Abstract
Beneficial plant–microbe associations play critical roles in plant health. Bacterial chemotaxis provides a competitive advantage to motile flagellated bacteria in colonization of plant root surfaces, which is a prerequisite for the establishment of beneficial associations. Chemotaxis signaling enables motile soil bacteria to sense and respond to gradients of chemical compounds released by plant roots. This process allows bacteria to actively swim towards plant roots and is thus critical for competitive root surface colonization. The complete genome sequences of several plant-associated bacterial species indicate the presence of multiple chemotaxis systems and a large number of chemoreceptors. Further, most soil bacteria are motile and capable of chemotaxis, and chemotaxis-encoding genes are enriched in the bacteria found in the rhizosphere compared to the bulk soil. This review compares the architecture and diversity of chemotaxis signaling systems in model beneficial plant-associated bacteria and discusses their relevance to the rhizosphere lifestyle. While it is unclear how controlling chemotaxis via multiple parallel chemotaxis systems provides a competitive advantage to certain bacterial species, the presence of a larger number of chemoreceptors is likely to contribute to the ability of motile bacteria to survive in the soil and to compete for root surface colonization.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
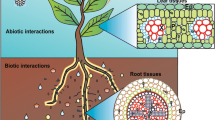
Plant growth and productivity depend on the soil type and architecture, and on the activity of diverse microbes associated with plant roots. A range of associations between microbes and plant roots, from pathogenic or symbiotic to commensals, can be established. For soil bacteria, which live in spatially and temporally heterogeneous environments, the ability to locate niches that support optimum growth in the rhizosphere is often critical to their survival. Abundant experimental evidence shows that chemotaxis, the ability of motile bacteria to direct their movement in gradients of chemorepellents and chemoattractants, enhances the ability of soil bacteria to colonize the roots of diverse plant hosts (Ames and Bergman 1981; Bais et al. 2006; Bauer and Caetano-Anollés 1990; Berendsen et al. 2012; Caetano-Anollés et al. 1988b; Dharmatilake and Bauer 1992; Gulash et al. 1984; Reinhold et al. 1985). Furthermore, the genomes of most soil bacteria analyzed to date contain chemotaxis and motility genes, an observation that emphasizes the competitive advantage of chemotaxis in this environment.
A nutrient gradient is formed in the soil by plant-released root exudates and rhizodeposits, which chemotactically attract diverse motile bacteria (Badri et al. 2009; Bais et al. 2004; Bais et al. 2006). Root colonization may be initiated at the root hair zones, the root tips and the points of emergence of secondary roots, suggesting that these sites release copious amount of exudates [e.g., (Gulash et al. 1984; McDougall and Rovira 1970; Vande Broek et al. 1998)]. Root exudates composition may vary with plant species, stage of development, or environmental conditions, leading to the recruitment and growth stimulation of different members of the rhizosphere microbial communities (Badri et al. 2009; Dennis et al. 2010; Lakshmanan et al. 2014). Recent evidence also indicates that plants may actively recruit specific microbes to the rhizosphere, including those supporting plant growth under conditions of stress, via modulation of root exudates composition (Berendsen et al. 2012; Lakshmanan et al. 2014; Walker et al. 2003).
Many motile soil bacteria recruited to the rhizosphere via chemotaxis toward root exudates are beneficial to plant productivity (Berendsen et al. 2012; Walker et al. 2003). Chemotaxis has been studied in detail and characterized at the molecular level in several beneficial soil bacteria. This review focuses on sensing and signaling during chemotaxis in the following widespread rhizosphere beneficial bacteria that are motile by flagella, namely Azospirillum brasilense, Rhizobium leguminosarum and Sinorhizobium meliloti. Their chemotaxis systems represent a range of structural characteristics that are widespread in soil and rhizosphere bacteria. Despite the importance of beneficial Pseudomonas species in the rhizosphere, their chemotaxis systems will not be covered here. Instead we refer to a recent review article by Sampedro et al. (2015). The present review illustrates the diversity of chemotaxis systems, including sensing, signal transduction and regulation, and their role in the establishment of beneficial associations between soil bacteria and plant roots.
Chemotaxis paradigm and diversity in beneficial plant associated bacteria
Chemotaxis in the model organism Escherichia coli
The chemotaxis signal transduction system (Che) was first described in Escherichia coli. A number of behavioral, genetic, biochemical and biophysical studies conducted over several decades on the E. coli Che system led to a detailed molecular understanding of how chemotaxis functions to navigate this bacterium in gradients of various chemicals (Hazelbauer 2012; Parkinson et al. 2015). Analysis of chemotaxis in several other bacterial species, followed by comparative genomics in recent years, established the conservation of basic chemotaxis principles identified in E. coli (Wadhams and Armitage 2004). In E. coli, chemoreceptors (called methyl-accepting chemotaxis proteins or chemotaxis transducers) are located in the polar regions of motile cells where they form large membrane-bound arrays that relay sensory information to interacting cytoplasmic proteins. The cytoplasmic chemotaxis signal transduction system comprises a histidine kinase named CheA, a scaffolding protein, CheW, that bridges CheA and the signaling domain of chemoreceptors, and the response regulator, CheY, which is phosphorylated upon phosphate transfer from phospho-CheA. The major signaling output of the chemotaxis pathway regulates the phosphorylation state of the CheY response regulator. CheY~P ultimately controls the direction of flagellar motor rotation via interaction with the flagellar switch complex, and thus the probability of switching the direction of rotation of the flagellar motors and the occurrence of tumbles that reorient the cell in a new swimming direction. In E. coli, environmental cues are sensed by five dedicated chemoreceptors (Parkinson et al. 2015). All chemoreceptors contain a highly conserved domain (HCD) flanked by two methylation helices comprising a cytoplasmic signaling module. Upon attractant or repellent binding to the periplasmic domain of transmembrane chemoreceptors, a conformational change occurs and is transmitted through the membrane to the cytoplasmic signaling domain. This induced conformational change modulates the kinase activity of CheA. In addition, the combined activity of a constitutively active methyltransferase, named CheR, and a methylesterase, which is activated by transfer of phosphate from CheA~P, reset the sensitivity of chemoreceptors following ligand-induced conformational changes. Methylation and demethylation of chemoreceptors by CheR and CheB~P is kinetically slower than phospho group transfer between CheA~P and CheY, introducing a short time delay between chemotaxis signaling excitation and adaptation and functions as the bacterial chemotaxis “memory” (Wadhams and Armitage 2004).
Additional chemotaxis components in beneficial plant associated bacteria
Most sequenced bacterial genomes, especially of species found in soils and sediments, possess an average of two or more chemotaxis pathways (Buchan et al. 2010) (Table 1). Chemotaxis signaling is thus prevalent in bacteria occupying these environments and likely to provide a competitive advantage. A recent phylogenomics analysis (Wuichet and Zhulin 2010) identified 18 classes of chemotaxis systems: 16 classes comprise Che systems that control flagellar-based motility (F1 through F16 classes), one class includes Che systems controlling type IV pili motility (Tfp class) and one includes Che systems that control cellular functions other than motility (alternative cellular function or ACF class), such as cyst differentiation (Berleman and Bauer 2005) or development (Kirby and Zusman 2003). Since orthologous che operons appear to perform different functions in various organisms, functional assignment of chemotaxis systems based on sequence alone is not straightforward: for example, the F5 chemotaxis pathway named Che1 regulates all flagellar-dependent taxis responses in Rhodospirillum centenum, while the orthologous F5 chemotaxis system in A. brasilense, also called Che1, has a minor role in controlling taxis responses (Bible et al. 2008, 2012; Hauwaerts et al. 2002). Interestingly, the F7 class of chemotaxis systems is prevalent in rhizosphere bacteria (Buchan et al. 2010; Wisniewski-Dyé et al. 2011), suggesting that specific characteristics of the F7 Che system provide enhanced chemotaxis in this environment. The genomes of A. brasilense (Wisniewski-Dyé et al. 2011), R. leguminosarum (Young et al. 2006) and S. meliloti (Galibert et al. 2001) encode two, two and four Che systems, respectively. All three species utilize a single, F7 class chemotaxis system as the major system controlling chemotaxis responses and competitive root colonization (see below) (Fig. 1).
Architecture of the che operons controlling chemotaxis responses. The gene content of che systems from Sinorhizobium meliloti (top), Rhizobium leguminosarum (middle) and Azospirillum brasilense (bottom) are shown as arrows indicating the direction of transcription. The operon flagellar class is indicated in brackets next to each operon’s name. See text for details on this classification
In addition to multiple chemotaxis systems, there is a significant variation in the number and type of chemotaxis proteins and receptors encoded in the genomes of plant-associated bacteria (Table 1). One significant variation is the occurrence of more than one homolog of CheY. For example, S. meliloti possesses two, R. leguminosarum has three, while A. brasilense has seven CheY homologs. Not all CheY homologs directly affect the flagellar motors. In S. meliloti, CheY2 controls flagellar motor rotation while CheY1, which also interacts with CheA, acts as a phosphate sink to promote signal termination (Sourjik and Schmitt 1996, 1998). In addition, there are differences in flagellar motor function: R. leguminosarum and S. meliloti possess a unidirectional flagellar motor while A. brasilense possesses a bidirectional motor, similar to that of E. coli. Motor directionality correlates with different effects of CheY~P, the output of the signaling pathway, on flagellar rotation. The output of the chemotaxis pathway triggers a change in the rotational speed of the flagellar motors in S. meliloti (Attmannspacher et al. 2005; Platzer et al. 1997) and R. leguminosarum (Miller et al. 2007). Therefore, tumbles are produced by asynchronous rotational speed of individual flagella in these species (Scharf 2002). In A. brasilense, chemotaxis signaling causes a change in both the swimming speed and the direction of flagellar rotation (Bible et al. 2012). In addition to the core chemotaxis components, genes encoding accessory proteins that are not found in the E. coli Che system are also present in the genome of many plant associated bacteria. Such proteins include homologs of the CheC phosphatase, which promotes signal termination by enhancing CheY~P dephosphorylation and the CheD deamidase, which modifies conserved residues in the signaling domain of chemoreceptors and modulates their activity, as experimentally shown in B. subtilis (Rosario et al. 1995; Kristich and Ordal 2002). CheD is found in the F7 class Che system and thus it is present in the genome of S. meliloti, R. leguminosarum and A. brasilense (Fig. 1), but only the two latter species possess CheC, which is encoded outside of the che operons. The diversity in chemotaxis systems of beneficial plant-associated bacteria thus includes a greater number of Che pathways, chemoreceptors and ancillary proteins for signal termination and adaptation. The increased complexity of chemotaxis signaling compared to the E. coli Che system is widespread in other soil and plant-associated bacteria, suggesting that additional chemotaxis components provide a specific competitive advantage in these environments.
Chemotaxis systems in S. meliloti
There are two chemotaxis systems. The chromosomal che1 operon (F7 class) contains ten genes and in-frame deletions in any of these genes result in abolished or diminished chemotaxis (Dogra et al. 2012; Meier et al. 2007; Sourjik and Schmitt 1996; Scharf, unpublished results). In addition to the canonical chemotaxis genes, the che1 operon contains a gene coding for the receptor-modifying deamidase CheD, the chemoreceptor gene, icpA, and two open reading frames coding for novel proteins, CheS and CheT. CheS and CheT have no counterparts in enteric bacteria, but display similarities to unassigned genes in other members of alpha-proteobacteria Ulrich and Zhulin 2009). CheS facilitates an efficient drainage of the phosphate sink by forming a tight complex with the kinase CheA, which allows an accelerated dephosphorylation of CheY1 (Dogra et al. 2012). A cheT deletion strain has the same phenotype as a cheY2 or cheA deletion strain, but its exact function is currently under investigation (Scharf, unpublished results).
The che2 operon (ACF class) is located on the pSymA plasmid and contains five core chemotaxis genes, namely cheR, cheW, mcpS, cheA-REC (encoding a hybrid protein with a REC domain fused to the C-terminus of CheA), and cheB (Barnett et al. 2001; Galibert et al. 2001; Meier et al. 2007). In-frame deletion of mcpS has no effect on chemotaxis (Meier et al. 2007). In addition, analysis of translational fusions of McpS with green fluorescent protein and transcriptional fusions of the upstream region of che2 with a lacZ reporter gene gave no indication of gene expression under liquid culture growth (Meier and Scharf 2009). Therefore, chemotaxis in S. meliloti appears to be mediated by one system, Che1, whereas a role of Che2 in chemotaxis can be excluded.
Chemotaxis systems in R. leguminosarum
Rhizobium leguminosarum also possesses two chemotaxis systems, Che1 (F7 class) and Che2 (F8 class) (Fig. 1). In contrast to S. meliloti, both che1 and che2 are expressed in liquid cultures of R. leguminosarum (Miller et al. 2007). The R. leguminosarum Che1 is orthologous to the S. meliloti Che1 and essential for chemotaxis because mutations in che1 yield a null phenotype (Miller et al. 2007). The role of Che2 in chemotaxis is more subtle and, likely, indirect: a mutation abolishing Che2 function has no significant effect on chemotaxis, but over-expressing cheB 2 from this pathway affects chemotaxis, probably via chemoreceptor modification. This suggests that Che2 has a role in fine-tuning the chemotaxis response mediated by Che1 signaling. However, the direct signaling output of Che2 remains unknown.
Chemotaxis systems in A. brasilense
The genome of A. brasilense encodes four Che pathways named Che1, Che2, Che3 and Che4. Phylogenetic analysis assigned Che1, Che2, and Che4 to flagellar motility Che pathways of the F5, F9, and F7 classes, respectively, while Che3 is an ACF-type pathway (Wisniewski-Dyé et al. 2011). Che1 comprises a full set of chemotaxis proteins, including CheA-REC (hybrid CheA harboring a C-terminal REC domain), CheY, CheW, CheB and CheR (Hauwaerts et al. 2002). Experimental evidence indicates that Che1 controls a transient increase in swimming speed in response to attractants (Bible et al. 2008, 2012) but does not change swimming direction directly (Bible et al. 2008), suggesting that another chemotaxis system provides this function. Consistently, the Che1 system has a minor role, if any, in plant root colonization (Siuti et al. 2011), though chemotaxis is essential for colonization of the rhizosphere by A. brasilense (Greer-Phillips et al. 2004). While the identity of the major chemotaxis pathway has not been experimentally demonstrated, Che4 is the most likely candidate for the following reasons: che2 is not expressed under laboratory conditions (Xie and Alexandre, unpublished results) and che3 is predicted to belong to the ACF class of chemotaxis systems (Wisniewski-Dyé et al. 2011). Therefore, at least two chemotaxis systems control chemotaxis responses in A. brasilense, which parallels the suggested regulation of chemotaxis in R. leguminosarum (Fig. 1). The underlying advantage provided by two chemotaxis systems is not obvious given that chemotaxis is efficient in species using only one system, such as E. coli or S. meliloti. Both A. brasilense and R. leguminosarum, possess a greater number of chemoreceptors than E. coli or S. meliloti, which could suggest a threshold in the number of chemoreceptors above which additional Che systems enhance chemotaxis responses.
Regulation of chemotaxis and flagellar gene expression in plant associated bacteria
Chemotaxis and motility provide a competitive advantage in colonization of the root surface. Thus, expression of flagellar and chemotaxis genes is strictly coordinated. In addition to the structural diversity of chemotaxis systems, plant-associated bacteria differ from E. coli in their expression patterns of flagellar and chemotaxis genes. These differences may be directly related to their metabolic versatility and rhizosphere lifestyle.
In S. meliloti, all che genes, with the exception of genes encoding for chemoreceptors, flagellar (fla, flg, flh, and fli), motility (mot), and regulatory genes (visN, visR, rem, flbT) are clustered in one contiguous chromosomal region, the flagellar regulon (Galibert et al. 2001; Sourjik et al. 1998). The expression of genes in the flagellar regulon is organized as a four-class hierarchy: class IA comprises the master regulatory genes, visN and visR; class IB, the transcription factor encoding rem; class II, controlled by Rem, includes flagellar assembly and motility genes; and class III contains flagellin and chemotaxis genes requiring class II for expression (Fig. 2a). The LuxR-type global transcription activator VisNR is constitutively expressed during liquid culture growth (Sourjik et al. 2000). In contrast, expression of the OmpR-like transcription factor Rem is confined to exponential growth and it thereby acts as a temporal determinant of swimming motility (Rotter et al. 2006). Motility in S. meliloti is also controlled by factors outside of the flagellar regulon. Upstream of VisNR, flagellar motility and chemotaxis is repressed by the Sin/ExpR quorum-sensing-based transcriptional regulation program (Hoang et al. 2008; McIntosh et al. 2008, 2009; Zatakia et al. 2014) and through the ExoR/ExoS/ChvI pathway (Yao et al. 2004). Cell density is thus a key controlling factor of motility and chemotaxis in this species. In addition, the Ros-like zinc finger protein MucR, a regulator of exopolysaccharide (EPS) production, inhibits expression of rem (Bahlawane et al. 2008), while the symbiosis regulator CbrA positively controls transcription of visN and visR (Gibson et al. 2007). Lastly, the small protein EmmA and the two-component system EmmB-EmmC are found to regulate motility, EPS production, and nodule formation. Interestingly, emm appears to not only affect motility through VisNR, but may have multiple targets in the motility pathway (Morris and González 2009). Thus, coupling of these regulatory systems provides a dynamic and precise control of cellular processes important for host interaction (Charoenpanich et al. 2013; Janczarek 2011). In particular, the integrated control of motility and chemotaxis by quorum sensing, master regulators of EPS and nodulation ensures that cells remain motile until they initiate symbiosis or establish a dense biofilm on the root surfaces of non-legumes.
Regulatory cascades controlling expression of chemotaxis and motility genes in S. meliloti (a) and R. leguminosarum (b). The stepwise assembly of flagella is reflected by a regulatory cascade of four classes. Operons are indicated as horizontal arrows, the corresponding gene products as ellipsoids. Positive and negative transcription controls are shown as black arrows and blunt ended lines, respectively, and translation to gene products as open arrows. Positive transcription controls are shown as vertical black arrows and translation to gene products as open arrows
The regulation of expression of chemotaxis and motility genes in R. leguminosarum exhibits a few differences from S. meliloti. While the basic VisN/R—Rem pathway is conserved and required for expression of the major flagellin genes and the primary (che1) chemotaxis operon, the secondary (che2) chemotaxis operons, some mcp genes, and the minor flagellin genes are not dependent on this regulatory cascade (Tambalo et al. 2010) (Fig. 2b). In contrast to S. meliloti, the expression of the Rem-dependent genes remains high in late exponential and stationary phase, and evidence for down-regulation through quorum sensing has not been found. All chemotaxis and motility genes examined, including the regulatory genes visN, visR and rem, are down-regulated in the nodule (Tambalo et al. 2010; Yost et al. 2004), although the nodulation stage at which regulation occurs remains unknown. This down-regulation was also observed in microarray studies of both S. meliloti (Becker et al. 2004; Capela et al. 2006) and R. leguminosarum (Karunakaran et al. 2009). Interestingly, the mechanism of down-regulation is independent of known symbiotic regulators.
Little information is available regarding the regulation of flagellar and chemotaxis gene expression in A. brasilense. This bacterium is motile by means of two types of flagella: a single polar flagellum mediates motility in liquid environments (swimming) and peritrichous lateral flagella are produced in response to growth on surfaces to promote motility under viscous conditions (Moens et al. 1995, 1996). A. brasilense cells are motile under all growth conditions tested, and loss of motility only occurs upon exposure to persistent and severe nutrient starvation and metabolic stress (Sadasivan and Neyra 1985) underscoring the role of motility in this species. This pattern of expression parallels that of the che1 and che4 chemotaxis systems, which are also constitutively expressed in A. brasilense (Xie and Alexandre, unpublished results). The A. brasilense genome lacks evidence of a complete quorum sensing system, making a regulation of motility and chemotaxis by cell density unlikely (Wisniewski-Dyé et al. 2011). While chemotaxis has been linked to swimming motility, there is no experimental evidence indicating that chemotaxis controls lateral-flagella dependent swarming. Since A. brasilense cells are motile under most conditions, motility and chemotaxis may play an even greater role in the ability of these bacteria to survive in the soil and the rhizosphere. A. brasilense is a versatile plant-associated bacterium that can establish in the rhizosphere of diverse plants (Steenhoudt and Vanderleyden 2000). Motility and chemotaxis under changing conditions could further enhance the competitive colonization abilities of A. brasilense in diverse rhizospheres.
A variety of cues are sensed by beneficial plant associated bacteria
In plant-associated bacteria, the cues that stimulate a chemotaxis response are expected to include compounds found in root exudates or on the root surfaces since chemotaxis toward crude root exudates has been demonstrated in several plant associated bacterial species (Caetano-Anollés et al. 1992; Dharmatilake and Bauer 1992; Caetano-Anollés et al. 1988a, b; Reinhold et al. 1985; Barbour et al. 1991; Caetano-Anollés et al. 1992; Gulash et al. 1984; Heinrich and Hess 1985; Mandimba et al. 1986). The composition of root exudates is highly complex and varies with environmental conditions and plant development stages, complicating their detailed characterization. However, numerous components, some being common and others rather unique to certain plants species have been identified (Bais et al. 2004, 2006; Caetano-Anollés et al. 1988a, b; Mandal et al. 2010; Uren 2000). Generally, the root mucilage that contains polysaccharides is abundantly produced by root caps and is also present in the exudates secreted by root tips. Meristems and elongation zones preferentially contain rapidly oxidized organic compounds such as sugars, organic acids and amino acids (Dennis et al. 2010; Walker et al. 2003). Sensing of organic acids is widespread in beneficial soil bacteria, including S. meliloti, R. leguminosarum, A. brasilense, and several species of Pseudomonas, for which specific receptors for a range of structurally different organic acids have been identified (Sampedro et al. 2015). S. meliloti (Bringhurst and Gage 2002), R. leguminosarum (Poole et al. 1994), and A. brasilense (Mukherjee and Ghosh 1987) use organic acids as catabolite repressors. Organic acids thus represent key metabolic regulators for adaptation to the rhizosphere and may explain the widespread occurrence of organic acid chemotaxis in plant-associated bacteria (Alexandre et al. 2000; Meier et al. 2007; Miller et al. 2007; Robinson and Bauer 1993). In addition to organic acids, these beneficial bacteria sense sugars and sugar alcohols (Alexandre et al. 2000; Bowra and Dilworth 1981; Burg et al. 1982; Meier et al. 2007; Miller et al. 2007), which are also present in root exudates of different plants. Sensing of these molecules typically occurs indirectly via the phosphotransferase system and/or periplasmic binding proteins, which then bind to corresponding chemoreceptors (Hazelbauer et al. 2008; Neumann et al. 2012; Wadhams and Armitage 2004). For example, the periplasmic binding protein ChvE specifically binds galactose and contributes to both metabolism and chemotaxis in A. brasilense (Van Bastelaere et al. 1999), but the interacting chemoreceptor is not known. Amino acids are relatively strong attractants for S. meliloti (Götz et al. 1982; Meier et al. 2007; Van Bastelaere et al. 1999) and R. leguminosarum (Miller et al. 2007) but are weak attractants for A. brasilense (Alexandre et al. 2000). S. meliloti and R. leguminosarum fix nitrogen only under symbiotic conditions and must rely on other nitrogen sources, including amino acids, under free-living conditions. In contrast, A. brasilense fixes nitrogen under free-living conditions, an ability that could explain the weak chemostimulatory effect of amino acids in this species. Chemotaxis toward flavonoids, which are host root phenolic compounds that stimulate expression of nodulation genes in bacteria, was proposed for S. meliloti (Caetano-Anollés et al. 1988a; Dharmatilake and Bauer 1992) and R. leguminosarum (Armitage et al. 1988). Chemotaxis towards flavonoids was reported to occur in the nanomolar concentration range, which would imply chemoreceptor-mediated sensing. However, these results could not be reproduced in later studies (Webb and Scharf, unpublished results; Miller, Hynes and Alexandre, unpublished results). Therefore, the role, if any, of flavonoids released in root exudates to specifically attract motile rhizobia to the legume rhizosphere remains to be demonstrated.
Chemoreceptors and ligand specificity for plant association
Signal transduction during bacterial chemotaxis is initiated by the detection of extracellular cues via dedicated receptors. Chemoreceptors are structurally and functionally modular: they possess an N-terminal sensory domain, typically exposed to the extracellular environment and a highly conserved C-terminal domain required for signal transduction (Hazelbauer and Lai 2010). The cues sensed by chemoreceptors thus determine the environment toward which a bacterium may move. Bacteria possessing a greater number of receptors would be predicted to navigate a greater variety of chemical gradients. The number of chemoreceptors encoded in the genomes of plant-associated bacteria varies, but how these differences are reflected in distinct behaviors in the rhizosphere or other environments has not been demonstrated experimentally (Table 1).
Sinorhizobium meliloti possesses nine chemoreceptors; eight of them are involved in sensing environmental stimuli to direct flagellar motor rotation. Similar to what is observed for other sequenced genomes, the majority of the receptor genes has a monocistronic organization and are scattered throughout the genome. Recently, McpU has been identified as a proline sensor mediating chemotaxis towards its alfalfa host (Webb et al. 2014). Further investigations also suggest that McpU functions as general amino acid receptor (Webb and Scharf, unpublished results). In contrast, the function of the remaining seven chemoreceptors is not known and subject of current investigations.
Although ligands for various chemoreceptors in R. leguminosarum have not been identified, it is clear through nodule competition assays that chemoreceptors have a biological role in establishing a successful association with the plant. Mutations in mcpB and mcpC genes cause significant decreases in the ability of the mutants to form nodules on peas when challenged with a wild-type competitor strain. This finding suggests that both receptors detect some specific compound(s) exuded by plant roots and regulate movement towards roots, and explicitly, appropriate infection sites (Yost et al. 1998). Intriguingly, it has been found that mcpC mutants do not have a competitive disadvantage when inoculated on some other legume hosts of R. leguminosarum (Yost and Hynes, unpublished results), indicating that a different attractant spectrum is present in the rhizosphere of various plant species. Furthermore, the study suggests that the symbiotic competition phenotype of mutants in other mcp genes (mcpG, mcpD) (Yost et al. 1998, 2003) should be reexamined using different host plant species and cultivars, to detect plant specific effects.
Similar to S. meliloti and R. leguminosarum, the ligand specificity of most chemoreceptors encoded within the genome of A. brasilense is not known. However, the critical role of Tlp1 in promoting plant–root colonization indicates that chemotaxis modulates plant–root surface colonization by A. brasilense. Experimental evidence shows that the expression pattern of chemoreceptors parallels their function in chemotaxis. In particular, receptor expression is being upregulated under conditions where the cue(s) it senses is prevalent (Xie et al. 2010; Russell and Alexandre, unpublished results). This could represent a strategy to enhance sensitivity to a particular cue and could function as a reinforcing signal for sustained chemotaxis. However, since transcriptomic studies performed in R. leguminosarum failed to identify any mcp genes significantly up-regulated in the rhizosphere (Karunakaran et al. 2009), this is probably not a universal strategy.
Conclusion: challenges and outlook
Bacterial chemotaxis promotes the recruitment of motile soil bacteria to the roots of plants and it is thus critical for the establishment of many associations of bacteria with the roots of plants. Sensing of specific chemoeffectors exuded by roots are likely chemostimulatory and identifying such cues could represent a strategy to specifically recruit beneficial bacteria to enhance plant growth. However, the ligand specificity of most chemoreceptors is unknown and identifying the active fraction(s) of compounds within root exudates that specifically attract bacteria remains challenging. Overcoming these limitations would provide tools for the rationale design of plants or manipulation of growth conditions for enhancing the recruitment of beneficial bacteria to the rhizosphere. This is one of the many approaches needed to achieve sustainable agriculture and ensure future food securities.
References
Alexandre G, Greer SE, Zhulin IB (2000) Energy taxis is the dominant behavior in Azospirillum brasilense. J Bacteriol 182:6042–6048
Ames P, Bergman K (1981) Competitive advantage provided by bacterial motility in the formation of nodules by Rhizobium meliloti. J Bacteriol 148:728–908
Armitage JP, Gallagher A, Johnston AW (1988) Comparison of the chemotactic behaviour of Rhizobium leguminosarum with and without the nodulation plasmid. Mol Microbiol 2:743–748
Attmannspacher U, Scharf B, Schmitt R (2005) Control of speed modulation (chemokinesis) in the unidirectional rotary motor of Sinorhizobium meliloti. Mol Microbiol 56:708–718
Badri DV, Weir TL, van der Lelie D, Vivanco JM (2009) Rhizosphere chemical dialogues: plant–microbe interactions. Curr Opin Biotechnol 20:642–650
Bahlawane C, McIntosh M, Krol E, Becker A (2008) Sinorhizobium meliloti regulator MucR couples exopolysaccharide synthesis and motility. Mol Plant Microbe Interact 21:1498–1509
Bais HP, Park SW, Weir TL, Callaway RM, Vivanco JM (2004) How plants communicate using the underground information superhighway. Trends Plant Sci 9:26–32
Bais HP, Weir TL, Perry LG, Gilroy S, Vivanco JM (2006) The role of root exudates in rhizosphere interactions with plants and other organisms. Ann Rev Plant Biol 57:233–266
Barbour WM, Hattermann DR, Stacey G (1991) Chemotaxis of Bradyrhizobium japonicum to soybean exudates. Appl Env Microbiol 57:2635–2639
Barnett MJ et al (2001) Nucleotide sequence and predicted functions of the entire Sinorhizobium meliloti pSymA megaplasmid. Proc Natl Acad Sci USA 98:9883–9888
Bauer WD, Caetano-Anollés G (1990) Chemotaxis, induced gene expression and competitiveness in the rhizosphere. Plant Soil 129:45–52
Becker A et al (2004) Global changes in gene expression in Sinorhizobium meliloti 1021 under microoxic and symbiotic conditions. Molec Plant-Microbe Interact 17:292–303
Berendsen RL, Pieterse CM, Bakker PA (2012) The rhizosphere microbiome and plant health. Trends Plant Sci 17:478–486
Berleman JE, Bauer CE (2005) Involvement of a Che-like signal transduction cascade in regulating cyst cell development in Rhodospirillum centenum. Mol Microbiol 56:1457–1466
Bible AN, Stephens BB, Ortega DR, Xie Z, Alexandre G (2008) Function of a chemotaxis-like signal transduction pathway in modulating motility, cell clumping, and cell length in the alphaproteobacterium Azospirillum brasilense. J Bacteriol 190:6365–6375
Bible A, Russell MH, Alexandre G (2012) The Azospirillum brasilense Che1 chemotaxis pathway controls swimming velocity, which affects transient cell-to-cell clumping. J Bacteriol 194:3343–3355
Bowra BJ, Dilworth MJ (1981) Motility and chemotaxis towards sugars in Rhizobium leguminosarum. J Gen Microbiol 126:231–235
Bringhurst RM, Gage DJ (2002) Control of inducer accumulation plays a key role in succinate-mediated catabolite repression in Sinorhizobium meliloti. J Bacteriol 184:5385–5392
Buchan A, Crombie B, Alexandre GM (2010) Temporal dynamics and genetic diversity of chemotactic-competent microbial populations in the rhizosphere. Environ Microbiol 12:3171–3184
Burg D, Guillaume J, Tailliez R (1982) Chemotaxis by Rhizobium meliloti. Arch Microbiol 133:162–163
Caetano-Anollés G, Crist-Estes DK, Bauer WD (1988a) Chemotaxis of Rhizobium meliloti to the plant flavone luteolin requires functional nodulation genes. J Bacteriol 170:3164–3169
Caetano-Anollés G, Wall LG, De Micheli AT, Macchi EM, Bauer WD, Favelukes G (1988b) Role of motility and chemotaxis in efficiency of nodulation by Rhizobium meliloti. Plant Physiol 86:1228–1235
Caetano-Anollés G, Wrobel-Boerner E, Bauer WD (1992) Growth and movement of spot inoculated Rhizobium meliloti on the root surface of alfalfa. Plant Physiol 98:1181–1189
Capela D, Filipe C, Bobik C, Batut J, Bruand C (2006) Sinorhizobium meliloti differentiation during symbiosis with alfalfa: a transcriptomic dissection. Molec Plant-Microbe Interact 19:363–372
Charoenpanich P, Meyer S, Becker A, McIntosh M (2013) Temporal expression program of quorum sensing-based transcription regulation in Sinorhizobium meliloti. J Bacteriol 195:3224–3236
Dennis PG, Miller AJ, Hirsch PR (2010) Are root exudates more important than other sources of rhizodeposits in structuring rhizosphere bacterial communities? FEMS Microbiol Ecol 72:313–327
Dharmatilake AJ, Bauer WD (1992) Chemotaxis of Rhizobium meliloti towards nodulation gene-inducing compounds from alfalfa roots. Appl Environ Microbiol 58:1153–1158
Dogra G et al (2012) Sinorhizobium meliloti CheA complexed with CheS exhibits enhanced binding to CheY1, resulting in accelerated CheY1 dephosphorylation. J Bacteriol 194:1075–1087
Galibert F et al (2001) The composite genome of the legume symbiont Sinorhizobium meliloti. Science 293:668–672
Gibson KE, Barnett MJ, Toman CJ, Long SR, Walker GC (2007) The symbiosis regulator CbrA modulates a complex regulatory network affecting the flagellar apparatus and cell envelope proteins. J Bacteriol 189:3591–3602
Götz R, Limmer N, Ober K, Schmitt R (1982) Motility and chemotaxis in two strains of Rhizobium with complex flagella. J General Microbiol 128:789–798
Greer-Phillips SE, Stephens BB, Alexandre G (2004) An energy taxis transducer promotes root colonization by Azospirillum brasilense. J Bacteriol 186:6595–6604
Gulash M, Ames P, Larosiliere RC, Bergman K (1984) Rhizobia are attracted to localized sites on legume roots. Appl Environ Microbiol 48:149–152
Hauwaerts D, Alexandre G, Das SK, Vanderleyden J, Zhulin IB (2002) A major chemotaxis gene cluster in Azospirillum brasilense and relationships between chemotaxis operons in α-proteobacteria. FEMS Microbiol Lett 208:61–67
Hazelbauer GL (2012) Bacterial chemotaxis: the early years of molecular studies. Ann Rev Microbiol 66:285–303. doi:10.1146/annurev-micro-092611-150120
Hazelbauer GL, Lai WC (2010) Bacterial chemoreceptors: providing enhanced features to two-component signaling. Curr Opin Microbiol 13:124–132
Hazelbauer GL, Falke JJ, Parkinson JS (2008) Bacterial chemoreceptors: high-performance signaling in networked arrays. Trends Biochem Sci 33:9–19
Heinrich D, Hess D (1985) Chemotactic attraction of Azospirillum lipoferum by wheat roots and characterization of some attractants. Can J Microbiol 31:26–31
Hoang HH, Gurich N, González JE (2008) Regulation of motility by the ExpR/Sin quorum-sensing system in Sinorhizobium meliloti. J Bacteriol 190:861–871
Janczarek M (2011) Environmental signals and regulatory pathways that influence exopolysaccharide production in rhizobia. Int J Molec Sci 12:7898–7933
Karunakaran R et al (2009) Transcriptomic analysis of Rhizobium leguminosarum biovar viciae in symbiosis with host plants Pisum sativum and Vicia cracca. J Bacteriol 191:4002–4014
Kirby JR, Zusman DR (2003) Chemosensory regulation of developmental gene expression in Myxococcus xanthus. Proc Natl Acad Sci USA 100:2008–2013
Kristich CJ, Ordal GW (2002) Bacillus subtilis CheD is a chemoreceptor modification enzyme required for chemotaxis. J Biol Chem 277:25356–25362
Lakshmanan V, Selvaraj G, Bais HP (2014) Functional soil microbiome: belowground solutions to an aboveground problem. Plant Physiol 166:689–700
Mandal SM, Chakraborty D, Dey S (2010) Phenolic acids act as signaling molecules in plant–microbe symbioses. Plant Signal Behav 5:359–368
Mandimba G, Heulin T, Bally R, Guckert A, Balandreau J (1986) Chemotaxis of free-living nitrogen-fixing bacteria towards maize mucilage. Plant Soil 90:129–139
McDougall BM, Rovira AD (1970) Sites of exudation of 14C-labelled compounds from wheat roots. New Phytol 69:999–1003
McIntosh M, Krol E, Becker A (2008) Competitive and cooperative effects in quorum-sensing-regulated galactoglucan biosynthesis in Sinorhizobium meliloti. J Bacteriol 190:5308–5317
McIntosh M, Meyer S, Becker A (2009) Novel Sinorhizobium meliloti quorum sensing positive and negative regulatory feedback mechanisms respond to phosphate availability. Mol Microbiol 74:1238–1256
Meier VM, Scharf BE (2009) Cellular localization of predicted transmembrane and soluble chemoreceptors in Sinorhizobium meliloti. J Bacteriol 191:5724–5733
Meier VM, Muschler P, Scharf BE (2007) Functional analysis of nine putative chemoreceptor proteins in Sinorhizobium meliloti. J Bacteriol 189:1816–1826
Miller LD, Yost CK, Hynes MF, Alexandre G (2007) The major chemotaxis gene cluster of Rhizobium leguminosarum bv. viciae is essential for competitive nodulation. Molec Microbiol 63:348–362
Moens S, Michiels K, Keijers V, Van Leuven F, Vanderleyden J (1995) Cloning, sequencing, and phenotypic analysis of laf1, encoding the flagellin of the lateral flagella of Azospirillum brasilense Sp7. J Bacteriol 177:5419–5426
Moens S, Schloter M, Vanderleyden J (1996) Expression of the structural gene, laf1, encoding the flagellin of the lateral flagella in Azospirillum brasilense Sp7. J Bacteriol 178:5017–5019
Morris J, González JE (2009) The novel genes emmABC are associated with exopolysaccharide production, motility, stress adaptation, and symbiosis in Sinorhizobium meliloti. J Bacteriol 191:5890–5900
Mukherjee A, Ghosh S (1987) Regulation of fructose uptake and catabolism by succinate in Azospirillum brasilense. J Bacteriol 169:4361–4367
Neumann S, Grosse K, Sourjik V (2012) Chemotactic signaling via carbohydrate phosphotransferase systems in Escherichia coli. Proc Natl Acad Sci USA 109:12159–12164
Parkinson JS, Hazelbauer GL, Falke JJ (2015) Signaling and sensory adaptation in Escherichia coli chemoreceptors: 2015 update. Trends Microbiol 23:257–266
Platzer J, Sterr W, Hausmann M, Schmitt R (1997) Three genes of a motility operon and their role in flagellar rotary speed variation in Rhizobium meliloti. J Bacteriol 179:6391–6399
Poole PS, Blyth A, Reid CJ, Walters K (1994) myo-Inositol catabolism and catabolite regulation in Rhizobium leguminosarum bv. viciae. Microbiology 140:2787–2795
Reinhold B, Hurek T, Fendrik I (1985) Strain-specific chemotaxis of Azospirillum spp. J Bacteriol 162:190–195
Robinson JB, Bauer WD (1993) Relationships between C4 dicarboxylic acid transport and chemotaxis in Rhizobium meliloti. J Bacteriol 175:2284–2291
Rosario MM, Kirby JR, Bochar DA, Ordal GW (1995) Chemotactic methylation and behavior in Bacillus subtilis: role of two unique proteins, CheC and CheD. Biochemistry 34:3823–3831
Rotter C, Mühlbacher S, Salamon D, Schmitt R, Scharf B (2006) Rem, a new transcriptional activator of motility and chemotaxis in Sinorhizobium meliloti. J Bacteriol 188:6932–6942
Sadasivan L, Neyra CA (1985) Flocculation in Azospirillum brasilense and Azospirillum lipoferum: exopolysaccharides and cyst formation. J Bacteriol 163:716–723
Sampedro I, Parales RE, Krell T, Hill JE (2015) Pseudomonas chemotaxis. FEMS Microbiol Rev 39:17–46
Scharf B (2002) Real-time imaging of fluorescent flagellar filaments of Rhizobium lupini H13-3: flagellar rotation and pH-induced polymorphic transitions. J Bacteriol 184:5979–5986
Siuti P, Green C, Edwards AN, Doktycz MJ, Alexandre G (2011) The chemotaxis-like Che1 pathway has an indirect role in adhesive cell properties of Azospirillum brasilense. FEMS Microbiol Lett 323:105–112
Sourjik V, Schmitt R (1996) Different roles of CheY1 and CheY2 in the chemotaxis of Rhizobium meliloti. Mol Microbiol 22:427–436
Sourjik V, Schmitt R (1998) Phosphotransfer between CheA, CheY1, and CheY2 in the chemotaxis signal transduction chain of Rhizobium meliloti. Biochemistry 37:2327–2335
Sourjik V, Sterr W, Platzer J, Bos I, Haslbeck M, Schmitt R (1998) Mapping of 41 chemotaxis, flagellar and motility genes to a single region of the Sinorhizobium meliloti chromosome. Gene 223:283–290
Sourjik V, Muschler P, Scharf B, Schmitt R (2000) VisN and VisR are global regulators of chemotaxis, flagellar, and motility genes in Sinorhizobium (Rhizobium) meliloti. J Bacteriol 182:782–788
Steenhoudt O, Vanderleyden J (2000) Azospirillum, a free-living nitrogen-fixing bacterium closely associated with grasses: genetic, biochemical and ecological aspects. FEMS Microbiol Rev 24:487–506
Tambalo DD, Del Bel KL, Bustard DE, Greenwood PR, Steedman AE, Hynes MF (2010) Regulation of flagellar, motility and chemotaxis genes in Rhizobium leguminosarum by the VisN/R-Rem cascade. Microbiology 156:1673–1685
Ulrich LE, Zhulin IB (2009) The MiST2 database: a comprehensive genomics resource on microbial signal transduction. Nucleic Acids Res 38:D401–407
Uren NC (2000) Types, amounts and possible functions of compounds released into the rhizosphere by soil-grown plants. In: Pinton R, Varanini Z, Nannipiero P (eds) The rhizosphere: Biochemistry and organic substances at the soil–plant interface. Marcel Dekker, New York, pp 19–20
Van Bastelaere E, Lambrecht M, Vermeiren H, Van Dommelen A, Keijers V, Proost P, Vanderleyden J (1999) Characterization of a sugar-binding protein from Azospirillum brasilense mediating chemotaxis to and uptake of sugars. Molec Microbiol 32:703–714
Vande Broek A, Lambrecht M, Vanderleyden J (1998) Bacterial chemotactic motility is important for the initiation of wheat root colonization by Azospirillum brasilense. Microbiology 144:2599–2606
Wadhams GH, Armitage JP (2004) Making sense of it all: bacterial chemotaxis. Nature Rev Mol Cell Biol 5:1024–1037
Walker TS, Bais HP, Grotewold E, Vivanco JM (2003) Root exudation and rhizosphere biology. Plant Physiol 132:44–51
Webb BA, Hildreth S, Helm RF, Scharf BE (2014) Sinorhizobium meliloti Chemoreceptor McpU mediates chemotaxis toward host plant exudates through direct proline sensing. Appl Environ Microbiol 80:3404–3415
Wisniewski-Dyé F et al (2011) Azospirillum genomes reveal transition of bacteria from aquatic to terrestrial environments. PLoS Genet 7:e1002430
Wuichet K, Zhulin IB (2010) Origins and diversification of a complex signal transduction system in prokaryotes. Sci Signal 3:ra50. doi:10.1126/scisignal.2000724
Xie Z, Ulrich LE, Zhulin IB, Alexandre G (2010) PAS domain containing chemoreceptor couples dynamic changes in metabolism with chemotaxis. Proc Natl Acad Sci USA 107:2235–2240
Yao SY et al (2004) Sinorhizobium meliloti ExoR and ExoS proteins regulate both succinoglycan and flagellum production. J Bacteriol 186:6042–6049
Yost CK, Rochepeau P, Hynes MF (1998) Rhizobium leguminosarum contains a group of genes that appear to code for methyl-accepting chemotaxis proteins. Microbiology 144:1945–1956
Yost CK, Clark KT, Del Bel KL, Hynes MF (2003) Characterization of the nodulation plasmid encoded chemoreceptor gene mcpG from Rhizobium leguminosarum. BMC Microbiol 3:1
Yost CK, Del Bel KL, Quandt J, Hynes MF (2004) Rhizobium leguminosarum methyl-accepting chemotaxis protein genes are down-regulated in the pea nodule. Arch Microbiol 182:505–513
Young JPW et al (2006) The genome of Rhizobium leguminosarum has recognizable core and accessory components. Genome Biol 7:R34. doi:10.1186/gb-2006-7-4-r34
Zatakia HM, Nelson CE, Syed UJ, Scharf BE (2014) ExpR coordinates the expression of symbiotically important, bundle-forming Flp pili with quorum sensing in Sinorhizobium meliloti. Appl Environ Microbiol 80:2429–2439
Acknowledgments
Research in our laboratories is funded by NSF-330344 (GMA), NSF-1253234 (BES) and NSERC Canada RGPIN 2015-03926 (MFH). Any opinion, findings, and conclusions or recommendations expressed in this material are those of the author(s) and do not necessarily reflect the views of the National Science Foundation.
Author’s contribution
BES, MFH and GMA wrote and revised the manuscript and designed the figures and tables.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Scharf, B.E., Hynes, M.F. & Alexandre, G.M. Chemotaxis signaling systems in model beneficial plant–bacteria associations. Plant Mol Biol 90, 549–559 (2016). https://doi.org/10.1007/s11103-016-0432-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-016-0432-4