Abstract
Fungal communities in soybean rhizosphere from reproductive growth stages R1 (beginning bloom) to R8 (full maturity) were studied based on the polymerase chain reaction (PCR)-denaturing gradient gel electrophoresis (DGGE) banding patterns of partial rDNA internal transcribed spacer regions (ITS1) and sequencing methods. Pot experiment subjecting three soybean genotypes grown in two soils (Mollisol and Alfisol) indicated that the soil type was the major factor in shaping the fungal communities in the soybean rhizosphere. Field experiment was conducted in an Alfisol field with three soybean genotypes, and both pot and field experiments showed that rhizosphere fungal communities shifted with growth stages, and more diversity of communities was found in early reproductive growth stages than later stages. No major difference among fungal communities of three soybean genotypes was detected at individual growth stage. BLAST search of ITS sequence data generated from excised DGGE bands showed that fungi belonging to Ascomycetes and Basidiomycetes predominantly inhabited in the soybean rhizosphere. In addition, a few bands had low similarity with database sequences inferred that unknown fungal groups existed in soybean rhizosphere.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The rhizosphere is the soil region influenced by plant roots and characterized with high microbial activities (Hiltner 1904). Rhizosphere microbial communities carry out fundamental processes that contribute to nutrient cycling, healthy root growth, and plant growth promotion (Buchenauer 1998; Atkinson and Watson 2000; Sylvia and Chellemi 2001). Plant roots release a wide variety of compounds into the rhizosphere, which form unique micro-environments for soil microorganisms. It is commonly recognized that the root exudates differ according to plant species, even cultivars, and plant growth stages (Rovira 1959; Nelson 1990; Whipps 2001; Rengel 2002). As different microbes respond differently to the compounds released by roots, different composition of root exudates is believed to explain the plant specific rhizosphere microbal communities (Marschner et al. 2001, 2002; Smalla et al. 2001; Kowalchuk et al. 2002). Soil type is another important factor for the determination of the rhizosphere microbal communities (Buyer et al. 1999; Dalmastri et al. 1999; Kowalchuk et al. 2000), as different soils show different particle size distribution, pH, aeration, and physico-chemical characteristics that can affect microbial communities either directly, e.g., by providing a specific habitat for selecting specific microbes, and indirectly, e.g., by affecting plant root exudation (Garbeva et al. 2004). Therefore, the interactions among soil types, plant species/genotypes and growth stages complicatedly affect microbial communities in the rhizosphere (Marschner et al. 2001). The effect of soil type on the community is greater than that of plant species in some cases (Dalmastri et al. 1999; Buyer et al. 2002; Singh et al. 2007), while in other cases plant species show greater effect on the community composition than the soil type (Grayston et al. 1998; Wieland et al. 2001). Although most of the previous studies focused on the rhizosphere bacterial communities, only a few reports were related to fungal communities in plant rhizosphere (Gomes et al. 2003; Singh et al. 2007).
Soybean (Glycine max (L.) Merrill) is a major crop in Northeast China. The acreage of soybean cultivation and its total yield in this region account for about 33% and 44% of nation's total, respectively (Liu and Herbert 2002). Since the first cultivar was released in 1923, more than 600 cultivars had been released in Northeast China by the end of the last century (Liu et al. 2008). Although soybean breeding and genotype improvement contributed primarily to increase soybean yield in this region, the influence of soybean genotype on microbial communities, especially on fungal communities in the rhizosphere is not clear.
Culture-dependent techniques were traditionally used for assessing fungal diversity. It has become increasingly know that only 5–10% of fungal community members can be detected by culture methods (Hawksworth and Rossman 1997; Hawksworth 2001). Recently, culture-independent techniques, such as analyses of the microbial DNA extracted directly from environments have pushed a remarkable progress in microbial ecology research (Marschner et al. 2001; Ebersberger et al. 2004; Yao et al. 2006). Our previous researches have focused on bacterial communities in soybean rhizosphere (Xu et al., unpublished data). In this work, we analyzed the potential effects of two soil types (Mollisol and Alfisol), and three soybean genotypes (Hefeng 25, Suinong 14, and D2003-1) on fungal community structures during soybean reproductive growth stages by the pot and field experiments, and the community structures were estimated by using denaturing gradient gel electrophoresis (DGGE) for community evaluation and sequencing of DGGE bands for phylogenetic determination of dominant fungal members.
Materials and methods
Pot experiment
A Black soil (Mollisol) and a Dark Brown soil (Alfisol) were collected from farming fields at Hailun Agro-Ecological Experimental Station, Chinese Academy of Sciences (47°26′N, 126°38′E) and Lanling village, Jixi City (45°17′N, 130°42′E), respectively, in March 2005 (Table 1). Both sites were located in Heilongjiang Province, which is the largest soybean cropping province in Northeast China. Fifteen kilograms of soil were transferred into each pot (25.5 cm diameter and 29.0 cm deep), and the moisture content was adjusted to 70% field capacity. Two soybean genotypes of Hefeng 25 (average yield: 2260 kg ha−1) and Suinong 14 (average yield: 2550 kg ha−1) that were released in 1984 and 1996, respectively, and a new line of D2003-1 with the yielding potential of 3000 kg ha−1 were chosen in this study. All three genotypes take about 120–125 days to maturity in Heilongjiang Province, China.
Three genotypes were sown to the Dark Brown soil, while Hefeng 25 and D2003-1 were sown in the Black soil (six seeds per pot). After the seed germination, seedlings were thinned to three plants per pot. All pots were placed in a glasshouse (daytime: 24–28°C, night time 16–20°C). Four replicate pots were prepared for each genotype, soil and sampling time combinations. At the soybean reproductive growth stages of R1 (beginning bloom), R3 (beginning pod), R4 (full pod), R5 (beginning seed) and R8 (full maturity), the shoot was cut off, and the roots were carefully separated from soils by inverting the pots. Only the soil adhering the roots was considered as rhizosphere soil (Nazih et al. 2001), and the rhizosphere soils were collected by shaking off from roots in the air. A portion of composite soil samples was placed into autoclaved microcentrifuge tubes (2 ml) immediately. The tubes were stored at −80°C until use.
Field experiment
A field experiment was conducted on a Dark Brown soil at Lanling village in 2005. Three genotypes of Hefeng 25, Suinong 14 and D2003-1 were sown with the density of 30 plants m−2 on 15 May. Randomized design with three replicates for each genotype was performed, and each plot contains ten rows with the 15 m long and 0.67 m wide. Fertilization was followed to the local recommendation with N 50 kg ha−1, P 45 kg ha−1 and K 60 kg ha−1. At the growth stages of R1, R3, R4, R5 and R8, ten plants were harvested, and the rhizosphere soils were collected and treated in the same way as the pot experiment.
DNA extraction and purification
DNA was extracted from the soil samples (0.5 g wet weight) by bead-beating method based on the method of Zhou et al. (1996) and Watanabe et al. (2004). DNA extracts were dissolved in 100 µl of TE buffer (10 mM Tris-HCl, 1 mM EDTA, pH 8.0) and purified with Sephadex G-200 (Cahyani et al. 2004).
PCR-DGGE
A nested PCR amplification was conducted targeting fungal rDNA internal transcribed spacer (ITS) regions (Bastias et al. 2006). Fungal-specific primers ITS1-F (Gardes and Bruns 1993) and ITS4 (White et al. 1990), GC-ITS1-F and ITS2 (Gardes and Bruns 1993) were used for the first and second round of PCR amplifications, respectively. PCR amplification was performed according to the description by Bastias et al. (2006). DGGE was performed by using 8% (w/v) acrylamide gel with a 20–60% denaturant gradient and run in 1×TAE (Tris-acetate-EDTA) buffer for 16 h under conditions at 60°C and 75 V. After the electrophoresis, the gel was stained in 1:3300 (v/v) GelRed (Biotium, USA) nucleic acid staining solution for 20 min. DGGE profiles were photographed by using Bio-Rad transilluminator (BIO-RAD Laboratories, Segrate, Italy) under UV light.
Analysis of the DGGE profile
Banding patterns of the DGGE profile were analyzed by the Quantity one software (version 4.5). The position and intensity of each band were determined automatically. The density value of each band was divided by the average band density of the lane in order to minimize the influence of loaded DNA concentrations among samples (Garland and Mills 1991; Graham and Haynes 2005). Normalized data were used for principal component analysis as described previously (Matsuyama et al. 2007).
Sequencing of DGGE bands
Several common bands and bands showing variations with soil types, growth stages and genotypes were carefully excised from the DGGE gel and subjected to sequencing. DNA extraction from DGGE bands, verification of the mobility of bands and sequencing were performed as described previously by Cahyani et al. (2004).
Phylogenetic analysis
Closest relatives and phylogenetic affiliations of the obtained sequences were determined by using the BLAST search program at the NCBI web site. All sequences determined in the present study were deposited in the DNA Data Bank of Japan (DDBJ) under accession numbers from AB438048 to AB438071.
Results
Pot experiment
DGGE profiles of fungal communities in the rhizosphere are shown in Fig. 1. In total, 56 bands with different mobility were detected. The average number of DGGE bands across growth stages from R1 to R8 for genotypes Hefeng 25, D2003-1 and Suinong 14 in Dark Brown soil were 35.6 ± 5.4, 37.2 ± 5.9 and 37.0 ± 3.8, respectively, and for genotypes Hefeng 25 and D2003-1 in Black soil were 29.8 ± 1.6 and 31.8 ± 3.3, respectively. Under both soil conditions, the DGGE band number was in the highest level at stage R1 for every genotype, and then it decreased with soybean development to the lowest level at growth stage R8 (Fig. 1).
DGGE profiles of fungal communities in rhizospheres of three soybean genotypes grown in the Black and the Dark Brown soils (pot experiment). R1, R3, R4, R5 and R8 represent soybean stages at beginning bloom, beginning pod, full pod, beginning seed and full maturity, respectively. H, D and S represent soybean genotype of Heifeng 25, D2003-1 and Suinong 14, respectively. Numbers below the profiles indicate the DGGE band abundances of the line, and arrows indicate bands excised from the gel for sequencing
DGGE banding patterns of rhizosphere fungal communities among all three genotypes were very similar to each other at the individual sampling time, but different between the two soil types (Fig. 1). For examples, bands P-SyRF4 and P-SyRF5 were specific to Dark Brown soil, band P-SyRF6 was restricted to Black soil, and more abundances of bands P-SyRF3 in Black soil, and P-SyRF7 in Dark Brown soil than those in counterpart soil.
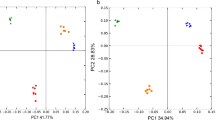
Excluding band P-SyRF19 (ITS region of soybean), principal component analysis clearly separated fungal communities in soybean rhizospheres between Black soil and Dark Brown soil, irrespective of genotypes (Fig. 2). In addition, the fungal communities in soybean early reproductive growth stages were different from those in later stages (Fig. 2), and the succession pattern of fungal communities from growth stages R1 to R8 was similar in Black soil and Dark Brown soil. No significant differences were found in the fungal community among the genotypes at individual growth stage.
Principal component analysis of DGGE profiles of fungal communities in rhizospheres of D2003-1 (∆), Hefeng 25 (○) and Suinong 14 (▫) (pot experiment). Numbers beside the symbols indicate soybean reproductive growth stages. Solid and open symbols represent the soybean grown in the Black and the Dark Brown soils, respectively
Field experiment
As shown in Fig. 3, DGGE profiles of fungal communities in the field experiment were very similar with each other among the genotypes, and the difference was only observed for a few bands or band density. For instance, band F-SyRF1 showed the highest intensity at stage R8, and band F-SyRF6 had relatively high intensity at stages R5 and R8 than preceding growth stages.
DGGE profiles of fungal communities in the rhizospheres of three soybean genotypes (field experiment). R1, R3, R4, R5 and R8 represent soybean stages at beginning bloom, beginning pod, full pod, beginning seed and full maturity, respectively. H, D and S represent soybean genotypes of Hefeng 25, D2003-1 and Suinong 14, respectively. Numbers below the profiles indicate the DGGE band abundances of the line, and arrows indicate bands excised from the gel for sequencing
In total, 74 bands with different mobility were observed in the field experiment, and average band numbers across growth stages for Hefeng 25, D2003-1 and Suinong 14 were 49.0 ± 6.5, 48.2 ± 6.2 and 46.4 ± 4.2, respectively. Similar to the pot experiment, the band number tended to be more numerous in the early reproductive growth stages than in the later stages (Fig. 3).
Excluding band F-SyRF7 (ITS region of soybean), principal component analysis divided the bacterial communities into three groups (A, B and C). Groups A, B and C contained all three soybean genotypes at stages R1 ~ R4, R5 and R8, respectively (Fig. 4). The succession of fungal communities of all three genotypes with growth stages had a similar tendency, and the significant effect of genotypes on fungal community changes was also not detected at individual growth stage.
Sequence analysis of DGGE bands
Nineteen common, dense and soil-specific bands in Fig. 1 were excised for sequencing. BLAST search indicated that all bands have closest relatives with fungal clones or isolates (Table 2), except for band P-SyRF19 come from the ITS region of soybean. Among the 18 fungal related bands, 11 bands affiliated to Ascomycota, five bands to Basidiomycota, one band to Oomycetes, and one band to an uncultured soil fungal clone.
In the field experiment, seven bands were excised and sequenced. Results showed that the nucleotide sequence of band F-SyRF7 was identical to P-SyRF19 in pot experiment, and it was the ITS region of soybean. Other four bands belonged to Ascomycota, and one band was closely related to uncultured soil fungal clone (Table 2). Comparison of nucleotide sequences among the bands elucidated that bands P-SyRF7, P-SyRF15 and P-SyRF17 in the pot experiment were identical to bands F-SyRF2, F-SyRF3 and F-SyRF4 in the field experiment, respectively.
Discussion
Although many primers have been designed for amplification of fungal environmental DNA, the fungal ITS region was found to take more information in taxonomy than other genomic regions (e.g. 18S rDNA) (Bruns et al. 1991). The primers ITS1 and ITS2 were tested suitable for DGGE analysis of fungal communities associated with the forest and the moorland ecosystems (Anderson et al. 2003), and used for a research on forest soils (Bastias et al. 2006). In the present research, we also used this primer set for analysis of soybean fungal community, and clear and sharp DGGE banding patterns were obtained (Figs. 1 and 3). In addition, four out of six bands in field experiment were sequenced with identical nucleotide sequences to those of counterpart bands in pot experiment, suggesting that the primers used in this study were suitable. However, BLAST search indicated that identical sequence of bands P-SyRF19 and F-SyRF7 at the lowest position in Figs. 1 and 3 were 100% similarity with the ITS region of soybean, inferred that those primers are not specific enough for fungal environmental DNA analysis. This finding was not discovered in previous studies, although some bands were sequenced in their researches (Anderson et al. 2003; Bastias et al. 2006).
Many researches on rhizosphere microbial community based on pot or microcosm container experiments (Wieland et al. 2001; Buyer et al. 2002; Marschner et al. 2002, 2004; Singh et al. 2007), only a few studies based on field sampling (Gomes et al. 2001, 2003; Costa et al. 2006). In this study, both pot and field experiments were used to reveal the fungal community in soybean rhizosphere. Although more bands were obtained in DGGE profiles from field experiment than pot experiment of soybean grown in Dark Brown soil, comparison of the two DGGE profiles indicated that the banding patterns of majority bands between the two experiments were similar (Figs. 1 and 3), suggesting that the pot experiment in this study can reflect the fungal community dynamics in field condition. The reasons for more bands in field experiment are not know, one might be the contamination of weeds grown nearby the soybean, in some extent, contributed to increase fungal diversity in soybean rhizosphere.
Principal component analysis in Fig. 2 clearly demonstrated that the fungal communities were separated into Black soil and Dark Brown soil groups, suggesting that the soil type is the major factor to determine the fungal community in the soybean rhizosphere. This finding was consistent with our previous research on bacterial community in soybean rhizosphere (Xu et al., unpublished data), and also consistent with other studies, indicating that the soil type was the most important factor in determining microbial community in the rhizospheres of various plants (Girvan et al. 2003; De Ridder-Duine et al. 2005; Singh et al. 2007). Marschner et al. (2004) reported that many factors contributed to change rhizosphere bacterial community, such as soil pH, nutrition and soil type. As the two soil types have difference soil properties (Table 1), we conclude that the difference of rhizosphere fungal community between two soils is related to soil characteristics.
Relative impact of soil type on microbial communities varies with soil properties such as soil texture and organic matter contents. Given that clayey soils exert a greater influence on microbial communities than sandy soils (Garbeva et al. 2004; Marschner et al. 2004), and the contents of clay is more and sand is less in Black soil than those in Dark Brown soil in this study (Table 1), higher similarity of microbial communities in soybean rhizosphere grown in Black soil than those in Dark Brown soil would be detected. However, this presumption was not observed in this study for analysis of fungal community (Fig. 2), but was detected in bacterial community (Xu et al., unpublished data). Those results inferred that the relative impact strength of soil type on fungal community in soybean rhizosphere was less than on bacterial community. In addition, more DGGE bands in Dark Brown soil than in Black soil suggested that more diversity of fungal communities in Dark Brown soil, although it had relative low organic matter content.
Gomes et al. (2001, 2003) reported that bacterial and fungal abundance in maize rhizosphere increased with growth development, and no relevant differences in TGGE/DGGE banding patterns of both bacterial and fungal communities were observed between two maize cultivars. Similar to their results, in this study, principal component analyses indicated the shifts of soybean rhizosphere fungal communities with growth stages, and the community structures among three soybean genotypes were similar (Figs. 2 and 4), suggesting that the growth stage is the second major factor in shaping fungal communities in the soybean rhizosphere. However, in contrast to the band abundance observed by Gomes et al. (2001, 2003), a decline tendency of DGGE band number from growth stages R1 to R8 was observed in both pot and field experiments (Figs. 1 and 3). This finding suggested that more complex fungal community in soybean early reproductive growth stages than that in later stages. Similar result was also observed in our previous study on bacterial communities (Xu et al., unpublished data). It should be noticed that the primers used in our research were difference with those used by Gomes et al. (2001, 2003), a future examination is necessary to elucidate the diversity of rhizosphere microbial communities between soybean and maize.
The succession of microbial communities with growth stage might be related with two mechanisms. One might be the environmental changes such as temperature and soil moisture with the growth stage (Nazih et al. 2001). However, this mechanism could be minor, because the temperature and water regime were relatively uniform throughout the growth stage in the pot experiment. The other mechanism might be ascribed to the changes in quality and quantity of root exudates/rhizodepositions with the growth stage (Marschner et al. 2002). Although root exudates were not measured in the present study, there are many evidences that root exudates are strongly affected by the growth stage, which in turn can affect rhizosphere microbial communities over time (Yang and Crowley 2000; Duineveld et al. 2001; Garbeva et al. 2004). Thus, the succession of fungal communities in the soybean rhizosphere is concluded due to the change in root exudates or rhizodepositions.
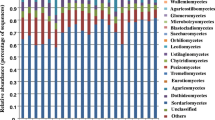
Eighteen DGGE bands were excised from pot experiment and their sequences were positively assigned to fungal ITS regions (Table 2). BLAST search showed that about 61% and 28% of bands belonged to Ascomycete and Basidiomycete, respectively. The fungi belonging to Ascomycete and Basidiomycete dominated in soil environments were also observed by Anderson et al. (2003) in transect ecosystems from moorland to forest and by Bastias et al. (2006) in a prescribed burning sclerophyll forest.
Although the majority of sequenced DGGE bands had high similarity (≥90%) with database sequences, four bands (i.e., P-SyRF1, P-SyRF11, P-SyRF14 and P-SyRF16) had low similarity values (50 ~ 82%) suggesting that some unknown fungal members exist in soybean rhizosphere. Gomes et al. (2003) reported that young maize plants seemed to select the Ascomycetes order Pleosporales, and senescent maize seemed to favor different members of the Ascomycetes and Basidiomycetes living in mazie rhizospheres. However, those phenomena were not detected in soybean rhizosphere, and bands P-SyRF3 and P-SyRF5 belonging to Basidiomycetes were evenly observed from soybean growth stages from R1 to R8 (Fig. 1). Furthermore, the sequences of bands P-SyRF2, P-SyRF6, P-SyRF7 and P-SyRF10 had 100% similarity with the fungal groups usually isolated from soybean rhizosphere, and some species, such as Fusarium oxysporum and Fusarium solani were identified as common pathogens to cause soybean root rot in Heilongjiang Province, China (Xin and Ma 1987).
In conclusion, the present study indicated that fungal communities in the soybean rhizosphere were primarily regulated by soil type, and also changed with growth stage. No significant differences of fungal communities among three genotypes were detected from both experiments. More diversity of fungal community observed in Dark Brown soil than in Black soil, and in early reproductive growth stages than in later stages. Sequence analysis of the DGGE bands revealed that fungal belonging to Ascomycetes and Basidiomycetes dominantly inhabited in the soybean rhizosphere.
References
Anderson IC, Campbell CD, Prosser JI (2003) Diversity of fungi in organic soils under a moorland-Scots pine (Pinus sylvestris L.) gradient. Environ Microbiol 5:1121–1132 doi:10.1046/j.1462-2920.2003.00522.x
Atkinson D, Watson CA (2000) The beneficial rhizosphere: a dynamic entity. Appl Soil Ecol 15:99–104 doi:10.1016/S0929-1393(00)00084-6
Bastias BA, Huang ZQ, Blumfield T, Xu Z, Cairney JWG (2006) Influence of repeated prescribed burning on the soil fungal community in an eastern Australian wet sclerophyll forest. Soil Biol Biochem 38:3492–3501 doi:10.1016/j.soilbio.2006.06.007
Bruns TD, White TJ, Taylor JW (1991) Fungal molecular systematics. Annu Rev Ecol Syst 22:4356–4360 doi:10.1146/annurev.es.22.110191.002521
Buchenauer H (1998) Biological control of soil-borne diseases by rhizobacteria. J Plant Dis Prot 105:329–348
Buyer JS, Roberts DP, Russek-Cohen E (1999) Microbial community structure and function in the spermosphere as affected by soil and seed type. Can J Microbiol 45:138–144 doi:10.1139/cjm-45-2-138
Buyer JS, Roberts DP, Russek-Cohen E (2002) Soil and plant effects on microbial community structure. Can J Microbiol 48:955–964 doi:10.1139/w02-095
Cahyani VR, Matsuya K, Asakawa S, Kimura M (2004) Succession and phylogenetic profile of methanogenic archaeal communities during the composting process of rice straw estimated by PCR-DGGE analysis. Soil Sci Plant Nutr 50:555–563
Costa R, Götz M, Mrotzek N, Lottmann J, Berg G, Smalla K (2006) Effects of site and plant species on rhizosphere community structure as revealed by molecular analysis of microbial guilds. FEMS Microbiol Ecol 56:236–249 doi:10.1111/j.1574-6941.2005.00026.x
Dalmastri C, Chiarini L, Cantale C, Bevivino A, Tabacchioni S (1999) Soil type and maize cultivar affect the genetic diversity of maize root-associated Burkholderia cepacia populations. Microb Ecol 38:273–284 doi:10.1007/s002489900177
De Ridder-Duine AS, Kowalchuk GA, Gunnewiek PJAK, Smant W, van Veen JA, de Boer W (2005) Rhizosphere bacterial community composition in natural stands of Carex arenaria (sand sedge) is determined by bulk soil community composition. Soil Biol Biochem 37:349–357 doi:10.1016/j.soilbio.2004.08.005
Duineveld B, Kowalchuck GA, Keijzer A, van Elsas JD, van Veen JA (2001) Analysis of bacterial communities in the rhizosphere of the chrysanthemum via denaturing gradient gel electrophoresis of PCR-amplified 16S rRNA as well as DNA fragments coding for 16S rRNA. Appl Environ Microbiol 67:172–178 doi:10.1128/AEM.67.1.172-178.2001
Ebersberger D, Wermbter N, Niklaus PA, Kandeler E (2004) Effect of long term CO2 enrichment on microbial community structure in calcareous grass land. Plant Soil 264:313–323 doi:10.1023/B:PLSO.0000047768.89268.8c
Garbeva P, Veen JAV, Elsas JDV (2004) Microbial diversity in soil: selection of microbial populations by plant and soil type and implications for disease suppressiveness. Annu Rev Phytopathol 42:243–270 doi:10.1146/annurev.phyto.42.012604.135455
Gardes M, Bruns TD (1993) ITS primers with enhanced specificity for basidiomycetes: application to the identification of mycorrhiza and rusts. Mol Ecol 2:113–118 doi:10.1111/j.1365-294X.1993.tb00005.x
Garland JL, Mills AL (1991) Classification and characterization of heterotrophic microbial communities on the basis of patterns of community-level sole-carbon-source utilization. Appl Environ Microbiol 57:2351–2359
Girvan MS, Bullimore J, Pretty JN, Osborn AM, Ball AS (2003) Soil type is the primary determinant of the composition of the total and active bacterial communities in arable soils. Appl Environ Microbiol 69:1800–1809 doi:10.1128/AEM.69.3.1800-1809.2003
Gomes NCM, Heuer H, Schönfeld J, Costa R, Hagler-Mendonca L, Smalla K (2001) Bacterial diversity of the rhizosphere of maize (Zea mays) grown in tropical soil studied by temperature gradient gel electrophoresis. Plant Soil 232:167–180 doi:10.1023/A:1010350406708
Gomes NCM, Fagbola O, Costa R, Rumjanek NG, Buchner A, Hagler-Mendonca L, Smalla K (2003) Dynamics of fungal communities in bulk and maize rhizosphere soil in the tropics. Appl Environ Microbiol 69:3758–3766 doi:10.1128/AEM.69.7.3758-3766.2003
Graham MH, Haynes RJ (2005) Catabolic diversity of soil microbial communities under sugarcane and other land uses estimated by Biolog and substrate-induced respiration methods. Appl Soil Ecol 29:155–164 doi:10.1016/j.apsoil.2004.11.002
Grayston SJ, Wang SQ, Campbell CD, Edwards AC (1998) Selective influence of plant species on microbial diversity in the rhizosphere. Soil Biol Biochem 30:369–378 doi:10.1016/S0038-0717(97)00124-7
Hawksworth DL, Rossman AY (1997) Where are all the undescribed fungi? Phytopthology 87:888–891 doi:10.1094/PHYTO.1997.87.9.888
Hawksworth DL (2001) The magnitude of fungal diversity: the 1.5 million species estimate revisited. Mycol Res 105:1422–1432 doi:10.1017/S0953756201004725
Hiltner L (1904) Uber neuere erfarungen und problem auf dem gebiet der bodenbakteriologie und unter besonderer berucksichtigung der grundung und brache. Arbeitent Dtsch Landwirtschafts-Gesellschaft 98:59–78
Kowalchuk GA, Buma DS, de Boer W, Klinkhamer PGL, van Veen JA (2002) Effects of above-ground plant species composition and diversity on the diversity of soil-borne microorganisms. Antonie Van Leeuwenhoek 81:509–520 doi:10.1023/A:1020565523615
Kowalchuk GA, Stienstra AW, Heilig GHJ, Stephen JR, Woldendorp JW (2000) Changes in the community structure of ammonia-oxidizing bacteria during secondary succession of calcareous grasslands. Environ Microbiol 2:99–110 doi:10.1046/j.1462-2920.2000.00080.x
Liu XB, Herbert SJ (2002) Fifteen years of research examining cultivation of continuous soybean in Northeast China. Field Crops Res 79:1–7 doi:10.1016/S0378-4290(02)00042-4
Liu XB, Jin J, Wang GH, Herbert SJ (2008) soybean yield physiology and development of high-yielding practices in Northeat China. Field Crops Res 105:157–171 doi:10.1016/j.fcr.2007.09.003
Marschner P, Crowley D, Yang CH (2004) Development of specific rhizosphere bacterial communities in relation to plant species, nutrition and soil type. Plant Soil 261:199–208 doi:10.1023/B:PLSO.0000035569.80747.c5
Marschner P, Neumann G, Kania A, Weiskopf L, Liebere R (2002) Spatial and temporal dynamics of the microbial community structure in the rhizosphere of cluster roots of white lupin (Lupinus albus L.). Plant Soil 246:167–174 doi:10.1023/A:1020663909890
Marschner P, Yang CH, Lieberei R, Crowley DE (2001) Soil and plant specific effects on bacterial community composition in the rhizosphere. Soil Biol Biochem 33:1437–1445 doi:10.1016/S0038-0717(01)00052-9
Matsuyama T, Nakajima Y, Matsuya K, Ikenaga M, Asakawa S, Kimura M (2007) Bacterial community in plant residues in a Japanese paddy field estimated by RFLP and DGGE analyses. Soil Biol Biochem 39:463–472 doi:10.1016/j.soilbio.2006.08.016
Nazih N, Finlay-Moore O, Hartel PG, Fuhrmann JJ (2001) Whole soil fatty acid methyl ester (FAME) profiles of early soybean rhizosphere as affected by temperature and matric water potential. Soil Biol Biochem 33:693–696 doi:10.1016/S0038-0717(00)00197-8
Nelson EB (1990) Exudate molecules initiating fungal responses to seeds and roots. Plant Soil 129:61–73 doi:10.1007/BF00011692
Rengel Z (2002) Genetic control of root exudation. Plant Soil 245:59–70 doi:10.1023/A:1020646011229
Rovira AD (1959) Root excretions in relation to the rhizosphere effect. IV. Influence of plant species, age of plant, light, temperature, and calcium nutrition on exudation. Plant Soil 11:53–64 doi:10.1007/BF01394753
Singh BK, Munro S, Potts JM, Millard P (2007) Influence of grass species and soil type on rhizosphere microbial community structure in grassland soils. Appl Soil Ecol 36:147–155 doi:10.1016/j.apsoil.2007.01.004
Smalla K, Wieland G, Buchner A, Zock A, Parzy J, Kaiser S, Roskot N, Heuer H, Berg G (2001) Bulk and rhizosphere soil bacterial communities studied by denaturing gradient gel electrophoresis: plant-dependent enrichment and seasonal shifts revealed. Appl Environ Microbiol 67:4742–4751 doi:10.1128/AEM.67.10.4742-4751.2001
Sylvia DM, Chellemi DO (2001) Interactions among root-inhabiting fungi and their implications for biological control of root pathogens. Adv Agron 73:1–33 doi:10.1016/S0065-2113(01)73003-9
Watanabe T, Asakawa S, Nakamura A, Nagaoka K, Kimura M (2004) DGGE method for analyzing 16S rDNA of methanogenic archaeal community in paddy field soil. FEMS Microbiol Lett 232:153–163 doi:10.1016/S0378-1097(04)00045-X
Whipps JM (2001) Microbial interactions and biocontrol in the rhizosphere. J Exp Bot 52:487–511
White TJ, Buns TD, Lee S, Taylor J (1990) Analysis of phylogenetic relationships by amplification and direct sequencing of ribosomal RNA genes. In: Innis MA, Gefland DH, Sninsky JJ, White TJ (eds) PCR protocols: A guide to methods and applications. Academic, New York, pp 315–322
Wieland G, Neumann R, Backhaus H (2001) Variation of microbial communities in soil, rhizosphere, and rhizoplane in response to crop species, soil type, and crop development. Appl Environ Microbiol 67:5849–5854 doi:10.1128/AEM.67.12.5849-5854.2001
Xin H, Ma H (1987) Occurrence and control for soybean root rot. Soybean Sci 6:189–196 in Chinese
Yang CH, Crowley DE (2000) Rhizosphere microbial community structure in relation to root location and plant iron nutritional status. Appl Environ Microbiol 66:345–351
Yao H, Jiao X, Wu F (2006) Effect of continuous cucumber cropping and alternative rotations under protected cultivation on soil microbial community diversity. Plant Soil 284:195–203 doi:10.1007/s11104-006-0023-2
Zhou J, Bruns MA, Tieduje JM (1996) DNA recovery from soils of diverse composition. Appl Environ Microbiol 62:316–322
Acknowledgment
The authors are grateful to Professor Makoto Kimura, Nagoya University Japan, for his comments and critical review of this manuscript. This research was supported by grants from Science and Technology Bureau of Heilongjiang Province (GA06B101-3-1) and National Natural Science Foundation of China (40671099).
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible Editor: Petra Marschner.
Rights and permissions
About this article
Cite this article
Wang, G., Xu, Y., Jin, J. et al. Effect of soil type and soybean genotype on fungal community in soybean rhizosphere during reproductive growth stages. Plant Soil 317, 135–144 (2009). https://doi.org/10.1007/s11104-008-9794-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-008-9794-y