Abstract
Pyruvate kinase (Pyk, EC 2.7.1.40) is a glycolytic enzyme that generates pyruvate and adenosine triphosphate (ATP) from phosphoenolpyruvate (PEP) and adenosine diphosphate (ADP), respectively. Pyk couples pyruvate and tricarboxylic acid metabolisms. Synechocystis sp. PCC 6803 possesses two pyk genes (encoded pyk1, sll0587 and pyk2, sll1275). A previous study suggested that pyk2 and not pyk1 is essential for cell viability; however, its biochemical analysis is yet to be performed. Herein, we biochemically analyzed Synechocystis Pyk2 (hereafter, SyPyk2). The optimum pH and temperature of SyPyk2 were 7.0 and 55 °C, respectively, and the Km values for PEP and ADP under optimal conditions were 1.5 and 0.053 mM, respectively. SyPyk2 is activated in the presence of glucose-6-phosphate (G6P) and ribose-5-phosphate (R5P); however, it remains unaltered in the presence of adenosine monophosphate (AMP) or fructose-1,6-bisphosphate. These results indicate that SyPyk2 is classified as PykA type rather than PykF, stimulated by sugar monophosphates, such as G6P and R5P, but not by AMP. SyPyk2, considering substrate affinity and effectors, can play pivotal roles in sugar catabolism under nonphotosynthetic conditions.
Key message
Glucose-6-phosphate and ribose-5-phosphate increased Synechocystis Pyk2 affinity for PEP, possibly contributing to PEP consumption under nonphotosynthetic conditions.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Cyanobacteria can utilize CO2 via photosynthesis to synthesize value-added metabolites for a low-carbon society (Hidese et al. 2020; Angermayr et al. 2014; Hasunuma et al. 2018). Synechocystis sp. PCC 6803 (hereafter, Synechocystis) is a unicellular, non-nitrogen fixing model cyanobacterium that is utilized for bioproduction (Ruffing 2011; Wang et al. 2012; Yu et al. 2013). Synechocystis produces carboxylic acids, such as D-lactate, and polyhydroxy-3-butyrate (PHB), as food additives and bioplastics; these two metabolites are derived from pyruvate (Osanai et al. 2015; Hidese et al. 2020; Ito et al. 2017; Carpine et al. 2017).
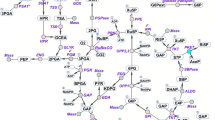
Various groups have widely studied primary carbon metabolism in Synechocystis, including metabolic regulation, pathway identification, and metabolic enzyme biochemistry (Fig. 1). Synechocystis has several glucose catabolic routes, such as the Embden–Meyerhof–Parnas (EMP) and oxidative pentose phosphate (OPP) pathways (You et al. 2014). Synechocystis has a unique tricarboxylic acid (TCA) cycle lacking 2-oxoglutarate dehydrogenase and possessing alternative pathways, such as a γ-butyric amino acid (GABA) shunt (Zhang and Bryant 2011; Xiong et al. 2014). The properties of the enzymes of the Synechocystis TCA cycle have been studied, including those of citrate synthase (CS encoded by gltA, sll0401) (Ito et al. 2019), aconitase (Aco encoded by acnB, slr0665) (Nishii et al. 2021; de Alvarenga et al. 2020), isocitrate dehydrogenase (IDH encoded by icd, slr1289) (Muro-Pastor and Florencio 1992), malate dehydrogenase (MDH encoded by citH, sll0891) (Takeya et al. 2018), malic enzyme (ME encoded by me, sll0721) (Katayama et al. 2022), and succinate dehydrogenase (SDH encoded by sdh, sll0823 and sll1625) (Cooley and Vermaas 2001). Two enzymes, phosphoenolpyruvate carboxylase (PEPC) and pyruvate kinase (Pyk), catalyze specific reactions that provide carbon sources to the TCA cycle. Synechocystis possesses one PEPC encoded by ppc (sll0920) and two Pyks encoded by pyk1 (sll0587) and pyk2 (sll1275) (Kaneko et al. 1996). Synechocystis PEPC exhibits a unique allosteric regulation uninhibited by several metabolic effectors, such as malate, aspartate, and fumarate (Takeya et al. 2017; Scholl et al. 2020). Pyk enzymatic activity from Synechocystis cell cultures has been measured and reported to be higher under nonphotosynthetic conditions than those under photoautotrophic and mixotrophic conditions (Knowles and Plaxton 2003).
Pathway map of Synechocystis sp. PCC 6803 (Synechocystis). The metabolic maps of the Embden–Meyerhof–Parnas (EMP) pathway/gluconeogenesis, oxidative pentose phosphate (OPP) pathway, pyruvate metabolism, and tricarboxylic acid (TCA) cycle. The gene names encoding metabolic enzymes in Synechocystis were obtained from the Kyoto Encyclopedia of Genes and Genomes database. The rounded rectangle indicated value-added metabolites from Synechocystis
The phylogenetic analysis of the Pyks of Synechocystis revealed bacterial Pyk and an evolutionary distance between the two isoforms of Pyks, Pyk1 (hereafter, SyPyk1) and Pyk2, in cyanobacteria (Haghighi 2021). Bacterial Pyk is classified as PykA and PykF, and sugar monophosphates, such as AMP, G6P, and R5P, stimulate PykA. PykF is activated by sugar diphosphates, such as FBP, in Escherichia coli (Kornberg and Malcovati 1973; Waygood et al. 1975, 1976). SyPyk2 is classified into pykF, and pyk2 knockout causes severe growth defects (Yao et al. 2020). In a previous study, the biochemical analysis of the Pyk of another cyanobacterium, Synechococcus sp. PCC 6301 (hereafter, Synechococcus Pyk), was reported to be homologous to that of Synechocystis pyk2, suggesting that ATP and TCA metabolism inhibited Synechococcus Pyk, thus indicating its roles under dark conditions (Knowles et al. 2001). pyk2 expression increases in the wild-type strain in the presence of glucose (Kaniya et al. 2013). pyk2 expression patterns remain unchanged during the day/night cycle (Saha et al. 2016). These reports indicate that SyPyk2 is essential during photosynthetic and nonphotoautotrophic conditions in Synechocystis. This study revealed the regulatory properties of SyPyk2 via biochemical analysis and demonstrated that sugar phosphates activated SyPyk2 activity.
Materials and methods
Construction of cloning vector for recombinant SyPyk2
The amino acid sequence of Pyk2 (sll1275) polypeptide was obtained from the Kyoto Encyclopedia of Genes and Genomes database (https://www.genome.jp/kegg/) and synthesized by Eurofins Genomics Japan (Tokyo, Japan). The synthesized fragment was inserted within the BamHI–XhoI site of the vector pGEX6P-1 (GE Healthcare Japan, Tokyo, Japan). The cloned expression vector was transformed into competent E. coli BL21 cells (Takara Bio, Shiga, Japan) and cultured in 6 mL of Luria–Bertani medium at 30 °C with shaking at 150 rpm. Recombinant E. coli BL21 cells from 1.2-L culture were suspended in 40 mL of phosphate-buffered saline/Tween (PBST) (1.37 M NaCl, 27 mM KCl, 81 mM Na2HPO4・12H2O, 14.7 mM KH2PO4, and 0.05% Tween 20) and sonicated (model VC-750; EYELA, Tokyo, Japan). This procedure was repeated 10 times at 20% intensity for 20 s. The lysed cells were centrifuged twice at 12,500 rpm for 15 min at 4 °C. The supernatant was transferred into a 50 mL tube, and 2 mL of Glutathione Sepharose 4 B resin (Cytiva Japan, Tokyo, Japan) was added. The tubes were shaken gently for 60 min on ice. The mixture was centrifuged at 8,000 rpm for 2 min at 4 °C to remove the supernatant. The resin was transferred to a 15 mL tube and washed using PBST. After washing, the recombinant protein was eluted five times using 650 μL of glutathione-S-transferase (GST) elution buffer (50 mM Tris–HCl, pH 9.6-, and 10-mM reduced glutathione) and incubated in Vivaspin 500 MWCO 50,000 device (Sartorius, Göttingen, Germany) for protein concentration. The protein concentrations were determined using Pierce BCA Protein Assay Kit (Thermo Fisher Scientific, Rockford, IL, USA). The solution was transferred to a 1.5 mL tube, and 40 units of PreScission Protease (equivalent to the purified recombinant protein) (Cytiva) were added and allowed to stand at 4 °C for 16 h to remove GST-tag from the recombinant proteins. Approximately 750 µL of Glutathione Sepharose 4B resin was added, rotated for 1 h at room temperature to remove the cleaved tag from the solution, at 11,000 rpm for 4 min at 4 °C, and the supernatant was collected. Sodium dodecyl sulfate–polyacrylamide gel electrophoresis was performed to confirm protein purification, and gels were stained using Quick Blue reagent (BioDynamics Inc. Tokyo, Japan).
Enzyme assay
All solutions were prepared using Milli-Q water; the SyPyk2 reaction was coupled with lactate dehydrogenase (LDH) from a pig heart (Wako Chemicals, Osaka, Japan) reaction and measured at 30 °C or 55 °C by monitoring NADH oxidation at 340 nm in a final volume of 1 mL. The experiments were performed using 1.88 pmol of the recombinant SyPyk2 proteins. We measured SyPyk2 activity at an intracellular condition of 30 °C and pH 7.8 (Inoue et al. 2001; Nakamura et al. 2021). Unless otherwise indicated, the assay conditions for SyPyk2 were 100 mM Tris–HCl (pH 7.8) or 100 mM MES-NaOH (pH 7.0), 15 mM MgCl2 or 5 mM MnCl2, 100 mM KCl, 0.2 mM NADH, 2 mM ADP-2Na, 15 units/mL desalted LDH from pig heart, and 5 mM PEP-Na. All measurements were performed using 15 mM Mg2+ except where indicated. One unit of SyPyk2 activity was defined as the oxidation of 1 μmol NADH per minute produced. Each effector was added 0.1 mM each: glucose-6-phosphate-2Na (G6P); fructose-6-phosphate-2Na (F6P); fructose-1, 6-bisphosphate-3Na (FBP); ribose-5-phosphate-2Na (R5P); 6-phospho-D-gluconate (6PG); adenosine monophosphate-Na (AMP); adenosine diphosphate-2Na (ADP); adenosine triphosphate-2Na (ATP); citrate-3Na; 2-oxoglutarete (2OG); succinate-2Na; fumarate; malate-Na. The results were plotted as a graph of the reaction rate to substrate and coenzyme concentration. Km and Vmax values were calculated by curve fitting using Kaleida Graph ver. 4.5 software and kcat were calculated from the Vmax.
Statistical analyses
Paired two-tailed Student’s t-tests were performed to calculate the P-values using Microsoft Excel for Windows (Redmond, WA, USA). All experiments were conducted independently in triplicate.
Results
Purification of SyPyk2 and determination of optimal temperature and pH
We expressed GST-tagged SyPyk2 proteins in E. coli BL21 and purified them using affinity chromatography (Fig. 2a). SyPyk2 activity for PEP was the highest in MES-NaOH buffer at pH 7.0 and temperature 55 °C (Fig. 2b and c). The experiments measured at pH 8.5 and 9.0 Tri-HCl using Mn2+ was precipitated (Fig. 2b). Following this, SyPyk2 activities for PEP were measured under optimal conditions (55 °C and pH 7.0) or intracellular conditions (30 °C and pH 7.8).
Sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) and optimal pH and temperature for Synechocystis pyruvate kinase 2 (SyPyk2). a Purified GST-tagged SyPyk2 (89 kDa) and untagged SyPyk2 (63 kDa) proteins. GST-Pyk2 indicated GST-tagged SyPyk2, and Pyk2 indicated untagged SyPyk2. The gel was prepared using 8% (w/v) acrylamide and stained with Quick Blue G250. Optimum pH and temperature for SyPyk2. b Effects of the pH on SyPyk2 activity. The circle and square represented Mg2+ and Mn2+, respectively. Blue and green represented the buffer MES-NaOH and Tris–HCl, respectively. The concentrations of phosphoenolpyruvate (PEP), adenosine diphosphate (ADP), and KCl were fixed at 5.0, 2.0, and 100 mM, respectively. The experiments of Mn2+ measured at pH 8.5 and 9.0 Tri-HCl was precipitated. The mean ± SD values were calculated from three independent experiments. c The effects of temperature on SyPyk2 activity. This experiment was measured in MES-NaOH buffer pH 7.0, and 15 mM Mg2+ of the cofactor was used. PEP, ADP, and KCl concentrations were fixed at 5.0, 2.0, and 100 mM, respectively. The mean ± SD values were calculated from three independent experiments
Dependence SyPyk2 cations for catalytic activity
Similar to the other bacterial Pyks (Waygood and Sanwal 1974; Kapoor and Venkitasubramanian 1983; Wu and Turpin 1992; Snášel and Pichová 2019), SyPyk2 activity was dependent on divalent cations such as Mg2+ or Mn2+, and the Vmax (maximum reaction velocity) of SyPyk2 activity in the presence of Mn2+ was half of that in the presence of Mg2+ (Figs. 3, 4a, and b). The activity of SyPyk2 was higher in the presence of divalent cations than in the presence of monovalent cations, and its activation by monovalent cations was not K+-specific (Fig. 3). We determined the kinetic parameters of SyPyk2 with respect to its dependence on Mg2+ and Mn2+. Under optimal conditions, the Km (half-saturation concentration) value of SyPyk2 for Mg2+ and Mn2+ dependence was 3.54 ± 0.61 and 0.296 ± 0.02 mM, respectively (Fig. 4a and b). Under intracellular conditions, the Km value of SyPyk2 for Mg2+ and Mn2+ dependence was 6.70 ± 0.26 and 2.18 ± 0.51 mM, respectively (Fig. 4a and b). The Km value of Synechococcus Pyk and Synechocystis PEPC for Mg2+ dependence were 2.9 and 4.27 ± 0.46 mM, respectively (Knowles et al. 2001; Scholl et al. 2020). Thus, we defined that the optimum conditions for SyPyk2 were 15 mM MgCl2 and 5 mM MnCl2.
Effects of cofactor monovalent and divalent cations for Synechocystis pyruvate kinase 2 (SyPyk2) activity. The monovalent and divalent cations were fixed at 100 and 5 mM, respectively, except for MgCl2 and ZnCl2 fixed at 15 and 0.5 mM. The experiment was performed using 100 mM MES-NaOH buffer (pH 7.0) at 55 °C. The concentrations of PEP and ADP were fixed at 5.0 and 2.0 mM, respectively. The mean ± SD values were calculated from three independent experiments. K, KCl; Na, NaCl2; NH4, NH4Cl; Mg, MgCl2·6H2O; Mn, MnCl2·4H2O; Ca, CaCl2; Zn, ZnSO4·7H2O
Synechocystis pyruvate kinase 2 (SyPyk2) activity at different concentrations of MgCl2 (a) and MnCl2 (b). These experiments were performed under optimum conditions at 55 °C and pH 7.0 in MES-NaOH buffer (blue) or intracellular conditions at 30 °C and pH 7.8 in Tris–HCl buffer (yellow). These experiments fixed the phosphoenolpyruvate (PEP), adenosine diphosphate (ADP), and KCl concentrations at 5.0, 2.0, and 100 mM, respectively. The mean ± SD values were calculated from three independent experiments
Determination of kinetic parameters of SyPyk2
We measured the kinetic parameters of SyPyk2 for PEP and ADP under optimal conditions. The saturation curves of SyPyk2 for PEP displayed a sigmoidal curve with Vmax and Km of 241 ± 10.5 unit/mg and 1.53 ± 0.07 mM, respectively, and a Hill coefficient of 3.10 ± 0.11, indicating positive homotropic cooperativity (Fig. 5a and Table 1). The saturation curves of SyPyk2 for ADP followed a hyperbolic (Michaelis–Menten) curve with Vmax and Km of 239 ± 6 unit/mg and 0.0527 ± 0.0075 mM, respectively (Fig. 5b and Table 1). The saturation curves of SyPyk2 for PEP and ADP under intracellular conditions were determined. The saturation curves of SyPyk2 for PEP showed a sigmoidal curve with Vmax and Km of 119 ± 7 unit/mg and 2.54 ± 0.12 mM, respectively, and a Hill coefficient of 2.60 ± 0.18, suggesting positive homotropic cooperativity (Fig. 5a and Table 1). The saturation curves of SyPyk2 for ADP exhibited a hyperbolic curve with Vmax and Km of 80.3 ± 5.3 Unit/mg and 0.0602 ± 0.0081 mM, respectively (Fig. 5b and Table 1). Similar to Synechococcus Pyk and other bacterial Pyks, the saturation curves of SyPyk2 for PEP showed sigmoidal curves (Knowles et al. 2001; Jetten et al. 1994; Abdelhamid et al. 2019; 2021), and for ADP, hyperbolic curves (Abdelhamid et al. 2019; 2021). Additionally, we measured the activity of SyPyk2 at 30 °C and pH 7.0; the conditions were optimal for Synechocystis PEPC (Takeya et al. 2017) and competed with SyPyk2 for PEP consumption. The saturation curves of SyPyk2 for PEP displayed a sigmoidal curve with Vmax and Km of 132 ± 5 unit/mg and 2.36 ± 0.2 mM, respectively, and a Hill coefficient of 2.61 ± 0.18, indicating positive homotropic cooperativity and intracellular conditions (Supplemental Fig. 1a and Table 2). The saturation curves of ADP followed a hyperbolic curve with Vmax and Km of 123 ± 9 unit/mg and 0.111 ± 0.017 mM, respectively (Supplemental Fig. 1b and Table 2). In these conditions, the Km value of SyPyk2 for PEP was approximately 30-, 40-, or 20-fold higher than that of SyPyk2 for ADP under optimum, intracellular, and optimum for Synechocystis PEPC conditions, respectively (Tables 1 and 2). The Km value of SyPyk2 for ADP was half of that of Synechococcus Pyk, and the Km value of SyPyk2 for PEP was 5-fold higher than that of Synechococcus Pyk (Knowles et al. 2001). Compared with the Km value of Synechocystis PEPC (0.34 mM: Takeya et al. 2017, 0.88 mM: Scholl et al. 2020) for PEP, SyPyk2 required more than 2-fold PEP (Tables 1 and 2).
Saturation curves of Synechocystis pyruvate kinase 2 (SyPyk2) for phosphoenolpyruvate (PEP) and adenosine diphosphate (ADP). a The saturation curves of SyPyk2 for PEP. These measurements were performed in an optimum condition at 55 °C and pH 7.0 in MES-NaOH buffer (blue) or an intracellular condition of 30 °C and pH 7.8 in Tris–HCl (yellow). The ADP concentration was 2.0 mM. The mean ± SD values were calculated from three independent experiments. b The saturation curves of SyPyk2 for ADP. These measurements were performed in an optimum condition at 55 °C and pH 7.0 in MES-NaOH buffer (blue) or an intracellular condition of 30 °C and pH 7.8 in Tris–HCl (yellow). The PEP concentration was 5 mM. The concentrations of KCl and MgCl2 were 100 and 15 mM, respectively. The mean ± SD values were calculated from three independent experiments
Activation and inhibition of SyPyk2 by sugar phosphates and organic acids
We measured the relative catalytic activity of SyPyk2 for PEP in the presence of sugar phosphates from the EMP/gluconeogenesis and OPP pathways at 1 mM, and PEP and ADP was fixed at Km (2.5 mM) and 2 mM respectively, under intracellular conditions. The effectors did not affect LDH (coupled enzyme) used in the experiment. Additionally, several effector metabolites known to inhibit Pyk from the TCA cycle, such as citrate and 2OG, were added at 1 mM under optimum conditions (Wu and Turpin 1992; Knowles et al. 2001). Under optimal conditions, SyPyk2 was activated by 1 mM G6P and R5P up to 200 and 150%, respectively, and inhibited down to 75% by ATP (Fig. 6). Under intracellular conditions, SyPyk2 was activated in the presence of 1 mM G6P and R5P by 150 and 125%, respectively, and inhibited in the presence of ATP by up to 75% (Fig. 6). Unlike Synechococcus Pyk and green alga, Chlamydomonas reinhardtii Pyk, SyPyk2 was not activated by AMP or F6P and not inhibited by the TCA cycle metabolites, such as citrate, 2OG, and malate (Fig. 6; Wu and Turpin 1992; Knowles et al. 2001).
Effects of effectors for Synechocystis pyruvate kinase 2 (SyPyk2) activity. The effects of various metabolites on the activity SyPyk2. These experiments measured optimum conditions at 55 °C and pH 7.0 in MES-NaOH buffer (left blue bar) and intracellular conditions at 30 °C and pH 7.8 in Tris–HCl buffer (right yellow bar). The mean ± SD values were calculated from three independent experiments. The concentration of PEP and ADP were fixed at Km (2.5 mM) and 2.0 mM, respectively. In the measurements of the saturation curves of SyPyk2, the concentrations of KCl and MgCl2 were 100 and 15 mM, respectively. The concentration of several effectors was 1.0 mM. G6P, glucose-6-phosphate-2Na; F6P, fructose-6-phosphate-2Na; FBP, fructose-1, 6-bisphosphate-3Na; R5P, ribose-5-phosphate-2Na; 6PG, 6-phospho-D-gluconate; AMP, adenosine monophosphate-Na; ADP, adenosine diphosphate-2Na; ATP, adenosine triphosphate-2Na; Cit, citrate-3Na; 2OG, 2-oxoglutarete; Suc, succinate-2Na; Fum: fumarate, Mal: malate-Na. The asterisks indicated significant differences between the absence and presence of the salt (Student’s t-test; *P < 0.01, **P < 0.005)
Furthermore, we calculated the kinetic parameters of SyPyk2 for PEP in the presence of activators or inhibitors under intracellular conditions. The saturation curves of SyPyk2 for PEP in the presence of G6P revealed a hyperbolic curve with Vmax and Km of 122 ± 5 unit/mg and 0.607 ± 0.01 mM, respectively, and a Hill coefficient of 2.0 ± 0.1, indicating G6P converting SyPyk2 from sigmoidal to hyperbolic kinetics (Fig. 7a and Table 1). The enzymatic activities of SyPyk2 for PEP in the presence of R5P displayed a hyperbolic curve with Vmax and Km of 125 ± 18 unit/mg and 0.548 ± 0.132 mM, respectively, and a Hill coefficient of 1.4 ± 0.2, indicating R5P altering SyPyk2 from sigmoidal to hyperbolic kinetics (Fig. 7a and Table 1). Similar to Synechococcus Pyk, G6P and R5P decreased the Km value of SyPyk2 for PEP to one-fifth of its value (Knowles et al. 2001 and Table 1). Moreover, the Km value of SyPyk2 was increased by ATP from 2.54 to 2.73 mM (Fig. 7b and Table 1). To demonstrate the effects of ATP for SyPyk2, we calculated the kinetic parameters of SyPyk2 for PEP in the presence of ATP and either G6P or R5P under intracellular conditions (Fig. 7a and b). G6P, R5P, and ATP concentrations were fixed at 1 mM. The saturation curves of SyPyk2 for PEP in the presence of ATP and G6P revealed a hyperbolic curve with Vmax and Km of 130 ± 29.5 unit/mg and 0.619 ± 0.022 mM, respectively, and a Hill coefficient of 1.73 ± 0.36 (Fig. 7a and b, and Table 1). The enzymatic activities of SyPyk2 for PEP in the presence of ATP and R5P displayed a hyperbolic curve with Vmax and Km of 84.0 ± 7.08 unit/mg and 0.572 ± 0.111 mM, respectively, and a Hill coefficient of 1.89 ± 0.41 (Fig. 7a and b and Table 1). G6P and R5P relieved the effects of ATP (Fig. 7b). Additionally, the IC50 (median inhibition concentration) of ATP for SyPyk2 was 4.1 mM (Supplemental Fig. 2), which was approximately 3-fold higher than that of Synechococcus Pyk (1.5 mM, Knowles et al. 2001). G6P and R5P increased the affinity of SyPyk2 for PEP and remained unaltered in the presence of AMP, F6P, or FBP (Figs. 6, 7a, and b, Table 1).
a Influence of several effectors for Synechocystis pyruvate kinase 2 (SyPyk2) activity. Circles (blue) indicated the phosphoenolpyruvate (PEP) saturation curve, squares (red and purple) indicated 1.0 mM of glucose-6-phosphate (G6P) and ribose-5-phosphate (R5P), diamonds (gray) indicated 1.0 mM of adenosine triphosphate (ATP) added, and red or purple diamonds indicated the presence of G6P and R5P with ATP added respectively. The mean ± SD values were calculated from three independent experiments. b This figure shows the Km value of SyPyk2 for PEP in the presence of G6P, R5P, and ATP, coexisting intracellular conditions of 30 °C and pH 7.8 in Tris–HCl buffer. The mean ± SD values were calculated from three independent experiments
Discussion
This study demonstrated the properties of SyPyk2 via biochemical analysis, with G6P and R5P increasing the affinity of SyPyk2 for PEP in vitro. The optimum pH and temperature of Pyks were discovered in bacteria (Chai et al. 2016; Kapoor and Venkitasubramanian 1983; Abbe and Yamada 1982). The optimum pH of Pyks displayed a wide peak range, from acidic to alkaline (Chai et al. 2016; Abbe and Yamada 1982). For Synechococcus Pyk, the optimal pH ranged from 6.0 to 7.8, and it was active in the dark (Knowles et al. 2001). The intracellular pH of Synechocystis under the photoautotrophic condition was 7.8 (Nakamura et al. 2021), and light to dark transition decreases the intracellular pH of other cyanobacteria from alkaline to neutral (Falkner et al. 1976; Mangan et al. 2016). Thus, the broad pH range of SyPyk2 indicated that SyPyk2 could act on PEP consumption under any cultivation conditions. The optimum temperature of SyPyk2 was 55 °C (Fig. 2c). Synechocystis grows at ~ 30 °C (Inoue et al. 2001), and the optimum temperature of SyPyk2 is higher than that of the cultivation conditions (Fig. 2c). Although for a short time 5 min, Synechocystis is viable up to 54 °C (Inoue et al. 2001). Under heat shock conditions, ATP plays a crucial role in protein maintenance through chaperones (Soini et al. 2005). The gene expression of pyk2 increases during heat shocks (Slabas et al. 2006), and hence, SyPyk2 may contribute to ATP production by increasing its enzymatic activity.
Pyks have been studied for their properties and primary sequences (Hunsley and Suelter 1969; Cottam et al. 1969; Abdelhamid et al. 2019; 2021). All Pyks require divalent cations, such as Mg2+ or Mn2+, and numerous Pyks require K+ for activity (Baek and Nowak 1982; Kachmar and Boyer 1953). SyPyk2 showed Mg2+- and Mn2+-dependent activity and other bacterial Pyks, and its activity was stimulated by monovalent cations, such as K+, Na+, or NH4+(Fig. 3). Bacterial Pyks are classified into two types: PykA, which is stimulated by sugar monophosphates, such as AMP, G6P, and R5P, and PykF, activated by sugar diphosphates, such as FBP in E. coli (Kornberg and Malcovati 1973; Waygood et al. 1975, 1976). Moreover, SyPyk2 is classified as PykF (Kaneko et al. 1996). In silico analysis suggests that two isozymes, Pyk1 and Pyk2 have the same allosteric sites for G6P, R5P, FBP, AMP, and ATP (Haghighi 2021). Pyks containing Pyk1 and Pyk2 from Synechocystis cells are activated by G6P, F6P, R5P, and AMP but not by FBP (Knowles and Plaxton 2003). Based on these findings, we demonstrated the regulation of SyPyk2 by adding various metabolites from the OPP pathway, EMP/gluconeogenesis pathway, and TCA cycle (Fig. 7a). Our findings reveal a decrease in the Km value of SyPyk2 with K-type characteristics, indicative of altered substrate affinity and allosteric activation by G6P and R5P (Fig. 7b and Table 1). Our results suggest that G6P and R5P may also activate SyPyk1. Therefore, SyPyk2 is dependent on divalent cations, such as Mg2+ and Mn2+, and is classified as PykA type rather than PykF, stimulated by sugar monophosphates, such as G6P and R5P, but not by AMP.
The Km values of SyPyk2 for PEP were 40-fold higher than ADP, indicating a higher requirement for PEP in its enzymatic reaction than ADP under intracellular conditions (Table 1). The Km value of Synechococcus Pyk for PEP is higher than that for ADP (Knowles et al. 2001). The Km value of SyPyk2 for PEP is higher than Synechococcus Pyk 5-fold (Table 1 and Knowles et al. 2001). Additionally, the absolute concentration of ADP in Synechocystis cells is three times higher than that of PEP (Dempo et al. 2014). These data indicate that these two Pyk enzymes, Synechocystis and Synechococcus, are limited by PEP concentration in their reactions under photosynthetic conditions. However, its mechanism is different. Synechococcus Pyk has high PEP affinity and allosteric inhibition by citrate and ATP (Knowles et al. 2001). Synechocystis has a low affinity and not inhibited by either citrate or ATP (Table 1 and Fig. 6). Thus, these findings suggested that the availability of PEP limited the enzymatic activity of SyPyk2 for the flux of PEP to pyruvate via SyPyk2 under photosynthetic conditions.
Rapid glycogen catabolism induces glucan polymer such as G6P and signaling metabolites such as 2OG occur during nitrogen depletion (Joseph et al. 2014), indicating that G6P may activate SyPyk2. Although Synechococcus Pyk is repressed by 2OG, SyPyk2 is not (Fig. 6). However, a previous study reveals that pyk1 expression is induced 3.5-fold, and pyk2 expression is reduced by half during nitrogen-deficient conditions (Osanai et al. 2006). Hence, these findings indicate that to provide a carbon source to the TCA cycle for 2OG production, SyPyk2 may act during the initial stages of nitrogen depletion through G6P and R5P activation and then be replaced with SyPyk1. Furthermore, pyk1 is regulated by several nitrogen-related regulators such as SigE, Rre37 thorough with NtcA (Giner-Lamia et al. 2017; Iijima et al. 2014; Osanai et al. 2005). Therefore, in Synechocystis, we consider SyPyk1 and SyPyk2 to mainly function as pyruvate kinase during the late and initial nitrogen depletion stages, respectively.
In comparison to the Km value of PEP for Synechocystis PEPC (0.34 mM: Takeya et al. 2017; 0.88 mM: Scholl et al. 2020 and Table 2), SyPyk2 (2.54 mM, Table 1) required more than 2-fold higher concentration of PEP (Tables 1 and 2). The Km value of SyPyk2 for PEP was decreased from 2.54 to 0.607 or 0.548 mM by G6P or R5P, respectively (Fig. 7a and b and Table 1). The higher PEP requirement and the enhanced affinity of SyPyk2 for PEP by G6P and R5P, suggesting a role for SyPyk2 in Synechocystis cells. In a previous study, Pyk from Synechocystis demonstrated a higher Pyk activity under heterotrophic conditions than under photoautotrophic and mixotrophic conditions (Knowles and Plaxton 2003). Recently, ME, which generates pyruvate from malate by the ME-dependent TCA cycle, was reportedly active under photoautotrophic conditions (Katayama et al. 2022), indicating that pyruvate is synthesized by ME and not by Pyk (Bricker et al. 2004; Qian et al. 2018). The pathway involving PEPC, MDH, and ME constitutes an alternate route for pyruvate formation in Synechocystis cells under photosynthetic conditions (You et al. 2014; Bricker et al. 2004). ATP functions as an inhibitor of Synechococcus Pyk, which is the homolog of SyPyk2 (Knowles et al. 2001). This observation suggests that the lack of Pyk flux under photosynthetic conditions can be attributed to ATP inhibition (Bricker et al. 2004). In E. coli, the intracellular concentration of ATP is suggested to be 0.6 mM (Boecker et al. 2019) and not much different from Synechocystis (Wan et al. 2017). G6P and ATP exhibit comparable concentrations, approximately 1.84*100 and 2.14*100 µmol/g-dry cell weight, respectively (Dempo et al. 2014). R5P is one-tenth of the concentration of ATP, amounting to 1.95*10–1 µmol/g-dry cell (Dempo et al. 2014). Hence, to demonstrate the in vivo effects of metabolites, we maintained uniform effector concentrations at 1 mM. Inhibition by 1 mM ATP decreased the Vmax and increased the Km value of SyPyk2 activity (Table 1 and Fig. 7b). The Vmax of SyPyk2 for PEP decreased from 119 ± 7 to 107 ± 5, and the Km value of SyPyk2 for PEP increased from 2.54 to 2.74 mM (1.07-fold) (Fig. 7b and Table 1). In comparison, the Km value of Synechococcus Pyk increased from 0.54 to 0.75 mM (1.37-fold) by 0.5 mM ATP (Knowles et al. 2001). Furthermore, compared to the IC50 of Synechococcus Pyk for ATP (1.5 mM: Knowles et al. 2001), SyPyk2 (4.1 mM: Supplemental Fig. 2) was approximately 3-fold higher, indicating that SyPyk2 is less affected by ATP than Synechococcus Pyk. Additionally, the presence of G6P and R5P alleviated the inhibitory effects of ATP, reducing the Km value of the substrate concentration to approximately one-fifth (Fig. 7a and b). Our results showed that the effects of ATP on SyPyk2 are less potent than those of G6P and R5P. Considering the IC50 and the slight increase in the Km value by ATP, it suggests that SyPyk2 is less influenced by ATP. Therefore, we conclude that the low flux of PEP to pyruvate via Pyk is due to its extremely low affinity for PEP (Tables 1 and 2; Knowles et al. 2001) and the absence of activators such as G6P and R5P under photosynthetic conditions.
The flux through the OPP pathway increases under nonphotosynthetic conditions (Wan et al. 2017) relative to photosynthetic conditions (Young et al. 2011), indicating a significant elevation in the levels of G6P and R5P under nonphotosynthetic conditions. Moreover, PEP increases and decreases under photosynthetic and nonphotosynthetic conditions, respectively (Werner et al. 2019). Consequently, the accumulation of G6P and R5P under nonphotosynthetic conditions upregulates the SyPyk2 reaction. Following SyPyk2 activation in the presence of G6P and R5P, PEP is consumed by SyPyk2, alleviating glucose-6-phosphate dehydrogenase (G6PDH encoded by zwf, slr1843) inhibition, a rate-limiting enzyme of the OPP pathway (Ito and Osanai 2020). Furthermore, PEP consumption via SyPyk2 activating relieving the inhibition of 6-phosphogluconate dehydrogenase (6PGDH encoded by gnd, sll0329) (Ito and Osanai, 2018), an enzyme involved in the third reaction of the OPP pathway (Fig. 8). In this feedforward regulation, SyPyk2 primarily acts as relieving the sugar catabolic repression under nonphotosynthetic conditions.
Model of glucose-6-phosphate (G6P), ribose-5-phosphate (R5P) and phosphoenolpyruvate (PEP) regulation of carbon flow under photosynthetic or nonphotosynthetic conditions in Synechocystissp. PCC 6803. EMP, Embden–Meyerhof–Parnas pathway; OPP, oxidative pentose phosphate pathway; CBB, Calvin Benson Bassham cycle; Pyr, pyruvate; G6PDH, glucose-6-phosphate dehydrogenase; 6PGDH, 6-phosphogluconate dehydrogenase; Pyk2, pyruvate kinase 2
This study offers valuable insights into the biosynthesis and fermentation of metabolites associated with pyruvate metabolism, particularly PEP consumption in Synechocystis.
This study demonstrated that the regulation of SyPyk2 is dependent on PEP accumulation, the presence of G6P, R5P, and divalent cations, such as Mg2+ and Mn2+, rather than pH and ATP. Therefore, our experiments indicated that SyPyk2 contributed less to PEP consumption under photosynthetic conditions and that it plays a pivotal role in sugar catabolism under nonphotosynthetic conditions in response to sugar phosphate accumulation.
Data availability
Not applicable.
References
Abbe K, Yamada T (1982) Purification and properties of pyruvate kinase from Streptococcus mutans. J Bacteriol 149:299–305. https://doi.org/10.1128/jb.149.1.299-305.1982
Abdelhamid Y, Brear P, Greenhalgh J, Chee X, Rahman T, Welch M (2019) Evolutionary plasticity in the allosteric regulator-binding site of pyruvate kinase isoform PykA from Pseudomonas aeruginosa. J Biol Chem 294:15505–15516. https://doi.org/10.1074/jbc.RA119.009156
Abdelhamid Y, Wang M, Parkhill SL, Brear P, Chee X, Rahman T, Welch M (2021) Structure, function and regulation of a second pyruvate kinase isozyme in Pseudomonas aeruginosa. Front Microbiol 12:790742. https://doi.org/10.3389/fmicb.2021.790742
Abernathy MH, Yu J, Ma F, Liberton M, Ungerer J, Hollinshead WD, Gopalakrishnan S, He L, Maranas CD, Pakrasi HB, Allen DK, Tang YJ (2017) Deciphering cyanobacterial phenotypes for fast photoautotrophic growth via isotopically nonstationary metabolic flux analysis. Biotechnol Biofuels 10:273. https://doi.org/10.1186/s13068-017-0958-y
Allison W, Corey DB, Ashok P, Christie AMP (2019) A comprehensive time-course metabolite profiling of the model cyanobacterium Synechocystis sp. PCC 6803 under diurnal light:dark cycles
Angermayr SA, van der Woude AD, Correddu D, Vreugdenhil A, Verrone V, Hellingwerf KJ (2014) Exploring metabolic engineering design principles for the photosynthetic production of lactic acid by Synechocystis sp. PCC6803. Biotechnol Biofuels 7:99. https://doi.org/10.1186/1754-6834-7-99
Baek YH, Nowak T (1982) Kinetic evidence for a dual cation role for muscle pyruvate kinase. Arch Biochem Biophys 217:491–497. https://doi.org/10.1016/0003-9861(82)90529-x
Boecker S, Zahoor A, Schramm T, Link H, Klamt S (2019) Broadening the scope of enforced ATP wasting as a tool for metabolic engineering in Escherichia coli. Biotechnol J 14:1800438. https://doi.org/10.1002/biot.201800438
Bricker TM, Zhang S, Laborde SM, Mayer PR 3rd, Frankel LK, Moroney JV (2004) The malic enzyme is required for optimal photoautotrophic growth of Synechocystis sp. strain PCC 6803 under continuous light but not under a diurnal light regimen. J Bacteriol 186:8144–8148. https://doi.org/10.1128/JB.186.23.8144-8148
Carpine R, Du W, Olivieri G, Pollio A, Hellingwerf KJ, Marzocchella A, dos Santos FB (2017) Genetic engineering of Synechocystis sp. PCC6803 for poly-β-hydroxybutyrate overproduction. Algal Res 25:117–127. https://doi.org/10.1016/j.algal.2017.05.013
Chai X, Shang X, Zhang Y, Liu S, Liang Y, Zhang Y, Wen T (2016) A novel pyruvate kinase and its application in lactic acid production under oxygen deprivation in Corynebacterium glutamicum. BMC Biotechnol 16:79. https://doi.org/10.1186/s12896-016-0313-6
Cooley JW, Vermaas WF (2001) Succinate dehydrogenase and other respiratory pathways in thylakoid membranes of Synechocystis sp. strain PCC 6803: capacity comparisons and physiological function. J Bacteriol 183:4251–4258. https://doi.org/10.1128/JB.183.14.4251-4258.2001
Cottam GL, Hollenberg PF, Coon MJ (1969) Subunit structure of rabbit muscle pyruvate kinase. J Biol Chem 244:1481–1486. https://doi.org/10.1016/S0021-9258(18)91785-0
de Alvarenga LV, Hess WR, Hagemann M (2020) AcnSP—a novel small protein regulator of aconitase activity in the cyanobacterium Synechocystis sp. PCC 6803. Front Microbiol 11:1445. https://doi.org/10.3389/fmicb.2020.01445
Dempo Y, Ohta E, Nakayama Y, Bamba T, Fukusaki E (2014) Molar-based targeted metabolic profiling of cyanobacterial strains with potential for biological production. Metabolites 4:499–516. https://doi.org/10.3390/metabo4020499
Falkner G, Horner F, Werdan K, Heldt H (1976) Concomitant changes in phosphate uptake and photophosphorylation in the blue-green Alga Anacystis nidulans during adaptation to phosphate deficiency. Plant Physiol 58:717–718. https://doi.org/10.1016/S0176-1617(11)80928-4
Giner-Lamia J, Robles-Rengel R, Hernández-Prieto MA, Muro-Pastor MI, Florencio FJ, Futschik ME (2017) Identification of the direct regulon of NtcA during early acclimation to nitrogen starvation in the cyanobacterium Synechocystis sp. PCC 6803. Nucleic Acids Res 45:11800–11820. https://doi.org/10.1093/nar/gkx860
Guerrero-Mendiola C, García-Trejo JJ, Encalada R, Saavedra E, Ramírez-Silva L (2017) The contribution of two isozymes to the pyruvate kinase activity of Vibrio cholerae: one K+-dependent constitutively active and another K+-independent with essential allosteric activation. PLoS ONE 12:e0178673. https://doi.org/10.1371/journal.pone.0178673
Haghighi O (2021) In silico study of the structure and ligand preference of pyruvate kinases from cyanobacterium Synechocystis sp. PCC 6803. Appl Biochem Biotechnol 193:3651–3671. https://doi.org/10.1007/s12010-021-03630-9
Hasunuma T, Matsuda M, Kondo A (2016) Improved sugar-free succinate production by Synechocystis sp. PCC 6803 following identification of the limiting steps in glycogen catabolism. Metab Eng Commun 3:130–141. https://doi.org/10.1016/j.meteno.2016.04.003
Hasunuma T, Matsuda M, Kato Y, Vavricka CJ, Kondo A (2018) Temperature enhanced succinate production concurrent with increased central metabolism turnover in the cyanobacterium Synechocystis sp. PCC 6803. Metab Eng 48:109–120. https://doi.org/10.1016/j.ymben.2018.05.013
Hauf W, Schlebusch M, Hüge J, Kopka J, Hagemann M, Forchhammer K (2013) Metabolic changes in Synechocystis PCC6803 upon nitrogen-starvation: excess NADPH sustains polyhydroxybutyrate accumulation. Metabolites 3:101–118. https://doi.org/10.3390/metabo3010101
Hidese R, Matsuda M, Osanai T, Hasunuma T, Kondo A (2020) Malic enzyme facilitates D-lactate production through increased pyruvate supply during anoxic dark fermentation in Synechocystis sp. PCC 6803. ACS Synth Biol 9:260–268. https://doi.org/10.1021/acssynbio.9b00281
Hunsley JR, Suelter CH (1969) Yeast pyruvate kinase. I: purification and some chemical properties. J Biol Chem 244:4815–4818. https://doi.org/10.1016/S0021-9258(18)94276-6
Iijima H, Watanabe A, Takanobu J, Hirai MY, Osanai T (2014) rre37 overexpression alters gene expression related to the tricarboxylic acid cycle and pyruvate metabolism in Synechocystis sp. PCC 6803. Sci World J 2014:921–976. https://doi.org/10.1155/2014/921976
Inoue N, Taira Y, Emi T, Yamane Y, Kashino Y, Koike H, Satoh K (2001) Acclimation to the growth temperature and the high-temperature effects on photosystem II and plasma membranes in a mesophilic cyanobacterium, Synechocystis sp. PCC6803. Plant Cell Physiol 42:1140–1148. https://doi.org/10.1093/pcp/pce147
Ito S, Osanai T (2018) Single amino acid change in 6-phosphogluconate dehydrogenase from Synechocystis conveys higher affinity for NADP+ and altered mode of inhibition by NADPH. Plant Cell Physiol 59:2452–2461. https://doi.org/10.1093/pcp/pcy165
Ito S, Osanai T (2020) Unconventional biochemical regulation of the oxidative pentose phosphate pathway in the model cyanobacterium Synechocystis sp. PCC 6803. Biochem J 477(1309–1321): https://doi.org/10.1042/BCJ20200038
Ito S, Takeya M, Osanai T (2017) Substrate specificity and allosteric regulation of a D-Lactate dehydrogenase from a unicellular cyanobacterium are altered by an amino acid substitution. Sci Rep 7:15052. https://doi.org/10.1038/s41598-017-15341-5
Ito S, Koyama N, Osanai T (2019) Citrate synthase from Synechocystis is a distinct class of bacterial citrate synthase. Sci Rep 9:6038. https://doi.org/10.1038/s41598-019-42659-z
Jazmin LJ, Xu Y, Cheah YE, Adebiyi AO, Johnson CH, Young JD (2017) Isotopically nonstationary 13C flux analysis of cyanobacterial isobutyraldehyde production. Metab Eng 42:9–18. https://doi.org/10.1016/j.ymben.2017.05.001
Jetten MS, Gubler ME, Lee SH, Sinskey AJ (1994) Structural and functional analysis of pyruvate kinase from Corynebacterium glutamicum. Appl Environ Microbiol 60:2501–2507. https://doi.org/10.1128/aem.60.7.2501-2507.1994
Joseph A, Aikawa S, Sasaki K, Teramura H, Hasunuma T, Matsuda F, Osanai T, Hirai MY, Kondo A (2014) Rre37 stimulates accumulation of 2-oxoglutarate and glycogen under nitrogen starvation in Synechocystis sp. PCC 6803. FEBS Lett 588:466–471. https://doi.org/10.1016/j.febslet.2013.12.008
Kachmar JF, Boyer PD (1953) Kinetic analysis of enzyme reactions, II: the potassium activation and calcium inhibition of pyruvic phosphoferase. J Biol Chem 200:669–682. https://doi.org/10.1016/S0021-9258(18)71413-0
Kaneko T, Sato S, Kotani H, Tanaka A, Asamizu E, Nakamura Y, Miyajima N, Hirosawa M, Sugiura M, Sasamoto S, Kimura T, Hosouchi T, Matsuno A, Akiko M, Nakazaki N, Naruo K, Okumura S, Shimpo S, Takeuchi C, Wada T, Watanabe A, Yamada M, Yasuda M, Tabata S (1996) Sequence analysis of the genome of the unicellular cyanobacterium Synechocystis sp. Strain PCC6803. ii. Sequence determination of the entire genome and assignment of potential protein-coding regions (supplement). DNA Res 3:185–209. https://doi.org/10.1093/dnares/3.3.185
Kaniya Y, Kizawa A, Miyagi A, Kawai-Yamada M, Uchimiya H, Kaneko Y, Nishiyama Y, Hihara Y (2013) Deletion of the transcriptional regulator cyAbrB2 deregulates primary carbon metabolism in Synechocystis sp. PCC 6803. Plant Physiol 162:1153–1163. https://doi.org/10.1104/pp.113.218784
Kapoor R, Venkitasubramanian TA (1983) Purification and properties of pyruvate kinase from Mycobacterium smegmatis. Arch Biochem Biophys 225:320–330. https://doi.org/10.1016/0003-9861(83)90036-X
Katayama N, Iwazumi K, Suzuki H, Osanai T, Ito S (2022) Malic enzyme, not malate dehydrogenase, mainly oxidizes malate that originates from the tricarboxylic acid cycle in cyanobacteria. Mbio 13:e0218722. https://doi.org/10.1128/mbio.02187-22
Kayne FJ (1973) 11 pyruvate kinase. Enzymes 8:353–382. https://doi.org/10.1016/S1874-6047(08)60071-2
Knoop H, Zilliges Y, Lockau W, Steuer R (2010) The metabolic network of Synechocystis sp. PCC 6803: systemic properties of autotrophic growth. Plant Physiol 154:410–422. https://doi.org/10.1104/pp.110.157198
Knowles VL, Plaxton WC (2003) From genome to enzyme: analysis of key glycolytic and oxidative pentose-phosphate pathway enzymes in the cyanobacterium Synechocystis sp. PCC 6803. Plant Cell Physiol 44:758–763. https://doi.org/10.1093/pcp/pcg086
Knowles VL, Smith CS, Smith CR, Plaxton WC (2001) Structural and regulatory properties of pyruvate kinase from the cyanobacterium Synechococcus PCC 6301. J Biol Chem 276:20966–20972. https://doi.org/10.1074/jbc.M008878200
Kornberg HL, Malcovati M (1973) Control in situ of the pyruvate kinase activity of Escherichia coli. FEBS Lett 32:257–259. https://doi.org/10.1016/0014-5793(73)80846-4
Mangan NM, Flamholz A, Hood RD, Milo R, Savage DF (2016) pH determines the energetic efficiency of the cyanobacterial CO2 concentrating mechanism. Proc Natl Acad Sci USA 113:E5354–E5362. https://doi.org/10.1073/pnas.1525145113
Muro-Pastor MI, Florencio FJ (1992) Purification and properties of NADP-isocitrate dehydrogenase from the unicellular cyanobacterium Synechocystis sp. PCC 6803. Eur J Biochem 203:99–105. https://doi.org/10.1111/j.1432-1033.1992.tb19833.x
Nakajima T, Kajihata S, Yoshikawa K, Matsuda F, Furusawa C, Hirasawa T, Shimizu H (2014) Integrated metabolic flux and omics analysis of Synechocystis sp. PCC 6803 under mixotrophic and photoheterotrophic conditions. Plant Cell Physiol 55:1605–1612. https://doi.org/10.1093/pcp/pcu091
Nakamura S, Fu N, Kondo K, Wakabayashi K, Hisabori T, Sugiura K (2021) A luminescent Nanoluc-GFP fusion protein enables readout of cellular pH in photosynthetic organisms. J Biol Chem 296:100134. https://doi.org/10.1074/jbc.RA120.016847
Nishii M, Ito S, Katayama N, Osanai T (2021) Biochemical elucidation of citrate accumulation in Synechocystis sp. PCC 6803 via kinetic analysis of aconitase. Sci Rep 11:17131. https://doi.org/10.1038/s41598-021-96432-2
Osanai T, Kanesaki Y, Nakano T, Takahashi H, Asayama M, Shirai M, Kanehisa M, Suzuki I, Murata N, Tanaka K (2005) Positive regulation of sugar catabolic pathways in the cyanobacterium Synechocystis sp. PCC 6803 by the group 2 σ factor SigE. J Biol Chem 280:30653–30659. https://doi.org/10.1074/jbc.M505043200
Osanai T, Imamura S, Asayama M, Shirai M, Suzuki I, Murata N, Tanaka K (2006) Nitrogen induction of sugar catabolic gene expression in Synechocystis sp. PCC 6803. DNA Res 13:185–195. https://doi.org/10.1093/dnares/dsl010
Osanai T, Oikawa A, Shirai T, Kuwahara A, Iijima H, Tanaka K, Ikeuchi M, Kondo A, Saito K, Hirai MY (2013) Capillary electrophoresis–mass spectrometry reveals the distribution of carbon metabolites during nitrogen starvation in Synechocystis sp. PCC 6803. Environ Biol 16:512–524. https://doi.org/10.1111/1462-2920.12170
Osanai T, Shirai T, Iijima H, Nakaya Y, Okamoto M, Kondo A, Hirai MY (2015) Genetic manipulation of a metabolic enzyme and a transcriptional regulator increasing succinate excretion from unicellular cyanobacterium. Front Microbiol 6:1064. https://doi.org/10.3389/fmicb.2015.01064
Qian X, Zhang Y, Lun DS, Dismukes GC (2018) Rerouting of metabolism into desired cellular products by nutrient stress: fluxes reveal the selected pathways in cyanobacterial photosynthesis. ACS Synth Biol 7:1465–1476. https://doi.org/10.1021/acssynbio.8b00116
Ruffing AM (2011) Engineered cyanobacteria: teaching an old bug new tricks. Bioeng Bugs 2:136–149. https://doi.org/10.4161/bbug.2.3.15285
Saha R, Liu D, Hoynes-O’Connor A, Liberton M, Yu J, Bhattacharyya-Pakrasi M, Balassy A, Zhang F, Moon TS, Maranas CD, Pakrasi HB (2016) Diurnal regulation of cellular processes in the cyanobacterium Synechocystis sp. strain PCC 6803: insights from transcriptomic, fluxomic, and physiological analyses. Mbio. https://doi.org/10.1128/mBio.00464-16
Sakai H, Suzuki K, Imahori K (1986) Purification and properties of pyruvate kinase from Bacillus stearothermophilus. J Biochem 99:1157–1167. https://doi.org/10.1093/oxfordjournals.jbchem.a135579
Scholl J, Dengler L, Bader L, Forchhammer K (2020) Phosphoenolpyruvate carboxylase from the cyanobacterium Synechocystis sp. PCC 6803 is under global metabolic control by PII signaling. Mol Microbiol 114:292–307. https://doi.org/10.1111/mmi.14512
Slabas AR, Suzuki I, Murata N, Simon WJ, Hall JJ (2006) Proteomic analysis of the heat shock response in Synechocystis PCC6803 and a thermally tolerant knockout strain lacking the histidine kinase 34 gene. Proteomics 6:845–864. https://doi.org/10.1002/pmic.200500196
Snášel J, Pichová I (2019) Allosteric regulation of pyruvate kinase from Mycobacterium tuberculosis by metabolites. Biochim Biophys Acta Proteins Proteom 1867:125–139. https://doi.org/10.1016/j.bbapap.2018.11.002
Soini J, Falschlehner C, Mayer C, Böhm D, Weinel S, Panula J, Vasala A, Neubauer P (2005) Transient increase of ATP as a response to temperature up-shift in Escherichia coli. Microb Cell Factories 4:9. https://doi.org/10.1186/1475-2859-4-9
Takahashi H, Uchimiya H, Hihara Y (2008) Difference in metabolite levels between photoautotrophic and photomixotrophic cultures of Synechocystis sp. PCC 6803 examined by capillary electrophoresis electrospray ionization mass spectrometry. J Exp Bot 59:3009–3018. https://doi.org/10.1093/jxb/ern157
Takeya M, Hirai MY, Osanai T (2017) Allosteric inhibition of phosphoenolpyruvate carboxylases is determined by a single amino acid residue in cyanobacteria. Sci Rep 7:41080. https://doi.org/10.1038/srep41080
Takeya M, Ito S, Sukigara H, Osanai T (2018) Purification and characterisation of malate dehydrogenase from Synechocystis sp. PCC 6803: biochemical barrier of the oxidative tricarboxylic acid cycle. Front Plant Sci 9:947. https://doi.org/10.3389/fpls.2018.00947
Tanaka K, Shirai T, Vavricka CJ, Matsuda M, Kondo A, Hasunuma T (2023) Dark accumulation of downstream glycolytic intermediates initiates robust photosynthesis in cyanobacteria. Plant Physiol 191:2400–2413. https://doi.org/10.1093/plphys/kiac602
Wan N, DeLorenzo DM, He L, You L, Immethun CM, Wang G, Baidoo EEK, Hollinshead W, Keasling JD, Moon TS, Tang YJ (2017) Cyanobacterial carbon metabolism: fluxome plasticity and oxygen dependence. Biotechnol Bioeng 114:1593–1602. https://doi.org/10.1002/bit.26287
Wang B, Wang J, Zhang W, Meldrum DR (2012) Application of synthetic biology in cyanobacteria and algae. Front Microbiol. https://doi.org/10.3389/fmicb.2012.00344
Waygood EB, Sanwal BD (1974) The control of pyruvate kinases of Escherichia coli. I: physicochemical and regulatory properties of the enzyme activated by fructose 1,6-diphosphate. J Biol Chem 249:265–274. https://doi.org/10.1016/S0021-9258(19)43120-7
Waygood EB, Rayman MK, Sanwal BD (1975) The control of pyruvate kinases of Escherichia coli. II: effectors and regulatory properties of the enzyme activated by ribose 5-phosphate. Can J Biochem 53:444–454. https://doi.org/10.1139/o75-061
Waygood EB, Mort JS, Sanwal BD (1976) The control of pyruvate kinase of Escherichia coli: binding of substrate and allosteric effectors to the enzyme activated by fructose 1,6-bisphosphate. Biochem 15:277–282. https://doi.org/10.1021/bi00647a006
Werner A, Broeckling CD, Prasad A, Peebles CAM (2019) A comprehensive time-course metabolite profiling of the model cyanobacterium Synechocystis sp. PCC 6803 under diurnal light: dark cycles. Plant J 99:379–388. https://doi.org/10.1111/tpj.14320
Wu HB, Turpin DH (1992) Purification and characterization of pyruvate kinase from the green alga Chlamydomonas reinhardtii. J Phycol 28:472–481. https://doi.org/10.1111/j.0022-3646.1992.00472.x
Xiong W, Brune D, Vermaas WF (2014) The γ-aminobutyric acid shunt contributes to closing the tricarboxylic acid cycle in Synechocystis sp. PCC 6803. Mol Microbiol 93:786–796. https://doi.org/10.1111/mmi.12699
Yao L, Shabestary K, Björk SM, Asplund-Samuelsson J, Joensson HN, Jahn M, Hudson EP (2020) Pooled CRISPRi screening of the cyanobacterium Synechocystis sp PCC 6803 for enhanced industrial phenotypes. Nat Commun 11:1666. https://doi.org/10.1038/s41467-020-15491-7
You L, Berla B, He L, Pakrasi HB, Tang YJ (2014) 13C-MFA delineates the photomixotrophic metabolism of Synechocystis sp. PCC 6803 under light- and carbon-sufficient conditions. Biotechnol J 9:684–692. https://doi.org/10.1002/biot.201300477
Young JD, Shastri AA, Stephanopoulos G, Morgan JA (2011) Mapping photoautotrophic metabolism with isotopically nonstationary 13C flux analysis. Metab Eng 13:656–665. https://doi.org/10.1016/j.ymben.2011.08.002
Yu Y, You L, Liu D, Hollinshead W, Tang YJ, Zhang F (2013) Development of Synechocystis sp. PCC 6803 as a phototrophic cell factory. Mar Drugs 11:2894–2916. https://doi.org/10.3390/md11082894
Zavřel T, Sinetova MA, Búzová D, Literáková P, Červeny J (2015) Characterization of a model cyanobacterium Synechocystis sp. PCC 6803 autotrophic growth in a flat-panel photobioreactor. Eng Life Sci 15:122–132. https://doi.org/10.1002/elsc.201300165
Zhang S, Bryant DA (2011) The tricarboxylic acid cycle in cyanobacteria. Science 334:1551–1553. https://doi.org/10.1126/science.1210858
Funding
Open Access funding provided by Meiji University. This work was supported by the following grants to TO: JSPS KAKENHI Grant-in-Aid for Scientific Research (B) (Grant Number 20H02905) and JST-ALCA of the Japan Science and Technology Agency (Grant Number JPMJAL1306).
Author information
Authors and Affiliations
Contributions
MK designed the study, performed the experiments, analyzed the data, and wrote the manuscript. NK analyzed the data. TO analyzed the data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Karikomi, M., Katayama, N. & Osanai, T. Pyruvate kinase 2 from Synechocystis sp. PCC 6803 increased substrate affinity via glucose-6-phosphate and ribose-5-phosphate for phosphoenolpyruvate consumption. Plant Mol Biol 114, 60 (2024). https://doi.org/10.1007/s11103-023-01401-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11103-023-01401-0