diversity at the plot scale increased with increasing neighbor tree richness. Secondly, we found that reducing redundant species and increasing distinct species led to the differentiation of nematode communities under different individual trees in high-richness plots. Finally, our data revealed that the functional diversity of the neighbor tree community had the largest effects on the diversity of the local nematode community.
Conclusion
Our study emphasizes that there is a significant influence from neighboring trees, mainly the functional traits of the trees, even though they are spaced at distances multitudes greater than the distance over which soil nematodes can move. Hence it is important to consider these broader spatial aspects when examining the plant and soil biotic interactions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plant diversity can affect the diversity and functioning of soil communities via host-specific effects through provisioning of diverse organic substrates that originate from root exudates and litter, and this can influence resource partitioning and niche differentiation of soil organisms (Gould et al. 2016; Wardle et al. 2004; Scherber et al. 2010; Schuldt et al. 2019). As measures of biodiversity, phylogenetic diversity (PD) and functional diversity (FD) connect the different evolutionary histories and trait-based aspects of species. Therefore, it can point out the deeper connections between the above- and belowground diversity, such as the co-evolutionary relationship between plants and soil organisms, and the relationship between plant functional traits and soil biological functions (Bardgett and van der Putten 2014; Flynn et al. 2011). However, most studies focus on taxonomic diversity (TD), while ignoring the phylogenetic diversity (PD) and functional diversity (FD) (Gundale and Kardol 2021; Le Bagousse-Pinguet et al. 2019). Understanding the relationship between the multiple facets diversity of both above- and belowground communities can better predict how the spatial distribution of trees influences belowground biota (Cameron et al. 2019; Korboulewsky et al. 2016; Thakur et al. 2020).
Theoretically, biological mechanisms altered by altering plant community diversity could be manifested at the individual tree scale through interactions between plants (Potvin and Dutilleul 2009), which have been demonstrated by finding that a positive correlation between tree diversity and productivity is driven by neighborhood tree diversity (Huang et al. 2018). Such an impact of biodiversity on the productivity of individual tree species is bound to further affect adjacent soil biodiversity through nutrient cycling processes such as litter decomposition. Recent studies have shown that the key factors that drive belowground biogeographic patterns differ with the spatial scale considered (Boraks et al. 2021; Thompson et al. 2017). At the plot scale, it is widely accepted that increased tree diversity can provide more niche opportunities, allowing for a positive correlation between tree diversity and species diversity of soil organisms (Chen et al. 2019; Korboulewsky et al. 2016). But, at the scale of plant individuals, studies have reported that the presence of neighbor plants can greatly reduce the diversity or abundance of soil macrofauna such as arthropods, and litter ants on individual plants (Kostenko et al. 2017; Skarbek et al. 2020). Interaction between plant species, such as competition for light or facilitation by common mycorrhizal networks, may directly or indirectly mediate the effect of plants on soil organisms (Setiawan et al. 2016; Bilas et al. 2021). Soil microfauna are also susceptible to varying local environmental conditions and have limited range of activity (Nielsen et al. 2015), hence they are expected to respond significantly to neighbor plant diversity. However, whether neighbor plant diversity exerts similar negative effects on belowground biota associated with a focal plant has remained unresolved so far. Revealing the differences in the impact of plant diversity changes on soil microfauna diversity at plot and plant individuals scale is of great significance for an in-depth understanding of the association mechanism between above- and belowground biodiversity.
Nematodes are one of the oldest group of organisms in the soil, with a high taxonomic and phylogenetic diversity, and the number of known species has exceeded 10,000 (van den Hoogen et al. 2019). Such diverse nematodes also perform various functions in the ecosystem. For example, they can be classified into different trophic groups based on their feeding habits, including bacterivores, fungivores, herbivores, omnivores and predators (Yeates et al. 1993). Nematodes are also widely distributed and highly responsive to environmental conditions (Xiong et al. 2021). Therefore, they are good indicators to investigate how soil biodiversity responds to tree diversity at different scales. The response of nematodes to tree diversity is related to the quantity and quality of soil resources provided by trees, and therefore is affected by neighbor trees (Cortois et al. 2017). Because of the presence of functionally or phylogenetically redundant species in a nematode community, the loss or gain of a single species may have only slight effects on PD and FD, or vice versa, specific species might have strong effects on PD and FD, leading to a divergence between changes in PD or FD and changes in TD (Baiser and Lockwood 2011; Jarzyna and Jetz 2017; Li et al. 2020). Evaluation of the changes in PD and FD and their congruence with changes in TD can therefore comprehensively point out the influence of the diversity of neighbor trees on the relative change of redundant species and distinct species in nematode communities (Singer et al. 2018; Wang et al. 2019). However, few studies to date have assessed the effects of neighbor tree diversity on the relative change of taxonomic, functional, and phylogenetic diversity.
Recent studies on plant diversity have demonstrated the importance of functional and phylogenetic diversity components in explaining the causal relationships between changes in plant species richness and biodiversity at higher trophic levels (Whitfeld et al. 2012; Staab et al. 2016). In addition, recent evidence indicates that plant functional traits and phylogenetic diversity are relevant drivers of the community structure of soil organisms (Pellissier et al. 2013; Wang et al. 2019). Compared with taxonomic diversity, functional and phylogenetic diversity can more accurately reflect specific characteristics of plant communities, which are often significantly correlated with soil biodiversity (Barberán et al. 2015). However, the relative importance of the effects of different tree diversity on soil biodiversity remains unclear, and it is uncertain whether key plant function indicators are relevant. Understanding the relationships between these aspects of diversity of above- and belowground communities can improve our understanding of the determinants of soil community composition.
Biodiversity-ecosystem functioning (BEF) experiments in forest ecosystems provide plots with controlled levels of tree diversity, and are therefore an ideal platform to evaluate the effects of tree diversity on different facets of soil biodiversity (Delgado-Baquerizo et al. 2020; Jochum et al. 2020). To understand the impact of neighbor tree diversity on soil biodiversity at different scales and to elucidate which facets of tree diversity have a stronger impact on soil biodiversity. We collected soil from underneath individuals of 16 tree species growing in plots with four different species richness levels in the BEF-China forest biodiversity experiment (Bruelheide et al. 2014). We first examined the impact of plot tree species richness on soil nematode diversity (TD, PD and FD) and community composition at the individual tree scale. Subsequently, we calculated TD, PD, and FD of the neighbor tree community to evaluate the contribution of different facets of neighbor tree diversity on soil nematode diversity. Specifically, we hypothesized that soil nematode TD, PD, and FD at the individual tree scale decrease with increasing neighbor tree species richness (H1); and PD and FD of the neighbor tree community are more strongly related to soil nematode diversity than TD of the tree community at the individual tree scale (H2).
Materials and methods
Study site and experiment design
The study site is part of the forest biodiversity-ecosystem functioning experiment in China (BEF-China), which was established near Xingangshan, Dexing, in Jiangxi province (29.08°-29.11° N, 117.90°-117.93° E). The region is characterized by a typical subtropical monsoon climate, with a mean annual precipitation of 1964 mm and a mean temperature of 15.1℃. The natural vegetation is subtropical forest with a mix of evergreen and deciduous species. The preceding pine and Chinese fir plantation were clear-cut in 2009. The total experiment used a pool of 42 tree species and 18 shrub species, planted in 566 plots (667 m2 each), which were distributed across two sites (A and B). The plot size is 25.8 m × 25.8 m and 400 tree individuals were regularly planted at a distance of 1.29 m in each plot. Species were randomly assigned to individual positions in each plot in the different treatments. The entire experiment consists of six tree species richness levels: 1, 2, 4, 8, 16 or 24 tree species, with 16, 8, 4, 2, 1 and 1 replicate plots for each richness level, respectively, at each site. In this design, each species is present in one specific community at each richness level at each site. The plots were distributed randomly in both sites. The soils of the region are mainly Cambisols, Regosols, Acrisols, Gleysols and Anthrosols (Scholten et al. 2017). A more detailed description of BEF-China can be found in Bruelheide et al. (2014).
Soil sampling
For the current study, soil sampling was conducted in September 2015. We selected 16 target tree species to sample soil nematodes: Castanea henryi, Nyssa sinensis, Liquidambar formosana, Sapindus saponaria, Quercus serrata, Castanopsis sclerophylla, Choerospondias axillaris, Triadica sebifera, Alniphyllum fortunei, Machilus leptophylla, Elaeocarpus chinensis, Machilus thunbergii, Betula luminifera, Castanopsis fargesii, Manglietia yuyuanensis and Quercus phillyreoides. Eight of these species were planted at site A, while the other eight species were planted at site B. For each tree species, we sampled soil under four tree individuals in plots with 1, 2, 4 and 8 species diversity levels. In total, there were 256 soil samples (16 tree species × 4 diversity levels × 4 tree individuals). For each soil sample, ten cores (0–10 cm in depth, 2.5 cm in diameter) were collected 10 cm around the center of the selected tree (Fig. 1). The ten cores were then pooled into one sample. Subsequently, the samples were taken to laboratory and stored at 4 °C for further analysis. Each soil sample was divided into two subsamples, one to test soil moisture and one for nematode extraction. Nematodes were extracted from 100 g of fresh soil using a cotton-wool filter method (Townshend 1963). After counting the total nematodes of each sample, 100 individuals per sample were randomly chosen and identified at genus level, according to nematode classification guides (Bongers 1994; Li et al. 2017).
Nematode phylogenetic tree
With the development of DNA sequencing and computing technologies, a substantial number of gene sequences from soil-dwelling nematodes has been included in databases and this has increased the accuracy of their phylogenetic relationships. The Open Tree of Life (OTL) project constructs a comprehensive, dynamic and digitally available tree of life by synthesizing published phylogenetic trees along with taxonomic data (Hinchliff et al. 2015). Here, we built a nematode phylogenetic tree for all identified genera in this study based on phylogenetic and taxonomic data from the OTL database using the “rotl” package (Michonneau et al. 2016). Polytomies in the phylogenetic tree were replaced with a random series of dichotomies with one or more branches, using the multi2di function in the “ape” package (Paradis and Schliep 2019). Since the synthetic tree does not have associated branch lengths for nematodes, we used Grafen’s method to compute branch lengths in which the length of each node is defined by the number of descendant tips minus one (Grafen and Hamilton 1989).
Nematode trait-based dendrogram
We used nematode feeding habits, colonizer-persister values of life-course strategies (c-p value) and genus body mass (µg) to estimate the functional characterization of the nematodes. Feeding habits reflect the trophic level of nematodes in the soil food web and reflect the role of nematodes in decomposition processes and nutrient cycling. Soil nematodes can be categorized into a 1–5 colonizer-persister series, ranging from extreme r- to k-strategists (c-p value 1–5). The body mass of nematodes was calculated based on the general morphology of each genus. Thus, it partly represents a measure of nematode activity. The selected traits are critical for nematodes to perform different functions in terrestrial ecosystems. Since there are categorical variables in our functional traits, we chose tree-based functional measures. All nematode traits were obtained from the Nemaplex database (http://nemaplex.ucdavis.edu). We first calculated the distance matrix (functional dissimilarity) between samples using Gower’s distance (1971), and then used the unweighted pair group method with arithmetic mean clustering (UPGMA) to transform the matrix into a trait-based dendrogram. Cluster analysis was performed using the “cluster” package (Maechler et al., 2019).
Nematode taxonomic, phylogenetic, and functional diversity
At the individual tree scale, alpha taxonomic diversity (αTD) was expressed as the total number of genera present in each sample. Faith’s phylogenetic diversity was used to calculate alpha phylogenetic diversity (αPD) and functional diversity (αFD) based on nematode phylogenetic trees and trait-based dendrogram, respectively (Tsirogiannis and Sandel 2016). For beta diversity, beta taxonomic diversity (βTD) was assessed using Sørensen’s dissimilarity between samples. Phylogenetic Sørensen’s dissimilarity was used to calculate beta phylogenetic diversity (βPD) and beta functional diversity (βFD). Functional redundancy and phylogenetic redundancy were calculated as described in Ricotta et al. (2018). Nestedness component and turnover component of the nematode community were calculated following the method of Baselga (2010). At the plot scale, nematode αTD, αPD and αFD was obtained by merging all samples in each plot and then calculated using the same method as the individual tree scale.
Tree taxonomic, phylogenetic, and functional diversity
We selected four functional traits related to belowground diversity to describe the functional diversity of trees: (a) leaf nitrogen content per leaf dry mass; (b) leaf carbon/nitrogen (C/N) ratio; (c) vegetative tree height; (d) specific leaf area (SLA). The functional traits of each tree species were obtained from the TRY plant trait database (Kattge et al. 2020). The tree alpha and beta diversities of each sampling plot were calculated using the same method as described for the nematode community.
Statistical analyses
All statistical analyses were conducted in the R statistical software version 3.6.1. (R Core Team 2021). Linear mixed-effects model was used to test the effects of tree species richness per plot and tree species identity and their interaction on nematode alpha and beta diversity. Plot identity was used as a random factor. The relationships between plot tree species richness, and nematode diversity at the individual tree scale were tested with a Spearman correlation analysis. We assess the impact of changes in the diversity of aboveground tree species on the assemblage of nematode communities by evaluating the relative change in nematode αTD, αFD and αPD (e.g. relative change in αTD = [αTD in a tree species mixture – αTD in the tree species monoculture] / αTD in the tree species monoculture) (Jarzyna and Jetz 2017). For example, a larger decrease in αTD compared to αFD or αPD would arise from species new to the assemblage that have redundant trait characteristics or phylogenetic positions. Similarly, the relative change in βTD, βFD, βPD (e.g. relative change in βTD = βTD of the tree species mixture – βTD of the tree species monoculture) can reflect changes in dissimilarity of soil nematode community function and phylogeny relationship between treatments (Baiser and Lockwood 2011). If the relative change in βTD and βFD or βPD increase at the same time, two assemblages must gain different functionally or phylogenetically distinct species. A Wald-test was used to test for differences among the slope of the relationship between any changes of nematode diversity and a 1:1 line.
In order to compare the effects of different facets tree diversity on nematode diversity, we constructed a structural equation model (SEM). We started with a simple conceptual model, for which we hypothesized that tree taxonomic, phylogenetic, and functional diversity drive nematode taxonomic, phylogenetic, and functional diversity. We also hypothesized a direct effect of taxonomic diversity on functional and phylogenetic diversity for both the tree and nematode communities. We fitted the hypothesized model to the data and evaluated the model based on χ2 value and p-value using the “lavaan” package (Rosseel 2012). The relationships between tree functional traits and alpha diversity were tested with a Spearman correlation analysis. The relationships between tree functional traits and beta diversity were tested with mantel test analysis.
Results
Treatment effects on nematode alpha and beta diversity
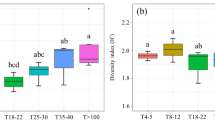
A total 140 genera from 256 soil samples were identified, including 39 bacterivores, 17 fungivores, 41 herbivores, and 43 omnivores and predators. Three bacterivore genera (Alaimus, Paramphidelus, Prismatolaimus), one fungivore (Filenchus) and one herbivore (Helicotylenchus) were dominant genera (Table S1). At the individual tree scale, nematode αTD, αPD and αFD were significantly affected by plot tree species richness, tree identity and their interaction (Table S2). Nematode αTD and αFD decreased with increasing plot tree richness (Fig. 2a, c). Nematode βTD, βPD and βFD were significantly affected by plot tree species richness, tree species identity and their interaction (Table S2), and increased with increasing plot tree richness (Fig. 2d, e, f). At the plot scale, the nematode αTD, αPD and αFD obtained by merging all samples in each plot increased with increasing plot tree richness (Fig. 3). Nematode phylogenetic redundancy decreased with increasing plot tree richness (Fig. S1b). The diversity pattern of soil nematodes was mainly explained by the turnover component (Fig. S1c, d). Bacterivores and herbivores were the most abundant among different trophic groups. Bacterivores and fungivores increased with increasing plot tree richness (Fig. S2).
Boxplots of nematode alpha and beta TD, PD, and FD values at the individual tree scale for the four tree species richness treatments. Correlation coefficients (r) are from a Spearman correlation analysis between nematode diversity and plot tree species richness. The significance of the slope of the regression is indicated by asterisks (*p < 0.05; **p < 0.01). The x-axes are log2-transformed
Boxplots of nematode αTD (a), αPD (b), and αFD (c) values at plot scale for the four tree species richness treatments. Correlation coefficients (r) are from a Spearman correlation analysis between nematode diversity and plot tree species richness. The significance of the slope of the regression is indicated by asterisks (*p < 0.01). The x-axes are log2-transformed
Changes in nematode alpha and beta diversity
A large relative change in nematode αTD was generally associated with a small relative change in both αPD and αFD (slopes significantly smaller than 1; Fig. 4a, b). Most nematode assemblages with a decrease in the relative change in αTD showed a higher decrease in the relative change in αTD than in the relative change in αPD or αFD (307 and 301, respectively; Fig. 4a, b). We found that the relative change in βTD of nematode assemblage between high richness tree plots and monoculture plots exhibited more increases than decreases (1057 and 649, respectively; Fig. 4d, e). Among those points that experienced increases in the relative change in βTD, most experienced an increase in the relative change in βPD and βFD (554 and 570, respectively; Fig. 4d, e).
Relationships between relative change in nematode TD, PD, and FD based on alpha (a, b) and beta diversity (d, e) at the individual tree scale. The insets show the number of points within the different quadrants with the largest value being shown in bold for each inset. The concept diagram (c, f) shows the potential consequences of changes in diversity for nematode community assemblages. + and - indicate gains and losses of taxa, respectively. Blue solid lines indicate the slope of the relationship between change in TD and change in FD or PD. Black dashed lines indicate the 1:1 line. The quadrant with gray background has the most points
Relationships between the tree communities and the soil nematode communities
In the SEM (Fig. 5a, χ2 = 0.71, DF = 2, p = 0.70), we did not find a significant effect of the tree αTD on nematode αTD and αFD. Instead, nematode αTD and αPD were directly driven by the tree αFD and αPD, respectively. Within the tree community or nematode community, αTD had a direct effect on αPD and αFD, and tree αFD was significantly correlated with tree αPD. Tree αFD was the decisive factor affecting the alpha diversity of nematode communities (Fig. 5b). SLA and tree height were negatively correlated with soil nematode alpha diversity. Leaf C/N ratio, SLA and tree height were positively correlated with soil nematode beta diversity (Fig. 6). Phylogeny and trait-based dendrogram of the soil nematodes and plant were shown in Figs. S3 and S4, respectively.
Structural equation model linking tree TD, PD, and FD to nematode TD, PD, and FD based on alpha (χ2 = 0.71, DF = 2, p = 0.70) diversity at the individual tree scale (a). Arrows indicate causal relationships among measured variables (solid arrows = significant relationships; dashed arrows = non-significant relationships; p > 0.05). Numbers in the arrows give the standardized coefficients. Arrow width was scaled by the standardized path coefficients. Standardized total effects of aboveground diversity to belowground diversity derived from the structural equation models (b)
Discussion
Our study provides insights into the linkages between neighbor tree diversity at the plot scale and the multiple facets of the soil nematode diversity at the individual tree scale. Our study shows that increased tree species richness at the plot level slightly decreased alpha diversity and increased beta diversity of the nematode community at the individual tree scale. Importantly, αFD of the tree community appeared as a decisive factor affecting the diversity of nematode communities. Our findings are relevant for the assessment of the impact of tree diversity on soil biodiversity and highlight the importance of multifaceted diversity, leading to a deeper understanding of the significance of plant biodiversity conservation (Delgado-Baquerizo et al. 2019; Schuldt et al. 2019).
Nematode diversity and community assemblages
Consistent to our first hypothesis, nematode αTD, αPD and αFD at the individual tree scale slightly decreased with tree diversity at the plot level (Fig. 2). Other studies have shown that the diversity of soil biota is usually positively related to tree diversity in a certain area, like a single plot, and this has often been explained to be a result of increased resource heterogeneity or an increase in the number of niche opportunities (Skarbek et al. 2020; Zhang et al. 2016). Our study showed that these contradictory results are caused by the differences in the study scale (Reid and Emery 2018). In our research, when all individual tree scale samples in one plot are combined into a plot-scale sample, soil alpha diversity was also strongly positively correlated with plot tree richness (Fig. 3). An increase in neighbor trees diversity can lead to positive or negative effects on individual trees by competition for light, amelioration of abiotic stress, or modification of soil microbes (Bilas et al. 2021). Therefore, these neighboring trees may indirectly affect the soil nematode community by affecting the amount of litter and root exudates of local individual trees. We found that tree species identity and plot tree species richness have a strong interaction effect on soil nematode diversity, which means that the impact of local tree species on soil nematode diversity is indeed regulated by neighboring species (Tedersoo et al. 2016). Under different soil conditions driven by tree diversity, the process of local extinction or persistence of nematodes may result in a decrease in local nematode diversity. Previous studies have found that the spatial variance of fast-growing bacterivores and fungivores is positively correlated with soil organic matter content (Mikola and Sulkava 2001; Quist et al. 2019), considering the increase of tree species richness can promote the accumulation of soil organic carbon at different soil depths (Li et al. 2019a) and positive relationship between bacterivores, fungivores and plot tree species richness, our results suggested that soil resources might be essential for neighbor tree diversity affecting local soil biodiversity. This speculation is also supported by the greater contribution of turnover components to nematode beta diversity, which is driven by potential ecological processes caused by environmental gradients, such as environmental filtering or competition (Fig. S1c, d) (Legendre 2014).
Important changes in nematode communities are not only brought about by changes at the taxonomic level, but also by those at the functional or phylogenetic level (Liu et al. 2019). We found that the relative changes in αPD and αFD of nematode communities among single tree species and high richness tree plots are weaker than the relative changes in αTD (Fig. 4a, b). This means that those nematodes lost in the community were functionally and phylogenetically redundant (Jarzyna and Jetz 2017). This is also supported by the results that functional and phylogenetic redundancy indices were lower in the high-richness plots (Fig. S1a, b). The high redundancy in the low-richness plots indicates that the nematode community coexists in a state of highly overlapping resource requirements (Matlack 2001; Neher 2010). At the same time, the relative change in βPD and βFD of soil nematode communities were more often positive than negative (Fig. 4d, e), which means that with different tree species in the neighborhood, functional and phylogenetic relationships of nematode communities became more differentiated (Baiser and Lockwood 2011). A decrease in redundant species or increase in distinct species caused differentiation of nematode community functions or phylogenetic relationships (Fig. 4f). The functions or phylogenetic differentiation in high-richness plots indicated that the diversification of adjacent tree species benefits the local community to maintain more nematodes with different functions and phylogenetic relationships (Li et al. 2020). Conversely, although the alpha diversity of nematodes under each tree slightly increased when neighbor tree diversity decreased, our results suggest that the increase was due to functionally or phylogenetically redundant species. More specifically, when considered at a relatively large scale, such as the plot scale, the similarity between soil nematode communities in different facets will increase with neighbor tree diversity decreased because some distinct species are lost when environmental conditions are similar. Nematode diversity in different facets at the plot scale is indeed higher in high-richness plots (Fig. 3), highlighting the importance of maintaining plant diversity, not only at the taxonomic level but also in functional and phylogenetic relationships for soil biodiversity.
Relative importance of tree multiple facets diversity
In agreement with our second hypothesis, our study shows that αFD of neighbor trees is a better factor explaining the alpha diversity of nematode communities among the three facets of tree diversity (Fig. 5). Tree height and the leaf economics spectrum (e.g., specific leaf area and leaf nutrient content) largely determined the quantity and quality of litter and detritus, and both litter and detritus have been demonstrated to have important effects on soil biotic communities (Fu et al. 2017; Li et al. 2019b). Therefore, functional diversity could be the most direct indicator reflecting the influence of plant communities on soil communities, rather than taxonomic diversity and phylogenetic diversity (Mahaut et al. 2020). The negative effect of neighbor trees αFD on nematode diversity of multiple facets at the individual tree scale indicates that diverse tree mixtures modulate nematode communities at the local scale, which may have far-reaching consequences, for example by influencing the nutrient cycling process in which soil nematodes participate (Bardgett and van der Putten 2014) and may affect aboveground diversity through plant-soil feedback effects (van der Putten et al. 2016). Further research should verify how different combinations of functional traits affect soil community, which have important implications for maximizing the effects of reforestation and plantation forestry and maintaining biodiversity.
Previous studies have found no significant relationship between plant functional traits and soil biota, and this may be explained by the absence of key traits that drive plant-soil organism associations (Leff et al. 2018; Marshall et al. 2011; Meyer et al. 2018). The three functional traits we selected in this study, including trophic groups, c-p value, and body mass, can well reflect the intensity, preference, and turnover rate of nematodes involved in nutrient cycling. The above-ground functional diversity represented by the leaf nutrient content significantly affected the nematode functional diversity, emphasized the importance of functional diversity in connecting above- and belowground relationships, and supported our speculation that the resource characteristics affected by neighbor tree diversity drive the assembly of soil nematode communities. Some studies also suggested that phylogenetic diversity may be a better predictor than functional diversity on certain conditions. However, we found that tree αPD did not have a strong impact on soil nematode diversity. First, it is possible that plant phylogenetic differences are associated with only a subset of the belowground communities due to coevolution with mutualists or pathogens, instead of being associated with the entire soil community (Anacker et al. 2014). Second, our experiment is still in the initial stage of recovery since it was sampled only six years after the trees had been planted, and the tree community and specific underground organisms may not have yet developed final co-evolutionary relationships. In general, our results suggest that the neighbor trees mainly regulate soil nematode diversity through functional traits, such as plant height and SLA and leaf C/N ratio (Fig. 6), but that the TD and PD also play a role (Milcu et al. 2013).
Conclusion
In this study, we assessed how soil nematode diversity at the individual tree scale responds to neighbor tree diversity, considering the taxonomic, phylogenetic, and functional facets of alpha and beta diversities in subtropical forest ecosystem. We found that the increased richness of neighbor trees slightly reduced alpha diversity of nematode community at plant scale, but significantly increased alpha diversity of nematode community at plot scale. The differential results revealed the different responses of redundant and rare soil nematode species to changes in aboveground plant diversity, suggesting that higher plant diversity is beneficial to the maintenance of phylogenetic relationships with specific functions in the nematode community, emphasizing the importance of aboveground diversity in maintaining the diversity of soil organisms in different dimensions. Furthermore, the finding that tree αFD had the largest effects on the soil nematode community opens new avenues of biodiversity and ecosystem functioning research. Future research could use controlled experiments to investigate which of the resources provided by trees have the greatest impact on soil communities and whether these effects are brought about by fallen leaves or root exudates.
Data availability
Data supporting the findings of this study will be deposited in the SCIENCE DATA BANK.
References
Anacker BL, Klironomos JN, Maherali H, Reinhart KO, Strauss SY (2014) Phylogenetic conservatism in plant-soil feedback and its implications for plant abundance. Ecol Lett 17:1613–1621. https://doi.org/10.1111/ele.12378
Baiser B, Lockwood JL (2011) The relationship between functional and taxonomic homogenization. Global Ecol Biogeogr 20:134–144. https://doi.org/10.1111/j.1466-8238.2010.00583.x
Barberán A, McGuire KL, Wolf JA, Jones FA, Wright SJ, Turner BL, Essene A, Hubbell SP, Faircloth BC, Fierer N (2015) Relating belowground microbial composition to the taxonomic, phylogenetic, and functional trait distributions of trees in a tropical forest. Ecol Lett 18:1397–1405. https://doi.org/10.1111/ele.12536
Bardgett RD, van der Putten WH (2014) Belowground biodiversity and ecosystem functioning. Nature 515:505. https://doi.org/10.1038/nature13855
Baselga A (2010) Partitioning the turnover and nestedness components of beta diversity. Global Ecol Biogeogr 19:134–143. https://doi.org/10.1111/j.1466-8238.2009.00490.x
Bilas RD, Bretman A, Bennett T (2021) Friends, neighbours and enemies: an overview of the communal and social biology of plants. Plant Cell Environ 44:997–1013. https://doi.org/10.1111/pce.13965
Bongers T (1994) De Nematoden van Nederland. Vormgeving en technische realisatie. Uitgeverij Pirola, Schoorl
Boraks A, Plunkett GM, Doro TM, Alo F, Sam C, Tuiwawa M, Ticktin T, Amend AS (2021) Scale-dependent influences of distance and vegetation on the composition of aboveground and belowground tropical fungal communities. Microb Ecol 81:874–883. https://doi.org/10.1007/s00248-020-01608-4
Bruelheide H, Nadrowski K, Assmann T, Bauhus J, Both S, Buscot F, Chen XY, Ding BY, Durka W, Erfmeier A et al (2014) Designing forest biodiversity experiments: general considerations illustrated by a new large experiment in subtropical China. Methods Ecol and Evol 5:74–89. https://doi.org/10.1111/2041-210X.12126
Cameron EK, Martins IS, Lavelle P, Mathieu J, Tedersoo L, Bahram M, Gottschall F, Guerra CA, Hines J, Patoine G et al (2019) Global mismatches in aboveground and belowground biodiversity. Conserv Biol 33:1187–1192. https://doi.org/10.1111/cobi.13311
Chen C, Chen HYH, Chen X, Huang Z (2019) Meta-analysis shows positive effects of plant diversity on microbial biomass and respiration. Nat Commun 10:1332. https://doi.org/10.1038/s41467-019-09258-y
Cortois R, Veen GF, Duyts H, Abbas M, Strecker T, Kostenko O, Eisenhauer N, Scheu S, Gleixner G, De Deyn GB et al (2017) Possible mechanisms underlying abundance and diversity responses of nematode communities to plant diversity. Ecosphere 8:e01719. https://doi.org/10.1002/ecs2.1719
Delgado-Baquerizo M, Bardgett RD, Vitousek PM, Maestre FT, Williams MA, Eldridge DJ, Lambers H, Neuhauser S, Gallardo A, García-Velázquez L et al (2019) Changes in belowground biodiversity during ecosystem development. Proc Natl Acad Sci USA 116:6891. https://doi.org/10.1073/pnas.1818400116
Delgado-Baquerizo M, Reich PB, Trivedi C, Eldridge DJ, Abades S, Alfaro FD, Bastida F, Berhe AA, Cutler NA, Gallardo A et al (2020) Multiple elements of soil biodiversity drive ecosystem functions across biomes. Nat Ecol Evol 4:210–220. https://doi.org/10.1038/s41559-019-1084-y
Flynn DFB, Mirotchnick N, Jain M, Palmer MI, Naeem S (2011) Functional and phylogenetic diversity as predictors of biodiversity–ecosystem-function relationships. Ecology 92:1573–1581. https://doi.org/10.1890/10-1245.1
Fu X, Guo D, Wang H, Dai X, Li M, Chen F (2017) Differentiating between root- and leaf-litter controls on the structure and stability of soil micro-food webs. Soil Biol Biochem 113:192–200. https://doi.org/10.1016/j.soilbio.2017.06.013
Gundale MJ, Kardol P (2021) Multi-dimensionality as a path forward in plant-soil feedback research. J Ecol 109:3446–3465. https://doi.org/10.1111/1365-2745.13679
Gould IJ, Quinton JN, Weigelt A, De Deyn GB, Bardgett RD (2016) Plant diversity and root traits benefit physical properties key to soil function in grasslands. Ecol Lett 19:1140–1149. https://doi.org/10.1111/ele.12652
Gower JC (1971) A general coefficient of similarity and some of its properties. Biometrics 27:857. https://doi.org/10.2307/2528823
Grafen A, Hamilton WD (1989) The phylogenetic regression. Phil Trans R Soc London B Biol Sci 326:119–157. https://doi.org/10.1098/rstb.1989.0106
Hinchliff CE, Smith SA, Allman JF, Burleigh JG, Chaudhary R, Coghill LM, Crandall KA, Deng J, Drew BT, Gazis R et al (2015) Synthesis of phylogeny and taxonomy into a comprehensive tree of life. Proc Natl Acad Sci 112:12764. https://doi.org/10.1073/pnas.1423041112
Huang Y, Chen Y, Castro-Izaguirre N, Baruffol M, Brezzi M, Lang A, Li Y, Härdtle W, von Oheimb G, Yang X et al (2018) Impacts of species richness on productivity in a large-scale subtropical forest experiment. Science 362:80–83. https://doi.org/10.1126/science.aat6405
Jarzyna MA, Jetz W (2017) A near half-century of temporal change in different facets of avian diversity. Global Change Biol 23:2999–3011. https://doi.org/10.1111/gcb.13571
Jochum M, Fischer M, Isbell F, Roscher C, Plas Fvd, Boch S, Boenisch G, Buchmann N, Catford JA, Cavender-Bares J et al (2020) The results of biodiversity–ecosystem functioning experiments are realistic. Nat Ecol Evol 4:1485–1494. https://doi.org/10.1038/s41559-020-1280-9
Kattge J, Bönisch G, Díaz S, Lavorel S, Prentice IC, Leadley P, Tautenhahn S, Werner GDA, Aakala T, Abedi M et al (2020) TRY plant trait database–enhanced coverage and open access. Global Change Biol 26:119–188. https://doi.org/10.1111/gcb.14904
Korboulewsky N, Perez G, Chauvat M (2016) How tree diversity affects soil fauna diversity: a review. Soil Biol Biochem 94:94–106. https://doi.org/10.1016/j.soilbio.2015.11.024
Kostenko O, Mulder PPJ, Courbois M, Bezemer TM (2017) Effects of plant diversity on the concentration of secondary plant metabolites and the density of arthropods on focal plants in the field. J Ecol 105:647–660. https://doi.org/10.1111/1365-2745.12700
Le Bagousse-Pinguet Y, Soliveres S, Gross N, Torices R, Berdugo M, Maestre FT (2019) Phylogenetic, functional, and taxonomic richness have both positive and negative effects on ecosystem multifunctionality. Proc Natl Acad Sci USA 116:8419. https://doi.org/10.1073/pnas.1815727116
Leff JW, Bardgett RD, Wilkinson A, Jackson BG, Pritchard WJ, De Long JR, Oakley S, Mason KE, Ostle NJ, Johnson D et al (2018) Predicting the structure of soil communities from plant community taxonomy, phylogeny, and traits. ISME J 12:1794–1805. https://doi.org/10.1038/s41396-018-0089-x
Legendre P (2014) Interpreting the replacement and richness difference components of beta diversity. Global Ecol Biogeogr 23:1324–1334. https://doi.org/10.1111/geb.12207
Li Q, Liang WJ, Zhang XK, Mahamood M (2017) Soil nematodes of grasslands in northern China. Academic, Cambridge
Li X, Zhu H, Geisen S, Bellard C, Hu F, Li H, Chen X, Liu M (2020) Agriculture erases climate constraints on soil nematode communities across large spatial scales. Global Change Biol 26:919–930. https://doi.org/10.1111/gcb.14821
Li Y, Bruelheide H, Scholten T, Schmid B, Sun ZK, Zhang NL, Bu WS, Liu XJ, Ma KP (2019) Early positive effects of tree species richness on soil organic carbon accumulation in a large-scale forest biodiversity experiment. J Plant Ecol 12:882–893. https://doi.org/10.1093/jpe/rtz026
Li YB, Bezemer TM, Yang JJ, Lü X, Li XT, Liang WJ, Han XG, Li Q (2019) Changes in litter quality induced by N deposition alter soil microbial communities. Soil Biol Biochem 130:33–42. https://doi.org/10.1016/j.soilbio.2018.11.025
Liu T, Hu F, Li H (2019) Spatial ecology of soil nematodes: perspectives from global to micro scales. Soil Biol Biochem 137:107565. https://doi.org/10.1016/j.soilbio.2019.107565
Maechler M, Rousseeuw P, Struyf A, Hubert M, Hornik K (2019) cluster: Cluster analysis basics and extensions. R package version 2.1.0. https://CRAN.R-project.org/package=cluster. Accessed June 2019
Mahaut L, Fort F, Violle C, Freschet GT (2020) Multiple facets of diversity effects on plant productivity: species richness, functional diversity, species identity and intraspecific competition. Funct Ecol 34:287–298. https://doi.org/10.1111/1365-2435.13473
Marshall CB, McLaren JR, Turkington R (2011) Soil microbial communities resistant to changes in plant functional group composition. Soil Biol Biochem 43:78–85. https://doi.org/10.1016/j.soilbio.2010.09.016
Matlack GR (2001) Factors determining the distribution of soil nematodes in a commercial forest landscape. For Ecol Manag 146:129–143. https://doi.org/10.1016/S0378-1127(00)00454-0
Meyer ST, Ptacnik R, Hillebrand H, Bessler H, Buchmann N, Ebeling A, Eisenhauer N, Engels C, Fischer M, Halle S et al (2018) Biodiversity-multifunctionality relationships depend on identity and number of measured functions. Nat Ecol Evol 2:44–49. https://doi.org/10.1038/s41559-017-0391-4
Michonneau F, Brown JW, Winter DJ (2016) Rotl: an R package to interact with the open tree of life data. Methods Ecol Evol 7:1476–1481. https://doi.org/10.1111/2041-210X.12593
Milcu A, Allan E, Roscher C, Jenkins T, Meyer ST, Flynn D, Bessler H, Buscot F, Engels C, Gubsch M et al (2013) Functionally and phylogenetically diverse plant communities key to soil biota. Ecology 94:1878–1885. https://doi.org/10.1890/12-1936.1
Mikola J, Sulkava P (2001) Responses of microbial-feeding nematodes to organic matter distribution and predation in experimental soil habitat. Soil Biol Biochem 33:811–817. https://doi.org/10.1016/S0038-0717(00)00229-7
Neher DA (2010) Ecology of plant and free-living nematodes in natural and agricultural soil. Annu Rev Phytopathol 48:371–394. https://doi.org/10.1146/annurev-phyto-073009-114439
Nielsen UN, Wall DH, Six J (2015) Soil biodiversity and the environment. Annu Rev Environ Resour 40:63–90. https://doi.org/10.1146/annurev-environ-102014-021257
Paradis E, Schliep K (2019) Ape 5.0: an environment for modern phylogenetics and evolutionary analyses. R Bioinf 35:526–528. https://doi.org/10.1093/bioinformatics/bty633
Pellissier L, Ndiribe C, Dubuis A, Pradervand JN, Salamin N, Guisan A, Rasmann S (2013) Turnover of plant lineages shapes herbivore phylogenetic beta diversity along ecological gradients. Ecol Lett 16:600–608. https://doi.org/10.1111/ele.12083
Potvin C, Dutilleul P (2009) Neighborhood effects and size-asymmetric competition in a tree plantation varying in diversity. Ecology 90:321–327. https://doi.org/10.1890/08-0353.1
Quist CW, Gort G, Mooijman P, Brus DJ, van den Elsen S, Kostenko O, Vervoort M, Bakker J, van der Putten WH, Helder J (2019) Spatial distribution of soil nematodes relates to soil organic matter and life strategy. Soil Biol Biochem 136:107542. https://doi.org/10.1016/j.soilbio.2019.107542
R Core Team (2021). R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. https://www.R-project.org/. Accessed July 2019
Reid ML, Emery SM (2018) Scale-dependent effects of Gypsophila paniculata invasion and management on plant and soil nematode community diversity and heterogeneity. Biol Conserv 224:153–161. https://doi.org/10.1016/j.biocon.2018.05.026
Ricotta C, Bacaro G, Caccianiga M, Cerabolini BEL, Pavoine S (2018) A new method for quantifying the phylogenetic redundancy of biological communities. Oecologia 186:339–346. https://doi.org/10.1007/s00442-017-4026-x
Rosseel Y (2012) lavaan: an R Package for Structural equation modeling. J Stat Softw 48:1–36. https://doi.org/10.18637/jss.v048.i02
Scherber C, Eisenhauer N, Weisser WW, Schmid B, Voigt W, Fischer M, Schulze ED, Roscher C, Weigelt A, Allan E et al (2010) Bottom-up effects of plant diversity on multitrophic interactions in a biodiversity experiment. Nature 468:553–556. https://doi.org/10.1038/nature09492
Scholten T, Goebes P, Kuhn P, Seitz S, Assmann T, Bauhus J, Bruelheide H, Buscot F, Erfmeier A, Fischer M et al (2017) On the combined effect of soil fertility and topography on tree growth in subtropical forest ecosystems-a study from SE China. J Plant Ecol 10:111–127. https://doi.org/10.1093/jpe/rtw065
Schuldt A, Ebeling A, Kunz M, Staab M, Guimarães-Steinicke C, Bachmann D, Buchmann N, Durka W, Fichtner A, Fornoff F et al (2019) Multiple plant diversity components drive consumer communities across ecosystems. Nat Commun 10:1460. https://doi.org/10.1038/s41467-019-09448-8
Setiawan NN, Vanhellemont M, Baeten L, Gobin R, De Smedt P, Proesmans W, Ampoorter E, Verheyen K (2016) Does neighbourhood tree diversity affect the crown arthropod community in saplings? Biodivers Conserv 25:169–185. https://doi.org/10.1007/s10531-015-1044-z
Singer D, Kosakyan A, Seppey CVW, Pillonel A, Fernández LD, Fontaneto D, Mitchell EAD, Lara E (2018) Environmental filtering and phylogenetic clustering correlate with the distribution patterns of cryptic protist species. Ecology 99:904–914. https://doi.org/10.1002/ecy.2161
Skarbek CJ, Noack M, Bruelheide H, Härdtle W, von Oheimb G, Scholten T, Seitz S, Staab M (2020) A tale of scale: plot but not neighbourhood tree diversity increases leaf litter ant diversity. J Anim Ecol 89:299–308. https://doi.org/10.1111/1365-2656.13115
Staab M, Bruelheide H, Durka W, Michalski S, Purschke O, Zhu CD, Klein AM (2016) Tree phylogenetic diversity promotes host–parasitoid interactions. Proc R Soc B Biol Sci 283:20160275. https://doi.org/10.1098/rspb.2016.0275
Tedersoo L, Bahram M, Cajthaml T, Põlme S, Hiiesalu I, Anslan S, Harend H, Buegger F, Pritsch K, Koricheva J et al (2016) Tree diversity and species identity effects on soil fungi, protists and animals are context dependent. ISME J 10:346–362. https://doi.org/10.1038/ismej.2015.116
Thakur MP, Phillips HRP, Brose U, De Vries FT, Lavelle P, Loreau M, Mathieu J, Mulder C, van der Putten WH, Rillig MC et al (2020) Towards an integrative understanding of soil biodiversity. Biol Rev 95:350–364. https://doi.org/10.1111/brv.12567
Thompson LR, Sanders JG, McDonald D, Amir A, Ladau J, Locey KJ, Prill RJ, Tripathi A, Gibbons SM, Ackermann G et al (2017) A communal catalogue reveals Earth’s multiscale microbial diversity. Nature 551:457–463. https://doi.org/10.1038/nature24621
Townshend JL (1963) A modifcation and evaluation of the apparatus for the Oostenbrink direct cotton wool flter extraction method. Nematologica 9:106–110
Tsirogiannis C, Sandel B (2016) PhyloMeasures: a package for computing phylogenetic biodiversity measures and their statistical moments. Ecography 39:709–714. https://doi.org/10.1111/ecog.01814
van den Hoogen J, Geisen S, Routh D, Ferris H, Traunspurger W, Wardle DA, De Goede RGM, Adams BJ, Ahmad W, Andriuzzi WS et al (2019) Soil nematode abundance and functional group composition at a global scale. Nature 572:194–198. https://doi.org/10.1038/s41586-019-1418-6
van der Putten WH, Bradford MA, Brinkman EP, van de Voorde TFJ, Veen GF (2016) Where, when and how plant-soil feedback matters in a changing world. Funct Ecol 30:1109–1121. https://doi.org/10.1111/1365-2435.12657
Wang MQ, Li Y, Chesters D, Anttonen P, Bruelheide H, Chen JT, Durka W, Guo PF, Hardtle W, Ma KP et al (2019) Multiple components of plant diversity loss determine herbivore phylogenetic diversity in a subtropical forest experiment. J Ecol 107:2697–2712. https://doi.org/10.1111/1365-2745.13273
Wardle DA, Bardgett RD, Klironomos JN, Setälä H, van der Putten WH, Wall DH (2004) Ecological linkages between aboveground and belowground biota. Science 304:1629. https://doi.org/10.1126/science.1094875
Whitfeld TJS, Novotny V, Miller SE, Hrcek J, Klimes P, Weiblen GD (2012) Predicting tropical insect herbivore abundance from host plant traits and phylogeny. Ecology 93:S211–S222. https://doi.org/10.1890/11-0503.1
Xiong D, Wei CZ, Wang XG, Lü XT, Fang S, Li YB, Wang XB, Liang WJ, Han XG, Bezemer TM et al (2021) Spatial patterns and ecological drivers of soil nematode β-diversity in natural grasslands vary among vegetation types and trophic position. J Anim Ecol 90:1367–1378. https://doi.org/10.1111/1365-2656.13461
Yeates GW, Bongers T, Degoede RGM, Freckman DW, Georgieva SS (1993) Feeding habits in soil nematode families and genera-an outline for soil ecologists. J Nematol 25:315–331
Zhang K, Lin S, Ji Y, Yang C, Wang X, Yang C, Wang H, Jiang H, Harrison RD, Yu DW (2016) Plant diversity accurately predicts insect diversity in two tropical landscapes. Mol Ecol 25:4407–4419. https://doi.org/10.1111/mec.13770
Acknowledgements
We are grateful to Prof. Keping Ma and the BEF-China consortium for support. We thank Xuefei Yang, Sabine Both, Xiaojuan Liu and Bo Yang for coordinating the platform. We are grateful for the help of Bo Yang and several local assistants for their helps in the sampling. This work was supported by the National Natural Science Foundation of China (U20A2083), the China Postdoctoral Science Foundation (2021T140697), K.C.Wong Education Foundation (GJTD-2019-10) and Major Program of Institute of Applied Ecology, Chinese Academy of Sciences (IAEMP202201). H.B. acknowledges funding from the German Research Foundation (DFG FOR 891/3).
Author information
Authors and Affiliations
Contributions
QL, HB, and MB conceived the ideas and designed the research; QL, XFD, ZWW and XLS collected the data; YBL and XH analyzed the data; YBL, QL, MB and WJL led the writing of the manuscript. All authors contributed critically to the drafts and gave final approval for publication.
Corresponding authors
Ethics declarations
Competing interests
All authors report no conflicts of interest.
Additional information
Responsible Editor: Manuel Delgado-Baquerizo.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Yingbin Li and Xiaofang Du should be considered joint first author.
Supplementary information
Below is the link to the electronic supplementary material.
ESM 1
(DOCX 2.50 MB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, Y., Du, X., Su, X. et al. Local-scale soil nematode diversity in a subtropical forest depends on the phylogenetic and functional diversity of neighbor trees. Plant Soil 486, 441–454 (2023). https://doi.org/10.1007/s11104-023-05882-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-023-05882-2