Abstract
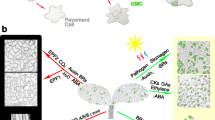
Plant leaves consist of three layers, including epidermis, mesophyll and vascular tissues. Their development is meticulously orchestrated. Stomata are the specified structures on the epidermis for uptake of carbon dioxide (CO2) while release of water vapour and oxygen (O2), and thus play essential roles in regulation of plant photosynthesis and water use efficiency. To function efficiently, stomatal formation must coordinate with the development of other epidermal cell types, such as pavement cell and trichome, and tissues of other layers, such as mesophyll and leaf vein. This review summarizes the regulation of stomatal development in three dimensions (3D). In the epidermis, specific stomatal transcription factors determine cell fate transitions and also activate a ligand-receptor- MITOGEN-ACTIVATED PROTEIN KINASE (MAPK) signaling for ensuring proper stomatal density and patterning. This forms the core regulation network of stomatal development, which integrates various environmental cues and phytohormone signals to modulate stomatal production. Under the epidermis, mesophyll, endodermis of hypocotyl and inflorescence stem, and veins in grasses secrete mobile signals to influence stomatal formation in the epidermis. In addition, long-distance signals which may include phytohormones, RNAs, peptides and proteins originated from other plant organs modulate stomatal development, enabling plants to systematically adapt to the ever changing environment.
Key message
Understanding the molecular regulation of stomatal development from a three-dimensional perspective.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plant leaves develop from primordia which consist of three distinct cell layers that give rise to the epidermis, mesophyll, and vascular tissues (Becraft 1999). The epidermis contains three main cell types including jigsaw-puzzle-shaped pavement cells, trichomes and stomata which are adjustable valves consisting of a pair of guard cells (GCs) flanking a pore. Stomata are essential for regulation of plant photosynthesis and water use efficiency, since their opening and closure driven by cell turgor are central for uptake of CO2 while release of water vapour and O2. The stomata of most plants are formed by a pair of kidney-shaped GCs (Fig. 1A). By contrast, grasses produce a pair of dumbbell-shaped GCs flanked by dome-shaped or triangular-shaped lateral subsidiary cells (SCs) which enable more rapid stomatal movement in response to intrinsic and extrinsic signals (Fig. 1B). In the dicot plant Arabidopsis and tomato, stomata are composed of a pair of kidney-shaped GCs and distributed on the leaf epidermis randomly (Fig. 1A). They are produced from a subset of protodermal cells which gain the meristemoid mother cell (MMC) fate and then initiate the stomatal development through undergoing asymmetric cell division, producing a meristemoid cell (M) and a stomatal lineage ground cell (SLGC). The M can self renew by asymmetric cell divisions, which produce more SLGCs, and then differentiates into a guard mother cell (GMC). The GMC undergoes symmetric cell division, producing paired GCs. The SLGCs usually differentiate into pavement cells, but the newly formed SLGCs can also undergo asymmetric division away from preexisting stomata or stomatal precursors to produce meristemoid cells which finally differentiate into stomata, resulting in two adjacent stomata are spaced by at least of one none-stomatal cell (Fig. 1C) (Bergmann and Sack 2007; Lau and Bergmann 2012; Pillitteri and Torii 2012; Pillitteri and Dong 2013; Han and Torii 2016; Ortega et al. 2019).
Stomata in the leaf epidermis of Arabidopsis thaliana and grasses. A Arabidopsis stomata are composed of paired, kidney-shaped GCs. They are scattered in the epidermis. B Grass stomata are composed of paired, dumbbell-shaped GCs flanked by dome-shaped or triangular-shaped lateral subsidiary cells. They are formed in cell files adjacent to veins. C Developmental trajectory of Arabidopsis stomata. Multiple, dispersed protodermal cells on the leaf obtained the MMC fate (light orange). MMC undergoes asymmetric cell division, producing a small M (orange) and a large SLGC. The M can self-renew and finally differentiates into GMC (yellow). GMC divides symmetrically once, producing paired GCs (green). The SLGCs can either differentiate into pavement cells or initiate stomatal development away from pre-existing stomata or stomatal precursors, making stomatal development obeys the one-cell-spacing rule. D Developmental trajectory of grass stomata. Certain cell files flanking leaf veins gain the fate of stomatal file (light orange). Cells within stomatal file divide asymmetrically, producing early GMCs (orange) toward leaf tip. Late GMCs (yellow) induce SMCs (light blue) to divide asymmetrically, producing SCs (blue) toward GMC. Then, late GMCs divide symmetrically once to produce paired GCs (green), finally forming the four-celled stomatal complex
In grasses, including Brachypodium, barley, wheat, rice and maize, stomata are differentiated toward leaf tip with the same orientation in specific files arranged parallelly on both sides of the leaf vein (Fig. 1B). Stomatal cell files are established with unknown mechanisms early in the leaf base and cells within them undergo asymmetric cell divisions to produce early GMCs toward leaf tip. Then, GMCs mature and induce their lateral subsidiary mother cells (SMCs) to divide asymmetrically, producing subsidiary cells (SCs). After recruitment of SCs, GMCs divide symmetrically to produce a pair of GCs (Fig. 1D) (McKown and Bergmann 2020; Nunes et al. 2020).
Epidermis originated signals regulate stomatal development
In Arabidopsis, the successive MMC–M–GMC–GC cell fate specifications are determined by three closely related basic helix–loop–helix (bHLH) transcription factors, SPEECHLESS (SPCH), MUTE and FAMA, respectively. They form heterodimer with two paralogous bHLH-leucine zipper transcription factors SCREAM (SCRM)/ICE1 (inducer of CBF expression 1) and SCRM2 and recruit RNA polymerase II (Pol II) complex to promote stomatal cell fate transitions (Zhou and Lau 2021; Han et al. 2021; Ohashi-Ito and Bergmann 2006; MacAlister et al. 2007; Pillitteri et al. 2007; Kanaoka et al. 2008; Chen et al. 2016a, 2021). The MYB transcription factor FOUR LIPS (FLP) is also proved to interact with SCRM and function redundantly with MUTE and FAMA in the differentiation of GMC and GC, respectively (Lai et al. 2005; Lee et al. 2014; Li et al. 2023) (Fig. 2A). Stomatal cell-fate transition and cell cycle progression are closely connected, and several specific cell cycle components have been found to play key roles in the process, such as CDKB1 (B-type Cyclin-Dependent Kinase 1), CDKA (A-type CDK), CYCA2 (A-type cyclin 2), CYCD3 (D-type cyclin 3), CYCD7 (D-type cyclin 7), SMR4 (SIAMESE RELATED 4) and CYCD5;1 (D-type cyclin 5;1) (Vanneste et al. 2011; Yang et al. 2014b, 2019; Boudolf et al. 2004; Han et al. 2018a, 2022; Weimer et al. 2018). In addition, stomatal transcriptome and chromatin accessibility are greatly changed during stomatal lineage progression (Adrian et al. 2015; Lee et al. 2019; Kim et al. 2022; Wallner et al. 2024; Liu et al. 2020). For a more comprehensive overview of the differences in stomatal cell identities and the linkage between stomatal development and cell cycle, readers may refer to the recent excellent reviews (Han and Torii 2019; Smit and Bergmann 2023).
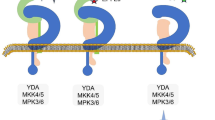
Molecular regulation of stomatal development in the epidermis of Arabidopsis thaliana and grasses. A In Arabidopsis, the successive MMC–M–GMC–GC cell fate specifications are determined by transcription factors, SPCH-SCRM, MUTE-SCRM-FLP and FAMA-SCRM-FLP, which form complex with Pol II. Upstream of these transcription factors is the YDA-MKK4/5/7/9-MPK3/6 cascade which is repressed by the receptor complexes TMM-ERf-SERKs, which recognizes EPFL peptides, and HSL1-SERKs, which recognizes CLE9/10 peptides. This constitutes the central signaling network of stomatal development, which integrates various plant hormone signals and environmental stimulus. The phytohormone signals and environmental cues integrated by the YDA-MAPK cascade influence each step of stomatal development, and they are not redundantly shown in the figure. The question mark indicates unidentified upstream signals mediated by YDA-MKK7/9-MPK3/6 that promote GMC differentiation. B Key transcription factors regulating grass stomatal cell fate transitions. OsSPCHs, OsSCRMs, OsSHRs, OsSCRs, BdSPCHs or BdSCRM functions in establishing the fate of stomatal file. OsSPCHs, OsSCRMs, BdSPCHs or BdSCRM functions in initiating the asymmetric cell divisions within stomatal file, producing early GMCs. OsMUTE, ZmMUTE, BdMUTE or BdFAMA promotes the differentiation of early GMCs into late GMCs, and initiates the asymmetric cell divisions of SMCs to produce SCs. OsFAMA, OsFLP, BdSCRM2 or BdFAMA promotes early GCs to differentiate into mature GCs. The question mark indicates unidentified regulators determining GMC to GC fate transition
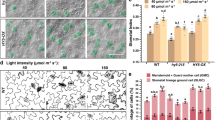
Shoot mesophyll and endodermis secrete regulators into epidermis for modulating stomatal development in Arabidopsis thaliana. The endodermis of hypocotyl and inflorescence stem secretes the peptide EPFL6 into the epidermis, which represses stomatal development mainly by activating ERf receptors. This process is dampened by the receptor TMM. The mesophyll secretes the peptide STOMAGEN into the epidermis, which promotes stomatal production by eliminating the suppression effects of EPF2. The mesophyll also secretes the transcription factor AN3 to promote the epidermal cell division. Both STOMAGEN and AN3 respond to light signals. The question mark indicates unidentified regulators secreted by mesophyll to regulate stomatal development in the epidermis
Vein may secrete signals to regulate stomatal development in grasses. Vein originated signals have long been proposed to be responsible for coordination of stomatal development with vein development and the grass SHR protein is a candidate that may move from vein into epidermis. The question mark indicates possible known or unknown regulators secreted by vein to regulate stomatal development in the epidermis
The old leaves or the root system may generate long-distance signals transmitted by vascular system for regulating stomatal development in young leaves, facilitating plant to systematically adapt to the ever changing environment. The possible long-distance signals include the glucose, transcription factors (e.g. HY5 and AN3), peptides (e.g. EPFs and CLEs), phytohormones (e.g. auxin, CK, BR, ABA and Jasmonate), and microRNAs. The question mark indicates possible known or unknown long-distance signals regulating stomatal development
SPCH is the most early regulator of stomatal development and acts as a molecular switch to initiate the stomatal lineage (Horst et al. 2015). SPCH activates the expression of itself and SCRMs, and then partners with SCRMs to further activate the expression of themselves, thus constituting a positive feedback loop for promoting the specification of stomatal-lineage fate (Lau et al. 2014; Horst et al. 2015). Concurrently, SPCH also activates the expression of EPIDERMAL PATTERNING FACTOR-LIKE (EPFL) peptide EPF2, leucine-rich repeat receptor-like protein TOO MANY MOUTHS (TMM) and the ERECTA (ER) family (ERf) of receptor-like kinases (Lau et al. 2014; Horst et al. 2015). EPF2 is co-expressed with SPCH in MMCs and early meristemoid cells and perceived by a receptor complex consisting of TMM, ERf and the SOMATIC EMBRYOGENESIS RECEPTOR KINASEs (SERKs). Genetic analysis indicated that EPF2 is mainly detected by ER, which activates a MAPK signal transduction cascade consisting of YODA (YDA)/MAPKKK, four MAPKKs (MKK4/5/7/9) and two MAPKs (MPK3/6). Activated MPK3/6 then phosphorylates SPCH-SCRMs, leading to the degradation of SPCH-SCRMs and thus inhibition of SPCH-SCRMs activity (Nadeau and Sack 2002; Shpak et al. 2005; Hara et al. 2009; Hunt and Gray 2009; Lee et al. 2012; Bergmann et al. 2004; Gray and Hetherington 2004; Wang et al. 2007; Lampard et al. 2008, 2009). Therefore, EPF2-TMM-ERf-MAPK cascade constitutes an autocrine negative feedback loop for repression of stomatal lineage initiation (Fig. 2A).
Another small secreted peptide EPF1, which is specifically expressed in late M, GMC and immature GC, is mainly recognized by ERf receptor ER-like 1 (ERL1) which is a direct target of MUTE and co-expressed with MUTE in late M and GMC (Hara et al. 2007; Lee et al. 2012). MUTE upregulates ERL1expression while EPF1-ERL1 suppress the activity of MUTE through the YDA MAPK cascade in turn, thus constituting a autocrine negative feedback loop for determination of GMC fate (Qi et al. 2017). Both EPF2 and EPF1 are also paracrine factors perceived by TMM-ERf-SERKs receptors expressed in the neighboring SLGC to repress stomatal formation. In addition, the peptide CLAVATA3/ESR-RELATED 9/10 (CLE9/10) which is also expressed in stomatal lineage cells is detected by the receptor HAESA-LIKE 1 (HSL1) which recruits SERKs as co-receptors to negatively regulate stomatal development by destabilizing SPCH (Qian et al. 2018) (Fig. 2A).
Plant hormones and environmental cues function on the epidermis to influence stomatal development by modulating the activity of stomatal signaling components (Chen 2023; Chen et al. 2020; Han et al. 2021; Le et al. 2014a, b) (Fig. 2A). The concentration of the auxin is dynamically changed during stomatal development and a decrease in auxin level in meristemoid cells is required for acquisition of GMC fate (Le et al. 2014a, b). Auxin also negatively regulates stomatal development by inhibiting AUXIN RESISTANT3 (AXR3) /IAA17 which is an auxin signaling repressor that promotes stomatal formation upstream of the YDA MAPK cascade (Balcerowicz et al. 2014). Cytokinin (CK) can induce the expression of SPCH and manipulation of CK levels affects epidermal cell division activity and stomatal production. The direct targets of SPCH also include ARABIDOPSIS RESPONSE REGULATOR 16 (ARR16), a type-A ARR that negatively regulates CK response, and CLE9/10, a stomatal lineage cell expressed small peptide that can repress type-A ARRs. ARR16/17 and CLE9/10 function counteractively to influence the CK sensitivity in meristemoid cells and SLGCs, thus determining the proliferative ability of these cells combined with SPCH activity (Vaten et al. 2018).
Abscisic acid (ABA) which accumulates during drought is sensed by its receptors including protein phosphatases type 2C (PP2Cs) and three SNF1-related protein kinases (SnRK2s), resulting in relieving the suppression of the SnRK2s by PP2C (Cutler et al. 2010). SnRK2s are present in the stomatal precursors and the ABA-activated SnRK2s then directly phosphorylate SPCH, leading to SPCH degradation and thus inhibiting stomatal production (Yang et al. 2022). Brassinosteroid (BR) regulates stomatal development by inactivation of BR INSENSITIVE2 (BIN2), a key negative regulator of BR signaling. BIN2 phosphorylates SPCH in the nucleus, triggering SPCH degradation (Gudesblat et al. 2012). BIN2 also activates YDA-MAPK cascade by phosphorylating YDA and MKK4 (Kim et al. 2012; Khan et al. 2013). In addition, BIN2 phosphorylates POLAR which functions as a scaffold that localizes polarly in the cell periphery and directs stomatal asymmetric cell division, leading to the stabilization of POLAR at the plasma membrane and subsequent polarized co-localization of BIN2 with POLAR. This thereby attenuates nuclear BIN2 mediated SPCH instability, thus promoting stomatal asymmetric cell division (Houbaert et al. 2018; Guo et al. 2021). Jasmonate represses stomatal production and its signaling functions upstream of stomatal key transcription factors (Han et al. 2018b).
Light signals promote stomatal formation by inhibiting its key signaling component RING E3 ubiquitin ligase CONSTITUTIVE PHOTOMORPHOGENIC 1 (COP1) which functions genetically upstream of YDA (Kang et al. 2009) and represses stomatal development through directly stimulating the degradation of SCRM in the dark (Lee et al. 2017). In addition, red light can induce the expression of GATA factors of the B-subfamily (B-GATA) transcription factors which directly promotes SPCH expression, thus positively regulating stomatal development (Klermund et al. 2016). Carbon dioxide (CO2) represses stomatal formation primarily by inducing the expression of CO2 RESPONSIVE SECRETED PROTEASE (CRSP) which promotes the maturation of EPF2 (Engineer et al. 2014). The heat-stress induces the accumulation of PHYTOCHROME-INTERACTING FACTOR 4 (PIF4) which directly repress SPCH expression and also enhances the activity of HEAT SHOCK PROTEINS 90 (HSP90s) which interacts with YDA and promotes the YDA MAPK cascade mediated phosphorylation and degredation of SPCH, thereby inhibiting stomatal production by suppressing SPCH at both transcriptional and translational level (Lau et al. 2018; Samakovli et al. 2020). Osmotic stress down-regulates SPCH protein level through the YDA MAPK cascade, resulting in reduced stomatal density (Kumari et al. 2014). Sucrose induces the accumulation of KIN10, one α-catalytic subunit of a central energy-sensor kinase SnRK1, in the nucleus of stomatal precursors, where KIN10 stabilizes SPCH by phosphorylation (Han et al. 2020). Therefore, sucrose promotes stomatal initiation. The molecules hydrogen peroxide (H2O2), nitric oxide (NO), carbon monoxide (CO) and hydrogen sulfide (H2S) have also been found to be involved in stomatal development. H2O2 is enriched in meristemoids and promotes stomatal development by inducing the nuclear localization of KIN10 (Shi et al. 2022). NO restricts GMC division by modulating the accumulation of aminocyclopropane-1-carboxylic acid (ACC), a ethylene precursor promoting the symmetric division of GMCs (Zhou et al. 2023; Yin et al. 2019). CO positively regulates stomatal formation by inhibiting EPF2 expression while H2S functions downstream of jasmonate signaling and upstream of SPCH to repress stomatal initiation (Weng et al. 2022; Deng et al. 2020).
It has been found that MicroRNAs (miRNAs) are involved in stomatal development and stomatal lineage cells express a large number of miRNAs (Kutter et al. 2007; Yang et al. 2014a; Zhu et al. 2020). However, their targets remain largely unknown. Stomatal pores are also main gates of plant for resisting pathogen invasion. Pathogens can release effectors to influence stomatal development for facilitating their invasion. For example, the pathogen Pseudomonas syringae releases the effector HopA1 which specifically inactivates MPK3/6, resulting in stomatal clustering (Kim et al. 2012; Zhang et al. 2007). The pathogen P. syringae pv. tomato (Pst), as well, releases two effector proteins, AvrPto and AvrPtoB, which disrupts the stomatal receptor SERKs, leading to defective stomatal development (Meng et al. 2015).
Compared with Arabidopsis, stomatal complexes in grasses are more sophisticated in composition and formation and very little is known about the mechanisms underlying their development. It has been found that some key stomatal regulators have conserved functions and some also obtained expanded roles. OsSPCHs, OsSCRMs, OsSHRs, OsSCRs, BdSPCHs or BdSCRM functions in establishing the fate of stomatal file. OsSPCHs, OsSCRMs, BdSPCHs or BdSCRM initiates the asymmetric cell divisions within stomatal file, producing early GMCs (Raissig et al. 2016; Wu et al. 2019). OsMUTE/ZmMUTE determines the GMC fate in rice or maize (Wang et al. 2019; Wu et al. 2019). Moreover, OsMUTE/ZmMUTE/BdMUTE which is specifically expressed in GMCs can move into lateral SMCs to induce SCs formation (Wang et al. 2019; Raissig et al. 2017). BdFAMA/OsFAMA and BdSCRM2 are found to control the GC specification in Brachypodium or rice (McKown et al. 2023; Liu et al. 2009; Wu et al. 2019). BdFAMA also functions redundantly with BdMUTE to promote GMC fate in Brachypodium (McKown et al. 2023). Additionally, OsFLP is involved in GC development (Wu et al. 2019; Zhang et al. 2022). It has been reported that BdYDA promotes correct stomatal spacing patterns and that the homologues of EPF1/2 and STOMAGEN in rice, barely, wheat and Brachypodium are involved in stomatal development (Lu et al. 2019; Jangra et al. 2021; Caine et al. 2019; Hughes et al. 2017; Abrash et al. 2018). However, whether they function via the mechanism similar to that of their Arabidopsis homologues is unknown.
Mesophyll originated signals regulate stomatal development
STOMAGEN, also named EPFL9, is a small peptide secreted by mesophyll cells and positively regulates stomatal development on the epidermis (Hunt et al. 2010; Kondo et al. 2010; Sugano et al. 2010). STOMAGEN competes with EPF2 for binding the ER-TMM receptors, which does not activate the downstream YDA MAPK cascade (Lee et al. 2015; Jewaria et al. 2013). Therefore, STOMAGEN promotes stomatal production by preventing the inhibitory effects of EPF2. To date, there is no other mesophyll cells secreted peptide like STOMAGEN has been identified, and STOMAGEN acts as a central mobile signaling molecule for integrating light and auxin signals in mesophyll to modulate stomatal development on the epidermis (Fig. 2A and Fig. 3). Light irradiation increases the accumulation of the bZIP transcription factor ELONGATED HYPOCOTYL 5 (HY5), a central light signal regulator promoting plant photomorphogenesis, by inactivation of COP1 which mediates HY5 ubiquitination and degradation. HY5 can directly bind to the promoter of STOMAGEN and promote the expression of STOMAGEN (Wang et al. 2021). Therefore, light signals also promote stomatal formation by triggering HY5 mediated activation of STOMAGEN transcription in mesophyll cells. In contrast, the expression of STOMAGEN is directly repressed by auxin response factor 5 (ARF5)/MONOPTEROS (MP), a key transcription factor of auxin signaling. Auxin binds to its receptors and subsequent triggers the ubiquitination and degradation of AUXIN/INDOLE-3-ACETIC ACID (AUX/IAAs) proteins, relieving the suppression of ARF5 by AUX/IAAs. Thus, auxin can also inhibit stomatal development by activating ARF5 in the mesophyll (Zhang et al. 2014).
Transcription factor ANGUSTIFOLIA3 (AN3) is another factor that enables mesophyll cells to modulate stomatal production on the epidermis (Fig. 3). AN3 is expressed in mesophyll cells but its protein AN3 moves into epidermal cells where it promotes stomatal production by directly inhibiting the expression of YDA (Kawade et al. 2013; Meng and Yao 2015). The expression of AN3 and the accumulation of AN3 protein are induced by light, and AN3 directly represses COP1expression (Meng et al. 2018), suggesting that AN3 is also a key point linking light signaling to stomatal development.
Endodermis originated signals regulate stomatal development in hypocotyls and stem
Mutations in the receptor TMM not only cause excess clustered stomata in leaves, but also confer the elimination of stomata from hypocotyls and stems (Yang and Sack 1995; Bhave et al. 2009), highlighting tissue specific regulation of stomatal development. Through a genetic screen of suppressors that could produce stomata in tmm background, the challah-1 (chal-1) mutation was identified (Abrash and Bergmann 2010). CHAL encodes EPFL6 and is specifically expressed in the endodermis of hypocotyls and inflorescence stem (Abrash and Bergmann 2010; Uchida et al. 2012). EPFL6 is secreted into epidermal cells to inhibit stomatal development by activating ERf receptors (Fig. 2A and Fig. 3). TMM dampens EPFL6 signaling (Abrash and Bergmann 2010; Abrash et al. 2011). Therefore, EPFL6 can also integrate both developmental and environmental cues in the inner tissues to modulate stomatal production on the epidermis.
Vein originated signals regulate stomatal development
Vein originated signals have long been hypothesized to be responsible for coordination of stomatal development with vein development, because, in grasses, stomata are not produced on the epidermis above veins but formed in cell files adjacent to veins. Vein may secrete an inhibitory signal to the epidermis overlying it or secrete an inductive signal to epidermal cell files that normally produce stomata (Hernandez et al. 1999). The SHORTROOT (SHR) protein which moves across cell layers in root is a possible candidate mediating layer communication between vein and epidermis (Fig. 4). In Arabidopsis, SHR moves from root stele into endodermis where it functions together with SCARECROW (SCR) to determine endodermis formation (Nakajima et al. 2001). In maize, ZmSHR moves at least eight cell layers from root endodermis into the cortex, controlling the formation of multi cortical layers (Ortiz-Ramírez et al. 2021). In the legume Medicago truncatula, MtSHR moves from hairy root stele into endodermis, cortex and even epidermis, regulating cortical cell division for nodulation (Dong et al. 2021). Interestingly, OsSHR is specifically expressed in leaf vasculature and its mutation greatly represses stomatal production (Wu et al. 2019; Liu et al. 2023; Schuler et al. 2018). In addition, OsSCR is specifically expressed in stomatal lineage cells and its mutation also sharply decreased stomatal density (Kamiya et al. 2003; Wu et al. 2019; Hughes and Langdale 2022). Moreover, over-expressing OsSHR or ZmSHR1 in rice produces extra stomatal files in more distant positions from the vein (Schuler et al. 2018; Liu et al. 2023). To date, it is not known whether SHR protein can move between grass leaf tissue layers, however, two possible mechanisms by which SHR regulates stomatal development in grasses have been proposed according to above findings. Firstly, grass SHR may move from vein into epidermis where it interacts with grass SCR to control stomatal development. Secondly, grass SHR may activate the expression of certain mobile factors such as small secreted peptides in veins to inhibit stomatal file formation on the epidermis immediately above it or induce stomatal file formation on the epidermis adjacent it (Schuler et al. 2018).
Long-distance signaling mediates stomatal development
Long-distance signaling between different plant organs enables plants to systematically adapt to the ever changing environment (Luo et al. 2022). For example, plant shoot, which uses light energy and CO2 for photosynthesis to obtain the energy required for plant growth and development, and plant root, which absorbs water and mineral nutrients from the soil, communicate broadly through long-distance signaling, facilitating plants to finally achieve overall coordination and better survival (Yang and Liu 2020). A number of mobile signals, mainly including phytohormones, RNAs, peptides and proteins, have been shown to move in vascular system which can permit proteins with a molecular weight that does not exceed 67 kDa to pass through (Stadler et al. 2005; Kitagawa et al. 2023; Song et al. 2021). They regulate several plant processes such as flowering, nodulation, stomatal closure, branching, pathogen responses, potato tuber formation and root development (Ryan and Pearce 2003; Yoo et al. 2004; Hilleary and Gilroy 2018; Lough and Lucas 2006; Oka-Kira and Kawaguchi 2006; Corbesier et al. 2007; Okamoto et al. 2016; Tsikou et al. 2018; Navarro et al. 2011).
It has been found that preexisting leaves can sense environmental cues, such as high light intensity and high CO2 concentrations, and generate long-distance signals to influence stomatal development in young leaves (Miyazawa et al. 2006). However, these long-distance signals remain elusive. It has been reported that light enhances the accumulation of HY5 which moves from mesophyll to root through the vascular system to regulate the elongation of the root system (van Gelderen et al. 2018; Chen et al. 2016b), suggesting that HY5 may act as a long-distance signal promoting stomatal development (Fig. 5). Other possible long-distance signals induced by light are STOMAGEN and AN3 which have been shown to move from mesophyll to epidermis to promote stomatal production (Kawade et al. 2013; Sugano et al. 2010). High CO2 concentrations induce CRSP mediated EPF2 maturation (Hara et al. 2009; Hunt and Gray 2009; Engineer et al. 2014), suggesting that CRSP and EPF2 may be candidates of long-distance signals inhibiting stomatal development> (Fig. 5). In addition, plants root can absorb phytohormones such as auxin, CK, BR, ABA and Jasmonate that are added in the Murashige and Skoog medium and transmit them to leaf epidermis to regulate stomatal formation (Le et al. 2014a, b; Zhang et al. 2014; Vaten et al. 2018; Gudesblat et al. 2012; Kim et al. 2012; Han et al. 2018b; Yang et al. 2022), indicating that these phytohormones may also act as long-distance signals (Fig. 5). Recently, glucose has been proved to be sensed in mature leaves to modulate stomatal development within young leaves through long-distance signaling (Bao et al. 2023) (Fig. 5).
Evidence is accumulating that long-distance movement of CLE small peptides plays an essential role in plant intercellular communication and abiotic stress response (Song et al. 2021). Water shortage induces the secretion of small peptide CLE25 by the root vascular tissue. CLE25 moves from the root to the leaf through the vascular system, and is further recognized by its receptors BARELYANY MERISTEM1 (BAM1) and BAM3, finally leading to the increase of ABA accumulation in leaves and thus stomatal closure (Takahashi et al. 2018). Interestingly, CLE25 can inhibit stomatal precursor MMC formation (Qian et al. 2018), suggesting that CLE25 is also a long-distance signal mediating stomatal development (Fig. 5). Other CLE small peptides including CLE9/10, CLE11, CLE12, CLE13, CLE14, CLE8, CLE45, CLE20, CLE26 and CLE2 can also repress MMC formation (Qian et al. 2018), suggesting that they are also possible candidates for long-distance signals involved in stoamtal production (Fig. 5).
Conclusion and perspectives
Stomatal development is tightly coordinated with the development of other plant tissues and organs. Increasing discoveries reveal the presence of stomatal regulators that can move within the epidermal layer or across tissue layers and organs, thus now making us began to understand stomatal development in 3D. The cell–cell communications within epidermis ensure the proper stomatal patterning, density and stomatal complex formation. However, in most cases, the exact regulatory mechanisms of those possible cell–cell communications remain unclear. For example, the small peptides EPF2 and EPF1 perceived by TMM-ERf-SERKs receptors mediate communications between stomatal lineage cells by acting as both autocrine and paracrine factors, however, how they contribute to establish asymmetric daughter cell fates is largely unknown, although it has been reported that BREAKING OF ASYMMETRY IN THE STOMATALLINEAGE (BASL), which is essential for the intrinsic polarity of meristemoid cells and asymmetric daughter cell fates, is improperly polarized in SLGCs in the absence of EPF1 (Dong et al. 2009). The small peptides CLE9 and CLE10 perceived by HSL1-SERKs receptors function parallelly with EPFs in repressing SPCH activity through YDA MAPK cascade, however, it is unknown about their concrete roles in stomatal cell–cell communications. To date, almost all studies focused the local roles of plant hormones and environmental cues in stomatal patterning and differentiation. It has not been proved clearly that plant hormones function as a long-distance signal and that which molecules could act as a long-distance signal for integrating the environmental stimulus from other organs to affect stomatal development. In grasses, close communication and coordination between the stomatal file, which produces paired GCs, and its bilateral cell files, which produce SCs, are crucial for the final formation of four-celled stomatal complex. To date, the mobile transcription factor MUTE is the only molecular proved to function in this process. The receptor-like proteins pangloss 1 (PAN1) and PAN2 which exhibit a polarized localization in SMCs at sites of GMC contact have long been proposed to recognize GMC-derived ligands for promoting asymmetric division of SMC, however, this is still an intriguing hypothesis to pursue.
The cell–cell communications between cell layers coordinate stomatal development with the differentiation of leaf mesophyll, vein and stem cortex for gas exchange and photosynthesis, and the cell–cell communications through long-distance signaling facilitate stomatal development to systematically response to the ever changing environment. However, compared with stomatal signals originated within the epidermis, few stomatal signals originated from the internal tissues and long distances have been identified with the exception of EPFL6, STOMAGEN and AN3. Whether grass SHR could move from vein to epidermis to modulate stomatal development, and whether transcription factors (e.g. HY5 and AN3), peptides (e.g. EPFs and CLEs), phytohormones (e.g. auxin, CK, BR, ABA and Jasmonate), and microRNAs could act as long-distance stomatal signals remain to be seen. With the advent of new technology, such as single-cell RNA sequencing and spatial transcriptomics, more and more spatiotemporal communications between cells can be visualized at high resolution, which will facilitate a deeper understanding about the regulation of stomatal development in 3D. Through manipulation of this 3D network of stomatal development, the stomatal density and patterns can be adjusted for creating “water-saver” crops in a predicted warmer future without sufficient water for agriculture.
Data availability
The original contributions in the study are included in the article. Further inquiries can be directed to the corresponding author.
References
Abrash E, Anleu Gil MX, Matos JL, Bergmann DC (2018) Conservation and divergence of YODA MAPKKK function in regulation of grass epidermal patterning. Development 145(14):165860
Abrash EB, Bergmann DC (2010) Regional specification of stomatal production by the putative ligand CHALLAH. Development 137(3):447–455
Abrash EB, Davies KA, Bergmann DC (2011) Generation of signaling specificity in Arabidopsis by spatially restricted buffering of ligand-receptor interactions. Plant Cell 23(8):2864–2879
Adrian J, Chang J, Ballenger CE, Bargmann BO, Alassimone J, Davies KA, Lau OS, Matos JL, Hachez C, Lanctot A, Vaten A, Birnbaum KD, Bergmann DC (2015) Transcriptome dynamics of the stomatal lineage: birth, amplification, and termination of a self-renewing population. Dev Cell 33(1):107–118
Balcerowicz M, Ranjan A, Rupprecht L, Fiene G, Hoecker U (2014) Auxin represses stomatal development in dark-grown seedlings via Aux/IAA proteins. Development 141(16):3165–3176
Bao QX, Mu XR, Tong C, Li C, Tao WZ, Zhao ST, Liu YX, Wang WN, Wei YT, Yu FH, Wang JW, Sun ZL, Fan BL, Sun J, Wang C, Loake GJ, Meng LS (2023) Sugar status in preexisting leaves determines systemic stomatal development within newly developing leaves. Proc Natl Acad Sci USA 120(24):e2302854120
Becraft PW (1999) Development of the leaf epidermis. Curr Top Dev Biol 45:1–40
Bergmann DC, Lukowitz W, Somerville CR (2004) Stomatal development and pattern controlled by a MAPKK kinase. Science 304(5676):1494–1497
Bergmann DC, Sack FD (2007) Stomatal development. Annu Rev Plant Biol 58:163–181
Bhave NS, Veley KM, Nadeau JA, Lucas JR, Bhave SL, Sack FD (2009) TOO MANY MOUTHS promotes cell fate progression in stomatal development of Arabidopsis stems. Planta 229(2):357–367
Boudolf V, Barrôco R, Engler Jde A, Verkest A, Beeckman T, Naudts M, Inzé D, De Veylder L (2004) B1-type cyclin-dependent kinases are essential for the formation of stomatal complexes in Arabidopsis thaliana. Plant Cell 16(4):945–955
Caine RS, Yin X, Sloan J, Harrison EL, Mohammed U, Fulton T, Biswal AK, Dionora J, Chater CC, Coe RA, Bandyopadhyay A, Murchie EH, Swarup R, Quick WP, Gray JE (2019) Rice with reduced stomatal density conserves water and has improved drought tolerance under future climate conditions. New Phytol 221(1):371–384
Chen L (2023) Emerging roles of protein phosphorylation in regulation of stomatal development. J Plant Physiol 280:153882
Chen L, Guan L, Qian P, Xu F, Wu Z, Wu Y, He K, Gou X, Li J, Hou S (2016a) NRPB3, the third largest subunit of RNA polymerase II, is essential for stomatal patterning and differentiation in Arabidopsis. Development 143(9):1600–1611
Chen L, Wu Z, Hou S (2020) SPEECHLESS speaks loudly in stomatal development. Front Plant Sci 11:114
Chen L, Zhao M, Wu Z, Chen S, Rojo E, Luo J, Li P, Zhao L, Chen Y, Deng J, Cheng B, He K, Gou X, Li J, Hou S (2021) RNA polymerase II associated proteins regulate stomatal development through direct interaction with stomatal transcription factors in Arabidopsis thaliana. New Phytol 230(1):171–189
Chen X, Yao Q, Gao X, Jiang C, Harberd NP, Fu X (2016b) Shoot-to-root mobile transcription factor HY5 coordinates plant carbon and nitrogen acquisition. Curr Biol 26(5):640–646
Corbesier L, Vincent C, Jang S, Fornara F, Fan Q, Searle I, Giakountis A, Farrona S, Gissot L, Turnbull C, Coupland G (2007) FT protein movement contributes to long-distance signaling in floral induction of Arabidopsis. Science 316(5827):1030–1033
Cutler SR, Rodriguez PL, Finkelstein RR, Abrams SR (2010) Abscisic acid: emergence of a core signaling network. Annu Rev Plant Biol 61:651–679
Deng G, Zhou L, Wang Y, Zhang G, Chen X (2020) Hydrogen sulfide acts downstream of jasmonic acid to inhibit stomatal development in Arabidopsis. Planta 251(2):42
Dong J, MacAlister CA, Bergmann DC (2009) BASL controls asymmetric cell division in Arabidopsis. Cell 137(7):1320–1330
Dong W, Zhu Y, Chang H, Wang C, Yang J, Shi J, Gao J, Yang W, Lan L, Wang Y, Zhang X, Dai H, Miao Y, Xu L, He Z, Song C, Wu S, Wang D, Yu N, Wang E (2021) An SHR-SCR module specifies legume cortical cell fate to enable nodulation. Nature 589(7843):586–590
Engineer CB, Ghassemian M, Anderson JC, Peck SC, Hu H, Schroeder JI (2014) Carbonic anhydrases, EPF2 and a novel protease mediate CO2 control of stomatal development. Nature 513(7517):246–250
Gray JE, Hetherington AM (2004) Plant development: YODA the stomatal switch. Curr Biol 14(12):R488-490
Gudesblat GE, Schneider-Pizon J, Betti C, Mayerhofer J, Vanhoutte I, van Dongen W, Boeren S, Zhiponova M, de Vries S, Jonak C, Russinova E (2012) SPEECHLESS integrates brassinosteroid and stomata signalling pathways. Nat Cell Biol 14(5):548–554
Guo X, Park CH, Wang ZY, Nickels BE, Dong J (2021) A spatiotemporal molecular switch governs plant asymmetric cell division. Nat Plants 7(5):667–680
Han C, Liu Y, Shi W, Qiao Y, Wang L, Tian Y, Fan M, Deng Z, Lau OS, De Jaeger G, Bai M-Y (2020) KIN10 promotes stomatal development through stabilization of the SPEECHLESS transcription factor. Nat Commun 11(1):4214
Han SK, Herrmann A, Yang J, Iwasaki R, Sakamoto T, Desvoyes B, Kimura S, Gutierrez C, Kim ED, Torii KU (2022) Deceleration of the cell cycle underpins a switch from proliferative to terminal divisions in plant stomatal lineage. Dev Cell 57(5):569–582
Han SK, Kwak JM, Qi X (2021) Stomatal lineage control by developmental program and environmental cues. Front Plant Sci 12:751852
Han SK, Qi X, Sugihara K, Dang JH, Endo TA, Miller KL, Kim ED, Miura T, Torii KU (2018a) MUTE directly orchestrates cell-state switch and the single symmetric division to create stomata. Dev Cell 45(3):303–315
Han SK, Torii KU (2016) Lineage-specific stem cells, signals and asymmetries during stomatal development. Development 143(8):1259–1270
Han SK, Torii KU (2019) Linking cell cycle to stomatal differentiation. Curr Opin Plant Biol 51:66–73
Han X, Hu Y, Zhang G, Jiang Y, Chen X, Yu D (2018b) Jasmonate Negatively Regulates Stomatal Development in Arabidopsis Cotyledons. Plant Physiol 176(4):2871–2885
Hara K, Kajita R, Torii KU, Bergmann DC, Kakimoto T (2007) The secretory peptide gene EPF1 enforces the stomatal one-cell-spacing rule. Genes Dev 21(14):1720–1725
Hara K, Yokoo T, Kajita R, Onishi T, Yahata S, Peterson KM, Torii KU, Kakimoto T (2009) Epidermal cell density is autoregulated via a secretory peptide, EPIDERMAL PATTERNING FACTOR 2 in Arabidopsis leaves. Plant Cell Physiol 50(6):1019–1031
Hernandez ML, Passas HJ, Smith LG (1999) Clonal analysis of epidermal patterning during maize leaf development. Dev Biol 216(2):646–658
Hilleary R, Gilroy S (2018) Systemic signaling in response to wounding and pathogens. Curr Opin Plant Biol 43:57–62
Horst RJ, Fujita H, Lee JS, Rychel AL, Garrick JM, Kawaguchi M, Peterson KM, Torii KU (2015) Molecular framework of a regulatory circuit initiating two-dimensional spatial patterning of stomatal lineage. PLoS Genet 11(7):e1005374
Houbaert A, Zhang C, Tiwari M, Wang K, de Marcos SA, Savatin DV, Urs MJ, Zhiponova MK, Gudesblat GE, Vanhoutte I, Eeckhout D, Boeren S, Karimi M, Betti C, Jacobs T, Fenoll C, Mena M, de Vries S, De Jaeger G, Russinova E (2018) POLAR-guided signalling complex assembly and localization drive asymmetric cell division. Nature 563(7732):574–578
Hughes J, Hepworth C, Dutton C, Dunn JA, Hunt L, Stephens J, Waugh R, Cameron DD, Gray JE (2017) Reducing stomatal density in barley improves drought tolerance without impacting on yield. Plant Physiol 174(2):776–787
Hughes TE, Langdale JA (2022) SCARECROW is deployed in distinct contexts during rice and maize leaf development. Development 149(7):dev200410
Hunt L, Bailey KJ, Gray JE (2010) The signalling peptide EPFL9 is a positive regulator of stomatal development. New Phytol 186(3):609–614
Hunt L, Gray JE (2009) The signaling peptide EPF2 controls asymmetric cell divisions during stomatal development. Curr Biol 19(10):864–869
Jangra R, Brunetti SC, Wang X, Kaushik P, Gulick PJ, Foroud NA, Wang S, Lee JS (2021) Duplicated antagonistic EPF peptides optimize grass stomatal initiation. Development 148(16):dev199780
Jewaria PK, Hara T, Tanaka H, Kondo T, Betsuyaku S, Sawa S, Sakagami Y, Aimoto S, Kakimoto T (2013) Differential effects of the peptides Stomagen, EPF1 and EPF2 on activation of MAP kinase MPK6 and the SPCH protein level. Plant Cell Physiol 54(8):1253–1262
Kamiya N, Itoh J, Morikami A, Nagato Y, Matsuoka M (2003) The SCARECROW gene’s role in asymmetric cell divisions in rice plants. Plant J 36(1):45–54
Kanaoka MM, Pillitteri LJ, Fujii H, Yoshida Y, Bogenschutz NL, Takabayashi J, Zhu JK, Torii KU (2008) SCREAM/ICE1 and SCREAM2 specify three cell-state transitional steps leading to Arabidopsis stomatal differentiation. Plant Cell 20(7):1775–1785
Kang CY, Lian HL, Wang FF, Huang JR, Yang HQ (2009) Cryptochromes, phytochromes, and COP1 regulate light-controlled stomatal development in Arabidopsis. Plant Cell 21(9):2624–2641
Kawade K, Horiguchi G, Usami T, Hirai MY, Tsukaya H (2013) ANGUSTIFOLIA3 signaling coordinates proliferation between clonally distinct cells in leaves. Curr Biol 23(9):788–792
Khan M, Rozhon W, Bigeard J, Pflieger D, Husar S, Pitzschke A, Teige M, Jonak C, Hirt H, Poppenberger B (2013) Brassinosteroid-regulated GSK3/Shaggy-like kinases phosphorylate mitogen-activated protein (MAP) kinase kinases, which control stomata development in Arabidopsis thaliana. J Biol Chem 288(11):7519–7527
Kim ED, Dorrity MW, Fitzgerald BA, Seo H, Sepuru KM, Queitsch C, Mitsuda N, Han SK, Torii KU (2022) Dynamic chromatin accessibility deploys heterotypic cis/trans-acting factors driving stomatal cell-fate commitment. Nat Plants 8(12):1453–1466
Kim TW, Michniewicz M, Bergmann DC, Wang ZY (2012) Brassinosteroid regulates stomatal development by GSK3-mediated inhibition of a MAPK pathway. Nature 482(7385):419–422
Kitagawa M, Tran TM, Jackson D (2023) Traveling with purpose: cell-to-cell transport of plant mRNAs. Trends Cell Biol 34(1):48–57
Klermund C, Ranftl QL, Diener J, Bastakis E, Richter R, Schwechheimer C (2016) LLM-domain B-GATA transcription factors promote stomatal development downstream of light signaling pathways in Arabidopsis thaliana hypocotyls. Plant Cell 28(3):646–660
Kondo T, Kajita R, Miyazaki A, Hokoyama M, Nakamura-Miura T, Mizuno S, Masuda Y, Irie K, Tanaka Y, Takada S, Kakimoto T, Sakagami Y (2010) Stomatal density is controlled by a mesophyll-derived signaling molecule. Plant Cell Physiol 51(1):1–8
Kumari A, Jewaria PK, Bergmann DC, Kakimoto T (2014) Arabidopsis reduces growth under osmotic stress by decreasing SPEECHLESS protein. Plant Cell Physiol 55(12):2037–2046
Kutter C, Schob H, Stadler M, Meins F Jr, Si-Ammour A (2007) MicroRNA-mediated regulation of stomatal development in Arabidopsis. Plant Cell 19(8):2417–2429
Lai LB, Nadeau JA, Lucas J, Lee EK, Nakagawa T, Zhao L, Geisler M, Sack FD (2005) The Arabidopsis R2R3 MYB proteins FOUR LIPS and MYB88 restrict divisions late in the stomatal cell lineage. Plant Cell 17(10):2754–2767
Lampard GR, Lukowitz W, Ellis BE, Bergmann DC (2009) Novel and expanded roles for MAPK signaling in Arabidopsis stomatal cell fate revealed by cell type-specific manipulations. Plant Cell 21(11):3506–3517
Lampard GR, Macalister CA, Bergmann DC (2008) Arabidopsis stomatal initiation is controlled by MAPK-mediated regulation of the bHLH SPEECHLESS. Science 322(5904):1113–1116
Lau OS, Bergmann DC (2012) Stomatal development: a plant’s perspective on cell polarity, cell fate transitions and intercellular communication. Development 139(20):3683–3692
Lau OS, Davies KA, Chang J, Adrian J, Rowe MH, Ballenger CE, Bergmann DC (2014) Direct roles of SPEECHLESS in the specification of stomatal self-renewing cells. Science 345(6204):1605–1609
Lau OS, Song Z, Zhou Z, Davies KA, Chang J, Yang X, Wang S, Lucyshyn D, Tay IHZ, Wigge PA, Bergmann DC (2018) Direct control of SPEECHLESS by PIF4 in the high-temperature response of stomatal development. Curr Biol 28(8):1273–1280
Le J, Liu XG, Yang KZ, Chen XL, Zou JJ, Wang HZ, Wang M, Vanneste S, Morita M, Tasaka M, Ding ZJ, Friml J, Beeckman T, Sack F (2014a) Auxin transport and activity regulate stomatal patterning and development. Nature Commun 5:3090
Le J, Zou J, Yang K, Wang M (2014b) Signaling to stomatal initiation and cell division. Front Plant Sci 5:297
Lee E, Lucas JR, Sack FD (2014) Deep functional redundancy between FAMA and FOUR LIPS in stomatal development. Plant J 78(4):555–565
Lee JH, Jung JH, Park CM (2017) Light inhibits COP1-mediated degradation of ICE transcription factors to induce stomatal development in Arabidopsis. Plant Cell 29(11):2817–2830
Lee JS, Hnilova M, Maes M, Lin YC, Putarjunan A, Han SK, Avila J, Torii KU (2015) Competitive binding of antagonistic peptides fine-tunes stomatal patterning. Nature 522(7557):439–443
Lee JS, Kuroha T, Hnilova M, Khatayevich D, Kanaoka MM, McAbee JM, Sarikaya M, Tamerler C, Torii KU (2012) Direct interaction of ligand-receptor pairs specifying stomatal patterning. Genes Dev 26(2):126–136
Lee LR, Wengier DL, Bergmann DC (2019) Cell-type-specific transcriptome and histone modification dynamics during cellular reprogramming in the Arabidopsis stomatal lineage. Proc Natl Acad Sci U S A 116(43):21914–21924
Li P, Chen L, Gu X, Zhao M, Wang W, Hou S (2023) FOUR LIPS plays a role in meristemoid-to-GMC fate transition during stomatal development in Arabidopsis. Plant J 114(2):424–436
Liu Q, Teng S, Deng C, Wu S, Li H, Wang Y, Wu J, Cui X, Zhang Z, Quick WP, Brutnell TP, Sun X, Lu T (2023) SHORT ROOT and INDETERMINATE DOMAIN family members govern PIN-FORMED expression to regulate minor vein differentiation in rice. Plant Cell 35(8):2848–2870
Liu T, Ohashi-Ito K, Bergmann DC (2009) Orthologs of Arabidopsis thaliana stomatal bHLH genes and regulation of stomatal development in grasses. Development 136(13):2265–2276
Liu Z, Zhou Y, Guo J, Li J, Tian Z, Zhu Z, Wang J, Wu R, Zhang B, Hu Y, Sun Y, Shangguan Y, Li W, Li T, Hu Y, Guo C, Rochaix JD, Miao Y, Sun X (2020) Global dynamic molecular profiling of stomatal lineage cell development by single-cell RNA sequencing. Mol Plant 13(8):1178–1193
Lough TJ, Lucas WJ (2006) Integrative plant biology: role of phloem long-distance macromolecular trafficking. Annu Rev Plant Biol 57:203–232
Lu J, He J, Zhou X, Zhong J, Li J, Liang YK (2019) Homologous genes of epidermal patterning factor regulate stomatal development in rice. J Plant Physiol 234–235:18–27
Luo XC, Liu H, Liu XY, Li XX, Liao H, Fu XD (2022) Long distance signal transduction in response to environmental changes in plants. Yi Chuan 44(7):556–566
MacAlister CA, Ohashi-Ito K, Bergmann DC (2007) Transcription factor control of asymmetric cell divisions that establish the stomatal lineage. Nature 445(7127):537–540
McKown KH, Anleu Gil MX, Mair A, Xu SL, Raissig MT, Bergmann DC (2023) Expanded roles and divergent regulation of FAMA in Brachypodium and Arabidopsis stomatal development. Plant Cell 35(2):756–775
McKown KH, Bergmann DC (2020) Stomatal development in the grasses: lessons from models and crops (and crop models). New Phytol 227(6):1636–1648
Meng LS, Li C, Xu MK, Sun XD, Wan W, Cao XY, Zhang JL, Chen KM (2018) Arabidopsis ANGUSTIFOLIA3 (AN3) is associated with the promoter of CONSTITUTIVE PHOTOMORPHOGENIC1 (COP1) to regulate light-mediated stomatal development. Plant Cell Environ 41(7):1645–1656
Meng LS, Yao SQ (2015) Transcription co-activator Arabidopsis ANGUSTIFOLIA3 (AN3) regulates water-use efficiency and drought tolerance by modulating stomatal density and improving root architecture by the transrepression of YODA (YDA). Plant Biotechol J 13(7):893–902
Meng X, Chen X, Mang H, Liu C, Yu X, Gao X, Torii KU, He P, Shan L (2015) Differential Function of Arabidopsis SERK Family Receptor-like Kinases in Stomatal Patterning. Curr Biol 25(18):2361–2372
Miyazawa S, Livingston NJ, Turpin DH (2006) Stomatal development in new leaves is related to the stomatal conductance of mature leaves in poplar (Populus trichocarpaxP. deltoides). J Exp Bot 57(2):373–380
Nadeau JA, Sack FD (2002) Control of stomatal distribution on the Arabidopsis leaf surface. Science 296(5573):1697–1700
Nakajima K, Sena G, Nawy T, Benfey PN (2001) Intercellular movement of the putative transcription factor SHR in root patterning. Nature 413(6853):307–311
Navarro C, Abelenda JA, Cruz-Oró E, Cuéllar CA, Tamaki S, Silva J, Shimamoto K, Prat S (2011) Control of flowering and storage organ formation in potato by FLOWERING LOCUS T. Nature 478(7367):119–122
Nunes TDG, Zhang D, Raissig MT (2020) Form, development and function of grass stomata. Plant J 101(4):780–799
Ohashi-Ito K, Bergmann DC (2006) Arabidopsis FAMA controls the final proliferation/differentiation switch during stomatal development. Plant Cell 18(10):2493–2505
Oka-Kira E, Kawaguchi M (2006) Long-distance signaling to control root nodule number. Curr Opin Plant Biol 9(5):496–502
Okamoto S, Tabata R, Matsubayashi Y (2016) Long-distance peptide signaling essential for nutrient homeostasis in plants. Curr Opin Plant Biol 34:35–40
Ortega A, de Marcos A, Illescas-Miranda J, Mena M, Fenoll C (2019) The tomato genome encodes SPCH, MUTE, and FAMA candidates that can replace the endogenous functions of their Arabidopsis orthologs. Front Plant Sci 10:1300
Ortiz-Ramírez C, Guillotin B, Xu X, Rahni R, Zhang S, Yan Z, Coqueiro Dias Araujo P, Demesa-Arevalo E, Lee L, Van Eck J, Gingeras TR, Jackson D, Gallagher KL, Birnbaum KD (2021) Ground tissue circuitry regulates organ complexity in maize and Setaria. Science 374(6572):1247–1252
Pillitteri LJ, Dong J (2013) Stomatal development in Arabidopsis. The Arabidopsis Book / American Society of Plant Biologists 11:e0162
Pillitteri LJ, Sloan DB, Bogenschutz NL, Torii KU (2007) Termination of asymmetric cell division and differentiation of stomata. Nature 445(7127):501–505
Pillitteri LJ, Torii KU (2012) Mechanisms of stomatal development. Annu Rev Plant Biol 63:591–614
Qi X, Han SK, Dang JH, Garrick JM, Ito M, Hofstetter AK, Torii KU (2017) Autocrine regulation of stomatal differentiation potential by EPF1 and ERECTA-LIKE1 ligand-receptor signaling. Elife 7(6):24102
Qian P, Song W, Yokoo T, Minobe A, Wang G, Ishida T, Sawa S, Chai J, Kakimoto T (2018) The CLE9/10 secretory peptide regulates stomatal and vascular development through distinct receptors. Nat Plants 4(12):1071–1081
Raissig MT, Abrash E, Bettadapur A, Vogel JP, Bergmann DC (2016) Grasses use an alternatively wired bHLH transcription factor network to establish stomatal identity. Proc Natl Acad Sci USA 113(29):8326–8331
Raissig MT, Matos JL, Anleu Gil MX, Kornfeld A, Bettadapur A, Abrash E, Allison HR, Badgley G, Vogel JP, Berry JA, Bergmann DC (2017) Mobile MUTE specifies subsidiary cells to build physiologically improved grass stomata. Science 355(6330):1215–1218
Ryan CA, Pearce G (2003) Systemins: a functionally defined family of peptide signals that regulate defensive genes in Solanaceae species. Proc Natl Acad Sci USA 100(Suppl 2):14577–14580
Samakovli D, Tichá T, Vavrdová T, Ovečka M, Luptovčiak I, Zapletalová V, Kuchařová A, Křenek P, Krasylenko Y, Margaritopoulou T, Roka L, Milioni D, Komis G, Hatzopoulos P, Šamaj J (2020) YODA-HSP90 Module Regulates Phosphorylation-dependent inactivation of SPEECHLESS to control stomatal development under acute heat stress in Arabidopsis. Mol Plant 13(4):612–633
Schuler ML, Sedelnikova OV, Walker BJ, Westhoff P, Langdale JA (2018) SHORTROOT-mediated increase in stomatal density has no impact on photosynthetic efficiency. Plant Physiol 176(1):757–772
Shi W, Wang L, Yao L, Hao W, Han C, Fan M, Wang W, Bai MY (2022) Spatially patterned hydrogen peroxide orchestrates stomatal development in Arabidopsis. Nat Commun 13(1):022–32770
Shpak ED, McAbee JM, Pillitteri LJ, Torii KU (2005) Stomatal patterning and differentiation by synergistic interactions of receptor kinases. Science 309(5732):290–293
Smit ME, Bergmann DC (2023) The stomatal fates: Understanding initiation and enforcement of stomatal cell fate transitions. Curr Opin Plant Biol 76:102449
Song XF, Hou XL, Liu CM (2021) CLE peptides: critical regulators for stem cell maintenance in plants. Planta 255(1):5
Stadler R, Wright KM, Lauterbach C, Amon G, Gahrtz M, Feuerstein A, Oparka KJ, Sauer N (2005) Expression of GFP-fusions in Arabidopsis companion cells reveals non-specific protein trafficking into sieve elements and identifies a novel post-phloem domain in roots. Plant J 41(2):319–331
Sugano SS, Shimada T, Imai Y, Okawa K, Tamai A, Mori M, Hara-Nishimura I (2010) Stomagen positively regulates stomatal density in Arabidopsis. Nature 463(7278):241–244
Takahashi F, Suzuki T, Osakabe Y, Betsuyaku S, Kondo Y, Dohmae N, Fukuda H, Yamaguchi-Shinozaki K, Shinozaki K (2018) A small peptide modulates stomatal control via abscisic acid in long-distance signalling. Nature 556(7700):235–238
Tsikou D, Yan Z, Holt DB, Abel NB, Reid DE, Madsen LH, Bhasin H, Sexauer M, Stougaard J, Markmann K (2018) Systemic control of legume susceptibility to rhizobial infection by a mobile microRNA. Science 362(6411):233–236
Uchida N, Lee JS, Horst RJ, Lai HH, Kajita R, Kakimoto T, Tasaka M, Torii KU (2012) Regulation of inflorescence architecture by intertissue layer ligand-receptor communication between endodermis and phloem. Proc Natl Acad Sci USA 109(16):6337–6342
Van Gelderen K, Kang C, Paalman R, Keuskamp D, Hayes S, Pierik R (2018) Far-red light detection in the shoot regulates lateral root development through the HY5 transcription factor. Plant Cell 30(1):101–116
Vanneste S, Coppens F, Lee E, Donner TJ, Xie Z, Van Isterdael G, Dhondt S, De Winter F, De Rybel B, Vuylsteke M, De Veylder L, Friml J, Inzé D, Grotewold E, Scarpella E, Sack F, Beemster GT, Beeckman T (2011) Developmental regulation of CYCA2s contributes to tissue-specific proliferation in Arabidopsis. EMBO J 30(16):3430–3441
Vaten A, Soyars CL, Tarr PT, Nimchuk ZL, Bergmann DC (2018) Modulation of asymmetric division diversity through cytokinin and SPEECHLESS regulatory interactions in the Arabidopsis stomatal lineage. Dev Cell 47(1):53–66
Wallner ES, Mair A, Handler D, McWhite C, Xu SL, Dolan L, Bergmann DC (2024) Spatially resolved proteomics of the Arabidopsis stomatal lineage identifies polarity complexes for cell divisions and stomatal pores. Dev Cell S1534–5807(24):00141–00142. https://doi.org/10.1016/j.devcel.2024.03.001
Wang H, Guo S, Qiao X, Guo J, Li Z, Zhou Y, Bai S, Gao Z, Wang D, Wang P, Galbraith DW, Song CP (2019) BZU2/ZmMUTE controls symmetrical division of guard mother cell and specifies neighbor cell fate in maize. PLoS Genet 15(8):e1008377
Wang H, Ngwenyama N, Liu Y, Walker JC, Zhang S (2007) Stomatal development and patterning are regulated by environmentally responsive mitogen-activated protein kinases in Arabidopsis. Plant Cell 19(1):63–73
Wang M, Yang K, Le J (2015) Organ-specific effects of brassinosteroids on stomatal production coordinate with the action of Too Many Mouths. J Integr Plant Biol 57(3):247–255
Wang S, Zhou Z, Rahiman R, Lee GSY, Yeo YK, Yang X, Lau OS (2021) Light regulates stomatal development by modulating paracrine signaling from inner tissues. Nature Commun 12(1):3403
Weimer AK, Matos JL, Sharma N, Patell F, Murray JAH, Dewitte W, Bergmann DC (2018) Lineage- and stage-specific expressed CYCD7;1 coordinates the single symmetric division that creates stomatal guard cells. Development 145(6):dev160671
Weng X, Zhu L, Yu S, Liu Y, Ru Y, Zhang Z, He Z, Zhou L, Chen X (2022) Carbon monoxide promotes stomatal initiation by regulating the expression of two EPF genes in Arabidopsis cotyledons. Front Plant Sci 13:1029703
Wu Z, Chen L, Yu Q, Zhou W, Gou X, Li J, Hou S (2019) Multiple transcriptional factors control stomata development in rice. New Phytol 223(1):220–232
Yang K, Jiang M, Le J (2014a) A new loss-of-function allele 28y reveals a role of ARGONAUTE1 in limiting asymmetric division of stomatal lineage ground cell. J Integr Plant Biol 56(6):539–549
Yang K, Wang H, Xue S, Qu X, Zou J, Le J (2014b) Requirement for A-type cyclin-dependent kinase and cyclins for the terminal division in the stomatal lineage of Arabidopsis. J Exp Bot 65(9):2449–2461
Yang K, Zhu L, Wang H, Jiang M, Xiao C, Hu X, Vanneste S, Dong J, Le J (2019) A conserved but plant-specific CDK-mediated regulation of DNA replication protein A2 in the precise control of stomatal terminal division. Proc Natl Acad Sci USA 116(36):18126–18131
Yang M, Sack FD (1995) The too many mouths and four lips mutations affect stomatal production in Arabidopsis. Plant Cell 7(12):2227–2239
Yang X, Gavya SL, Zhou Z, Urano D, Lau OS (2022) Abscisic acid regulates stomatal production by imprinting a SnRK2 kinase-mediated phosphocode on the master regulator SPEECHLESS. Sci Adv 8(40):eadd2063
Yang Y, Liu H (2020) Coordinated shoot and root responses to light signaling in arabidopsis. Plant Commun 1(2):100026
Yin J, Zhang X, Zhang G, Wen Y, Liang G, Chen X (2019) Aminocyclopropane-1-carboxylic acid is a key regulator of guard mother cell terminal division in Arabidopsis thaliana. J Exp Bot 70(3):897–908
Yoo BC, Kragler F, Varkonyi-Gasic E, Haywood V, Archer-Evans S, Lee YM, Lough TJ, Lucas WJ (2004) A systemic small RNA signaling system in plants. Plant Cell 16(8):1979–2000
Zhang C, Zhang J, Liu H, Qu X, Wang J, He Q, Zou J, Yang K, Le J (2022) Transcriptomic analysis reveals the role of FOUR LIPS in response to salt stress in rice. Plant Mol Biol 110(1–2):37–52
Zhang J, Shao F, Li Y, Cui H, Chen L, Li H, Zou Y, Long C, Lan L, Chai J, Chen S, Tang X, Zhou J-M (2007) A Pseudomonas syringae effector inactivates MAPKs to suppress PAMP-induced immunity in plants. Cell Host Microbe 1(3):175–185
Zhang JY, He SB, Li L, Yang HQ (2014) Auxin inhibits stomatal development through MONOPTEROS repression of a mobile peptide gene STOMAGEN in mesophyll. Proc Natl Acad Sci USA 111(29):E3015-3023
Zhou L, Yu S, Liu Y, Wang Y, Wen Y, Zhang Z, Ru Y, He Z, Chen X (2023) Nitric oxide is involved in the regulation of guard mother cell division by inhibiting the synthesis of ACC. Plant Cell Environ. https://doi.org/10.1111/pce.14734
Zhou Z, Lau OS (2021) Dissecting the developmental roles of Pol II-associated proteins through the stomatal pores. New Phytol 230(1):11–13
Zhu J, Park JH, Lee S, Lee JH, Hwang D, Kwak JM, Kim YJ (2020) Regulation of stomatal development by stomatal lineage miRNAs. Proc Natl Acad Sci USA 117(11):6237–6245
Acknowledgements
Thanks to Prof. Suiwen Hou of the School of Life Science, Lanzhou University, for his strong support and the anonymous reviewers for improving this article. This work was supported by Young Elite Scientists Sponsorship Program by Gansu Association for Science and Technology (GXH20210611-14), the Fundamental Research Funds for the Central Universities (lzujbky-2022-kb03), National Key Research & Development Program of China (2022YFD1201801), Grant Foundation of Science and Technology of Gansu Province (22ZD6NA049 and 21ZD10NF003-2), and Central University outstanding youth team project (lzujbky-2023-eyt02).
Author information
Authors and Affiliations
Contributions
LC conceived the topic, wrote the manuscript and drew figures.
Corresponding author
Ethics declarations
Competing interests
The authors have not disclosed any competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chen, L. Regulation of stomatal development by epidermal, subepidermal and long-distance signals. Plant Mol Biol 114, 80 (2024). https://doi.org/10.1007/s11103-024-01456-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11103-024-01456-7