Abstract
Monovalent cations such as potassium (K+) and ammonium \({\left( {{\text{NH}}^{ + }_{4} } \right)}\) are essential to plant growth and development whereas others, such as sodium (Na+), can greatly inhibit plant growth. To understand and potentially manipulate the roles of these monovalent cations, a detailed knowledge is required about how they move in, out, and throughout plants, processes that are mediated by membrane transporters. The application of genomics and post-genomics methods has provided an overall idea of the genes that encode membrane transporters and in combination with functional analyses we now have a more detailed picture of which proteins are involved in particular biological functions. This review will give an overview of the gene families that are involved in monovalent cation transport, their members and their broad functional classification. Subsequently, an update will be provided on the identity and the roles of specific isoforms with regard to important physiological processes such as the uptake, long distance transport and compartmentation of K+ and Na+ and the uptake from the soil of \({ {{\text{NH}}^{ + }_{4} } }\).
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plants need nutrients to grow and develop and most of these are found as inorganic minerals in the soil. However, most soils show deficiency for one or more essential minerals and optimal growth conditions are therefore rarely achieved. Throughout our agricultural history, man has tried to improve crop growth conditions through the use of organic or chemical fertilisers and currently agriculture relies on the input of an astonishing 150 million tonnes of chemical fertiliser to maintain production. Fertilisers add considerable costs to farming and also greatly impact on the environment: Fertiliser production and transport are highly energy consuming and (over) application can easily lead to greenhouse gas release and pollution of surface waters.
In contrast, many environments contain essential and non-essential minerals in concentrations high enough to cause toxicity and thus negatively affect plant growth. Examples include natural and man-made biotopes that contain toxic levels of heavy metals, arsenic, salt, protons and many other inorganics. Dealing with excessive amounts of inorganics is far less amenable to simple solutions and often the affected areas are therefore avoided for agronomic activities.
Plants have developed sophisticated mechanisms to cope with the low abundance of nutrients and also to provide tolerance against excessive levels of harmful inorganics: High affinity transport systems in roots ensure adequate nutrient uptake from dilute soil solutions. Often multiple uptake mechanisms with varying substrate affinities are available that respond to fluctuations in supply and suit a range of external substrate concentrations. Buffer mechanisms are present such as vacuolar accumulation during excess supply and subsequent mobilisation when supply becomes limited. Tolerance mechanisms include the restricted uptake and extrusion of harmful ions, their compartmentation into vacuoles and storage in particular tissues.
Nutrient acquisition and distribution and tolerance mechanisms inevitably rely on proteins that are embedded in membranes that are responsible for the transmembrane movement of ions. Not surprisingly, the study of such membrane transporters has featured heavily in general plant physiology as a discipline to understand plant nutrition and plant tolerance to abiotic stress. Post genomics data have shown the extent to which genomes encode membrane transporters and their phylogenetic relationships within and across species. It is clear that a significant proportion of the genome (>5%) encodes membrane transport proteins and establishing which particular transporters participate in a certain process is a daunting task. Nevertheless, the achieved and ongoing characterisation of specific transporters, combined with the large amount of post-genomics data that is now available allows a more detailed dissection of the role of transporters and transporter families in plant nutrition.
In this review I will update the recent progress that has been made in the functional annotation and characterisation of membrane transporters involved in the movement of monovalent cations. I will evaluate what the contribution of specific families and isoforms of transporters is to important physiological processes such as the uptake of nutrients, ion extrusion, and the compartmentation and long distance transport of minerals.
Monovalent cation transporters
Several plant genomes have been sequenced and this has generated ample insights into the composition of membrane transporter families. Although a significant proportion of putative membrane proteins is still annotated as ‘hypothetical’ or ‘unknown,’ most membrane proteins have been classified into specific gene families. In a small number of cases this classification is based on functional data but more often it is on the basis of inter- and intraspecies phylogenetic relationships. For the monovalent cations around 12 families (Table 1) can be distinguished that comprise transporters with various basic mechanisms (Fig. 1): (1) Ion channels, or uniporters, are integral membrane proteins, which form regulated, substrate specific pores in the membrane. Ion channels do not directly consume energy and exclusively catalyse ‘down-hill’ or ‘passive’ transport, i.e. ions moving down their electrochemical gradient. The opening and closing, or ‘gating,’ of ion channels is subject to many factors but often sensitive to the membrane potential (‘voltage gated channels’) whereas in other cases the presence of ligands is required to activate channels (‘ligand gated channels’), e.g. in the case of cyclic nucleotide gated channels. A third activating mechanism of ion channels is through sensing physical stress in the membrane itself in the case of so-called mechanosensitive ion channels.
Major classes of membrane proteins involved in the transport of monovalent cations. ‘Antiporters’ are secondary active mechanisms that use the proton motive force (‘H+ gradient’) to drive extrusion or vacuolar loading of cations. The main substrate and H+ move in opposite direction across the membrane. ‘Symporters’ similarly use the proton motive force for the accumulation or vacuolar release of monovalent cations but both substrates move in the same direction across the membrane. The large differences in cytoplasmic, apoplasmic and vacuolar pH constitute the necessary H+ gradients which act as an electrochemical ‘battery’ and can be used to energise antiport and symport activity. ‘Uniporters’ or ion channels are passive mechanisms. In essence, these are pores in the membrane whose opening and closing (gating) is strictly controlled by factors such as the membrane potential, the presence of ligands or physical forces on the membrane. Depending on the local electrical and chemical gradients, ion channels can facilitate either the uptake or the loss of cations from the cytosol. PM: plasma membrane; TO: tonoplast
Channels typically exhibit substrate affinities in the millimolar range and can catalyse the transport of millions of ions per second. The latter makes them ideally suited to function in the bulk movement of ions, for example during rapid adjustment to osmotic changes, large scale mineral uptake or in rapid signalling events. However, the substrate specificity of ion channels may be low. (2) In contrast to ion channels, antiporters or exchangers are a type of ‘carrier’ which can accumulate substrate. The energy for movement of the main substrate is derived from coupling its transport to the ‘down-hill’ movement of a second substrate. In plants, the second substrate is typically a proton (H+) and substantial H+ gradients exist across the plasma membrane and the vacuolar membrane to energise H+-coupled antiporters. (3) Symporters, or cotransporters, are also carrier type mechanisms that, like antiporters, use energy stored in the electrochemical H+ gradient to accumulate substrates against an electrochemical gradient. However, in the case of symporters, the main substrate and the H+ are moved in the same direction. Anti- and symporters can have very high substrate affinities and generally are highly selective. This inevitably means their turn over rates are considerably lower than those of ion channels and usually are limited to hundreds or thousands of substrates per second. All three mechanisms are found in the plasma membrane and endomembranes such as the tonoplast. A short description of the various gene families that encode monovalent cation transporters follows below.
Ion channels
On the basis of structure and homology to their animal counterparts, four main cation channel families can be distinguished that include voltage and ligand gated channel types (Fig. 2). Two families comprise K+ selective ion channels whereas the remaining two encode non-selective ion channels that do not or only weakly discriminate between monovalent cations.
Generalised secondary structures of ion channel subunits. a Shaker type voltage dependent K+ channels comprise 6 TMD with a voltage sensing region in the S4 domain that controls channel gating, and a ‘GYGD’ motif in the pore (P) region that confers K+ selectivity to the channel. Functional channels contain four 6 TMD subunits that form either hetero- or homotetramer. b Two pore K+ channels (TPKs) have 4 TMD with two GYGD motifs. The C-terminus often contains two Ca2+ binding ‘EF hand’ motifs (not shown). Two 4 TMD subunits form a functional channel, probably as a homodimer. c Cyclic nucleotide gated channels (CNGCs), like Shaker type channels, contain 6 TMD but do not have a voltage sensor. The pore region does not have a GYGD sequence and in plants, varies considerably from animal CNGCs. The C-terminus contains a calmodulin (CaM) and a cyclic nucleotide binding domain (CNBD) which partly overlap and control channel gating. Four 6 TMD subunits form either hetero- or homotetramers channels. d Glutamate like receptors (GLRs) contain 3 TMDs, two putative extracellular substrate binding domains (S) and functional GLRs may consist of four or five subunits
Shaker type channels are the most thoroughly characterised both in planta and using heterologous expression systems (see Very and Sentenac 2002 for review). Shaker type channels are predominantly expressed in the plasma membrane, exhibit a high selectivity for K+ and have a basic subunit structure of 6 transmembrane domains (TMDs) with a pore region between the 5th and 6th TMD (Fig. 2a). The pore region contains the selectivity filter with the canonical GYGD motif, a hall mark for K+ selectivity. The 4th TMD contains the voltage sensor and the entire protein contains 4 identical (homotetramer) or different (heterotetramer) subunits. Channel gating is activated in response to changes in the membrane potential and the family is subdivided accordingly: Depolarisation activated Shaker channels increase their open probability when the membrane becomes less polarised and this class of channel typically conducts outward K+ currents, i.e. K+ moving from the cytosol to either the external medium or into cellular compartments. In addition to these outward rectifying K+ channels, plasma membranes contain inward rectifying K+ channels that respond to hyperpolarisation of the membrane. Both inward and outward rectifying K+ channels show a strong dependence on membrane potential whereas a third subgroup only shows weak (inward) rectification. Shaker type channels fulfil a large number of functions but are major contributors to K+ uptake and K+ homeostasis and are main K+ conduits during cellular movement exemplified by stomatal function.
The two pore K + (TPK) channel family contains 5 members in Arabidopsis. TPKs show a 4-TMD, 2 pore structure with both pores containing the GYGD motif (Fig. 2b) and active channels presumably function as dimers (Voelker et al. 2006). No voltage sensor is present and TPK activity is not or very weakly affected by membrane voltage (Gobert et al. 2007). Four out of five TPKs contain two clearly defined ‘EF hand’ domains that are involved in Ca2+ binding, pointing to a regulatory function of this divalent cation. Only a few TPKs have been characterised either in heterologous systems or in planta (Becker et al. 2004; Gobert et al. 2007) and indicate a role in membrane potential regulation and in K+ homeostasis.
Cyclic nucleotide gated channels (CNGCs) show a very similar basic structure to Shaker type channels with 6 TMDs and a pore between the 5th and 6th TMD (Fig. 2c). Their gating is not, or only weakly, dependent on membrane voltage but does require the binding of ligand in the form of cGMP or cAMP. CNGC selectivity is likely to be low and this class of channel is believed to be able to conduct K+, Na+, Ca2+ and possibly other monovalent and divalent cations. In Arabidopsis, the CNGC family comprises 20 isoforms (see Talke et al. 2003 for review) and a similar number is present in rice. Some isoforms have been studied in detail and it appears that CNGCs function in the two broadly defined areas of plant pathogen response signalling and cation homeostasis. For example, CNGC2, 4, 11 and 12 all play roles in response to bacterial pathogens, particularly in the signal transduction that leads to the hypersensitive response (Clough et al. 2000; Balague et al. 2003; Yoshioka et al 2006). Other CNGC isoforms such as CNGC3 and CNGC10 have been shown to contribute to monovalent cation uptake in plant roots (Li et al. 2005; Gobert et al. 2006). A further cluster of CNGCs, CNGC7, 8, 16 and 18, is preferentially or exclusively expressed in pollen (Bock et al. 2006) although their function in this tissue is unknown.
The glutamate like receptor (GLR) gene family encodes another class of putative ligand gated ion channels. The functioning channel complex of GLRs is likely to consist of a tetra- or pentamer with each subunit containing three transmembrane segments and a pore region that includes the selectivity filter (Fig. 2d). Ligand binding is believed to take place at two extracellular sites. The Arabidopsis genome sequence revealed a gene family with 20 members encoding putative GLRs containing three subfamilies (Davenport 2002) and this arrangement is largely mirrored in rice. Presently very little is known regarding the physiological role of most GLRs: Overexpression of AtGLR3;2 led to Ca2+ deficiency symptoms and hypersensitivity to Na+ and K+ stress (Kim et al. 2001).
Apart from these four families, a two pore channel (TPC) with a structure that resembles two fused Shaker subunits has been found in Arabidopsis and other species. AtTPC1 mediates the vacuolar SV (slow vacuolar) current and may be relevant in trans-tonoplast monovalent cation fluxes or in Ca2+ signalling (Peiter et al. 2005). One putative K+ selective channel that resembles half the structure of TPK channels is also encoded by the Arabidopsis genome.
Ammonium transporters
Isoforms of the ammonium transporter (AMT) family constitute a class of ubiquitous integral membrane proteins that mediate movement of ammonia/ammonium \({\left( {{{\text{NH}}_{3} } \mathord{\left/ {\vphantom {{{\text{NH}}_{3} } {{\text{NH}}^{ + }_{4} }}} \right. \kern-\nulldelimiterspace} {{\text{NH}}^{ + }_{4} }} \right)}\) across cell membranes. AMTs possess 11 or 12 TMDs and have been found in all kingdoms of life. In plants, AMTs are expressed exclusively in the plasma membrane and their expression can be induced by substrate availability or by nitrogen deficiency. AMTs form homo- or heterotrimers, in which each subunit contains a narrow pore through which substrate transport occurs. The C-terminus of AMTs may be involved in allosteric regulation of the oligomer activity via phosphorylation of a threonine (Loque et al. 2007). Different AMT isoforms have diverging affinities that range between micromolar and millimolar and this property may vary in a tissue dependent manner (Ludewig et al. 2003). The AMT family in Arabidopsis has 5 to 6 members but is much bigger in plants that prefer \({\text{NH}}^{ + }_{4} \) as a nitrogen source such as rice which contains at least 11 AMT isoforms.
There is currently much debate about the exact mechanism employed by AMT transporters: In bacteria some AMT transporters can function as ‘gas channels’ facilitating transmembrane diffusion of ammonia (NH3) whereas plant AMT transporters are primarily involved in ammonium uptake. In plants, the high affinity system is thought to function as a uniport, i.e. thermodynamically equivalent to an ion channel. However, since it is virtually impossible to determine the exact nature of the permeating substrate, it can not be ruled out that some high affinity AMT proteins function as NH3:H+ symporters. The mechanism of low affinity \({\text{NH}}^{ + }_{4} \) uptake is unclear.
Cation exchangers
Plants have four major gene families that encode monovalent cation exchangers. Members of some of these families have been scrutinised for their substrate properties and physiological roles. The Na + :H + exchangers (NHXs) have 12 TMDs and a conserved domain that binds the inhibitor amiloride. The NHX family totals 6–8 members in Arabidopsis (Yokoi et al. 2002) and can be subdivided into a ‘vacuolar membrane’ and an ‘endomembrane’ subfamily with the former being capable of transporting Na+ and K+ with equal affinity whereas the other subclass has a greater affinity for K+ (Pardo et al. 2006). At the tonoplast, the capacity of NHXs to move Na+ into the vacuole makes them important plant salt tolerance determinants since vacuolar deposition of Na+ is deemed to be a major salinity tolerance mechanism. However, other functions include acidification of trafficking vesicles and endomembrane K+:H+ exchange. The latter is an important constituent of overall K+ homeostasis. A further function appears to be in vacuolar pH control: For example in the petals of morning glory, NHX1 is responsible for alkalisation of the vacuole thereby changing the petal colour from purplish red to sky-blue (Yoshida et al. 2005). Expression of NHXs occurs in all studied tissues with some isoforms showing tissue specific patterns. Expression can be constitutive but often responds to environmental stimuli, particularly the onset of salinity stress (Apse et al. 1999). An NHX-like protein, SOS1, is expressed at the plasma membrane where it participates in cytosolic Na+ extrusion (Zhu 2002).
A second family of cation H + exchangers (CHXs) is much larger and less well characterised (Sze et al. 2004). Around 30 isofoms are found in the Arabidopsis genome and around 20 in rice. CHXs have 10 to 12 TMDs and probably transport K+ and/or Na+ in exchange with H+ and are thus functionally and structurally very similar to the NHXs. Interestingly, the expression of around 10 Arabidopsis CHX proteins is pollen specific (Bock et al. 2006) and not detected in other tissues though the physiological relevance of this remains unknown.
Further putative plant Na+:H+ antiporters show high similarity to bacterial NhaD (Na+:H+ antiport family D) proteins which are 400–500 aa long and exhibit 10–13 TMDs. In Arabidopsis two isoforms (NHD1 and NHD2) are distinguished which may be primarily plastidic. The role of a similar protein in the salt resistant tree Populus euphratice (PeNhaD1) was recently reported on (Ottow et al. 2005).
The K + efflux antiporter (KEA) family was identified on the basis of homology to similar proteins in gram-negative bacteria. The Arabidopsis genome contains 6 KEA isoforms and although KEAs may function as K+ antiport, none of them has been studied in any detail (Maser et al. 2001).
Cation symporters
Three families of cation symporters are present in plant genomes that are involved in the movement of monovalent cations. The first family comprises the K + uptake permeases (KUPs) and is also annotated as the high affinity K + (HAK) or the K + transporter (KT) family (Ahn et al. 2004; Rodriguez-Navarro and Rubio 2006). The secondary structure of KUPs contains 12–14 TMDs and KUPs are highly selective for K+. Substrate affinities range from low micromolar to millimolar levels with some isoforms apparently capable of both low and high affinity K+ transport. In analogy with their bacterial orthologues, plant KUPs are assumed to mediate K+ transport via coupling to cotransported H+ but conclusive evidence for this mechanism has yet to be provided. There are around 12 KUPs in Arabidopsis and 17 in rice and targeting is likely to include both the plasma membrane and the tonoplast although often the exact membrane localisation is unknown. Many KUPs show tissue specific expression patterns and transcriptional regulation in response to K+ conditions. For example, the expression of several KUP isoforms is induced by K+ starvation (Armengaud et al. 2004; Gierth et al. 2005). The physiological roles of KUPs include high affinity K+ uptake at the root:soil boundary, intracellular K+ distribution, turgor driven growth (Rigas et al. 2001; Elumalai et al. 2002) and general K+ homeostasis (Ahn et al. 2004; Rodriguez-Navarro and Rubio 2006).
The high affinity K + transporter (HKT) family has only one member in Arabidopsis but 8 or 9 in rice and homologues have been found in species such as wheat, barley, eucalyptus and Mesembryanthemum (Platten et al. 2006). Structurally, plant HKTs are similar to fungal Trk K+ transporters with an 8 TMD, 4 pore architecture. The first plant HKT was identified using a cDNA library form starved wheat and TaHKT1 was capable of high affinity K+ transport when expressed in yeast (Rubio et al. 1995), hence its nomenclature. Yeast data also showed that, mechanistically, TaHKT1 mediated high affinity K+ uptake was coupled to Na+ transport and thus might constitute a Na+-gradient driven K+:Na+ symport. However, there is no evidence that such a system operates in planta (Maathuis et al. 1996) and subsequent studies have shown that Na+ coupled K+ transport may be an artefact (Haro et al. 2005) and that most HKTs are Na+ selective and likely to function as low affinity (e.g. Laurie et al. 2002; Berthomieu et al. 2003) or high affinity (Garciadeblas et al 2003; Haro et al. 2005) Na+ transporters in plants.
The cation-chloride-cotransporters (CCCs) were identified in plants on the basis of their homologues in animals. CCCs contain 12 TMDs and are well characterised in animals where they play essential roles in electrolyte homeostasis. Animal CCCs cotransport one Cl− with one K+, one Cl− with one Na+ or two Cl− with one K+ and one Na+ and thus operate electroneutrally. The number of plant CCCs is small, with only one isoform present in the Arabidopsis genome and two isoforms in rice. AtCCC functions as a 2Cl−:K+:Na+ cotransporter, is expressed in root and shoot tissues and affects chloride root-shoot partitioning (Colmenero-Flores et al. 2007). However, the potential role of CCCs in cation movement will need further study.
Potassium nutrition
The monovalent cation K+ is an essential nutrient for all living organisms and crucial for many metabolic reactions via its specific activation of enzymes. Its low charge:mass ratio also makes K+ well suited as a counter ion for cytoplasmic polyanions. Plants further need K+ in the vasculature as counter ion for long distance transport of sugars and nitrate and K+ is a predominant contributor to cell turgor via its osmotic effects in the vacuole. Thus, major physiological functions require high K+ concentrations and K+ can comprise up to 10% of plant dry weight. K+ deficiency on the other hand, has a large negative effect on plant health and biomass production.
K+ uptake from the soil
The mechanisms for K+ uptake into plant roots include high- and low-affinity components with Km values in the μM and mM range and were extensively studied using 86Rb+ as a K+ tracer (see Maathuis and Sanders 1996; Very and Sentenac 2003; Rodriguez-Navarro and Rubio 2006 for reviews). Low affinity uptake can proceed through passive mechanisms and is believed to be driven by the membrane potential whereas high affinity uptake may require additional mechanisms for energisation. Electrophysiological studies, particularly during the 1990s, showed that specific K+ ion channels and H+:K+ cotransport systems underlie these components of K+ uptake (Maathuis and Sanders 1994) and during the same period the molecular identity of the main contributing mechanisms was also established. A main component of the low affinity K+ uptake pathway in roots has been identified as an inward K+ selective channel of the Shaker type called AKT1 in Arabidopsis (Fig. 3; Sentenac et al. 1992; Hirsch et al. 1998) and with homologues in many species such as rice (OsAKT1), potato (StSKT1), carrot (DcDKT1) and maize (ZmKZM1). This type of channel is predominantly expressed in the plasma membrane of root cortical and epidermal cells (including root hairs) where it can be recorded as an ion channel with around 5 pS conductance and a Km for K+ of around 10 mM (Maathuis and Sanders 1995). Interestingly, its selectivity for K+ is several times higher than for Rb+, casting some doubt on the interpretation of many low affinity flux data particularly where the values of kinetic parameters such as Vmax and Km are concerned. Other Shaker type channels such as AtKC1 (also called AKT4/KAT3), although not functional as homomers, can modify K+ uptake by interacting with AKT1 (Reintanz et al. 2002; Very and Sentenac 2003). AKT1/KC1 heteromers show altered voltage dependence compared to AKT1 homomers. A low K+ induced signalling cascade that involves Ca2+ binding proteins and kinases may control the switch between AKT1 and AKT1/KC1 mediated K+ uptake (Li et al. 2006).
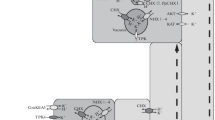
Various transporters involved in the uptake, long distance transport and compartmentation of K+. K+ uptake from the soil is predominantly mediated by HAK5 and AKT1 type transporters. AKT1 activity is modulated through heteromerisation with KC1 (AKT4) depending on K+ supply. Members of the CNGC family (e.g. CNGC10) may also play a role in K+ uptake. SKOR, and other non-identified channels, is responsible for xylem K+ loading whereas recycling of K+ through the phloem probably involves AKT3. The exact mechanism of vacuolar K+ loading is not understood but probably relies on H+:K+ antiporters from the CHX and/or NHX family. Vacuolar K+ release can be mediated by TPK1 in K+ replete conditions whereas KUP/HAK type symporters perform energised translocation of K+ to the cytoplasm when vacuolar K+ levels are diminished
Non selective, ligand gated channels from the CNGC family are also implicated as K+ uptake pathways. For example, CNGC10 which is relatively highly expressed in root tissue, was able to complement the K+ uptake deficient phenotype of the akt1–1 loss of function mutant, showing it can form a root K+ uptake pathway and considerably augment K+ uptake when overexpressed in the akt1–1 genotype (Li et al. 2005).
When external [K+] becomes increasingly low, K+ uptake needs to be energised. This presumably occurs through H+ coupled systems that have been shown to operate in root plasma membranes. Through a 1:1 coupling stoichiometry, such K+:H+ symports can drive 106 fold K+ accumulation (Maathuis and Sanders 1994). Many members of the KUP/HAK family show high selectivity for K+ and Km values in the micromolar range (Rodriguez-Navarro and Rubio 2006). Typically, HAKs are expressed ubiquitously making it difficult to find proteins that specifically participate in K+ uptake. However, loss of function studies and transcriptomics studies led to the identification of AtHAK5 as a major component of high affinity K+ uptake in roots (Armengaud et al. 2004; Gierth et al. 2005). This HAK isoform has a Km of around 25 μM which is in excellent agreement with that measured for unidirectional fluxes or using electrophysiology.
Thus, it appears that AKT1/KC1- type channels provide the major pathway for low affinity K+ uptake while H+ coupled KUP/HAK proteins, particularly HAK5, mediate high affinity K+ uptake. However, this demarcation between the two kinetic phases is probably less defined: deletion of AtAKT1 reduces low affinity K+ uptake but, interestingly, also reduced high affinity K+ uptake (Hirsch et al. 1998). The latter phenomenon was only observed in the presence of millimolar levels of \({\text{NH}}^{ + }_{4} \), which inhibit KUP/HAK type mechanisms, but suggests there is a large degree of plasticity in overall uptake. Studies with the akt1 loss of function mutant in the presence of \({\text{NH}}^{ + }_{4} \) also provide clear evidence that at least one other mechanism must be present in plant roots to mediate the residual K+ uptake in these conditions. The identity of this system has yet to be discovered and its characterisation would greatly help in completing our picture for K+ uptake from the soil.
Intercellular K+ distribution and long distance transport
High levels of K+ are required throughout the plant and a sophisticated mechanism is in place to ensure delivery of this ion to every cell. K+ uptake at the root soil boundary occurs through mechanisms discussed above and its distribution around the root symplast most likely depends on bulk flow through interconnecting plasmodesmata. Delivery to non-root tissues requires K+ loading into the xylem apoplast a process largely controlled by outward rectifying K+ channels originally identified in Arabidopsis as AtSKOR (Fig. 3; Gaymard et al. 1998). SKOR is a Shaker type K+ channel activated when the membrane depolarises. A SKOR knockout mutant showed a reduced shoot K+ content and a reduced K+ content in the xylem sap and SKOR transcription is inhibited by ABA (Lagarde et al. 1996). The latter ensures reduced K+ loading into the xylem to maintain adequate root turgor when soils dry out. All these properties suggest an important role for SKOR in K+ xylem loading and hence its delivery to shoot tissue. However, using patch clamp studies on xylem parenchyma cells, several other outward rectifying, K+ conducting channels have been observed (Roberts and Tester 1995; Wegner and deBoer 1999) but further research is needed to establish the molecular identity of these transporters.
Long distance K+ transport is by no means unidirectional and a large proportion of shoot K+ is recycled to the root through the phloem (Marschner et al. 1997). One reason for this apparently futile cycle is the role K+ plays as counterion for root to shoot translocation of \({\text{NO}}^{ - }_{{\text{3}}} \) and the phloem K+ influx that accompanies assimilate loading. The K+ channel AKT3 is weakly inward rectifying and has been shown to express in the phloem of Arabidopsis leaves (Marten et al. 1999). AKT3 is therefore a prime candidate for the release of K+ into the phloem.
Cellular K+ partitioning
The prominent role of K+ as turgor provider means it is deposited in the vacuole in high concentrations. However, in many conditions vacuolar K+ may need to be released, for example when K+ becomes deficient (Walker et al. 1996), when significant osmotic adjustment is necessary or when turgor driven movement is required such as during stomatal closure. The processes of vacuolar deposition and vacuolar release are mediated by tonoplast transporters involved in the bidirectional transfer of K+. Vacuolar loading of K+ may to some extent be mediated by cation channels such as TPC1 and TPK1 but must rely on energised mechanisms to reach K+ concentrations that are equal or higher than those in the cytoplasm. It is generally assumed that K+:H+ exchangers, particularly from the CHX and NHX family, drive such fluxes ((Fig. 3; Cellier et al. 2004; Sze et al. 2004; Pardo et al. 2006). Hard evidence for this assumption remains elusive and further research is urgently needed to identify specific members of the CHX or other exchanger families that are responsible for this important mechanism.
Release of vacuolar K+ is largely thermodynamically ‘down hill’ and thus likely to be through ion channels. Particularly TPK1 appears to be a main contributor to this process since its expression was shown to impact on overall K+ homeostasis and on stomatal closure in particular (Gobert et al. 2007). However, after prolonged K+ starvation, vacuolar K+ concentrations may become significantly lower than cytoplasmic ones (Walker et al. 1996) and thus vacuolar K+ release requires energisation. The participating transporter(s) is unknown but interestingly, proteomics studies suggest that several members of the HAK/KUP family are localised at the tonoplast (e.g. Jaquinod et al. 2007). Driven by the trans-tonoplast proton motive force, such systems could facilitate ‘up hill’ K+ release from the vacuole.
Salinity and sodium transport
Non-essential inorganic ions can be present in soils in toxic amounts. The most significant for agriculture is Na+ accumulation, a rapidly increasing problem in many areas around the globe. Salt-toxicity in plants comprises osmotic and ionic components, both causing diminished growth and a main question associated with this stress concerns the identity and regulation of transporters involved in the uptake and distribution of Na+. Although slow progress is being made in this area, most studies have been carried out using glycophytic models which to some extent may challenge the validity of the obtained results since they may not necessarily pertain to tolerance mechanisms but rather reflect general stress responses. On the other hand, most crops are glycophytic and thus it is important to understand mechanisms in this plant category before we can improve its salt tolerance.
Na+ uptake from the soil
The inward electrochemical gradient for Na+ is large in most conditions and it is generally accepted that its influx into the plant symplast occurs to a large extent via ion channels (see Demidchik and Maathuis 2007 for review). Unlike animals, plants do not appear to have Na+ selective ion channels and the selectivity of most K+ channels is sufficiently high to preclude significant Na+ conductance. However, a large number of studies has shown that non-selective cation channels are responsible for a large portion of Na+ uptake. With the large gene families encoding this class of ion channel, identification of specific gene products has been difficult. A particular, predominantly root expressed, CNGC isoform was recently identified as contributing to short term Na+ uptake and CNGC3 loss of function marginally improved plant salt tolerance (Fig. 4; Gobert et al. 2006). Other CNGCs such as CNGC10 may also play a role in Na+ uptake (Li et al. 2005).
Transporters involved in the uptake, long distance transport and compartmentation of Na+. The identity of the transporters that mediate Na+ uptake from the soil is largely unknown but probably includes non-selective ion channels such as CNGC3 and HKT1;1-type mechanisms for low affinity uptake. Although probably species dependent, plants may also use HKT type transporters for high affinity Na+ uptake, particularly when K+ is deficient. Salt tolerance in several species relies on Na+ extrusion via antiport systems such as SOS1 at the plasma membrane and NHX1 at the vacuolar membrane. Loading of Na+ into the xylem via SOS1 and CHX type transporters, its removal from the transpirational stream by HKT1;5 and its loading into the phloem by HKT1;1 all contribute to the overall long distance movement and shoot root partitioning of Na+
Another important factor for Na+ influx are the HKT transporters. In Arabidopsis the HKT family contains only one member but particularly in cereals larger families are present with 9 members in rice (Platten et al. 2006). In wheat, the use of TaHKT1 antisense led to reduced Na+ influx in the low affinity range and improved salt tolerance (Laurie et al. 2002). In cereals such as rice HKT transporters are likely to contribute to both low and high affinity Na+ uptake: OsHKT2;1 exhibited a Km of around 20 μM and is believed to mediate high affinity Na+ uptake whereas another isoform OsHKT1;1 showed an apparent Km of around 4 mM (Garciadeblas et al. 2003). High affinity Na+ uptake is not of relevance in the context of salt stress but does become important when K+ becomes severely limited. Both flux data and transcriptional data derived from cereals suggest that in those conditions, HKT transporters augment plant survival by mediating Na+ uptake that can substitute the lost K+ (Rodriguez-Navarro and Rubio 2006). Indeed, growth of K+ depleted plants is typically improved when moderate concentrations of Na+ are supplied and mechanisms such as OsHKT2;1 are likely to facilitate this.
Long distance transport and distribution of Na+
Many plants, including most crops, avoid excessive build up of salt in their photosynthetic tissues. Na+ translocation to shoots is therefore often limited in contrast to K+ translocation and this apparent K+/Na+ selectivity is often used as an important determinant of salt resistance. It remains to be shown how root symplastic Na+ enters the xylem. The K+ release channel SKOR is virtually impermeable to Na+ so can be ruled out. Root xylem parenchyma cells do contain low selectivity, outward rectifying, cation channels that can release Na+ into the xylem (Roberts and Tester 1995; Wegner and deBoer 1999) but their genes are not known. The NHX-like antiporter SOS1 is expressed in root xylem parenchyma cells and has been suggested to play a role in Na+ xylem loading during moderate salt stress (Fig. 4; Shi et al. 2002). Another H+ cation exchanger, AtCHX21, is primarily expressed in the root endodermis. Loss of function in this gene leads to diminished Na+ levels in the xylem but does not affect phloem Na+ levels. These results suggest that CHX21 is another important contributor to Na+ xylem loading (Hall et al. 2006).
Several HKTs have also been implicated in root to shoot translocation of Na+. In rice, HKT1;5 was found to be instrumental in controlling shoot K+/Na+ ratios. This plasma membrane Na+ transporter is expressed predominantly in xylem parenchyma cells where it is believed to retrieve Na+ from the sap to prevent Na+ accumulation in the shoot (Ren et al. 2005). TmHKT7 appears to fulfil a comparable role in durum wheat (Huang et al. 2006).
In Arabidopsis, distribution of Na+ between root and shoot tissues is to a large extent controlled by AtHKT1 (Berthomieu et al. 2003; Rus et al. 2004). AtHKT1 is mainly expressed in the phloem and in shoots is believed to load Na+ into this conductive tissue for retranslocation to the root. Since, as with many transporters, the activity of HKT1 is essentially reversible, it may further contribute to relieving the shoot from excessive Na+ load by extracting Na+ from the root phloem, possibly for extrusion into the root apoplast. However, the notion that retranslocation of shoot Na+ contributes significantly to salt tolerance has been challenged. Work carried out on wheat showed no correlation between shoot to root Na+ retranslocation and salt tolerance in a comparative study between two varieties of durum wheat known to differ in salt tolerance and Na+ accumulation (Davenport et al. 2005).
Efflux of Na+
In contrast to K+, Na+ can easily accumulate to toxic levels within the cell cytosol and various mechanisms are present to prevent this. Na+ compartmentation into the vacuole occurs in all tissues and is a main strategy to detoxify Na+ while still retaining its contribution as a ‘cheap’ osmolyte to lower the water potential. NHX1 is a H+ driven antiport mechanism at the tonoplast that plays a major role in this process (Fig. 4). AtNHX1 overexpression significantly improved salinity tolerance in Arabidopsis (Apse et al. 1999) and subsequent manipulation of NHX1 ortholog expression in other species such as wheat (Xue et al. 2004), rice (Fukuda et al. 2004) and tomato (Zhang and Blumwald 2001) showed the fundamental role this protein plays in salt tolerance and explains why it is a major focus for genetic engineering. Other NHX isoforms localising to the tonoplast (Yokoi et al. 2002) may also contribute to vacuolar Na+ sequestration.
Apart from improving vacuolar Na+ deposition, NHX1 could also mitigate the K+ deficiency that frequently accompanies salinity stress. The increasing evidence that NHX antiporters are main constituents of overall K+ homeostasis (Pardo et al. 2006) appears to be substantiated by the study of Xue et al., (2004) where increased NHX1 expression led to improved wheat salt tolerance but augmented shoot K+ levels rather than Na+.
Cytosolic Na+ is further controlled by H+ driven antiporters that are expressed at the plasma membrane. For example, SOS1 is expressed in xylem parenchyma cells where it may participate in loading and retrieval of xylem Na+. However, SOS1 is also prominent in root tip cells. Since these cells are predominantly evacuolate and therefore lack the option of vacuolar Na+ compartmentation, SOS1 is believed to be essential in this tissue for the extrusion of cytoplasmic Na+ into the apoplast. Na+ extrusion into the apoplast is assumed to take place in most plant tissues, particularly at the root–soil boundary. However, the participating proteins have yet to be identified and whether such a mechanism can substantially contribute to salt tolerance, particularly in field conditions, has yet to be established.
Ammonium nutrition
Plants predominantly use \({\text{NO}}^{ - }_{{\text{3}}} \) and \({\text{NH}}^{ + }_{4} \) as nitrogen source. As a reduced form of nitrogen \({\text{NH}}^{ + }_{4} \) is preferred by many plants over \({\text{NO}}^{ - }_{{\text{3}}} \) when both are available. Nevertheless, due to its tendency to dissipate transmembrane H+ gradients, \({\text{NH}}^{ + }_{4} \) readily leads to toxicity at high concentrations. Thus its transport and distribution are tightly controlled.
At the root–soil boundary members of the AMT family are responsible of \({\text{NH}}^{ + }_{4} \) uptake. In tomato root hairs LeAMT1;1 functions as a membrane potential driven uniport system with a Km for \({\text{NH}}^{ + }_{4} \) in the micromolar region (Ludewig et al. 2002). In the same organism, LeAMT1;2 functions in a similar way but with a lower substrate affinity.
AMT transporters not only move \({\text{NH}}^{ + }_{4} \) as a substrate but may simultaneously act as \({\text{NH}}^{ + }_{4} \) sensors via a cytosolic domain that is part of the C-terminus (Loque et al. 2007). Phosphorylation of this domain catalyses its interaction with cytoplasmic loops of the AMT protein providing rapid inactivation of \({\text{NH}}^{ + }_{4} \) transport.
Vacuolar deposition of \({\text{NH}}^{ + }_{4} \) may constitute a storage and detoxification mechanism and levels of 2–45 mM have been reported (Miller et al. 2001). Although \({\text{NH}}^{ + }_{4} \) permeable channels have been recorded at the tonoplast their role in \({\text{NH}}^{ + }_{4} \) compartmentation and molecular identity remains to be revealed. In addition, aquaporins may mediate transtonoplast movement of NH3. Similarly, an \({\text{NH}}^{ + }_{4} \) permeable channel in the rhizobium peribacteroid membrane has been implicated in the trans-symbiont nitrogen transfer (Tyerman et al. 1995) but it has not been identified at the gene level.
Concluding remarks
In their natural environment plants experience multiple and complex combinations of nutrient supply and also the presence of potentially detrimental minerals. Understanding how plants execute a coordinated response to fluctuating conditions and how they generate tolerance against harmful ions has been greatly improved by high throughput approaches such as genome sequencing and transcriptomics. Results from such studies show that the vast majority of plant transporters is encoded by multigene families to which broad functions can be assigned. However, it is not immediately obvious which members of a family contribute to specific processes such as the uptake or distribution of a particular nutrient. Thus, although bioinformatics has generated a platform for the selection of candidate genes, to elucidate the roles of specific transporters, reverse genetics, overexpression and functional analysis studies are still required.
The combination of genome based methods and functional analyses of individual transporter proteins has so far been very successful in assigning physiological functions to particular transporters but less so in understanding the complement of transporters that contributes to a particular process. In Arabidopsis for example, members of the HAK/KUP transporter family, of the KAT/AKT channel family, and CNGC family all play roles in K+ uptake from the soil, a process that is now fairly well understood. In many other cases, such as Na+ uptake or K+ partitioning, this information is far more limited.
It is also good to point out that much of the evidence for the role of particular proteins in physiological processes has yet to be confirmed outside the laboratory. Indeed, real field conditions could perceivably lead to expression of hitherto unknown proteins or reveal properties that otherwise remain obscured. With the advent of better model species such as rice and the application of more realistic conditions the ‘real’ physiological value of specific proteins will be easier to assess.
Thus, although there is still a long way to go and the laborious nature of ‘gene by gene’ functional analyses hampers rapid advancement, the continuing expansion of our knowledge regarding membrane transporters will lead to synergistic benefits and therefore speed up progress.
References
Ahn SJ, Shin R, Schachtman DP (2004) Expression of KT/KUP genes in Arabidopsis and the role of root hairs in K+ uptake. Plant Physiol 134:1135–1145
Apse MP, Aharon GS, Snedden WA, Blumwald E (1999) Salt tolerance conferred by overexpression of a vacuolar Na+/H+ antiport in Arabidopsis. Science 285:1256–1258
Armengaud P, Breitling R, Amtmann A (2004) The potassium-dependent transcriptome of Arabidopsis reveals a prominent role of jasmonic acid in nutrient signaling. Plant Physiol 136:2556–2576
Balague C, Lin BQ, Alcon C, Flottes G, Malmstrom S, Kohler C, Neuhaus G, Pelletier G, Gaymard F, Roby D (2003) HLM1, an essential signaling component in the hypersensitiveresponse, is a member of the cyclic nucleotide-gated channel ion channel family. Plant Cell 15:365–379
Becker D, Geiger D, Dunkel M, Roller A, Bertl A, Latz A, Carpaneto A, Dietrich P, Roelfsema MRG, Voelker C, Schmidt D, Mueller-Roeber B, Czempinski K, Hedrich R (2004) AtTPK4, an Arabidopsis tandem-pore K+ channel, poised to control the pollen membrane voltage in a pH- and Ca2+-dependent manner. Proc Natl Acad Sci USA 101:15621–15626
Berthomieu P, Conejero G, Nublat A, Brackenbury WJ, Lambert C, Savio C, Uozumi N, Oiki S, Yamada K, Cellier F, Gosti F, Simonneau T, Essah PA, Tester M, Very AA, Sentenac H, Casse F (2003) Functional analysis of AtHKT1 in Arabidopsis shows that Na+ recirculation by the phloem is crucial for salt tolerance. EMBO J 22:2004–2014
Bock KW, Honys D, Ward JM, Padmanaban S, Nawrocki EP, Hirschi KD, Twell D, Sze H (2006) Integrating membrane transport with male gametophyte development and function through transcriptomics. Plant Physiol 140:1151–1168
Cellier F, Conejero G, Ricaud L, Luu DT, Lepetit M, Gosti F, Casse F (2004) Characterization of AtCHX17, a member of the cation/H+ exchangers, CHX family, from Arabidopsis thaliana suggests a role in K+ homeostasis. Plant J 39:834–846
Clough SJ, Fengler KA, Yu I, Lippok B, Smith RK, Bent A (2000) The Arabidopsis dnd1 “defense, no death” gene encodes a mutated cyclic nucleotide-gated ion channel. Proc Natl Acad Sci USA 97:9323–9328
Colmenero-Flores JM, Martinez G, Gamba G, Vazquez N, Iglesias DJ, Brumos J, Talon M (2007) Identification and functional characterization of cation-chloride cotransporters in plants. Plant J 50:278–292
Davenport R (2002) Glutamate receptors in plants. Annals Bot 90:549–557
Davenport R, James RA, Zakrisson-Plogander A, Tester M, Munns R (2005) Control of sodium transport in durum wheat. Plant Physiol 137:807–818
Demidchik V, Maathuis FJM (2007) Physiological roles of non-selective cation channels in plants: from salt stress to signalling and development. New Phytol 175:387–404
Elgavish A, Elgavish GA (1985) In vitro ethanol effects on the transport properties of isolated renal brush-border membrane vesicles. J Membr Biol 88:123–130
Elumalai RP, Nagpal P, Reed JW (2002) A mutation in the Arabidopsis KT2/KUP2 potassium transporter gene affects shoot cell expansion. Plant Cell 14:119–131
Fukuda A, Nakamura A, Tagiri A, Tanaka H, Miyao A, Hirochika H, Tanaka Y (2004) Function, intracellular localization and the importance in salt tolerance of a vacuolar Na+/H+ antiporter from rice. Plant Cell Physiol 45:146–159
Garciadeblas B, Senn ME, Banuelos MA, Rodriguez-Navarro A (2003) Sodium transport and HKT transporters: the rice model. Plant J 34:788–801
Gaymard F, Pilot G, Lacombe B, Bouchez D, Bruneau D, Boucherez J, Michaux-Ferriere M, Thibaud J, Sentenac H (1998) Identification and disruption of a plant Shaker-like outward channel involved in K+ release into the xylem sap. Cell 94:647–655
Gierth M, Maser P, Schroeder JI (2005) The potassium transporter AtHAK5 functions in K+ deprivation- induced high-affinity K+ uptake and AKT1 K+ channel contribution to K+ uptake kinetics in Arabidopsis roots. Plant Physiol 137:1105–1114
Gobert A, Park G, Amtmann A, Sanders D, Maathuis FJM (2006) Arabidopsis thaliana Cyclic Nucleotide Gated Channel 3 forms a non-selective ion transporter involved in germination and cation transport. J Exp Bot 57:791–800
Gobert A, Isayenkov S, Voelker C, Czempinski K, Maathuis FJM (2007) The two pore channel TPK1 gene encodes the vacuolar K+ (VK) conductance and plays a role in K+ homeostasis. Proc Natl Acad Sci USA 104:10726–10731
Hall D, Evans AR, Newbury HJ, Pritchard J (2006) Functional analysis of CHX21: a putative sodium transporter in Arabidopsis. J Exp Bot 57:1201–1210
Haro R, Banuelos MA, Senn MAE, Barrero-Gil J, Rodriguez-Navarro A (2005) HKT1 mediates sodium uniport in roots. Pitfalls in the expression of HKT1 in yeast. Plant Physiol 139:1495–1506
Hirsch RE, Lewis BD, Spalding EP, Sussman MR (1998) A role for the AKT1 potassium channel in plant nutrition. Science 280:918–921
Huang SB, Spielmeyer W, Lagudah ES, James RA, Platten JD, Dennis ES, Munns R (2006) A sodium transporter (HKT7) is a candidate for Nax1, a gene for salt tolerance in durum wheat. Plant Physiol 142:1718–1727
Jaquinod M, Villiers F, Kieffer-Jaquinod S, Hugouvieu V, Bruley C, Garin J, Bourguignon J (2007) A proteomics dissection of Arabidopsis thaliana vacuoles isolated from cell culture. Mol Cell Prots 6:394–412
Kim SA, Kwak JM, Jae SK, Wang MH, Nam HG (2001) Overexpression of the AtGluR2 gene encoding an arabidopsis homolog of mammalian glutamate receptors impairs calcium utilization and sensitivity to ionic stress in transgenic plants. Plant Cell Physiol 42:74–84
Lagarde D, Basset M, Lepetit M, Conegero G, Gaymard F, Astruc S, Grignon C (1996) Tissue-specific expression of Arabidopsis AKT1 gene is consistent with a role in K+ nutrition. Plant J 9:195–203
Laurie S, Feeney KA, Maathuis FJM, Heard PJ, Brown SJ, Leigh RA (2002) A role for HKT1 in sodium uptake by wheat roots. Plant J 32:139–149
Li XL, Borsics T, Harrington HM, Christopher DA (2005) Arabidopsis AtCNGC10 rescues potassium channel mutants of E-coli, yeast and Arabidopsis and is regulated by calcium/calmodulin and cyclic GMP in E-coli. Function Plant Biol 32:643–653
Li LG, Kim BG, Cheong YH, Pandey GK, Luan S (2006) A Ca2+ signaling pathway regulates a K+ channel for low-K response in Arabidopsis. Proc Natl Acad Sci USA 103:12625–12630
Loque D, Lalonde S, Looger LL, von Wiren N, Frommer WB (2007) A cytosolic trans-activation domain essential for ammonium uptake. Nature 446:195–198
Ludewig U, von Wiren N, Frommer WB (2002) Uniport of \({\text{NH}}^{ + }_{4} \) by the root hair plasma membrane ammonium transporter LeAMT1;1. J Biol Chem 277:13548–13555
Ludewig U, Wilken S, Wu BH, Jost W, Obrdlik P, El Bakkoury M, Marini AM, Andre B, Hamacher T, Boles E, von Wiren N, Frommer WB (2003) Homo- and hetero-oligomerization of ammonium transporter-1 \({\text{NH}}^{ + }_{4} \) uniporters. J Biol Chem 278:45603–45610
Maathuis FJM, Sanders D (1994) Mechanism of high affinity potassium uptake in roots of Arabidopsis thaliana. Proc Natl Acad Sci USA 91:9272–9276
Maathuis FJM, Sanders D (1995) Contrasting roles in ion transport of two K+ channel types in root cells of Arabidopsis thaliana. Planta 197:456–464
Maathuis FJM, Sanders D (1996) Mechanisms of potassium absorption by higher plant roots. Physiol Plant 96:158–168
Maathuis FJM, Verlin D, Smith FA, Sanders D, Fernandez JA, Walker NA (1996) The physiological relevance of Na+-coupled K+-transport. Plant Physiol 112:1609–1616
Marschner H (1995) Mineral nutrition in higher plants. In Mineral nutrition of higher plants. Academic Press, London
Marschner H, Kirkby EA, Engels C (1997) Importance of cycling and recycling of mineral nutrients within plants for growth and development. Bot Act 110:265–273
Marten I, Hoth S, Deeken R, Ache P, Ketchum KA, Hoshi T, Hedrich R (1999) AKT3, a phloem-localized K+ channel, is blocked by protons. Proc Natl Acad Sci USA 96:7581–7586
Maser P, Thomine S, Schroeder JI, Hirschi K, Ward JM, Sze H, Amtmann A, Maathuis FJM, Talke I, Sanders D, Persans MW, Salt DE, Kim SA, Guerinot ML, Gribskov M (2001) Phylogenetic relationships within cation-transporter families of Arabidopsis thaliana. Plant Physiol 126:1646–1667
Miller AJ, Cookson SJ, Smith SJ, Wells DM (2001) The use of microelectrodes to investigate compartmentation and the transport of metabolized inorganic ions in plants. J Exp Bot 52:541–549
Ottow EA, Polle A, Brosche M, Kangasjarvi J, Dibrov P, Zorb C, Teichmann T (2005) Molecular characterization of PeNhaD1: the first member of the NhaD Na+/H+ antiporter family of plant origin. Plant Mol Biol 58:75–88
Padmanaban S, Chanroj S, Kwak JM, Li X, Ward JM, Sze H (2007) Participation of the endomembrane cation/H+ exchanger AtCHX20 in osmoregulation of guard cells. Plant Physiol 144:82–93
Pardo JM, Cubero B, Leidi EO, Quintero FJ (2006) Alkali cation exchangers: roles in cellular homeostasis and stress tolerance. J Exp Bot 57:1181–1199
Peiter E, Maathuis FJM, Mills LN, Knight H, Pelloux M, Hetherington AM, Sanders D (2005) The vacuolar Ca2+-activated channel TPC1 regulates germination and stomatal movement. Nature 434:404–408
Platten JD, Cotsaftis O, Berthomieu P, Bohnert H, Davenport RJ, Fairbairn DJ, Horie T, Leigh RA, Lin HX, Luan S, Maser P, Pantoja O, Rodriguez-Navarro A, Schachtman DP, Schroeder JI, Sentenac H, Uozumi N, Very AA, Zhu JK, Dennis ES, Tester M (2006) Nomenclature for HKT transporters, key determinants of plant salinity tolerance. Trends Plant Sci 11:372–374
Reintanz B, Szyroki A, Ivashikina N, Ache P, Godde M, Becker D, Palme K, Hedrich R (2002) AtKC1, a silent Arabidopsis potassium channel alpha-subunit modulates root hair K+ influx. Proc Natl Acad Sci USA 99:4079–4084
Ren ZH, Gao JP, Li LG, Cai XL, Huang W, Chao DY, Zhu MZ, Wang ZY, Luan S, Lin HX (2005) A rice quantitative trait locus for salt tolerance encodes a sodium transporter. Nature Genetics 37:1141–1146
Rigas S, Debrosses G, Haralampidis K, Vicente-Agullo F, Feldmann KA, Grabov A, Dolan L, Hatzopoulos P (2001) TRH1 encodes a potassium transporter required for tip growth inArabidopsis root hairs. Plant Cell 13:139–151
Roberts SK, Tester M (1995) Inward and outward K+-selective currents in the plasma membrane of protoplasts from maize root cortex and stele. Plant J 8:811–825
Rodriguez-Navarro A, Rubio F (2006) High-affinity potassium and sodium transport systems in plants. J Exp Bot 57:1149–1160
Rubio F, Gassmann W, Schroeder JI (1995) Sodium-driven potassium uptake by the plant potassium transporter HKT1 and mutations conferring salt tolerance. Science 270:1660–1663
Rus A, Lee BH, Munoz-Mayor A, Sharkhuu A, Miura K, Zhu JK, Bressan RA, Hasegawa PM (2004) AtHKT1 facilitates Na+ homeostasis and K+ nutrition in planta. Plant Physiol 136:2500–2511
Sentenac H, Bonneaud N, Minet M, Lacroute F, Salmon JM, Gaymard F, Grignon C (1992) Cloning and expression in yeast of a plant potassium ion transport system. Science 256:663–665
Shi HZ, Quintero FJ, Pardo JM, Zhu JK (2002) The putative plasma membrane Na+ /H+ antiporter SOS1 controls long-distance Na+ transport in plants. Plant Cell 14:465–477
Sze H, Padmanaban S, Cellier F, Honys D, Cheng NH, Bock KW, Conejero G, Li XY, Twell D, Ward JM, Hirschi KD (2004) Expression patterns of a novel AtCHX gene family highlight potential roles in osmotic adjustment and K+ homeostasis in pollen development. Plant Physiol 136:2532–2547
Talke IN, Blaudez D, Maathuis FJM, Sanders D (2003) CNGCs: prime targets of plant cyclic nucleotide signalling? Trends Plant Sci 8:286–293
Tyerman SD, Whitehead LF, Day DA (1995) A channel-like transporter for \({\text{NH}}^{ + }_{4} \) on the symbiotic interface of N2-fixing plants. Nature 378:629–632
Very AA, Sentenac H (2002) Cation channels in the Arabidopsis plasma membrane. Trends Plant Sci 7:168–175
Very AA, Sentenac H (2003) Molecular mechanisms and regulation of K+ transport in higher plants. Ann Rev Plant Biol 54:575–603
Voelker C, Schmidt D, Mueller-Roeber B, Czempinski K (2006) Members of the Arabidopsis AtTPK/KCO family form homomeric vacuolar channels in planta. Plant J 48:296–306
Walker DJ, Leigh RA, Miller AJ (1996) Potassium homeostasis in vacuolate plant cells. Proc Natl Acad Sci USA 93:10510–10514
Wegner LH, deBoer AH (1999) Activation kinetics of the K+ outward rectifying conductance KORC in xylem parenchyma cells from barley roots. J Membr Biol 170:103–119
Xue ZY, Zhi DY, Xue GP, Zhang H, Zhao YX, Xia GM (2004) Enhanced salt tolerance of transgenic wheat (Tritivum aestivum L.) expressing a vacuolar Na+ /H+ antiporter gene with improved grain yields in saline soils in the field and a reduced level of leaf Na+. Plant Sci 167:849–859
Yokoi S, Quintero FJ, Cubero B, Ruiz MT, Bressan RA, Hasegawa PM, Pardo JM (2002) Differential expression and function of Arabidopsis thaliana NHX Na+/H+ antiporters in the salt stress response. Plant J 30:529–539
Yoshida K, Kawachi M, Mori M, Maeshima M, Kondo M, Nishimura M, Kondo T (2005) The involvement of tonoplast proton pumps and Na+ (K+)/H+ exchangers in the change of petal color during flower opening of morning glory, Ipomoea tricolor cv. Heavenly Blue. Plant Cell Physiol 46:407–415
Yoshioka K, Moeder W, Kang HG, Kachroo P, Masmoudi K, Berkowitz G, Klessig DF (2006) The chimeric Arabidopsis Cyclic Nucleotide-Gated Ion Channel11/12 activates multiple pathogen resistance responses. Plant Cell 18:747–763
Zhang HX, Blumwald E (2001) Transgenic salt-tolerant tomato plants accumulate salt in foliage but not in fruit. Nature Biotech 19:765–768
Zhu JK (2002) Salt and drought stress signal transduction in plants. Ann Rev Plant Biol 53:247–273
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible Editor: Yongguan Zhu.
Rights and permissions
About this article
Cite this article
Maathuis, F.J.M. Monovalent cation transporters; establishing a link between bioinformatics and physiology. Plant Soil 301, 1–15 (2007). https://doi.org/10.1007/s11104-007-9429-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-007-9429-8