Abstract
Salt stress is one of the major factors limiting plant growth and productivity. Many studies have shown that serine hydroxymethyltransferase (SHMT) gene play an important role in growth, development and stress response in plants. However, to date, there have been few studies on whether SHMT3 can enhance salt tolerance in plants. Therefore, the effects of overexpression or silencing of CsSHMT3 gene on cucumber seedling growth under salt stress were investigated in this study. The results showed that overexpression of CsSHMT3 gene in cucumber seedlings resulted in a significant increase in chlorophyll content, photosynthetic rate and proline (Pro) content, and antioxidant enzyme activity under salt stress condition; whereas the content of malondialdehyde (MDA), superoxide anion (H2O2), hydrogen peroxide (O2·−) and relative conductivity were significantly decreased when CsSHMT3 gene was overexpressed. However, the content of chlorophyll and Pro, photosynthetic rate, and antioxidant enzyme activity of the silenced CsSHMT3 gene lines under salt stress were significantly reduced, while MDA, H2O2, O2·− content and relative conductivity showed higher level in the silenced CsSHMT3 gene lines. It was further found that the expression of stress-related genes SOD, CAT, SOS1, SOS2, NHX, and HKT was significantly up-regulated by overexpressing CsSHMT3 gene in cucumber seedlings; while stress-related gene expression showed significant decrease in silenced CsSHMT3 gene seedlings under salt stress. This suggests that overexpression of CsSHMT3 gene increased the salt tolerance of cucumber seedlings, while silencing of CsSHMT3 gene decreased the salt tolerance. In conclusion, CsSHMT3 gene might positively regulate salt stress tolerance in cucumber and be involved in regulating antioxidant activity, osmotic adjustment, and photosynthesis under salt stress.
Key message
CsSHMT3 gene may positively regulate the expression of osmotic system, photosynthesis, antioxidant system and stress-related genes in cucumber.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Salt stress is one of the major environmental stresses that limit crop growth, development and productivity, which poses a major threat to global food security (He et al. 2022). Plants have developed various strategies to adapt to salt stress (Yamaguchi-Shinozaki et al. 2006). Cucumber (Cucumis sativus L.) is a universal vegetable and one of the most popular vegetables in the world (Turan et al. 2022). It is grown in more than 80 countries (Yildirim et al. 2008). It is very sensitive to adverse environmental conditions and is classified as a moderately salt-sensitive vegetable (Hamedalla et al. 2022). Therefore, it is very important to explore the potential salt-tolerant genes in cucumber and improve its tolerance to salt stress.
Serine hydroxymethyltransferases (SHMTs) are one of the most important families of enzymes in the single-carbon pathway and photorespiration in plant cells (Ho and Saito 2001). To date, the SHMT gene family has been reported in many species, including Arabidopsis thaliana (Liu et al. 2019), rice (Wang et al. 2015), soybean (Lakhssassi et al. 2019). The SHMT gene family plays an important role in plant growth, development and stress response (Fang et al. 2020). It has been shown that salt-tolerant rice varieties had higher SHMT expression than salt-sensitive varieties, which was associated with a strong accumulation of serine in response to salt stress (Engel et al. 2011). In A. thaliana, SHMT1 played a key role in the photorespiratory cycle and was involved in abiotic and biotic stress responses (Moreno et al. 2005). The activity of the SHMT enzyme played a key role in salt stress-induced cellular damage by supporting photorespiration as part of plant scavenging mechanism in A. thaliana, thereby reducing the production of reactive oxygen species (ROS) and minimizing oxidative damage (Liu et al. 2019).
The role of ROS in plants is twofold: on the one hand, they act as positive regulators and signaling molecules in plants; on the other hand, they cause oxidative damage to plant tissues (Sheikh-Mohamadi et al. 2022). In order to better cope with stress, plants have developed antioxidant systems to scavenge ROS (Mohi-Ud-Din et al. 2021). Antioxidant systems include antioxidant enzyme systems and non-enzymatic systems (He et al. 2017). Superoxide dismutase (SOD), peroxidase (POD), catalase (CAT), and ascorbate peroxidase (APX) enzymes are the main components of the antioxidant enzyme system (Li et al. 2023). The known non-enzymatic antioxidants include L-Glutathione (GSH) and ascorbic acid (AsA) (Ozturk et al. 2021). Photosynthesis is an essential biochemical process in plants. Salt stress in plants leads to water deficit and poor gas exchange. As a result, the stomatal conductance (Gs) is reduced, the net photosynthetic rate (Pn) is decreased, and transpiration rate (Tr) is inhibited, so the photosynthesis is reduced (Zhang et al. 2022a). Due to photosynthesis is inhibited, and plants have difficulty fixing CO2 as organic matter, resulting in the increases of intercellular CO2 concentration (Ci) (Zahra at al. 2022). The SHMT1 has been shown to be an important photorespiratory enzyme that is involved in the conversion of glycine to serine (Mishra et al. 2019).
In cucumber, six members of the SHMT gene family were identified (Gao et al. 2022). The promoter region of cucumber SHMT gene family contains a variety of cis-acting elements related to hormone response, stress response, plant growth, and metabolism. SlSHMT3 gene has been shown to be a significant inducer of salt stress (Liu et al. 2022). Therefore, it is important to further investigate the roles of CsSHMT3 gene in salt stress response. In this study, we transiently silenced and transiently overexpressed CsSHMT3 gene in cucumber to investigate its role in salt stress resistance. Therefore, our results may provide evidence for CsSHMT3 gene in salt resistance studies and reveal a novel regulatory mechanism of CsSHMT in cucumber resistance to abiotic stresses.
Materials and methods
Plant material and growth conditions
Cucumber (Cucumis sativus L. ‘Chinese long inbred line 9930’) seeds were provided by the Institute of Vegetables and Flowers, Chinese Academy of Agricultural Sciences (Beijing, China). Full and uniform seeds were selected, sterilized with 5% NaClO for 10 min, rinsed with deionized water and placed in a culture room at a constant temperature of 25 °C for germination. After 2 d germination, cucumber seeds were sown in a mixed medium of vermiculite and perlite (3:1; v/v) and transferred to the hydroponic box for cultivation. The seedlings were grown under a cycle of 16 h of light with an irradiance of 250 µmol·m− 2 s− 1 photons at 26 °C and 8 h of darkness at 20 °C and 60% relative humidity. The nutrient solution was Hoagland solution. The cucumber seedlings were treated when the two-leaf stage was reached. After 7 d infected with Agrobacterium tumefaciens, cucumber seedlings were treated with Hoagland solution (the control) and 150 mM NaCl containing Hoagland solution for 3 d. Photographs were taken to record the phenotypes of the cucumber seedlings. The samples were immediately frozen in liquid nitrogen and stored at -80 °C in a refrigerator.
RNA extraction and quantitative RT-PCR analysis
Total RNA was extracted using a modified TRIzol (Takara Bio, Japan) method (Hou et al. 2021). The sample was ground into powder in liquid nitrogen, and 1 mL TRIzol was added and incubated for 10 min at 4 °C. Two hundred µL of chloroform were added to the mixture and incubated for 5 min. The supernatant was collected after centrifugation at 16,000 g, for 15 min at 4 °C. Then,an equal volume of isopropanol was added and incubated at -20 °C for more than 1 h. The supernatant was collected into the adsorption column and washed with 75% ethanol. The RNA was dissolved in the RNase-Free ddH2O.Approximately 400–500 ng of RNA from different samples were reverse transcribed using Evo M-MLV Reverse Transcription Kit (AG, China) according to the instructions. SYBR Premix Ex Taq II (AG, China) and Roche LightCycler Fluorescence Quantifier were used for quantitative real-time PCR analysis. The primer sequences of stress-related genes are listed in Table S1. The relative gene expression of each treatment was calculated using the 2−ΔΔCt method with three replicates for each treatment (Vandesompele et al. 2002).
Silencing and overexpression of CsSHMT3 gene in cucumber seedlings
In order to silence CsSHMT3 gene in cucumber, the TRV-mediated virus-induced gene silencing (VIGS) vector was constructed in this study. An approximately 457 bp fragment of the CsSHMT3 gene (Gene ID CsaV3_4G007440.1) was cloned using gene-specific primers (TRV-CsSHMT3-F, aaggttaccgaattctctagaTCTGTGTGTTTCCGCAGCAG; TRV-CsSHMT3-R, gagacgcgtgagctcggtaccACCTCGAAGTTAGCAGGCGA) with Xba 1 and Kpn 1 restriction sites. This fragment was then inserted into the TRV2 vector (Fig. S1A). The construct of pTRV2-CsSHMT3 was confirmed by sequencing and the corresponding gel images are shown in Fig. S2. The vector plasmids of pTRV1, pTRV2 and pTRV2-CsSHMT3 were then transferred into Agrobacterium tumefaciens GV3101. The mixed solution of pTRV1 and pTRV2 (1:1, v/v, OD600 = 0.4) was used to infect the cotyledons of cucumber seedlings as empty pTRV vector. Meanwhile, to silence the CsSHMT3 gene, the mixed solution of pTRV1 and pTRV2-CsSHMT3 (1:1, v/v, OD600 = 0.4) was used to infect the cotyledons of cucumber seedlings. Approximately 60 cucumber seedlings were infested at a time. Plants were photographed 7 d after infection, and silencing efficiency was measured by qRT-PCR using the first and second true leaves of cucumber. Only silenced CsSHMT3 gene lines with a 60% reduction in the level of CsSHMT3 gene transcripts were used in our experiment (Fig. S3).
To overexpress the CsSHMT3 gene in cucumber, an overexpression vector was constructed in this study. The CDS fragment of the CsSHMT3 gene (Gene ID CsaV3_4G007440.1) was cloned using gene-specific primers with Xba 1 and Kpn 1 restriction sites (pSUPER1300-CsSHMT3-F, ccaaatcgactctagtctagaATGCAGGCTACAAG; pSUPER1300-CsSHMT3-R, gcccttgctcaccatggtaccCAGGCCCGGTATTGGGAATTG). The fragment was then inserted into the pSUPER1300 vector (Fig. S1B). The construction of pSUPER1300-CsSHMT3-GFP (35 S: CsSHMT3) was confirmed by sequencing, and the corresponding gel image is shown in Fig. S4. The empty 35 S: GFP and 35 S: CsSHMT3 vector plasmids were transfected into Agrobacterium tumefaciens GV3101. Solutions of empty 35 S: GFP (35 S: 00) and 35: CsSHMT3 were used to infect the cotyledons of cucumber seedlings when the cotyledons were fully expanded. Each infestation involved approximately 60 cucumber seedlings. Plants were imaged 7 d after infection and gene overexpression efficiency was assessed by qRT-PCR using the first and two true leaves. Our experiments utilized only those CsSHMT3 gene overexpressing lines with 20 times increase in CsSHMT3 transcript levels (Fig. S5).
Subcellular localization analysis
The coding sequence of the CsSHMT3 gene (without stop codon) was amplified by PCR. The purified PCR products were inserted into the pSUPER1300 vector and sequenced to give the plasmid 35 S: CsSHMT3. The 35 S: CsSHMT3 vector plasmids were transfected into Agrobacterium tumefaciens GV3101. To identify the subcellular localization of CsSHMT3 protein, the 35 S: GFP: CsSHMT3 fusion vector and the 35 S: GFP empty vector were transiently expressed in Nicotiana benthamiana (Zhang et al. 2022a). The Agrobacterium tumefaciens was infected into N. benthamiana. Then, N. benthamiana was cultivated under normal conditions. After 48–72 h cultivation, the subcellular localization of the GFP signal was observed by laser confocal microscopy (Zeiss LSM710, Zeiss, German).
Measurement of relative conductivity, MDA and Pro content
Relative electrolytic leakage measurement was performed using the method of Huang et al. (2010) with slight modification. Fresh cucumber leaves (0.3 g) were placed in a test tube containing 30 ml distilled water and the samples were incubated in a water bath at 30 °C for 2 h. The initial conductivity (EC1) was measured using a conductivity meter (DSS-307, Shanghai, China). The samples were boiled for 15 min to induce maximum leakage. The electrolyte conductivity (EC2) was measured after cooling to room temperature and the relative electrolyte leakage was calculated.
The thiobarbituric acid (TBA) reaction method (Borgohain et al. 2019) was used for measuring MDA content. The frozen cucumber leaves (0.5 g) were placed in a mortar, then 5 mL of 10% trichloroacetic acid (TCA) solution and quartz sand were added and ground to a homogenate. The homogenate was centrifuged at 4,000 g for 15 min at 4 °C. Then, 2 mL 0.5% TBA solution was added to 1 mL supernatant. The mixture was heated in a water bath at 90 °C for 15 min, immediately cooled on ice and then centrifuged at 4000 g at 4 °C for 15 min. Then, the absorbance was measured at 450, 532, and 600 nm, respectively.
The ninhydrin reaction method was used to determine the Pro content (Nguyen et al. 2021). The 0.5 g cucumber leaves were mixed with 5 mL of 3% sulfosalicylic acid solution. The mixture was followed by a water bath at 100 °C for 10 min. The homogenate was centrifuged at 4000 g for 10 min at 4 °C. A combination of glacial acetic acid, ninhydrin and supernatant were mixed in a 1:1:1 ratio. The tubes were incubated in boiling water for 30 min. After cooling, 5 mL of toluene was added to the solution and mixed thoroughly. The absorbance was recorded at 520 nm using toluene as a blank control.
Measurement of photosynthetic pigment content and photosynthetic parameters
The chlorophyll a, b, and carotenoids were extracted according to the method of He et al. (2022). Fresh cucumber leaves were placed in test tubes containing 10 mL of 80% acetone. The tubes were placed in the dark for 48 h. When the leaves turned white, the absorbance was recorded at 663, 645, and 440 nm using 80% acetone as blank control.
The photosynthetic parameters, including Ci, Gs, Pn, and Tr, were determined using a Yaxin-1105 portable photosynthetic fluorometer (Beijing, China). Measurements were taken from 9:00 to 11:00 am on a sunny day. The photosynthetic apparatus was adjusted and preheated before measurement. Three plants were randomly selected from each strain. The same parts of the leaves were selected. After stabilisation of the measurements, the data were recorded. Each leaf was recorded 5 times and each replicate was averaged.
ROS level measurement and staining
Histochemical staining with nitroblue tetrazolium (NBT) and 3,3’-diaminobenzidine (DAB) were used to determine in situ accumulation of O2·− and H2O2, respectively (Shi et al. 2010). Preparation of 0.5 mg·mL− 1 NBT solution: 5.95 g HEPES was dissolved in 1 L distilled water (obtaining 25 mM HEPES), then 0.5 g NBT was added to 25 mM HEPES and the mixed solution is 0.5 mg·mL− 1 NBT. Preparation of 0.1% DAB solution: 6.057 g Tris was dissolved in 1 L distilled water (obtaining 50 mM Tris), then 0.5 g DAB was added to 50 mM Tris, the mixed solution is 0.1% DAB. Fresh cucumber leaves were immersed in 0.5 mg·mL− 1 NBT solution (pH 7.8). Incubation was carried out in the dark for approximately 6 h to detect in situ accumulation of O2·−. Fresh leaves were soaked in 0.1% DAB solution (pH 3.8). Light incubation was carried out for about 6 h and brown spots appeared on the leaves. To detect H2O2 accumulation in situ the stained leaves were bleached with 95% anhydrous ethanol. Then photos were taken with a digital camera.
The slightly modified method of Sagisaka (1976) was used to measure the H2O2 content. The frozen cucumber leaves (0.5 g) were soaked in 5 mL of acetone solution and crushed into a homogenous paste using a mortar and pestle. The homogenate was centrifuged at 10,000 g at 4 °C for 15 min. Then, 100 µL of the supernatant, 100 µL of 5% Ti (SO4)2 and 200 µL of NH3·H2O were mixed. The mixed solution was centrifuged at 3000 g for 10 min at room temperature and the supernatant was poured off to retain the precipitate. The precipitate was dissolved with 500 µL of 2 mol·L− 1 H2SO4. The absorbance was measured at 415 nm.
The O2·− content was measured using the method of Elstner and Heupel (1976). The frozen cucumber leaves (0.5 g) were extracted with 5 mL of 65 mM potassium phosphate buffer. The homogenate was centrifuged at 5000 g for 10 min at 4 °C. Then 1 mL of the supernatant was mixed with 0.9 mL of 65 mM phosphate buffer (pH 7.8) and 0.1 mL of 10 mM hydroxylamine hydrochloride. The mixture was incubated at 25 °C for 20 min before the adding of 1 mL of 17 mM sulphanilamide and 1 mL of 7 mM α-naphthyl. It was incubated at 25 °C for 20 min and the absorbance was measured at 530 nm.
Measurement of antioxidant enzyme activity
The method of Ji et al. (2022) with modifications was used to measure antioxidant enzyme activity. The cucumber leaves were extracted in 5 mL phosphate buffer (pH 7.8). The homogenate was centrifuged at 10,000 g for 15 min at 4 °C. The supernatant was collected for enzyme activity analysis. Preparation of phosphate buffer: approximately 23.279 g of Na2HPO4-7H2O was dissolved in 1 L of deionized water, which is the mixed solution (A) Approximately 10.14 g of NaH2PO4-H2O was dissolved in 1 L of deionized water, which is the mixed solution (B) The 917.5 mL of solution A and 116 mL of solution B were mixed as phosphate buffer.
The activity of SOD was measured according to the method described by Ji et al. (2022). In a test tube, 0.20 mL supernatant, 1.5 mL phosphate buffer, 0.3 mL methionine solution, 0.3 mL riboflavin, 0.3 mL EDTA-Na solution, 0.28 mL distilled water and 0.3 mL NBT were mixed. The tubes were placed under a 4000 1x lamp for 30 min. and the absorbance was measured at 560 nm.
The guaiacol method was used to determine POD activity (Shi et al. 2010). First, 0.1 mL of supernatant and 2.6 mL of 0.3% guaiacol were added to the cuvette. Then, 0.3 mL of 0.6% H2O2 was added to the cuvette after it was placed in the apparatus. The PBS was used as a blank. Finally, the absorbance of the solution was measured at 470 nm.
CAT activity was determined using Turan et al. (2022) method. First, 0.2 mL of supernatant was added to the cuvette. Next, after placing the cuvette in the apparatus, 2.8 mL of 0.067 mM H2O2 was added into the cuvette. Finally, the change in absorbance at 240 nm within 2 min was recorded immediately.
The method of Ramzan et al. (2023) was used to detect APX activity. Add 0.2 mL of supernatant, 1.75 mL of phosphate buffer (0.05 mM pH 7.0), 0.3 mL of 1 mM EDTA, 0.3 mL of 5 mM AsA, and 0.3 mL of distilled water inside the cuvette. The cuvette was then placed in the apparatus with the addition of 0.3 mL of 1 mM H2O2. The change in absorbance within 1 min at 290 nm was recorded immediately.
Measurement of leaf area, fresh weight, dry weight
Fresh weight in cucumber seedlings was obtained by cutting off the above-ground portion of the plants and weighing them on an electronic analytical balance. The leaves were placed in an envelope and baked in an oven at 105 °C for 30 min, then at 80 °C for 48 h until constant weight. The dry weight of cucumber seedlings was determined using an electronic analytical balance.
Cucumber seedling leaves were scanned with a scanner (YMJ-C, Zhejiang Top Co. LTD. China) and the images were saved, and the images of the leaves were batch analyzed with Win RHIZO software (Pro 2.0 VerSlon 2005; Regent Instruments, Quebec, QC, Canada) to obtain the leaf area.
Statistical analysis
Microsoft Excel 2017 was used to collect and analyze experimental data. The statistical analysis was performed using SPSS statistical software 22.0 (SPPS Inc. Chicago, IL, USA). The data (mean ± SE) are the average of three independent experiments (n = 3). Different letters indicate significant differences by Duncan’s multiple range test (p < 0.05). Graphs were created using GraphPad Prism 9.0.0 (GraphPad Software, San Diego, CA).
Results
CsSHMT3 gene was localized in the chloroplasts
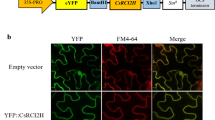
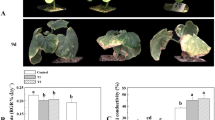
As shown in Fig. 1, the signal of the control was distributed in the cell membrane and nucleus, while the green fluorescence of the CsSHMT3-GFP fusion protein overlapped with the red auto-fluorescence of chloroplasts, which indicated that the CsSHMT3 protein was located in chloroplasts.
Subcellular localization analysis of CsSHMT3. The 35 S: GFP empty vector and 35 S: CsSHTM3: GFP fusion vector were transiently expressed in N. benthamiana. Confocalimages were collected 48–72 h after infestation and analysed by fluorescence microscopy for chloroplasts, bright field (bright), dark field (GFP) and merged field (merge). Scale bar = 20 μm
CsSHMT3 gene increased fresh weight, dry weight, and leaf area under salt stress
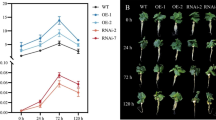
Salt stress significantly slowed seedling growth and leaf greening compared to the control (Fig. 2A, B). In the control, there were no significant differences in leaf area, fresh weight, and dry weight among WT seedlings, empty 35 S infected-seedlings, and 35 S: CsSHMT3 infected-seedlings (Fig. 2C-E). However, under salt stress, leaf area, fresh weight, and dry weight of 35 S: CsSHMT3 infected-seedlings were significantly higher than those of WT seedlings and empty 35 S infected-seedlings (Fig. 2C-E). Therefore, overexpression of CsSHMT3 gene under salt stress increased the leaf area, fresh weight, and dry weight in cucumber seedlings.
The CsSHMT3 gene enhance plant phenotypes and increased fresh weight, dry weight, and leaf area under salt stress. Cucumber seedlings were infested after complete cotyledon expansion. After 7 d of transient overexpression and transient silencing, cucumber seedlings were treated with Hoagland solution (the control) and 150 mM NaCl with Hoagland solution for 3 d. The photographs of whole plants phenotype (A) and leaf phenotype (B) were taken 3 d after treatment. The leaf area (C, F), fresh weight (D, G) and dry weight (E, H) were determined 3 d after treatment. The empty 35 S infected-seedlings were abbreviated as 35 S:00 in figures. Values [mean ± standard error (SE)] were the means of three independent experiments (n = 3). Different letters indicate significant differences by Duncan’s multiple range test (p < 0.05)
In the control, there were no significant differences in leaf area, fresh weight, and dry weight among WT seedlings, empty TRV infected-seedlings, and TRV: CsSHMT3 infected-seedlings (Fig. 2F-H). Under salt stress, TRV: CsSHMT3 infected-seedlings exhibited more attenuated chlorosis and slower growth compared to WT seedlings and empty TRV infected-seedlings. The leaf area, fresh weight, and dry weight of TRV: CsSHMT3 infected-seedlings were significantly lower than those of WT seedlings and empty TRV infected-seedlings (Fig. 2F-H). Therefore, transient silencing of CsSHMT3 gene reduced leaf area, fresh weight, and dry weight under salt stress.
CsSHMT3 gene reduced ROS accumulation under salt stress
Compared to the control, salt stress significantly increased ROS accumulation (Fig. 3). In the control treatment, there was no significant difference in H2O2 and O2·- content among WT seedlings, empty 35 S infected-seedlings, and 35 S: CsSHMT3 infected-seedlings (Fig. 3A, B, E, F). However, under salt stress, the H2O2 and O2·- content in 35 S: CsSHMT3 infected-seedlings was significantly lower than that in WT seedlings and empty 35 S infected-seedlings. For example, the H2O2 and O2·- content in 35 S: CsSHMT3 infected-seedlings was 18.90% and 28.04% below than that in WT seedlings, respectively (Fig. 3A, B, E, F). The results suggested that over-expression of CsSHMT3 gene reduced ROS accumulation under salt stress.
CsSHMT3 gene reduced ROS accumulation under salt stress. Cucumber seedlings were infested after complete cotyledon expansion. After 7 d of transient overexpression and transient silencing, cucumber seedlings were treated with Hoagland solution (the control) and 150 mM NaCl with Hoagland solution for 3 d. In situ accumulation of H2O2 and O2− was detected by histochemical staining with DAB (A, C) and NBT (B, D), respectively. The content of H2O2 (E, G) and O2·− (F, H) was measured 3 d after treatment. The empty 35 S infected-seedlings were abbreviated as 35 S:00 in figures. Values [mean ± standard error (SE)] are the means of three independent experiments (n = 3). Different letters indicate significant differences by Duncan’s multiple range test (p < 0.05)
WT seedlings, empty TRV infected-seedlings, and TRV: CsSHMT3 infected-seedlings did not show significant different ROS level in the control treatment (Fig. 3C, D, G, H). Compared with WT seedlings and empty TRV infected-seedlings, TRV: CsSHMT3 infected-seedlings had higher levels of H2O2 and O2·- content under salt stress (Fig. 3C, D, G, H). Therefore, CsSHMT3 gene silencing could accelerate ROS accumulation under salt stress.
CsSHMT3 gene increased proline content and reduces MDA content and relative conductivity under salt stress
In the control, there were no significant differences in MDA content, relative conductivity, and Pro content among WT seedlings, empty 35 S infected-seedlings, and 35 S: CsSHMT3 infected-seedlings (Fig. 4A-C). Under salt stress, MDA content and relative conductivity in 35 S: CsSHMT3 infected-seedlings were considerably lower than those in empty 35 S infected-seedlings and WT seedlings; while Pro content was markedly higher in 35 S: CsSHMT3 infected-seedlings than in empty 35 S infected-seedlings and WT seedlings. For example, MDA content and relative conductivity in 35 S: CsSHMT3 infected-seedlings were 28.68% and 44.51% lower than those in WT seedlings, respectively, and Pro content in 35 S: CsSHMT3 infected-seedlings was 50.38% higher than that in WT seedlings (Fig. 4A-C). Thus, the over-expression of CsSHMT3 gene under salt stress obviously decreased MDA content and relative conductivity, and enhanced Pro level.
CsSHMT3 gene increases proline content and reduces MDA content and relative conductivity under salt stress. Cucumber seedlings were infested after complete cotyledon expansion. After 7 d of transient overexpression and transient silencing, cucumber seedlings were treated with Hoagland solution (the control) and 150 mM NaCl with Hoagland solution for 3 d. The content of Pro (A, D) and MDA (B, E), and the relative conductivity were determined 3 d after treatment. The relative conductivity was determined 3 d after treatment (C, F). The empty 35 S infected-seedlings were abbreviated as 35 S:00 in figures. Values [mean ± standard error (SE)] are the means of three independent experiments (n = 3). Different letters indicate significant differences by Duncan’s multiple range test (p < 0.05)
In the control, WT seedlings, empty TRV infected-seedlings, and TRV: CsSHMT3 infected-seedlings did not show significant difference in Pro and MDA content, and relative conductivity (Fig. 4D-F). However, under salt stress, TRV: CsSHMT3 infected-seedlings significantly reduced Pro content compared to WT seedlings and empty TRV infected-seedlings. TRV: CsSHMT3 infected-seedlings had markedly higher MDA content and relative conductivity than empty TRV-infected seedlings and WT seedlings. For example, the MDA content and the relative conductivity in TRV: CsSHMT3 infected-seedlings were increased by 66.23% and 37.19% than in WT seedlings, respectively (Fig. 4D-F). Thus, CsSHMT3 gene might decreased Pro content while increased MDA content and relative conductivity under salt stress.
CsSHMT3 gene enhanced the activity of antioxidant enzymes under salt stress
Compared with the control, salt stress significantly increased the antioxidant enzyme activity (Fig. 5). There was no significant difference in SOD, POD, CAT, and APX activity among WT seedlings, empty 35 S infected-seedlings and 35 S: CsSHMT3 infected-seedlings in the control (Fig. 5A-D). However, under salt stress, SOD, POD, CAT, and APX activity in 35 S: CsSHMT3 infected-seedlings was higher than that in WT seedlings and empty 35 S infected-seedlings (Fig. 5A-D). These results indicated that the over-expression of CsSHMT3 gene increased the activity of SOD, POD, CAT and APX under salt stress.
CsSHMT3 gene enhanced the activity of antioxidant enzymes under salt stress. Cucumber seedlings were infested after complete cotyledon expansion. After 7 d of transient overexpression and transient silencing, cucumber seedlings were treated with Hoagland solution (the control) and 150 mM NaCl with Hoagland solution for 3 d. The activity of SOD (A, E), POD (B, F), CAT (C, G) and APX (D, H) was determined 3 d after treatment (A-H). The empty 35 S infected-seedlings were abbreviated as 35 S:00 in figures. Values [mean ± standard error (SE)] are the means of three independent experiments (n = 3). Different letters indicate significant differences by Duncan’s multiple range test (p < 0.05)
In the control, there were no significant differences in SOD, POD, CAT, and APX activity among WT seedlings, empty TRV infected-seedlings and TRV: CsSHMT3 infected-seedlings (Fig. 5E-H). However, under salt stress, the SOD, POD, CAT, and APX activity in TRV: CsSHMT3 infected-seedlings was lower than that in WT seedlings (Fig. 5E-H). Thus, transient silencing of the CsSHMT3 gene expression decreased the activity of SOD, POD, CAT and APX under salt stress.
CsSHMT3 gene enhanced photosynthetic pigment content and photosynthetic parameters under salt stress
In the control, 35 S: CsSHMT3 infected-seedlings did not show differences in chlorophyll content, carotenoid content, Pn, Tr, Gs, and Ci in comparison with WT seedlings and empty 35 S infected-seedlings (Fig. 6A-D, I-L). However, under salt stress, the chlorophyll a, b and a + b content in 35 S: CsSHMT3 infected-seedlings was significantly higher than that in WT seedlings; while the carotenoid content was markedly lower in 5 S: CsSHMT3 infected-seedlings than in WT seedlings (Fig. 6A-D). Compared with WT seedlings, 35 S: CsSHMT3 infected-seedlings showed 30.26%, 17.03%, and 27.59% higher Pn, Tr, and Gs. However, the Ci in 35 S: CsSHMT3 infected-seedlings was significantly lower than that in WT seedlings (Fig. 6I-L). Thus, the over-expression of CsSHMT3 gene increased chlorophyll content and photosynthetic efficiency, and reduced carotenoid accumulation under salt stress.
CsSHMT3 enhanced photosynthetic pigments and photosynthetic efficiency in cucumber under salt stress. Cucumber seedlings were infested after complete cotyledon expansion. After 7 d of transient overexpression and transient silencing, cucumber seedlings were treated with Hoagland solution (the control) and 150 mM NaCl with Hoagland solution for 3 d. The content of photosynthetic pigments was measured 3 d after treatment (A-H). The photosynthetic efficiency was measured 3 d after treatment (I-P). The empty 35 S infected-seedlings were abbreviated as 35 S:00 in figures. Values [mean ± standard error (SE)] are the means of three independent experiments (n = 3). Different letters indicate significant differences by Duncan’s multiple range test (p < 0.05)
In the control, the chlorophyll and carotenoid content, Pn, Tr, Gs, and Ci in TRV: CsSHMT3 infected-seedlings were not significantly different from those in WT and empty TRV infected-seedlings (Fig. 6E-H, M-P). Under salt stress, chlorophyll a, b, and a + b content was dramatically lower in TRV: CsSHMT3 infected-seedlings than in WT seedlings. TRV: CsSHMT3 infected-seedlings had significantly higher carotenoid level compared to WT seedlings (Fig. 6E-H). Pn, Tr, and Gs in TRV: CsSHMT3 infected-seedlings were considerably lower than those in WT seedlings; while Ci in TRV: CsSHMT3 infected-seedlings was significantly higher than that in WT seedlings (Fig. 6M-P). Thus, CsSHMT3 gene under salt stress decreased chlorophyll content, increased carotenoid accumulation and reduced photosynthetic efficiency.
CsSHMT3 gene increased expression of stress-related genes
In the control, WT seedlings, empty 35 S infected-seedlings, and 35 S: CsSHMT3 infected-seedlings did not show significant difference expression of stress-related genes SOD, CAT, SOS1, SOS2, NHX, and HKT (Fig. 7A-C, G-I). However, under NaCl stress, the expression of SOD, CAT, SOS1, SOS2, NHX, and HKT genes was significantly up- regulated by 88.73%, 72.20%, 57.98%, 41.91%, 94.42%, and 30.92% in 35 S: CsSHMT3 infected-seedlings compared with WT seedlings, respectively (Fig. 7A-C, G-I). Thus, over-expression of the CsSHMT3 gene could promote the expression of stress-related genes under salt stress.
CsSHMT3 gene increases expression of stress-related genes. Cucumber seedlings were infested after complete cotyledon expansion. After 7 d of transient overexpression and transient silencing, cucumber seedlings were treated with Hoagland solution (the control) and 150 mM NaCl with Hoagland solution for 3 d. The expression of SOS1 (A, D), SOS2 (B, E), SOD (C, F), CAT (G, J), NHX (H, K), and HKT (I, L) genes were measured 3 d after treatment. The empty 35 S infected-seedlings were abbreviated as 35 S:00 in figures. Values [mean ± standard error (SE)] are the means of three independent experiments (n = 3). Different letters indicate significant differences by Duncan’s multiple range test (p < 0.05)
Supporting Information
There were no significantly difference in the stress-related genes SOD, CAT, SOS1, SOS2, NHX, and HKT among WT seedlings, empty TRV infected-seedlings, and TRV: CsSHMT3 infected-seedlings in the control (Fig. 7D-F, J-L). However, the expression of SOD, CAT, SOS1, SOS2, NHX, and HKT genes was significantly decrease in TRV: CsSHMT3 infected-seedlings than in WT seedlings under NaCl stress (Fig. 7D-F, J-L). Thus, transient silencing of CsSHMT3 gene could reduce the expression of stress-related genes under salt stress.
Discussion
The SHMT genes are widely distributed in the plant kingdom and play an important role in plant growth and development as well as stress response (García-Cañaveras et al. 2021). It has been found that 18, 7, 12, 7, and 5 members of the SHMT gene family are present in soybean (Lakhssassi et al. 2019), A. thaliana (Nogués et al. 2022), Medicago truncatula (Ruszkowski et al. 2018), tomato (Liu et al. 2022), and rice (Fang et al. 2020), respectively. The SHMT genes were divided into 4 subfamilies that reflect their subcellular localization: mitochondria, cytoplasm, chloroplasts, and nuclei (Lakhssassi et al. 2019). It has been shown that seven SHMT genes (AtSHM1-7) are localized in mitochondria (AtSHM1, AtSHM2), chloroplasts (AtSHM3), cytosol (AtSHM4, AtSHM5), and nucleus (AtSHM6, AtSHM7) in A. thaliana (Nogués et al. 2022). In the study, CsSHMT3 gene was localized to chloroplasts, which is the same as AtSHMT3 gene localization (Fig. 1). This suggests that CsSHMT3 gene might play its role in chloroplast and has a similar function to AtSHM3 gene.
Salt stress damages plants mainly through osmotic stress, ionic stress and oxidative stress. Therefore, MDA content and relative conductivity are always used to reflect the integrity of cell membranes. Plants alleviate osmotic stress by producing osmotic regulators such as: Pro, sugars and sugar alcohols (Song et al. 2022). For example, overexpression of the ThASR3 gene in A. thaliana and Tamarix hispida increased Pro content and betaine biosynthesis and decreased MDA content and relative conductivity, thereby increasing the tolerance of A. thaliana and T. hispida to high salt and osmotic stress (Zhang et al. 2022b). Under salt stress, overexpression of the CsSHMT3 gene decreased relative conductivity and MDA content, and increased Pro content; while silencing of the CsSHMT3 gene increased relative conductivity and MDA content and decreased Pro content (Fig. 4). This suggests that CsSHMT3 gene may reduce osmotic stress by accumulating more Pro and alleviate excessive damage to cell membranes under salt stress. Thereby, CsSHMT3 gene might play a crucial role in plant resistance to salinity. This result is consistent with the finding that overexpression ApSHMT1 in Escherichia coli increased glycine betaine level and enhanced salt resistance (Waditee-Sirisattha et al. 2012).
Cellular metabolism normally produces ROS at low levels as a product of normal plant growth and metabolism. However, salt stress always causes a significant increase in ROS in plants, leading to disorganized metabolism, cell damage, and premature senescence or necrosis (Hossain and Dietz 2016). To protect plants from ROS damage, antioxidant enzymes such as SOD, POD, CAT, and APX play important roles in plants. It has been found that overexpression of IbTLD in tobaccos improved the ROS scavenging capacity, resulting in high salt tolerance of transgenic tobacco (Chen et al. 2022). In the present study, overexpression of the CsSHMT3 gene resulted in a significant increase in antioxidant enzyme activity under salt stress and a decrease in H2O2, O2·− level (Figs. 3 and 5). Silencing of the CsSHMT3 gene resulted in a decrease in antioxidant enzyme activity and an increase in H2O2, O2·− content (Figs. 3 and 5) under salt stress. This suggests that the CsSHMT3 gene might be involved in the ROS scavenging system under stress conditions. The OsSHMT was found to enhance cold tolerance in rice by interacting with APX, Hsp70, ATP-synα, ATP-synβ, and MSCP to remove excess H2O2 (Fang et al. 2020). The AtSHMT1 gene was involved in reducing ROS production and oxidative damage and played a critical role in abiotic stress-induced cell damage and cell death (Moreno et al. 2005). Thus, our results provide new evidence that CsSHMT3 gene may enhance salt stress tolerance in cucumber by increasing ROS-scavenger ability.
Under salt stress, an imbalance in osmoregulation leads to stomatal closure and an insufficient supply of carbon dioxide in plants, ultimately resulting in a reduction or cessation of photosynthesis. Ye et al. (2022) found that overexpressing the MpSnRK2.10 gene achieved stomatal opening and increased photosynthetic capacity in apple under salt stress. However, knockdown of StTST3.1 in potato resulted in reduced chlorophyll content and impaired photosynthesis, which interfered with plant growth (Liu et al. 2023). We found that silencing of CsSHMT3 gene caused cucumber seedlings to be more sensitive to salt stress, with a significant decrease in photosynthetic rate and chlorophyll content. However, overexpression of CsSHMT3 gene in cucumber seedlings resulted in plants with higher chlorophyll content and greater photosynthetic capacity (Fig. 6). It has been shown that overexpression of OsSHMT3 gene in A. thaliana slowed the decline in chlorophyll content under salt stress (Mishra et al. 2019). Additionally, overexpression of AtSHMT1 gene in tobacco increased the photosynthetic capacity of leaves under HCHO stress (Zhao et al. 2021). Thus, CsSHMT3 gene may positively regulate salt tolerance in cucumber through regulating photosynthetic system.
It is hypothesized that the enhanced abiotic stress tolerance is mainly due to a significant increase in the expression of abiotic stress-responsive genes. Overexpression of chrysanthemum transcription factor gene DgWRKY3 in tobacco enhanced the expression of stress-related genes involved in osmoregulation and membrane protection (NtP5CS, NtLEA5, and NtERD10D) and oxidative stress response (NtSOD, NtPOD, NtCAT, and NtAPX) under salt stress (Liu et al. 2013). However, the expression of Cat1, Cat2, ER5, TAS14, APX1, APX2, ERF1, P5CS, and GME2 genes was decreased by silencing SlNAC4 gene in tomato (Zhu et al. 2014), suggesting that SlNAC4 gene may regulate stress-related genes expression and enhance salt tolerance in tomato. In the present study, several stress-related genes (SOD, CAT, SOS1, SOS2, NHX, and HKT) were also selected to evaluate the protective effects of CsSHMT3 gene against abiotic stress. We found that overexpression of CsSHMT3 gene caused a significant increase in the expression of SOD, CAT, SOS1, SOS2, NHX, and HKT genes under salt stress, and silencing of CsSHMT3 gene resulted in a significant decrease in the expression of the genes (Fig. 7). It was shown that overexpression of OsSHMT3 in A. thaliana increased the expression of SOS1, HKT1-5, NHX1, SAMS1, PGDH1, and PGDH1 genes, thereby improving its salt tolerance (Mishra et al. 2019). Du et al. (2022) also found that the expression of MdSOS1 and MdNHX1 genes was significantly correlated with the overexpression of MdMYB108L in apple fruit under salt stress. Under salt stress, the overexpression of the MdMYB108L gene increased the expression of MdSOS1 and MdNHX1 genes, whereas silencing MdMYB108L gene resulted in the decrease of MdSOS1 and MdNHX1 gene expression (Du et al. 2022). FvMYB114 gene promoted the expression of salt stress-related genes AtSOS1/3, AtNHX1, and AtLEA3 under salt stress, thereby enhancing salt resistance in A. thaliana (Li et al. 2023). Thus, CsSHMT3 gene may regulate the SOD, CAT, SOS1, SOS2, NHX, and HKT gene expression to respond to salt stress in cucumber.
Conclusions
In summary, the CsSHMT3 protein was localized in chloroplasts, and the CsSHMT3 gene might positively regulate cucumber salt stress tolerance by regulating osmotic system, photosynthesis, antioxidant system, and the stress-related gene expression. These results provide a basis for further understanding of the role of CsSHMT3 gene under salt stress.
Data availability
All data generated or analyzed during this study are included in this manuscript.
References
Borgohain P, Saha B, Agrahari R, Chowardhara B, Sahoo S, van der Vyver C, Panda SK (2019) SlNAC2 overexpression in Arabidopsis results in enhanced abiotic stress tolerance with alteration in glutathione metabolism. Protoplasma 256:1065–1077. https://doi.org/10.1007/s00709-019-01368-0
Chen TC, Chou SY, Chen MC, Lin JS (2022) IbTLD modulates reactive oxygen species scavenging and DNA protection to confer salinity stress tolerance in tobacco. Plant Sci 323:111415. https://doi.org/10.1016/j.plantsci.2022.111415
Du B, Liu H, Dong K, Wang Y, Zhang Y (2022) Over-expression of an R2R3 MYB gene, MdMYB108L, enhances tolerance to salt stress in transgenic plants. Int J Mol Sci 23:9428. https://doi.org/10.3390/ijms23169428
Elstner EF, Heupel A (1976) Inhibition of nitrite formation from hydroxylammoniumchloride: a simple assay for superoxide dismutase. Anal Biochem 70:616–620. https://doi.org/10.1016/0003-2697(76)90488-7
Engel N, Ewald R, Gupta KJ, Zrenner R, Hagemann M, Bauwe H (2011) The presequence of Arabidopsis serine hydroxymethyltransferase SHM2 selectively prevents import into mesophyll mitochondria. Plant Physiol 157:1711–1720. https://doi.org/10.1104/pp.111.184564
Fang C, Zhang P, Li L, Yang L, Mu D, Yan X, Li Z, Lin W (2020) Serine hydroxymethyltransferase localised in the endoplasmic reticulum plays a role in scavenging H2O2 to enhance rice chilling tolerance. BMC Plant Biol 20:236. https://doi.org/10.1186/s12870-020-02446-9
Gao R, Luo Y, Pan X, Wang C, Liao W (2022) Genome-wide identification of SHMT family genes in cucumber (Cucumis sativus L.) and functional analyses of CsSHMTs in response to hormones and abiotic stresses. 3 Biotech 12:305. https://doi.org/10.1007/s13205-022-03378-x
García-Cañaveras JC, Lancho O, Ducker GS, Ghergurovich JM, Xu X, da Silva-Diz V, Minuzzo S, Indraccolo S, Kim H, Herranz D, Rabinowitz JD (2021) SHMT inhibition is effective and synergizes with methotrexate in T-cell acute lymphoblastic leukemia. Leukemia 35:377–388. https://doi.org/10.1038/s41375-020-0845-6
Hamedalla AM, Ali MM, Ali WM, Ahmed MAA, Kaseb MO, Kalaji HM, Gajc-Wolska J, Yousef AF (2022) Increasing the performance of cucumber (Cucumis sativus L.) seedlings by LED illumination. Sci Rep 12:852. https://doi.org/10.1038/s41598-022-04859-y
He L, He T, Farrar S, Ji L, Liu T, Ma X (2017) Antioxidants maintain cellular redox homeostasis by elimination of reactive oxygen species. Cell Physiol Biochem 44:532–553. https://doi.org/10.1159/000485089
He X, Wan Z, Jin N, Jin L, Zhang G, Lyu J, Liu Z, Luo S, Yu J (2022) Enhancement of cucumber resistance under salt stress by 2, 4-epibrassinolide lactones. Front Plant Sci 13:1023178. https://doi.org/10.3389/fpls.2022.1023178
Ho CL, Saito K (2001) Molecular biology of the plastidic phosphorylated serine biosynthetic pathway in Arabidopsis thaliana. Amino Acids 20:243–259. https://doi.org/10.1007/s007260170042https://doi.org/10.1007/s007260170042
Hossain MS, Dietz KJ (2016) Tuning of redox regulatory mechanisms, reactive oxygen species and redox homeostasis under salinity stress. Front Plant Sci 7:548. https://doi.org/10.3389/fpls.2016.00548
Hou X, Qi N, Wang C, Li C, Huang D, Li Y, Wang N, Liao W (2021) Hydrogen-rich water promotes the formation of bulblets in Lilium davidii var. Unicolor through regulating sucrose and starch metabolism. Planta 254:106. https://doi.org/10.1007/s00425-021-03762-6
Huang XS, Liu JH, Chen XJ (2010) Overexpression of PtrABF gene, a bZIP transcription factor isolated from Poncirus trifoliata, enhances dehydration and drought tolerance in tobacco via scavenging ROS and modulating expression of stress-responsive genes. BMC Plant Biol 10:230. https://doi.org/10.1186/1471-2229-10-230
Ji X, Tang J, Zhang J (2022) Effects of salt stress on the morphology, growth and physiological parameters of juglansmicrocarpa L. seedlings. Plants-Basel 11:2381. https://doi.org/10.3390/plants11182381
Lakhssassi N, Patil G, Piya S, Zhou Z, Baharlouei A, Kassem MA, Lightfoot DA, Hewezi T, Barakat A, Nguyen HT, Meksem K (2019) Genome reorganization of the GmSHMT gene family in soybean showed a lack of functional redundancy in resistance to soybean cyst nematode. Sci Rep 9:1506. https://doi.org/10.1038/s41598-018-37815-w
Li W, Li P, Chen H, Zhong J, Liang X, Wei Y, Zhang L, Wang H, Han D (2023) Overexpression of a Fragaria vesca 1R-MYB transcription factor gene (FvMYB114) increases salt and cold tolerance in Arabidopsis thaliana. Int J Mol Sci 24:5261. https://doi.org/10.3390/ijms24065261
Liu QL, Zhong M, Li S, Pan YZ, Jiang BB, Jia Y, Zhang HQ (2013) Overexpression of a chrysanthemum transcription factor gene, DgWRKY3, in tobacco enhances tolerance to salt stress. Plant Physiol Biochem 69:27–33. https://doi.org/10.1016/j.plaphy.2013.04.0160
Liu Y, Mauve C, Lamothe-Sibold M, Guérard F, Glab N, Hodges M, Jossier M (2019) Photorespiratory serine hydroxymethyltransferase 1 activity impacts abiotic stress tolerance and stomatal closure. Plant Cell Environ 42:2567–2583. https://doi.org/10.1111/pce.13595
Liu Z, Pan X, Wang C, Yun F, Huang D, Yao Y, Gao R, Ye F, Liu X, Liao W (2022) Genome-wide identification and expression analysis of serine hydroxymethyltransferase (SHMT) gene family in tomato (Solanum lycopersicum). PeerJ 10:e12943. https://doi.org/10.7717/peerj.12943
Liu T, Kawochar MA, Liu S, Cheng Y, Begum S, Wang E, Zhou T, Liu T, Cai X, Song B (2023) Suppression of the tonoplast sugar transporter, StTST3.1, affects transitory starch turnover and plant growth in potato. Plant J 113:342–356. https://doi.org/10.1111/tpj.16050
Mishra P, Jain A, Takabe T, Tanaka Y, Negi M, Singh N, Jain N, Mishra V, Maniraj R, Krishnamurthy SL, Sreevathsa R, Singh NK, Rai V (2019) Heterologous expression of serine hydroxymethyltransferase-3 from rice confers tolerance to salinity stress in E. Coli and Arabidopsis. Front Plant Sci 10:217. https://doi.org/10.3389/fpls.2019.00217
Mohi-Ud-Din M, Siddiqui MN, Rohman MM, Jagadish SVK, Ahmed JU, Hassan MM, Hossain A, Islam T (2021) Physiological and biochemical dissection reveals a trade-off between antioxidant capacity and heat tolerance in bread wheat (Triticum aestivum L). Antioxidants-Basel 10:351. https://doi.org/10.3390/antiox10030351
Moreno JI, Martín R, Castresana C (2005) Arabidopsis SHMT1, a serine hydroxymethyltransferase that functions in the photorespiratory pathway influences resistance to biotic and abiotic stress. Plant J 41:451–463. https://doi.org/10.1111/j.1365-313X.2004.02311.x
Nguyen HTT, Das Bhowmik S, Long H, Cheng Y, Mundree S, Hoang LTM (2021) Rapid accumulation of proline enhances salinity tolerance in australian wild rice oryza australiensis domin. Plants 10:2044. https://doi.org/10.3390/plants10102044
Nogués I, Sekula B, Angelaccio S, Grzechowiak M, Tramonti A, Contestabile R, Ruszkowski M (2022) Arabidopsis thaliana serine hydroxymethyltransferases: functions, structures, and perspectives. Plant Physiol Biochem 187:37–49. https://doi.org/10.1016/j.plaphy.2022.07.025
Ozturk M, Turkyilmaz Unal B, García-Caparrós P, Khursheed A, Gul A, Hasanuzzaman M (2021) Osmoregulation and its actions during the drought stress in plants. Physiol Plant 172:1321–1335. https://doi.org/10.1111/ppl.13297
Ramzan M, Gillani M, Shah AA, Shah AN, Kauser N, Jamil M, Ahmad RT, Ullah S (2023) Triticum aestivum: antioxidant gene profiling and morpho-physiological studies under salt stress. Mol Biol Rep 50:2569–2580. https://doi.org/10.1007/s11033-022-07990-1
Ruszkowski M, Sekula B, Ruszkowska A, Dauter Z (2018) Chloroplastic serine hydroxymethyltransferase from Medicago truncatula: a structural characterization. Front Plant Sci 9:584. https://doi.org/10.3389/fpls.2018.00584
Sagisaka S (1976) The occurrence of peroxide in a perennial plant, Populus Gelrica. Plant Physiol 57:308–309. https://doi.org/10.1104/pp.57.2.308
Sheikh-Mohamadi MH, Etemadi N, Aalifar M, Pessarakli M (2022) Salt stress triggers augmented levels of Na+, K + and ROS alters salt-related gene expression in leaves and roots of tall wheatgrass (Agropyron elongatum). Plant Physiol Biochem 183:9–22. https://doi.org/10.1016/j.plaphy.2022.04.022
Shi J, Fu XZ, Peng T, Huang XS, Fan QJ, Liu JH (2010) Spermine pretreatment confers dehydration tolerance of citrus in vitro plants via modulation of antioxidative capacity and stomatal response. Tree Physiol 30:914–922. https://doi.org/10.1093/treephys/tpq030
Song C, Acuña T, Adler-Agmon M, Rachmilevitch S, Barak S, Fait A (2022) Leveraging a graft collection to develop metabolome-based trait prediction for the selection of tomato rootstocks with enhanced salt tolerance. Hortic Res 9:uhac061. https://doi.org/10.1093/hr/uhac061
Turan M, Ekinci M, Kul R, Boynueyri FG (2022) Mitigation of salinity stress in cucumber seedlings by exogenous hydrogen sulfide. J Plant Res 135:517–529. https://doi.org/10.1007/s10265-022-01391-y
Vandesompele J, De Preter K, Pattyn F, Poppe B, Van Roy N, De Paepe A, Speleman F (2002) Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol 3:RESEARCH0034. https://doi.org/10.1186/gb-2002-3-7-research0034
Waditee-Sirisattha R, Sittipol D, Tanaka Y, Takabe T (2012) Overexpression of serine hydroxymethyltransferase from halotolerant cyanobacterium in Escherichia coli results in increased accumulation of choline precursors and enhanced salinity tolerance. FEMS Microbiol Lett 333:46–53. https://doi.org/10.1111/j.1574-6968.2012.02597.x
Wang D, Liu H, Li S, Zhai G, Shao J, Tao Y (2015) Characterization and molecular cloning of a serine hydroxymethyltransferase 1 (OsSHM1) in rice. J Integr Plant Biol 57:745–756. https://doi.org/10.1111/jipb.12336
Yamaguchi-Shinozaki K, Shinozaki K (2006) Transcriptional regulatory networks in cellular responses and tolerance to dehydration and cold stresses. Annu Rev Plant Biol 57:781–803. https://doi.org/10.1146/annurev.arplant.57.032905.105444
Ye Y, Jia X, Xue MY, Gao YC, Yue H, Ma FW, Gong XQ (2022) MpSnRK2.10 confers salt stress tolerance in apple via the ABA signaling pathway. https://doi.org/10.1016/j.scienta.2022.110998. Sci Hortic-amsterdan 298
Yildirim E, Turan M, Guvenc I (2008) Effect of foliar salicylic acid applications on growth, chlorophyll, and mineral content of cucumber grown under salt stress. J Plant Nutr 31:593–612. https://doi.org/10.1080/01904160801895118
Zahra N, Al Hinai MS, Hafeez MB, Rehman A, Wahid A, Siddique KHM, Farooq M (2022) Regulation of photosynthesis under salt stress and associated tolerance mechanisms. Plant Physiol Biochem 178:55–69. https://doi.org/10.1016/j.plaphy.2022.03.003
Zhang Q, Liu Y, Jiang Y, Li A, Cheng B, Wu J (2022a) OsASR6 enhances salt stress tolerance in rice. Int J Mol Sci 23:9340. https://doi.org/10.3390/ijms23169340
Zhang Q, Liu Y, Jiang Y, Li A, Cheng B, Wu J (2022b) ThASR3 confers salt and osmotic stress tolerances in transgenic Tamarix and Arabidopsis. BMC Plant Biol 22:586. https://doi.org/10.1186/s12870-022-03942-w
Zhao X, Zeng Z, Cao W, Khan D, Ikram M, Yang K, Chen L, Li K (2021) Co-overexpression of AtSHMT1 and AtFDH induces sugar synthesis and enhances the role of original pathways during formaldehyde metabolism in tobacco. Plant Sci 305:110829. https://doi.org/10.1016/j.plantsci.2021.110829
Zhu M, Chen G, Zhang J, Zhang Y, Xie Q, Zhao Z, Pan Y, Hu Z (2014) The abiotic stress-responsive NAC-type transcription factor SlNAC4 regulates salt and drought tolerance and stress-related genes in tomato (Solanum lycopersicum). Plant Cell Rep 33:1851–1863. https://doi.org/10.1007/s00299-014-1662-z
Funding
This work was supported by the National Natural Science Foundation of China (Nos. 32072559, 31860568, 32360743); the Fostering Foundation for the Excellent PH.D. Dissertation of Gansu Agricultural University (YB2022004); the Key Research and Development Program of Gansu Province, China (No. 21YF5WA096); the National Key Research and Development Program (2018YFD1000800); the Research Fund of Higher Education of Gansu, China (No. 2018 C-14 and 2019B-082); the Natural Science Foundation of Gansu Province, China (Nos. 1606RJZA073, 1606RJZA077, and 1606RJYA252).
Author information
Authors and Affiliations
Contributions
The authors WBL contributed to the study conception and design. Material preparation, data collection was performed by ZHZ, XMH, YHL, ZQD, YH, KDY, YDY, and CL. ZHZ and RG were involved in the experiments and data analysis. ZHZ wrote the first draft of the manuscript. WBL revised the manuscript to present form. All authors read and approved the same for publication.
Corresponding author
Ethics declarations
Conflict of interest
the authors declare no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, Z., Hou, X., Gao, R. et al. CsSHMT3 gene enhances the growth and development in cucumber seedlings under salt stress. Plant Mol Biol 114, 52 (2024). https://doi.org/10.1007/s11103-024-01451-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11103-024-01451-y