Abstract
Aims
The colonization pattern of three grapevine endophytes (families Sphingomonadaceae and Enterobacteriaceae) and their putative metabolic signature in plants were analyzed on Vitis vinifera L. cv. Pinot noir to determine the behavior of endophytic strains inside plants as well as how plants respond to such microsymbionts.
Methods
Strains Enterobacter ludwigii EnVs6, Pantoea vagans PaVv7 and Sphingomonas phyllosphaerae SpVs6, were root inoculated on micropropagated grapevine plantlets and colonization was determined by double labeling of oligonucleotide probes-fluorescence in situ hybridization (DOPE-FISH) coupled with confocal microscopy. After inoculation, the metabolic signature in plants colonized by Enterobacter ludwigii EnVs6 was further studied using UPLC//tandem mass spectrometry analysis.
Results
E. ludwigii EnVs6 and P. vagans PaVv7 colonized the plantlets and were both observed on the root surfaces and as endophytes in the cortex and inside the central cylinder up to xylem vessels, but not in the systemic plant parts. Strain SpVs6 also efficiently colonized the root surface, but not the endorhiza and was therefore not detected as an endophyte. A metabolic signature in plants inoculated with E. ludwigii EnVs6 was depicted, resulting in a significant increase in vanillic acid and a decrease in the concentration of catechin, esculin, arbutin, astringin, pallidol, ampelopsin, D-quadrangularin and isohopeaphenol. Changes in the concentration of epicatechin, procyanidin 1, taxifolin and the sum of quercetin-3-glucoside and quercetin-3-galactoside, in roots and stems were also detected, showing that the effect of colonization of plants is most prominent in the stems.
Conclusions
Colonization patterns in endophytes are divergent according to the strains used. A metabolic signature suggests the activation of pathways involved in plant defense but also modulation of the production of metabolites that are keys for colonization.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bacteria use multiple mechanisms for colonization of plant internal tissues (the endosphere), albeit each of these mechanisms is associated with three fundamental steps: attachment through adhesins (including strain-specific fimbriae, pili, surface polysaccharides and flagella), penetration, with disruption of natural barriers in the host (using mechanisms like lactic acid production, protease and lipase activity and receptor-mediated manipulation of the host cell) and eventually establishment, that supposes a strong interaction with the biotic and abiotic surroundings (Wilson et al. 2002). When bacteria approach the plant in the soil, they are attracted by the root’s exudates before attachment (Bacilio-Jiménez et al. 2003; Huang et al. 2014). Binding to and colonization of aerial plant parts can also occur following dispersal mechanisms such as rain, air flow and biological vectors (Bock et al. 2012). Penetration is usually achieved through secondary root emergence sites (Hallmann et al. 1997) or in wounded zones of the epidermis. Bacteria can also penetrate at sites without any apparent sign of disruption (Huang 1986). Penetration also supposes the activation of a dedicated group of genes used by the microorganism to evade the immune system of the host, while progressing towards the plant’s inside (Iniguez et al. 2005; Malfanova et al. 2013). Establishment is by far the most interesting process of colonization, since a strong interaction with the plant takes place. To establish, bacteria can form aggregation structures (micro-colonies, aggregates or biofilms) that facilitate living in the plant (Coombs and Franco 2003; Germaine et al. 2006). Ultimately, these structures activate defense pathways but also allow bacteria to gradually modify the microenvironment. Moreover, establishment occurs in confined environments (tissues or tissue-derived structures) that increase the probability of physical contact and molecular interactions with the plant (Bogino et al. 2013). Among these interactions, the stimulation by transcription activator-like effectors (TALEs) and microbe-associated molecular patterns (MAMPs) is linked to the suppression of host’s immune response through the activation of alternative metabolic pathways (Bittel and Robatzek 2007; Erbs and Newman 2012; Ji et al. 2014; Munoz Bodnar et al. 2013). The final outcome of this stimulation is the modulation of gene expression and the concomitant changes in metabolite production (Schmidt et al. 2014).
Colonization mechanisms have been described in groups of symbiotic and free-living bacteria including rhizobia (Jones et al. 2007), agrobacteria (Winans 1992) and Frankia (Capoen et al. 2009; Perrine-Walker et al. 2010). In these three scenarios, the importance of metabolic crosstalk has proven to be crucial for symbiosis (Kozyrovska 2013) and for understanding how colonization occurs, how bacteria adapt and how the plant responds to the symbiosis (Ferluga and Venturi 2009; Gurich and Gonzalez 2009).
In the case of other endophytic bacteria colonizing inner plant tissues and that can be either commensal or beneficial (Reinhold-Hurek and Hurek 2011; Schulz and Boyle 2006), colonization requires entry into the plant through either passive or active mechanisms and results in the assembly of complex microbial communities (Hardoim et al. 2008) that can be indispensable for the host (Calvaruso et al. 2006; Turner et al. 2013). The result of this association is key to the plant’s well-being and it has been postulated that individuals depleted from their endophytic microbiota might be less resilient to stress and more prone to pathogen infection than those colonized by beneficial bacteria (Gilbert et al. 2010; Partida-Martinez and Heil 2011).
Some studies show changes on the transcriptomic profiles in the bacterial partner during endophytic colonization (Hauberg-Lotte et al. 2012; Shidore et al. 2012) but only few works have focused on metabolic plant responses. For example colonization by diazotrophic bacteria and formation of nitrogen fixing structures show variation in transcriptomic profiles (Boscari et al. 2013) hinting at a possible role of the metabolic constituents of the plant as drivers of endophytic symbiosis. Moreover, most of the information that comes from transcriptomic analysis makes difficult defining actual candidate molecules driving the symbiosis. This points makes the metabolic endophyte-plant interplay an outstanding phenomenon that may be crucial for colonization and establishment.
Experimental evidence suggests that plant colonization by particular bacterial endophytes is marked by a change in the expression of key genes of plant central metabolic pathways (Bordiec et al. 2011). This is supported by observations showing that endophytic bacteria of the genus Paenibacillus generate a metabolic signature (a recurrent change in the metabolic profile of plants after inoculation with the same strain in repeated experiments) when artifically inoculated in poplar plants (Scherling et al. 2009). Surprisingly, other endophytic microorganisms (especially fungi) can also influence plant’s metabolism by increasing the content of antioxidant compounds in the host and this seems to be a trademark for some types of endophytic symbiosis (Torres et al. 2012). Thus, the metabolic signature might be a widespread characteristic in bacteria-plant interactions and especially in endophytic colonization.
Metabolic profiling of bacterial colonization is a new and interesting area of research (Allwood et al. 2008), given the advances on metabolite detection techniques including Matrix Assisted Laser Desorption/Ionization-Time of Flight Mass spectrometry (MALDI-ToF MS), Ultraperformance Liquid Chromatography-tandem mass spectrometry (UPLC-MS/MS) and MALDI/MS assisted imaging (Bajad et al. 2006; Ye et al. 2013; Ziegler et al. 2012). Analysis of metabolites with these techniques permits the quick assessment of molecules accumulated during host-symbiont interactions. Such an approach can depict the final outcome of a set of regulatory events in metabolic pathways, including those involved in plant growth and in colonization and adaptation in bacteria. Therefore metabolomics as well as colonization studies can enable to better understand how endophyte establish inside the plants.
To study colonization of plants by bacteria, the introduction of plasmids bearing auto-fluorescent proteins (AFPs) like the green (GFP) or red (dsRED) fluorescent proteins (Bloemberg et al. 2000; Tombolini et al. 1999) have been done. However, transformation of environmental strains with recombinant plasmids can sometimes be challenging, and unknown molecular mechanisms for plasmid compatibility might make customization of plasmids a time-consuming process. Alternatively, fluorescence in-situ hybridization (FISH) or derivates, can be used to track microbes on and inside plants. The advantages of this technique as compared to other types of approaches (such as immuno-staining, and chromosomal banding) include the good spatial resolution along with the possibility of analyzing a wide range of organisms with varying taxonomy (Levsky and Singer 2003). For plant-associated bacteria, the use of FISH or derivate has proved to be a good option to visualize interactions in the rhizosphere and the phyllosphere, as well as for the detection of organisms inside the plant (Compant et al. 2008a, b), which has led to the discovery of host adaptation and co-evolution phenomena (Campisano et al. 2014a).
In this work we follow re-establishment in grapevine plants of three endophytes isolated from Vitis vinifera L. using double labeling of oligonucleotide probes-fluorescence in situ hybridization (DOPE-FISH). We were able to detect bacteria in the plants after artificial inoculation and demonstrate that strains EnVs6 and PaVv7 colonize the plant root as endophytes, while strain SpVs6 was observed on the root surface but not in the endosphere. We also report a shift in the concentration of several metabolites in micropropagated grapevine, following inoculation with endophytic strain EnVs6, suggesting a metabolic signature of bacterial colonization. The pathways involved in this type of association are related the metabolism of phenylpropanoids in the plant, suggesting the activation of plant defense mechanisms during colonization and hinting to a link with symbiosis pathways previously described for endophytic bacteria.
Materials and methods
Bacterial strains and growth conditions
Pantoea vagans strain PaVv7, Enterobacter ludwigii strain EnVs6 and Sphingomonas phyllosphaerae strain SpVs6 were isolated from stems of both Vitis vinifera L. cv. sylvestris (EnVs6 and SpVs6) and V. vinifera L. cv. vinifera (PaVv7) and characterized previously (Campisano et al. 2014b). Bacteria were grown in LB broth at 28 °C with shaking (180 r.p.m.) until stationary phase (for approximately 10 h) and the growth was monitored by measuring the optical density on a Nanodrop 8000 (Thermo Fisher Scientific, U.S.A) at 600 nm (OD600). To link counts of viable cell density with OD600, several dilutions were plated on LB and incubated for 24–48 h until colonies were visible. Colony forming units per milliliter (CFU/ml) were counted. Bacterial suspensions were collected by centrifugation, washed twice and resuspended at the appropriate concentration in 1X phosphate-buffered saline (PBS) pH 7.2 before inoculation.
Artificial inoculation of in vitro plant material
In vitro micropropagated plants of Vitis vinifera L. cv Pinot noir (clone I-SMA 185) were prepared for inoculation as described before (Compant et al. 2005). Briefly, the plants were micropropagated in cylindrical glass tubes on complete Murashige-Skoog (MS) medium pH 5.6 (Duchefa biochemie, The Netherlands) supplemented with 3 % sucrose and 0.6 % microagar (Duchefa biochemie, The Netherlands). This clone was chosen because in our collection it appeared free of bacterial contaminants and other bacterial endophytes, when tested in PCR using primers 799 F/1520R as described elsewhere (Campisano et al. 2014c).
Explants with one node and internode were incubated in a growth chamber for 51 days at 21 °C, 16/8 h (light/dark) photoperiod and a photon irradiance of 50 μm*m−2*s−1. Healthy plantlets with no less than 3 leaves and no signs of microbial contaminations were used for experiments. All the plants’ basal leaves were aseptically pruned to avoid contamination from bacteria in the medium. Then plants were transferred to sterile plastic boxes containing 40 ml of MS agar inoculated with 100 μl of a bacterial cell suspension at a concentration of 3x108 CFU/ml (corresponding to the OD600 value of 0.1). Plants were then kept in the inoculation chamber and incubated for 10 days using the same photoperiod and temperature conditions described above.
Double labeling of oligonucleotide probes-Fluorescence in situ hybridization (DOPE-FISH)
DOPE-FISH was performed on bacterial pure culture alone and on plants inoculated with bacteria. For pure culture of bacteria, strains were cultured in LB medium at 27 °C and 120 r.p.m. on an orbital shaker until they reached exponential growth phases. Cells were harvested by centrifugation at 4500 x g for ten minutes and washed several times with PBS pH 7.2. Following washing, cells were fixed in a 4 % v/v paraformaldehyde (in PBS) at 4 °C overnight and then treated with a lysozyme solution (1 mg/ml) for 10 min at 37 °C. Cells were then rinsed three times with PBS and centrifuged at 4500 x g for 10 min in every washing step. Later cells were dehydrated in increasing concentrations of ethanol solutions (25, 50, 75 and 99 %), and stored at 4 °C until further use. Cells were poured into teflon-coated microscope slides (Immuno-cell, Belgium), air dried and hybridized according to Compant et al (2005) using probes EUB338, EUB338II, EUB338III (EUBmix) labelled with FLUOS and Gam42a labelled with Cy5 (Amann et al. 1990; Daims et al. 1999; Manz et al. 1992; Wallner et al. 1993) with fluorochromes at both 5’ and 3’ ends. NONEUB coupled with Cy5 was used a control of the experiments. Hybridization step was carried out with 20 μl of hybridization buffer (NaCl 0.9 M; Tris–HCl 0.02 M; 0.01 % SDS, 35 % formamide and probes at a concentration of 5 ng/μL) and slides were placed in 50 ml moisture chambers filled with 5 ml hybridization buffer. Hybridization was carried out for 2 h at 46 °C in the dark followed by a post-hybridization step at 48 °C during 30 min using a prewarmed solution (20 mM Tris–HCl pH 8.0; 0.01%SDS; 5 mM EDTA and NaCl corresponding to the formamide concentration used). Samples were then rinsed with distilled water and air dried overnight in the dark.
To observe colonization by endophytic bacteria, plantlets were aseptically dissected into roots, stem and leaves as described by Compant et al. (2005). Samples were then cut in small parts (5 mm), fixed and prepared for DOPE-FISH as described above for bacterial cells. Then the plant material was transferred to teflon-coated microscope slides (Immuno-cell, Belgium) and hybridized with probes EUB Mix and Gam42a as described above. Some samples were sectioned transversally using razor blades. The hybridization and post hybridization were carried out as described above for bacterial cells and five replicate plants were analyzed for each strain under study. Another five replicate plants inoculated with sterile PBS 1X pH 7.2 were prepared as control as well. Finally, five plants per strain were used to test probe specificity with the NONEUB probe. Samples were rinsed with distilled water before being air dried overnight in the dark and analyzed under confocal microscope (Olympus Fluoview FV1000 with multi-line laser FV5-LAMAR-2 HeNe(G) laser FV10-LAHEG230-2). Pictures were taken at 405, 488, 633 nm wavelengths and under normal light and then merged (RGB) using image J software. Pictures were also analyzed using Imaris 8 software (BITPLANE, UK). Z-stacks were then used to generate whole-stack pictures, these pictures were sharpened (removing convolution by built-in microscope software), and the light/contrast balance was adjusted to improve detail visualization as seen when samples were observed in the dark conditions under the microscope.
Metabolic profiling of inoculated grapevine plants
Twenty-four micropropagated plantlets of Vitis vinifera L. cv. Pinot noir clone I-SMA 185 were cultured and used in this experiment. Three replicates of four plantlets each were inoculated with strain E. ludwigii EnVs6 and three replicates consisting of four plants each were inoculated with E. coli strain DH5α (a non-endophytic, non-pathogenic laboratory strain that served as control) and kept in growth chambers under the same incubation conditions adopted for plants used for microscopic observation of tissue colonization.
After 10 days from inoculation, all plantlets were frozen in liquid nitrogen, crushed in a Retsch MM200 tissue lyser (Qiagen, The Netherlands) for 2 min in screw-cap steel capsules containing steel beads, at a frequency of 25 herz. Finally, the four grapevine plantlets that formed a replicate were pooled.
In a second replicate experiment, the identical procedure as described above was followed, but below- and above-ground plant organs were aseptically separated before the freezing step, with the purpose of confirming the distribution of secondary metabolites between plant organs.
The crushed plant material was analyzed according to previously established methods (Vrhovsek et al. 2012). Briefly, 0.1 g of crushed material were extracted in 2 ml eppendorf tubes with 5 ml of a water/methanol/chloroform (1:2:2) mixture. Additionally, 20 μl of internal standards (gentisic and rosmarinic acids 50 mg/l) were added. Samples were mixed by vortexing for 1 min and incubated in an orbital shaker for 15 min at room temperature. Samples were centrifuged at 15,000 × g at 4 °C for 5 min, and the aqueous phase was collected. Extraction from the pellet was repeated time using 600 μl of water/methanol (1:2) and 400 μl of chloroform, by shaking for 15 min. After centrifugation, the two aqueous phases were pooled, dried under a nitrogen stream and resuspended in 500 μl of methanol/water (2:1). Samples were transferred to glass vials and stored at −20 °C before injection.
Ultraperformance liquid chromatography was performed was performed as reported in Vrhovsek et al. (2012) on a Waters Acquity UPLC system (Milford, USA) consisting of a binary pump, an online vacuum degasser, an autosampler, and a column compartment. Separation of the phenolic compounds was achieved on a Waters Acquity HSS T3 column1.8 μm, 100 mm × 2.1 mm (Milford, USA), kept at 40 °C. The mobile phase A was water containing 0.1 % formic acid, the mobile phase B was acetonitrile containing 0.1 % formic acid. The flow was 0.4 ml/min, and the gradient profile was: 0 min, 5 % B; from 0 to 3 min, linear gradient to 20 % B; from 3 to 4.3 min, isocratic 20 % B; from 4.3 to 9 min, linear gradient to 45 % B; from 9 to 11 min, linear gradient to 100 % B; from 11 to 13 min, wash at 100 % B; from 13.01 to 15 min, back to the initial conditions of 5 % B. The injection volume of both the standard solutions and the samples was 2 μl. After each injection, the needle was rinsed with 600 μl of weak wash solution (water/methanol 90:10) and 200 μl of strong wash solution (methanol/water 90:10).
Mass spectrometry detection was performed on a Waters Xevo TQMS (Milford, USA) instrument equipped with an electrospray (ESI) source. Analysis was done in positive and negative mode. Flow injections of each individual metabolite were used to optimize the MRM conditions.
Statistical analysis
Quantitative results of UPLC-MS analysis of metabolites were studies by univariate and multivariate methods.
Principal component analysis (PCA) was performed by PAST 3.05 software (Hammer et al. 2001) on data from experiments one and two independently and on the entire dataset. Raw data were transformed to row percentages before analysis.
To find differences in the concentration of particular metabolites in plantlets treated with strain EnVs6, multiple t-tests were performed by correcting for false discovery rate (FDR) at q = 0.01, using the GraphPad Prism software version 6.00 for windows (GraphPad Software, San Diego California, USA, www.graphpad.com). The two experimental replicates were analysed together.
A two-way ANOVA was performed to determine the effect of treatment in both experiments, using metabolite and treatment as factors, at α = 0.05.
To reveal the effects of plant organ and treatment on the concentration of each metabolite, a generalized linear model (GLM) was implemented, in which the response variable was the concentration of each metabolite and the model tested differences for the treatment, organ and the interaction organ*treatment at α = 0.05. Analysis was performed in MINITAB release 14 for Windows (MINITAB, State College Pennsylvania, USA, www.minitab.com).
Results
Colonization of plants by endophytic bacteria
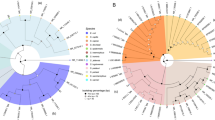
DOPE-FISH was performed using probes that were suitable for the recognition of the strains used. Such probes were selected according to the phylogenetical relationships elucidated by the sequencing of their 16SrDNA gene (Fig. 1). DOPE-FISH microscopy allowed visualizing E. ludwigii EnVs6 using EUBmix, Gam42a or probe combination (Fig. 2a–c). In plantlets, strain EnVs6 was observed on the rhizoplane of the main root (Fig. 2d), in secondary roots (Fig. 2e), as well as at the root tip level (Fig. 2f). Strain EnVs6 was detected as an endophyte in the cortex (Fig. 2g–i), as well as inside the central cylinder up to the xylem vessels (Fig. 2j–l) while no bacterial colonization was recorded in the aerial plant parts (data not shown). Similar experiments resulted in detection of P. vagans PaVv7 in pure cultures (Fig. 3a–c) as well as on the rhizoplane of grapevine plantlets (Fig. 3d–j) and inside plant tissues (Fig. 3k–q). Strain PaVv7 was detected particularly as colonizing the rhizoplane of plantlets at the main root level (Fig. 3d–f), secondary root (Fig. 3g–h) as well as at the root tip level (Fig. 3i–j) and bacteria were visualized as single cells (Fig. 3e–j) or colonizing the whole outline of some rhizodermal cells (Fig. 3d, and f–i). Bacterial aggregates as well as microcolonies were recorded in some plant parts (see Fig. 3f and h–i). Endophytism by strain PaVv7 was observed in the cortex (Fig. 3k–m), as well as inside the central cylinder (Fig. 3n–q) and up to the xylem vessels (Fig. 3p–q). Similarly to strain EnVs6, no endophytic colonization in the aerial plant parts was detected on all the examined plantlets (data not shown).
Phylogenetic relationships of strains EnVs6, PaVv7 and SpVs6. The tree was inferred using the Neighbor-Joining method. The bootstrap consensus tree is replicated 1000 times. The evolutionary distances were computed using the Kimura 2-parameter model; α-proteobacteria and γ-proteobacteria are highlighted in green and orange respectively, according to the fluorescence emission detected under the microscope
visualization of Enterobacter ludwigii EnVs6 in cell cultures (a–c) and in Vitis vinifera L. (d–l) by DOPE-FISH. Cultured bacterial cells of strain EnVs6 tagged with a the EUBmix probes b the Gam42a probe c a cocktail of probes EUBmix and Gam42a. Bacterial cells of strain EnVs6 hybridized with the EUBmix and Gam42a probes colonizing Vitis vinifera L. on d main root e secondary root f secondary root tip g exodermis, cortex and endodermis h cortex with disseminated bacterial cells i cortex with aggregated cells j xylem with few bacterial cells k aggregated cells in xylem vessels l bacterial cells on tracheids
visualization of Pantoea vagans PaVv7 in cell cultures (a–c) and in Vitis vinifera L. (d–l) by DOPE-FISH. Cultured bacterial cells of strain PaVv7 tagged with a Mix EUB probes b Gam42a probe c a mix of probes EUB and Gam42a. Bacterial cells of strain PaVv7 tagged with EUBmix and Gam42a probes colonizing V. Vinifera L. on d–f main root g secondary root emergence site h secondary root showing biofilm structures i root tip j root tip close-up k–m cortex with aggregated cells n endodermis with microcolony o cortex parenchyma and central cylinder p tracheids with dispersed bacterial cells q microcolony on xylem
Strain S. phyllosphaerae SpVs6 was also hybridized and examined as pure culture or during the colonization of grapevine plantlets except that only EUBmix probes were used. Figure 4a shows the pure culture of strain SpVs6. Interestingly, strain SpVs6 was detected as colonizing the rhizoplane of the plantlets at the main root level (Fig. 4b–c), secondary root (Fig. 4d–e) or slightly on root tips (Fig. 4f) but was never detected as an endophyte inside roots as well as inside tissues of shoots.
visualization of Sphingomonas phyllosphaerae SpVs6 in cells cultures a and in Vitis vinifera L. b–f by DOPE-FISH Cultured bacterial cells of strain SpVs6 tagged with EUBmix probes a Bacterial cells of strain SpVs6 tagged with Mix EUB colonizing V. Vinifera L. on b–c main root, d secondary root emergence site, e secondary root showing dispersed bacterial cells f root tip
As expected, in uninoculated control plants, no bacteria were visualized on the root surface (Fig. 5a–c) as well as inside root internal tissues (Fig. 5d). Additional controls with NONEUB probe did not result in visualization of bacteria on samples of grapevine inoculated with EnVs6 (Fig. 6a–b), PaVv7 (Fig. 6c–d), SpVs6 (Fig. 6e–f) or control treatment (Fig. 6g–h).
Probe specificity for DOPE-FISH on bacterial cells of strains EnVs6, PaVv7 or SpVs6 tagged with NONEUB probes colonizing V. vinifera L. a–b strain EnVs6 colonizing the main root of Vitis vinifera c–d strain PaVv7 colonizing main root or e the root tip of Vitis vinifera f strain SpVs6 colonizing main root of Vitis vinifera. Note in all sequences, the absence of bacterial cells and the autofluorescence of cells wall on the main roots of the plant
Metabolome analysis
We screened for the production of 56 secondary metabolites in inoculated grapevines in two independent experiments. Of these compounds, 46.4 % belonged to the flavonoid group while 26.8 % were stilbenes, including several stilbenoids and related compounds. The remaining ca.27 % are organic (hydroxycinnamic) acids (supplementary table 1). Several metabolites were present in concentrations below the detection limit before and after inoculation with strain EnVs6, including vanillin, vanillic acid and esculin (Fig. 7). Contrastingly, metabolites like caftaric, fertaric and trans-coutaric acids reached higher concentrations.
Multivariate analysis was used to test the outcomes of the two replicate experimental rounds. PCA of the entire dataset indicated that data points representing the samples in the two experiments clustered independently (data not shown). A one-way analysis of similarity (ANOSIM) confirmed the difference between experiments to be statistically significant (p = 0.01). PCA on samples from each of the experiments indicated that treated samples clustered separately from controls (Figs. 8 and 9).
Figure 8 shows that plants treated with EnVs6 and control plants clustered differentially, with points representing samples from the same treatment separated along the main component that explained 79.2 % of the total variance (Fig. 8). Despite the visualized clustering using PCA, ANOSIM did not detect significant differences between the treatment and control group (R = 0.074). The second experiment confirmed the effects of endophytic colonization on grapevine’s secondary metabolites. PCA (using Principal Components 1 and 4, which explain 44.9 and 5.4 % of the total variance, respectively) indicated separation of the treated samples from the controls as shown in Fig. 9.
A two-way ANOVA for both experiments revealed a significant difference between control plants and those inoculated with strain EnVs6 (F(1.560) = 38.87; p < 0.0001). Multiple t-tests outlined a significant difference in the concentration of specific metabolites and suggested selective accumulation or depletion of metabolites following colonization. While vanillic acid accumulated in the EnVs6-inoculated plants (p = 0.00003661525), esculin (p = 0.000007356803), catechin (p = 0.000100992), kampferol (p = 0.000000002935455), arbutin (p = 0.00001696334), astringin (p = 0.00134368), pallidol (p = 0.00130449), ampelopsin D-quadrangularin A (p = 0.000439665) and isohopeaphenol (p = 0.00015217) were significantly less concentrated after the treatment with this bacterial strain.
Differential distribution of metabolites in roots and stems
Post inoculation, several metabolites were differentially distributed in the roots and in the stems as shown in Fig. 10. Roots and stems did show significant differences in the concentration of metabolites between control and treated plants. This effect was more evident in stems. Epicatechin gallate (p = 0.0131), Procyanidin B1 (p = 0.0029), taxifolin (p = 0.0424) and the sum of quercetin-3- glucoside and quercetin-3-galactoside (p = 0.0474) suffered a significant decrease in above ground parts after inoculation with the enterobacterium EnVs6, as compared to the control plants.
Heatmap showing distribution and shift of concentrations of metabolites in roots and stems of V. Vinifera L. Concentration of metabolites are depicted from below detection limit (red) through highest concentration (green). Asterisk denote significant differences in the concentration of control and treated organs, in a Two way ANOVA at α = 0.05. CTRL: Plants inoculated with E. coli DH5α. R: Roots. S: Stems. Replicates are denoted by numbers 1–3
A general linear model (GLM) put in evidence the effect of plant organ and treatment in the shifts of concentration for particular metabolites. Treatment affected differently epicatechin gallate (p = 0.009) accumulation in roots and stems, with major shifts in stems of plants inoculated with EnVs6; for procyanidin B1 (p < 0.005) treatment affected both root and stem separately, showing its differential accumulation in both organs independently; treatment also had an effect on taxifolin (p = 0.024) the sum of quercetin-3-glucoside and quercetin-3-galactoside (p = 0.024) and E-cis-miyabenol (p = 0.043) accumulation in stems.
Analysis also suggested an accumulation of procyanidin B3 depending of plant organ and treatment (although this observation was not supported by a significant p = 0.051). Analysis of the concentrations in the organs suggested that both root and stem differentially accumulate the metabolite after treatment with strain EnVs6. Concentrations of caftaric acid, luteolin-7-O-glucoside and trans-coutaric acid suffered minor changes in concentration after inoculation with EnVs6. As expected, the concentration of metabolites in roots and stems was significantly different, with a higher concentration of metabolites in the stems as shown in Fig. 10.
Discussion
In this investigation, the colonization pattern of three endophytic strains was evaluated. Our results revealed the strategies for root penetration and provided a high resolution view of their tropism for root tissues. Our data also confirm the existence of a metabolic signature associated with the inoculation of grapevine with strain Enterobacter ludwigii EnVs6. Previously, this strain was characterized in terms of plant growth promotion properties, antibiotic resistance, quorum sensing activity, enzyme production and biocontrol against grapevine pathogens, performing very well as compared to a collection of endophytic bacteria (Campisano et al. 2014b). We chose strain EnVs6 for our experiments since analysis of plant protection properties have shown a remarkable performance as a biocontrol agent and as a plant growth promoter. Moreover, genomic studies on this strain have revealed that it possesses several symbiosis determinants including type III secretion systems, adhesins and cell wall degrading enzymes (Lòpez-Fernàndez et al 2015). Here we show that inoculation of grapevine with strain EnVs6 ends up in colonization of the plant endosphere and is associated with important changes in the metabolic profile of the host, adding more evidence to singularities that can be regarded as traits specific for endophytism.
Our observations under the fluorescence microscope showed how the three strains originally isolated from the grapevine endosphere were capable of colonizing the surface of plant roots, although each one with a specific pattern. We showed that two of the endophytes used in our study (EnVs6 and PaVv7) were competent for root endophytism in the conditions tested, while one (strain SpVs6) was able to colonize only root surfaces. Being unable to observe re-colonization of the endosphere in the limited range of conditions tested in this study is not sufficient to classify this organism as a non-competent root endophyte. Additionally, strain SpVs6 was originally isolated from the endosphere of wild grapevine, and previous research has shown that endophytic bacteria might have selectivity for wild and domesticated varieties of the same host species (Elbeltagy et al. 2001). Genome analysis of the root endophytic strains EnVs6 and PaVv7 was used to link phenotypic and genetic traits and showed how these two strains are able to utilize a plethora of mechanisms to efficiently adapt to the host and establish as symbionts (Lòpez-Fernàndez et al. 2015). The question whether strain SpVs6 may be a free living diazotroph remains unresolved, since it efficiently colonized root surfaces and appeared to promote plant growth with good efficiency (Campisano et al. 2014b).
Colonization by endophytes has been studied before and the candidate entry points in the plant have been established (Sturz et al. 2000). While entry to the aerial parts of the plant can happen in the stomata and the hydathodes (Huang 1986), colonization of the inner tissues of roots is usually linked to the emergence sites of secondary roots, root tip, root hairs, and by passing between the cells at other zones (Hardoim et al. 2008). Our observations corroborate the involvement of these entry points in colonization by our strains and further reinforce the notion that these sites are common colonization zones for endophytes. Attachment to secondary root emergence sites and to root-tip by the endophytic bacterium EnVs6 points at the existence of tissue tropism. Higher cell densities at specific locations suggest that the root has a strong influence on bacterial habitat selection. Our observations raise questions about plant selectivity for its endobiota occurring at specific points of the roots. We believe that, as previously suggested (Burdman et al. 1999), bacterial surface-associated molecules and plant receptors play a key role in the attachment to and recognition of these colonization sites. Our observations show that bacteria were consistently present in the mentioned areas confirming that colonization occurs mainly through emergence sites of roots. We observed the root endophytes colonizing also parts of the xylem, which suggests that the endophytes might move from the sites of attachment to internal plant tissues, and eventually spread through the plant organs as shown in Figs. 2g–l and 3k–q. We did not find however such movements in our trials. Our experiments confirm also, the colonization of the xylem vessels as it has been long been known for other endophytic bacteria. James and co-workers (1997) showed for instance how plants of Sorghum bicolor are colonized by the endophytic bacterium Herbaspirillum seropedicae, finding a considerable amount of bacteria in the protoxylem and inside the vessels. Furthermore, Gyaneshwar and co-workers (2001) showed that rice is vastly colonized by the endophytic diazotroph Serratia marcescens strain IRBG500. This bacterium is able to colonize roots and move towards the above-ground parts by efficiently colonizing the xylem vessels. Compant et al. in 2005 showed that the endophytic bacterium Burkholderia phytofirmans PsJN is capable of colonizing xylem vessels, cortical cells and the endodermis in primary roots of Vitis vinifera.
Colonization time was also an important factor that was indirectly corroborated during DOPE-FISH experiments. Previous research has shown that colonization of the grapevine rhizosphere can occur between the first and the third hour post inoculation, and that the bacteria can reach xylem during the first days in in vitro plantlets and spread relatively quickly to aerial plant parts as demonstrated using a systemic colonizer (Compant et al. 2005). Our data are in agreement with these findings since visualization of bacteria in the rhizosphere and inside the endorhiza was possible 10 days post inoculation with bacterial cells colonizing the exodermis, the endodermis and part of the xylem. However we did not observe systemic colonization at that time of experiment. We speculate that either colonization of above-ground plant organs may not occur when these endophytes enter through the roots, or that the time required for such endophytes to move to the upper parts of the plantlets might be longer than the timeframe used in our experiments.
Our questions about colonization were further addressed by assessing the effect of bacteria once they have established inside the plants. We inquired whether inoculation of plants with bacterial endophytes might lead to characteristic changes in the plant. It is well known that the endophyte-plant relationship is surrounded by a number of questions regarding what determines the commensalistic nature of this association. Niche overlapping between pathogens and endophytes makes virulence unpredictable and raises the question of whether it is possible to single out a specific characteristic of endophytism. Only a small set of investigations have pursued answering this issue and surprisingly, information on bacterial endophytes has been lagging behind that of their fungal counterparts. Traits activated during fungal colonization of plants have been extensively investigated and they are currently considered a fingerprint for the endophyte-plant association (Torres et al. 2012).
The symbiosis between fungal endophytes and plants relies on the production of fungal reactive oxygen species (ROS). Hyphal tip growth and interkingdom crosstalk are predominantly driven by ROS and ultimately affect how the plant keeps the endophyte in a symbiotic, non pathogenic state (Kogel et al. 2006; Tanaka et al. 2006). During the interaction with endophytic fungi, the content of antioxidant substances (mostly phenolic compounds) in the plant can be altered as shown previously (Malinowski et al. 1998), because plants respond to the ROS produced by the endophytes but also because the plant itself initiates an oxidative burst on the colonizing microorganism (White and Torres 2010). We searched for such a distinct pattern of metabolite shifting in grapevine during colonization by endophytic bacteria under the assumption that such patterns may arise following bacterial colonization as they do following colonization by fungal endophytes. We found that in an analogous fashion, the metabolic profile of plants inoculated with an endophytic bacterium changes upon colonization. In our experiments, after inoculation of micropropagated plantlets, UPLC-MS profiling showed a plausible metabolic signature in which the concentration of a specific set of phenolic compounds changes after endophytic colonization of plants. Interestingly, some of the metabolites whose concentration shifts during the interaction are phenylpropanoids, whose antioxidant activity has been demonstrated previously.
As suspected, changes in the concentration and distribution of phenylpropanoids in inoculated grapevine tend to be divergent, with increase or decrease of different metabolites. This somehow resembles previous observations where a decrease in the content of several aminoacid and molecules related to the central metabolism of the plant and an increase of a small subset of metabolites were recorded post-inoculation with an endophytic strain of Paenibacillus sp. producing a signature on the primary metabolism of poplar plants (Scherling et al. 2009).
Other symbiotic associations in non-endophytic organisms are also characterized by a strong chemical cross-talk between the host and the symbiont. For example, the Sinorhizobium-alfalfa symbiosis is characterized by the increase in the content of dicarboxylic acids and aminoacids like proline and 4-aminobutyrate whose accumulation befalls the nodules (Barsch et al. 2006) together with a strong shift in the concentration of molecules including flavonoids that act as messengers between the host and the bacterium (Jones et al. 2007). The outcome of this cross-talk (i.e., an effective symbiosis) is achieved by a finely-tuned regulation in the gene expression of both partners (Long 1996) in part due to the metabolic cross-talk.
Our findings suggest that the association between endophytic bacteria and grapevine is also characterized by a strong metabolic cross-talk where the plant responds to bacterial colonization by shifting the concentration of specific metabolites. We highlight the importance of the metabolites whose concentration change due to bacterial inoculation. The fact that the increase or depletion of several metabolites, did not affect the colonization by bacterial endophytes, suggests that this shifts in the metabolic profile of the plant favor the colonization by these microorganisms. Further investigations should be done in which mutant plants in key genes for some of the pathways involved in the metabolic signature, interact with endophytes to deeply evaluate the meaning of this phenomenon in the symbiosis process.
In our experimental setting, we observed mainly an increase of vanillic acid while the compounds esculin, catechin, kampferol, arbutin, astringin, pallidol, ampelopsin and isohopeaphenol decreased in concentration after endophytic inoculation. This evidence reinforces the existence of a trademark metabolic signature that accompanies the endophytic colonization of plants. We hypothesize that the metabolic signature found might be related to the overproduction or depletion of metabolites that play key roles in the colonization process, as previous evidence suggests for non-endophytic associations. In agrobacteria-plant symbiosis, microorganisms are able to recognize signals from the host which include a number of chemical entities like vanillin, guaiacol, sinapinic acid and several other phenolic compounds (Winans 1992). In the colonization process, agrobacteria are able to manipulate the plant cells to such an extent that growth of agrobacterial populations (and no other population of co-inoculating microorganism) will be supported by the bacterially induced synthesis in plants cells of secondary amines known as opines (McCullen and Binns 2006). Finally, in actinorhizal symbiosis, a third major archetype for colonization, Frankia species and dicotyledonous plants are subject to an intense chemical exchange (Capoen et al. 2009; Perrine-Walker et al. 2010). Moreover, the bacterium is capable of producing auxins and to sense isoflavonols to achieve colonization of the plant (Hocher et al. 2011).
Phenolic compounds are hitherto the most important class of antioxidant molecules present in plants (Duthie and Crozier 2000). Pathogen attack is characterized by the accumulation of pterocarpans, isoflavans, prenylated isoflavonoids, stilbenes, coumarins, furanocoumarins, 3-deoxyanthicyanidins, flavonols and aurones (Dixon and Paiva 1995). How the concentration of these classes of compounds may shift in other contexts (such as colonization by endophytes) is still poorly understood, especially in the case of bacterial endosymbionts.
In this study we detected changes in the concentration of secondary metabolites of the phenylpropanoid group during endophytic colonization of grapevine. For these molecules, fundamental evidence show diverse roles in plant protection against pathogens but to our knowledge, no information is available regarding their role in endosymbiosis.
Most of these compounds function as phytoalexins, substances involved in protection and antibiosis against plant pathogens. It is therefore plausible that these molecules are related to the non-self recognition of colonizing organisms in different symbiotic scenarios. For example, our findings show that vanillic acid accumulates in the plant after inoculation with the endophytic strain EnVs6. Previous work has demonstrated the role of vanillic acid in the activation of root microbiota. The proposed mechanisms for this substance includes and enhancement of the production of antifungal substances in the bacterial community of the rhizosphere (Jousset et al. 2010), thus we presume that the accumulation of the metabolite during endophytic colonization might be related with an activation mechanism whereby the colonizing endophyte recognizes the molecule and initiates colonization, as is the case for nodule-forming rhizobacteria. This last biological question should be later on addressed through experimental approaches.
On the other hand, some of the metabolites were less concentrated in plants treated with the enterobacterial strain EnVs6. Although the interpretation of this phenomenon might be difficult given the multiple purposes of such molecules in the plant, our results are in agreement with several experimental evidence showing that these metabolites are acting not only as antibiotics but also as symbiosis mediators. In one such case, kampferol has been studied as an anti-nodulation substance given that plants treated with this molecule show reduced numbers of nodules after inoculation with Azospirillum (Zhang et al. 2009). We question whether the decrease in the concentration of this metabolite during the endophytic colonization might be related to an effect on plants that leads to reduction of anti-symbiosis molecules during colonization, as a means of facilitating endo-symbiosis. In a similar way, previous work highlighted the role played by arbutin and esculin (whose concentration was reduced in plants treated with strain EnVs6) as inducers of the syrB gene in Pseudomonas syringae pv syringae (Mo and Gross 1991). The shifts in concentration of these metabolites, is in agreement with previous experiments showing similar trends in poplar. Although we don’t have certainty on whether these substances affect the symbionts directly or have indirect mechanisms, we propose a possible scenario in which endophytic colonization might trigger a plant response that leads to the depletion of particular metabolites, enhancing thus the “balanced antagonism” that has been described before for other endophytic organisms (Schulz et al. 1999) . We suggest that the accumulation of vanillic acid might be associated with an intrinsic response of the plant to colonization of endophytes while the lowering of other metabolites might be involved in the entrance of the bacterium to the plant in a scenario where the host probably holds back its defense mechanisms so the symbiosis can be established.
Other possible roles for these metabolites are the involvement in defense mechanisms against plant pathogens, as it has been shown for catechin, that confers resistance to attack by Pseudomonas syringae pv tomato in Arabidopsis thaliana (Prithiviraj et al. 2007) and astringin, whose accumulation in Picea abies occurs after infection with the insect-transmitted fungus Ceratocystis polonica (Hammerbacher et al. 2011).
A role of the accumulation of ampelopsin D and quadrangularin A in the plant against Plasmopara viticola has been recently established suggesting that increase in its concentration acts as a defense mechanism against infection (Malacarne et al. 2011). Similarly, for pallidol, evidence exists of its role in resistance of grapevine against P. viticola (Pezet et al. 2004). The accumulation of these metabolites in inoculated grapevines could explain the ability of E. ludwigii EnVs6 to control P. viticola infection on grapevine leaf discs (Campisano et al. 2014b).
Our experiments also show a systemic effect of endophytic colonization by strain EnVs6 in micropropagated grapevine. The fact that inoculation with the strain induces changes mostly in the stems of plants suggests the existence of an extensive modulation of plant metabolism affecting not only the bacterial attachment and entry points in the plant but also the upper roots and the above-ground organs. This is an important finding since it implies that the re-introduction of endophytes in plants as a tool for plant protection might be pharmacologically designed to achieve systemic effects.
Finally we claim that our approach to depict the metabolic signature is a novel method for metabolite analysis. It is a multitarget analytical method and includes compounds of different classes in order to be able to analyze a wide range of different matrices. The first screen was done on all compounds included in the method (more than 150). Of those, 56 compounds were reported in the study that correspond to those above the limit of detection in a given matrix. Thus, we are certain that the use of this technology is valuable tool in ecological studies of plant-bacteria interactions.
In conclusion, two of the three endophytes isolated from grapevine can recolonize grapevine plantlets as endophytes. E. ludwigii EnVs6 left a metabolic signature that is characterized by the accumulation of hydroxycinnamic acids and flavonoids and a decrease in phytoalexins. This metabolic signature may reflect the process of colonization and suggests the existence of a possible biological marker associated with endophytism. The mentioned metabolites are expected to counter the penetration of pathogens in grapevine plants when simultaneously accumulated. In our experiments their accumulation is associated to the successful and non-symptomatic colonization of the root endosphere.
References
Allwood JW, Ellis DI, Goodacre R (2008) Metabolomic technologies and their application to the study of plants and plant-host interactions. Physiol Plant 132:117–135. doi:10.1111/j.1399-3054.2007.01001.x
Amann RI, Binder BJ, Olson RJ, Chisholm SW, Devereux R, Stahl DA (1990) Combination of 16S rRNA-targeted oligonucleotide probes with flow cytometry for analyzing mixed microbial populations. Appl Environ Microbiol 56:1919–1925
Bacilio-Jiménez M, Aguilar-Flores S, Ventura-Zapata E, Pérez-Campos E, Bouquelet S, Zenteno E (2003) Chemical characterization of root exudates from rice (Oryza sativa) and their effects on the chemotactic response of endophytic bacteria. Plant Soil 249:271–277. doi:10.1023/A:1022888900465
Bajad SU, Lu W, Kimball EH, Yuan J, Peterson C, Rabinowitz JD (2006) Separation and quantitation of water soluble cellular metabolites by hydrophilic interaction chromatography-tandem mass spectrometry. J Chromatogr A 1125:76–88. doi:10.1016/j.chroma.2006.05.019
Barsch A, Tellstrom V, Patschkowski T, Kuster H, Niehaus K (2006) Metabolite profiles of nodulated alfalfa plants indicate that distinct stages of nodule organogenesis are accompanied by global physiological adaptations. Mol Plant Microbe Interact 19:998–1013. doi:10.1094/mpmi-19-0998
Bittel P, Robatzek S (2007) Microbe-associated molecular patterns (MAMPs) probe plant immunity. Curr Opin Plant Biol 10:335–341. doi:10.1016/j.pbi.2007.04.021
Bloemberg GV, Wijfjes AH, Lamers GE, Stuurman N, Lugtenberg BJ (2000) Simultaneous imaging of Pseudomonas fluorescens WCS365 populations expressing three different autofluorescent proteins in the rhizosphere: new perspectives for studying microbial communities. Mol Plant Microbe Interact 13:1170–1176. doi:10.1094/mpmi.2000.13.11.1170
Bock CH, Cook AZ, Parker PE, Gottwald TR, Graham JH (2012) Short-distance dispersal of splashed bacteria of Xanthomonas citri subsp. citri from canker-infected grapefruit tree canopies in turbulent wind. Plant Pathol 61:829–836. doi:10.1111/j.1365-3059.2011.02588.x
Bogino PC, Oliva Mde L, Sorroche FG, Giordano W (2013) The role of bacterial biofilms and surface components in plant-bacterial associations. Int J Mol Sci 14:15838–15859. doi:10.3390/ijms140815838
Bordiec S, Paquis S, Lacroix H, Dhondt S, Ait Barka E, Kauffmann S, Jeandet P, Mazeyrat-Gourbeyre F, Clement C, Baillieul F, Dorey S (2011) Comparative analysis of defence responses induced by the endophytic plant growth-promoting rhizobacterium Burkholderia phytofirmans strain PsJN and the non-host bacterium Pseudomonas syringae pv. pisi in grapevine cell suspensions. J Exp Bot 62:595–603. doi:10.1093/jxb/erq291
Boscari A, Del Giudice J, Ferrarini A, Venturini L, Zaffini AL, Delledonne M, Puppo A (2013) Expression dynamics of the Medicago truncatula transcriptome during the symbiotic interaction with Sinorhizobium meliloti: which role for nitric oxide? Plant Physiol 161:425–439. doi:10.1104/pp. 112.208538
Burdman S, Jurkevitch E, Schwartsburd B, Okon Y (1999) Involvement of outer-membrane proteins in the aggregation of Azospirillum brasilense. Microbiology 145(Pt 5):1145–1152
Calvaruso C, Turpault M-P, Frey-Klett P (2006) Root-associated bacteria contribute to mineral weathering and to mineral nutrition in trees: a budgeting analysis. Appl Environ Microb 72:1258–1266
Campisano A, Ometto L, Compant S, Pancher M, Antonielli L, Yousaf S, Varotto C, Anfora G, Pertot I, Sessitsch A, Rota-Stabelli O (2014a) Interkingdom transfer of the acne-causing agent, Propionibacterium acnes, from human to grapevine. Mol Biol Evol 31:1059–1065. doi:10.1093/molbev/msu075
Campisano A, Pancher M, Puopolo G, Puddu A, Lòpez-Fernàndez S, Biagini B, Yousaf S, Pertot I (2014b) Diversity in Endophytic Populations Reveals Functional and Taxonomic Diversity between Wild and Domesticated Grapevines. Am J Enol Viticult. doi:10.5344/ajev.2014.14046
Campisano A, Antonielli L, Pancher M, Yousaf S, Pindo M, Pertot I (2014c) Bacterial Endophytic Communities in the Grapevine Depend on Pest Management. PLoS One 9:e112763. doi:10.1371/journal.pone.0112763
Capoen W, Den Herder J, Sun J, Verplancke C, De Keyser A, De Rycke R, Goormachtig S, Oldroyd G, Holsters M (2009) Calcium Spiking Patterns and the Role of the Calcium/Calmodulin-Dependent Kinase CCaMK in Lateral Root Base Nodulation of Sesbania rostrata. Plant Cell 21:1526–1540. doi:10.1105/tpc.109.066233
Compant S, Reiter B, Sessitsch A, Nowak J, Clement C, Ait Barka E (2005) Endophytic colonization of Vitis vinifera L. by plant growth-promoting bacterium Burkholderia sp. strain PsJN. Appl Environ Microbiol 71:1685–1693. doi:10.1128/aem.71.4.1685-1693.2005
Compant S, Kaplan H, Sessitsch A, Nowak J, Ait Barka E, Clement C (2008a) Endophytic colonization of Vitis vinifera L. by Burkholderia phytofirmans strain PsJN: from the rhizosphere to inflorescence tissues. FEMS Microbiol Ecol 63:84–93. doi:10.1111/j.1574-6941.2007.00410.x
Compant S, Nowak J, Coenye T, Clement C, Ait Barka E (2008b) Diversity and occurrence of Burkholderia spp. in the natural environment. FEMS Microbiol Rev 32:607–626. doi:10.1111/j.1574-6976.2008.00113.x
Coombs JT, Franco CMM (2003) Visualization of an Endophytic Streptomyces Species in Wheat Seed. Appl Environ Microbiol 69:4260–4262. doi:10.1128/AEM.69.7.4260-4262.2003
Daims H, Bruhl A, Amann R, Schleifer KH, Wagner M (1999) The domain-specific probe EUB338 is insufficient for the detection of all Bacteria: development and evaluation of a more comprehensive probe set. Syst Appl Microbiol 22:434–444. doi:10.1016/s0723-2020(99)80053-8
Dixon RA, Paiva NL (1995) Stress-induced phenylpropanoid metabolism. Plant Cell 7:1085–1097
Duthie G, Crozier A (2000) Plant-derived phenolic antioxidants. Curr Opin Clin Nutr 3:447–451
Elbeltagy A, Nishioka K, Sato T, Suzuki H, Ye B, Hamada T, Isawa T, Mitsui H, Minamisawa K (2001) Endophytic colonization and in planta nitrogen fixation by a Herbaspirillum sp. isolated from wild rice species. Appl Environ Microbiol 67:5285–5293. doi:10.1128/aem.67.11.5285-5293.2001
Erbs G, Newman MA (2012) The role of lipopolysaccharide and peptidoglycan, two glycosylated bacterial microbe-associated molecular patterns (MAMPs), in plant innate immunity. Mol Plant Pathol 13:95–104. doi:10.1111/j.1364-3703.2011.00730.x
Ferluga S, Venturi V (2009) OryR is a LuxR-family protein involved in interkingdom signaling between pathogenic Xanthomonas oryzae pv. oryzae and rice. J Bacteriol 191:890–897. doi:10.1128/jb.01507-08
Germaine KJ, Liu X, Cabellos GG, Hogan JP, Ryan D, Dowling DN (2006) Bacterial endophyte-enhanced phytoremediation of the organochlorine herbicide 2,4-dichlorophenoxyacetic acid. FEMS Microbiol Ecol 57:302–310. doi:10.1111/j.1574-6941.2006.00121.x
Gilbert SF, McDonald E, Boyle N, Buttino N, Gyi L, Mai M, Prakash N, Robinson J (2010) Symbiosis as a source of selectable epigenetic variation: taking the heat for the big guy. Philos T R Soc B 365:671–678
Gurich N, Gonzalez JE (2009) Role of quorum sensing in Sinorhizobium meliloti-Alfalfa symbiosis. J Bacteriol 191:4372–4382. doi:10.1128/jb.00376-09
Gyaneshwar P, James EK, Mathan N, Reddy PM, Reinhold-Hurek B, Ladha JK (2001) Endophytic colonization of rice by a diazotrophic strain of Serratia marcescens. J Bacteriol 183:2634–2645
Hallmann J, Quadt-Hallmann A, Mahaffee WF, Kloepper JW (1997) Bacterial endophytes in agricultural crops. Can J Microbiol 43:895–914. doi:10.1139/m97-131
Hammer Ø, Harper D, Ryan P (2001) Past: Paleontological Statistics Software Package for education and data analysis. Paleontología Electrónica 4:1–9, http://palaeo-electronicaorg/2001_1/past/issue1_01 html
Hammerbacher A, Ralph SG, Bohlmann J, Fenning TM, Gershenzon J, Schmidt A (2011) Biosynthesis of the major tetrahydroxystilbenes in spruce, astringin and isorhapontin, proceeds via resveratrol and is enhanced by fungal infection. Plant Physiol 157:876–890. doi:10.1104/pp. 111.181420
Hardoim PR, van Overbeek LS, Elsas JD (2008) Properties of bacterial endophytes and their proposed role in plant growth. Trends Microbiol 16:463–471. doi:10.1016/j.tim.2008.07.008
Hauberg-Lotte L, Klingenberg H, Scharf C, Bohm M, Plessl J, Friedrich F, Volker U, Becker A, Reinhold-Hurek B (2012) Environmental factors affecting the expression of pilAB as well as the proteome and transcriptome of the grass endophyte Azoarcus sp. strain BH72. PLoS One 7:e30421. doi:10.1371/journal.pone.0030421
Hocher V, Alloisio N, Bogusz D, Normand P (2011) Early signaling in actinorhizal symbioses. Plant Signal Behav 6:1377–1379
Huang J-S (1986) Ultrastructure of bacterial penetration in plants. Annu Rev Phytopathol 24:141–157
Huang X-F, Chaparro JM, Reardon KF, Zhang R, Shen Q, Vivanco JM (2014) Rhizosphere interactions: root exudates, microbes, and microbial communities. Botany 92:267–275. doi:10.1139/cjb-2013-0225
Iniguez AL, Dong Y, Carter HD, Ahmer BMM, Stone JM, Triplett EW (2005) Regulation of enteric endophytic bacterial colonization by plant defenses. Mol Plant Microbe Interact 18:169–178. doi:10.1094/MPMI-18-0169
James E, Olivares F, Baldani J, Döbereiner J (1997) Herbaspirillum, an endophytic diazotroph colonizing vascular tissue 3Sorghum bicolor L. Moench J Exp Bot 48:785–798
Ji Z-Y, Xiong L, Zou L-F, Li Y-R, Ma W-X, Liu L, Zakria M, Ji G-H, Chen G-Y (2014) AvrXa7-Xa7 Mediated Defense in Rice Can Be Suppressed by Transcriptional Activator-Like Effectors TAL6 and TAL11a from Xanthomonas oryzae pv. oryzicola. Mol Plant-Microbe Interact 27:983–995. doi:10.1094/mpmi-09-13-0279-r
Jones KM, Kobayashi H, Davies BW, Taga ME, Walker GC (2007) How rhizobial symbionts invade plants: the Sinorhizobium-Medicago model. Nat Rev Microbiol 5:619–633. doi:10.1038/nrmicro1705
Jousset A, Rochat L, Lanoue A, Bonkowski M, Keel C, Scheu S (2010) Plants Respond to Pathogen Infection by Enhancing the Antifungal Gene Expression of Root-Associated Bacteria. Mol Plant Microbe In 24:352–358. doi:10.1094/MPMI-09-10-0208
Kogel KH, Franken P, Huckelhoven R (2006) Endophyte or parasite--what decides? Curr Opin Plant Biol 9:358–363. doi:10.1016/j.pbi.2006.05.001
Kozyrovska NO (2013) Crosstalk between endophytes and a plant host within information processing networks. Biopolymer Cell 29:234–243
Levsky JM, Singer RH (2003) Fluorescence in situ hybridization: past, present and future. J Cell Sci 116:2833–2838. doi:10.1242/jcs.00633
Long SR (1996) Rhizobium symbiosis: nod factors in perspective. Plant Cell 8:1885–1898. doi:10.1105/tpc.8.10.1885
Lòpez-Fernàndez S, Sonego P, Moretto M, Pancher M, Engelen K, Pertot I, Campisano A (2015) Whole-genome comparative analysis of virulence genes unveils similarities and differences between endophytes and other symbiotic bacteria. Fron Microbiol 6. doi: 10.3389/fmicb.2015.00419
Malacarne G, Vrhovsek U, Zulini L, Cestaro A, Stefanini M, Mattivi F, Delledonne M, Velasco R, Moser C (2011) Resistance to Plasmopara viticola in a grapevine segregating population is associated with stilbenoid accumulation and with specific host transcriptional responses. BMC Plant Biol 11:114. doi:10.1186/1471-2229-11-114
Malfanova N, Lugtenberg BJJ, Berg G (2013) Bacterial Endophytes: Who and Where, and What are they doing there? In: de Bruijn F (ed) Molecular Microbial Ecology of the Rhizosphere. John Wiley & Sons, Inc. Ch 36
Malinowski D, Alloush G, Belesky D (1998) Evidence for chemical changes on the root surface of tall fescue in response to infection with the fungal endophyte Neotyphodium coenophialum. Plant Soil 205:1–12. doi:10.1023/A:1004331932018
Manz W, Amann R, Ludwig W, Wagner M, Schleifer K-H (1992) Phylogenetic oligodeoxynucleotide probes for the major subclasses of proteobacteria: problems and solutions. Syst Appl Microbiol 15:593–600
McCullen CA, Binns AN (2006) Agrobacterium tumefaciens and plant cell interactions and activities required for interkingdom macromolecular transfer. Annu Rev Cell Dev Biol 22:101–127
Mo Y-Y, Gross DC (1991) Plant signal molecules activate the syrB gene, which is required for syringomycin production by Pseudomonas syringae pv. syringae. J Bacteriol 173:5784–5792
Munoz Bodnar A, Bernal A, Szurek B, Lopez CE (2013) Tell me a tale of TALEs. Mol Biotechnol 53:228–235. doi:10.1007/s12033-012-9619-3
Partida-Martinez LP, Heil M (2011) The microbe-free plant: fact or artifact? Front Plant Sci 2:100. doi:10.3389/fpls.2011.00100
Perrine-Walker F, Doumas P, Lucas M, Vaissayre V, Beauchemin NJ, Band LR, Chopard J, Crabos A, Conejero G, Peret B, King JR, Verdeil JL, Hocher V, Franche C, Bennett MJ, Tisa LS, Laplaze L (2010) Auxin carriers localization drives auxin accumulation in plant cells infected by Frankia in Casuarina glauca actinorhizal nodules. Plant Physiol 154:1372–1380. doi:10.1104/pp. 110.163394
Pezet R, Gindro K, Viret O, Richter H (2004) Effects of resveratrol, viniferins and pterostilbene on Plasmopara viticola zoospore mobility and disease development. Vitis 43:145–148
Prithiviraj B, Perry LG, Badri DV, Vivanco JM (2007) Chemical facilitation and induced pathogen resistance mediated by a root-secreted phytotoxin. New Phytol 173:852–860. doi:10.1111/j.1469-8137.2006.01964.x
Reinhold-Hurek B, Hurek T (2011) Living inside plants: bacterial endophytes. Curr Opin Plant Biol 14:435–443. doi:10.1016/j.pbi.2011.04.004
Scherling C, Ulrich K, Ewald D, Weckwerth W (2009) A metabolic signature of the beneficial interaction of the endophyte Paenibacillus sp. isolate and in vitro-grown poplar plants revealed by metabolomics. Mol Plant Microbe Interact 22:1032–1037. doi:10.1094/mpmi-22-8-1032
Schmidt R, Köberl M, Mostafa A, Ramadan EM, Monschein M, Jensen KB, Bauer R, Berg G (2014) Effects of bacterial inoculants on the indigenous microbiome and secondary metabolites of chamomile plants. Front Microbiol. doi:10.3389/fmicb.2014.00064
Schulz B, Boyle C (2006) What are endophytes? In: Microbial Root Endophytes. Springer, pp 1–13
Schulz B, Römmert A-K, Dammann U, Aust H-J, Strack D (1999) The endophyte-host interaction: a balanced antagonism? Mycol Res 103:1275–1283
Shidore T, Dinse T, Ohrlein J, Becker A, Reinhold-Hurek B (2012) Transcriptomic analysis of responses to exudates reveal genes required for rhizosphere competence of the endophyte Azoarcus sp. strain BH72. Environ Microbiol 14:2775–2787. doi:10.1111/j.1462-2920.2012.02777.x
Sturz AV, Christie BR, Nowak J (2000) Bacterial endophytes: potential role in developing sustainable systems of crop production. CRC CR Rev Plant Sci 19:1–30. doi:10.1080/07352680091139169
Tanaka A, Christensen MJ, Takemoto D, Park P, Scott B (2006) Reactive oxygen species play a role in regulating a fungus-perennial ryegrass mutualistic interaction. Plant Cell 18:1052–1066. doi:10.1105/tpc.105.039263
Tombolini R, van der Gaag DJ, Gerhardson B, Jansson JK (1999) Colonization pattern of the biocontrol strain Pseudomonas chlororaphis MA 342 on barley seeds visualized by using green fluorescent protein. Appl Environ Microbiol 65:3674–3680
Torres MS, White JF Jr, Zhang X, Hinton DM, Bacon CW (2012) Endophyte-mediated adjustments in host morphology and physiology and effects on host fitness traits in grasses. Fungal Ecol 5:322–330. doi:10.1016/j.funeco.2011.05.006
Turner TR, James EK, Poole PS (2013) The plant microbiome. Genome Biol 14:209. doi:10.1186/gb-2013-14-6-209
Vrhovsek U, Masuero D, Gasperotti M, Franceschi P, Caputi L, Viola R, Mattivi F (2012) A versatile targeted metabolomics method for the rapid quantification of multiple classes of phenolics in fruits and beverages. J Agric Food Chem 60:8831–8840. doi:10.1021/jf2051569
Wallner G, Amann R, Beisker W (1993) Optimizing fluorescent in situ hybridization with rRNA-targeted oligonucleotide probes for flow cytometric identification of microorganisms. Cytometry 14:136–143. doi:10.1002/cyto.990140205
White JF Jr, Torres MS (2010) Is plant endophyte-mediated defensive mutualism the result of oxidative stress protection? Physiol Plant 138:440–446. doi:10.1111/j.1399-3054.2009.01332.x
Wilson M, Rod McNab, Henderson. B (2002) Bacterial invasion as a virulence mechanism. In: Bacterial Disease Mechanisms. Cambridge University Press. pp 405-465
Winans SC (1992) Two-way chemical signaling in Agrobacterium-plant interactions. Microbiol Rev 56:12–31
Ye H, Gemperline E, Venkateshwaran M, Chen R, Delaux PM, Howes‐Podoll M, Ané JM, Li L (2013) MALDI mass spectrometry‐assisted molecular imaging of metabolites during nitrogen fixation in the Medicago truncatula–Sinorhizobium meliloti symbiosis. Plant J 75:130–145
Zhang J, Subramanian S, Stacey G, Yu O (2009) Flavones and flavonols play distinct critical roles during nodulation of Medicago truncatula by Sinorhizobium meliloti. Plant J 57:171–183. doi:10.1111/j.1365-313X.2008.03676.x
Ziegler D, Mariotti A, Pflüger V, Saad M, Vogel G, Tonolla M, Perret X (2012) In Situ Identification of Plant-Invasive Bacteria with MALDI-TOF Mass Spectrometry. PLoS One 7:e37189. doi:10.1371/journal.pone.0037189
Acknowledgments
We would like to thank Domenico Masuero from FEM for support in the processing of samples for the metabolite analysis in UPLC/MS.
Funding
This work was funded by Provincia Autonoma di Trento, progetto PAT - Call 2 Team 2009 - Incoming-Mecagrafic. We thank the European Cooperation in Science and Technology (COST) Action FA1103 for providing network activities and funding.
Conflict of interest
We state the following conflict of interest: manuscript authors A. Campisano and S. Compant serve as guest editors for this special issue
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible Editor: Kari Saikkonen.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 355 kb)
Rights and permissions
About this article
Cite this article
Lòpez-Fernàndez, S., Compant, S., Vrhovsek, U. et al. Grapevine colonization by endophytic bacteria shifts secondary metabolism and suggests activation of defense pathways. Plant Soil 405, 155–175 (2016). https://doi.org/10.1007/s11104-015-2631-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-015-2631-1